Abstract
Genetic deletions and duplications known as copy number variants have been strongly implicated in genetic susceptibility to schizophrenia, autism, attention deficit hyperactivity disorder and epilepsy. The overall rate of copy number variants in the University College London (UCL) bipolar disorder sample was found to be slightly lower than the rate in controls. This finding confirms the results from other studies that have also shown no increased rate of copy number variants in bipolar disorder. However, some rare duplications and deletions were observed only in bipolar disorder cases and not in controls, these included some that had previously been detected only in rare cases of bipolar disorder. We conclude that copy-number variant analysis shows no obvious sharing of the same genetic susceptibility between schizophrenia and bipolar disorder. Copy number variants do not seem to have an important role in susceptibility to bipolar disorder, they may, however, still represent a rare cause of the disease, although the evidence for this is far from clear.
Similar content being viewed by others
Introduction
Bipolar affective disorder (BPAD) is a common mental health disorder that is characterised by periods of mania and depression. BPAD is associated with increased rates of unipolar affective disorder in relatives and with comorbid alcohol abuse,1, 2 as well as with very high rates of suicide.3 Family, twin and adoption studies have shown that genetic heritability to BPAD is in excess of 80%, which makes bipolar disorder one of the most heritable of the psychiatric disorders.4, 5, 6 Several BPAD genome-wide association studies (GWASs) using up to 1 000 000 single nucleotide polymorphism (SNP) markers have been performed and these data can also be used to detect copy number variants (CNVs).7, 8, 9, 10, 11, 12 The GWASs data have implicated two genes ANK3 and CACNA1C with genome-wide levels of statistical significance.7 Studies of copy-number variation causing diseases are complicated by the presence of a high background rate of duplications and deletions found across the human genome that are not associated with any disease.13, 14, 15, 16 CNVs seem to have developed through a variety of processes that seem to be dependent and the size of the structural variant.16 CNVs in specific chromosomal regions have been reported that are appreciably increased in samples of schizophrenia, particularly in early age of onset.8, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33 Copy number variants that are shared with schizophrenia but not with bipolar disorder have been found in epilepsy.29, 34, 35, 36, 37 CNVs have also been found to be strongly associated with autism, attention deficit hyperactivity disorder and learning disability.38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50
There is limited evidence for the involvement of CNVs in susceptibility to bipolar disorder. Zhang et al51 have reported that singleton deletions over 100 Kb in length are more frequent in BPAD cases, a finding which was not replicated by Grozeva et al.52 Lachman et al53 reported an increase in GSK3β CNVs in BPAD, but this finding was also not confirmed by Grozeva et al.52 However, there have been reports of rare cases of BPAD with CNVs32, 51, 52, 53, 54 and it remains the case that specific rare CNVs may contribute to susceptibility to bipolar disorder. The aim of this study was to assess the frequency of CNVs in the UCL bipolar case control sample and to investigate the location of rare CNVs with reference to bipolar disorder-specific CNVs reported by Zhang et al51 and Grozeva et al.52
Subjects and methods
Bipolar research subjects
BPAD cases and controls included in the analyses comprised DNA from individuals from the University College London (UCL) bipolar disorder sample collection (see Sklar et al10 and Ferreira et al7 for a description of the case and control samples). Additional UCL Bipolar II samples collected using the same ascertainment criteria as described above were included in the analysis presented in this study.
The UCL Bipolar sample consisted of 97% bipolar 1 cases with psychotic symptoms according to Research Diagnostic Criteria (RDC) categories.55 The comparison subjects were 510 screened normal volunteers with no personal or first degree family history of any mental disorder. The cases and comparison subjects were selected if both their parents and all four grandparents were of Irish, Welsh, Scottish or English ancestry as defined by an ancestry checklist. Subjects were also included if one of the four grandparents was of European ancestry before the 2004 EU enlargement. U.K. National Health Service (NHS) multicenter and local research ethics committee approval was obtained. All subjects signed an approved consent form after reading an information sheet and after a description of the study had been given to each subject. All subjects were interviewed by a psychiatrist who used the lifetime version of the Schizophrenia and Affective Disorders Schedule.55 All the bipolar patients were also rated with the 90-item OPCRIT checklist.56 Family pedigree diagrams were drawn and drug-treatment response was recorded. DNA was extracted from whole blood using standard nuclear lysis phenol chloroform methods.
Genotyping
Genotyping was performed on the Affymetrix GeneChip Human Mapping 500 K Array Set (Santa Clara, CA, USA) by the Genetic Analysis Platform at the Broad Institute of Harvard and MIT as described previously.10 The method uses two arrays according to whether the genomic DNA from research subjects was digested with the restriction enzymes NspI or StyI before adaptor nucleotides are ligated with T4 DNA ligase. Because each array can replicate evidence for copy number variants, only cases where data for both arrays was available were analysed for CNVs (546 cases and 517 controls).
CNV Calling
CNV calling was performed using PennCNV software (version 2008 June 26; http://www.openbioinformatics.org/penncnv/).57 PennCNV uses the total fluorescent intensity signals for both alleles (log R ratio (LRR)) and the B allele frequency (BAF) for each SNP on the genotyping arrays. Standard PennCNV quality control checks are designed for Illumina arrays and are therefore not appropriate for the Affymetrix GeneChip Human Mapping 500 K Array used here (http://www.openbioinformatics.org/penncnv/penncnv_tutorial_affy_gw6.html). Samples, in which more than 100 CNVs were detected by PennCNV were excluded from further analysis (8 BPAD; 46 Controls). CNVs that were smaller than 50 Kb in length or comprised fewer than 10 SNPs were then removed from the analysis, as these were likely to be unreliable CNV calls. CNVs that spanned a centromere or that overlapped at least 50% of their length with regions previously described as being prone to false positives due to somatic mutations were also removed.22 A further 15 CNVs were also removed that were comprised of SNPs from only one of the two arrays. Finally, for the purpose of rare CNV burden analysis CNVs found in more than 1% of cases and controls were not considered further. A custom track for visualisation of CNVs detected in this paper on the UCSC Genome Browser (University of California, Santa Cruz, CA, USA; http://genome.ucsc.edu/), is available at http://www.ucl.ac.uk/~rejuamc/UCLBPCO.cnv.bed.
Analysis of Rare and singleton CNVs
CNV association analyses were performed with PLINK58 (version 1.07, http://pngu.mgh.harvard.edu/~purcell/plink/). Empirical significance values (two tailed) were derived with the use of 10 000 permutations.
Genomic locations presented here are from the March 2006 human genome sequence assembly (UCSC Hg18, National Center for Biotechnology Information build 36). Analyses of CNVs >50 and >100 kb were performed to allow comparison of results from previous studies that only investigated the larger class of CNV.
Results
Global burden of rare CNVs
The numbers of rare CNVs in each size group, their frequencies in cases and controls and the corresponding empirical significance values (where <0.05) are shown in Table 1. Two of the comparisons yielded nominally significant results. In the CNV size range between 200 and 500 Kb, there were significantly fewer deletions (0.183 CNV per control sample vs 0.115 per BPAD case; P=0.039; see Table 1) and when both deletion and duplication CNVs >100 Kb in size were considered there were also significantly fewer CNVs in the bipolar cases (0.818 per sample) compared with the controls (1.013 per sample; P=0.036; see Table 1). Analyses of CNV burden excluding CNVs <100 Kb were performed to allow comparison of the data with that presented in previous studies.
Singleton CNV analysis
The results of singleton CNV analysis are shown in Table 2. This is an attempt to replicate the findings of Zhang et al.51 In contrast to the study of Zhang et al, no significant over representation of bipolar singleton deletions were observed. We did however, observe a nominally significant increase in the frequency of control duplication CNVs over 100 Kb in size (P=0.03). This finding does not survive correction for the multiple tests carried out. It is noteworthy that there was also a nonsignificant increase in the rate of singleton duplications in the controls of Grozeva et al.52 We also compared the rate of singleton CNVs in cases of BPAD where a first episode of mania occurred at age 18 or below with BPAD samples with a later first episode of mania. No significant differences were observed in the comparison groups (data not shown). However, the rate of singleton deletions among the later age of onset case was higher (0.339) compared with the early onset cases (0.277). This finding is in the opposite direction to that of Zhang et al,51 but in agreement with the findings of Grozeva et al.52
Bipolar disorder case only CNVs
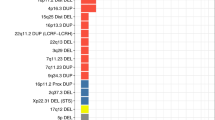
Although the overall burden of CNVs in cases was not found to be significantly different from that found in controls, we were interested to investigate CNVs found only in two or more bipolar disorder cases that disrupted genes. A total of 26 such CNVs regions were identified (see Table 3). Two of these overlapped with genes that were either duplicated or deleted by CNVs in BPAD cases only in the Grozeva et al52 study, one overlapped with genes also disrupted by the Zhang et al51 BPAD case CNVs, one CNV overlapped with genes also disrupted by ISC8 schizophrenia case only CNVs, and one CNV included NDE1 a gene previously found deleted in schizophrenia27 (see Table 3). Three of the genes in the CNV regions had also previously been found to be associated with neuropsychiatric phenotypes (see Table 3).
Discussion
The results of rare copy-number variation analysis presented here are similar to those of Grozeva et al52 and indicate that the larger type of CNVs that are currently detectable do not have a major role in susceptibility to BPAD. In agreement with Grozeva et al,52 we paradoxically report a small decrease in rare CNV burden in our BPAD cases compared with controls. Also in agreement with Grozeva et al,52 we do not find evidence for a previously reported increase51 in singleton deletion CNVs in our bipolar disorder cases nor did we find significant age of onset effects with singleton deletion CNVs in our sample. The numbers of samples used in this study are smaller than those of Grozeva et al52 and Zhang et al.51 The microarray technology used in this study was the same as in the study by Grozeva et al,52 however, the calling algorithms and quality control procedures differ slightly. Zhang et al51 used a higher density microarray platform, which might explain the difference between our own and the Grozeva et al52 compared with the study by Zhang et al.51
There was an overall increase in the rate of CNVs called in our study compared with the rate reported by Grozeva et al52 and Zhang et al.51 Call rates may be very sample dependent with differences in thresholds for CNV calling and the relatively small sample sizes influencing these rates. Grozeva et al52 detected CNVs separately on the two arrays and then combined the CNV calls. Their calling threshold required that 10 SNPs from one array showed consistent evidence for the presence of a CNV. We used a different approach where the intensity (LRR) and allele frequency data (BAF) from the two arrays were combined before CNV calling. Thus, despite the fact that both studies required 10 consecutive SNPs to show consistent evidence for a CNV to be called, the approach used here is likely to be more sensitive. We further required that all CNVs were made up of SNP data from both of the arrays such that localised microarray hybridisation artefacts would be unlikely to produce aberrant CNV calls. Zhang et al51 used a higher density microarray compared with the arrays used in this study therefore, one would expect that their study would have been capable of detecting more CNVs than we report in this study. Again the calling algorithms are different and this may account for the differences in call rates and indeed for the lack of replication in the frequency of singleton deletions in BPAD cases.
The evidence from our own study and from the literature for rare case only CNVs having a role in BPAD is not clear. However, we present a list of CNVs that occurred in two or more cases and indicate those for whom previous bipolar disorder or schizophrenia studies have also found evidence for genes being deleted or duplicated by CNVs or where there is evidence for genetic association with bipolar disorder or related neuropsychiatric phenotypes. Although some of these overlapping findings might be due to chance, future studies will be able to replicate these rare occurrences with greater power.
References
Frye MA, Salloum IM : Bipolar disorder and comorbid alcoholism: prevalence rate and treatment considerations. Bipolar Disord 2006; 8: 677–685.
Regier DA, Farmer ME, Rae DS et al: Comorbidity of mental disorders with alcohol and other drug abuse. Results from the Epidemiologic Catchment Area (ECA) Study. Jama 1990; 264: 2511–2518.
Chen YW, Dilsaver SC : Lifetime rates of suicide attempts among subjects with bipolar and unipolar disorders relative to subjects with other Axis I disorders. Biol Psychiatry 1996; 39: 896–899.
Blackwood DH, Visscher PM, Muir WJ : Genetic studies of bipolar affective disorder in large families. Br J Psychiatry Suppl 2001; 41: s134–s136.
Craddock N, Jones I : Molecular genetics of bipolar disorder. Br J Psychiatry 2001; 178: S128–S133.
Rifkin L, Gurling H : Genetic aspects of affective disorders; in: Horton R, Katona C (eds).: Biological Aspects of Affective Disorders. London: Academic Press, 1991, pp 305–329.
Ferreira MA, O’Donovan MC, Meng YA et al: Collaborative genome-wide association analysis supports a role for ANK3 and CACNA1C in bipolar disorder. Nat Genet 2008; 40: 1056–1058.
ISC: Rare chromosomal deletions and duplications increase risk of schizophrenia. Nature 2008; 455: 237–241.
Scott LJ, Muglia P, Kong XQ et al: Genome-wide association and meta-analysis of bipolar disorder in individuals of European ancestry. Proc Natl Acad Sci USA 2009; 106: 7501–7506.
Sklar P, Smoller JW, Fan J et al: Whole-genome association study of bipolar disorder. Mol Psychiatry 2008; 13: 558–569.
Smith EN, Bloss CS, Badner JA et al: Genome-wide association study of bipolar disorder in European American and African American individuals. Mol Psychiatry 2009; 14: 755–763.
WTCCC : Genome-wide association study of 14 000 cases of seven common diseases and 3000 shared controls. Nature 2007; 447: 661–678.
Feuk L, Carson AR, Scherer SW : Structural variation in the human genome. Nat Rev Genet 2006; 7: 85–97.
Iafrate AJ, Feuk L, Rivera MN et al: Detection of large-scale variation in the human genome. Nat Genet 2004; 36: 949–951.
Lupski JR : Genomic rearrangements and sporadic disease. Nat Genet 2007; 39: S43–S47.
Conrad DF, Pinto D, Redon R et al: Origins and functional impact of copy number variation in the human genome. Nature 2010; 464: 704–712.
Stefansson H, Rujescu D, Cichon S et al: Large recurrent microdeletions associated with schizophrenia. Nature 2008; 455: 232–236.
Kirov G, Grozeva D, Norton N et al: Support for the involvement of large copy number variants in the pathogenesis of schizophrenia. Hum Mol Genet 2009; 18: 1497–1503.
Kirov G, Gumus D, Chen W et al: Comparative genome hybridization suggests a role for NRXN1 and APBA2 in schizophrenia. Hum Mol Genet 2008; 17: 458–465.
McCarthy SE, Makarov V, Kirov G et al: Microduplications of 16p11.2 are associated with schizophrenia. Nat Genet 2009; 41: 1223–1227.
Ikeda M, Aleksic B, Kirov G et al: Copy number variation in schizophrenia in the Japanese population. Biol Psychiatry 2010; 67: 283–286.
Need AC, Ge D, Weale ME et al: A genome-wide investigation of SNPs and CNVs in schizophrenia. PLoS Genet 2009; 5: e1000373.
Walsh T, McClellan JM, McCarthy SE et al: Rare structural variants disrupt multiple genes in neurodevelopmental pathways in schizophrenia. Science 2008; 320: 539–543.
Kirov G, Rujescu D, Ingason A, Collier DA, O’Donovan MC, Owen MJ : Neurexin 1 (NRXN1) deletions in schizophrenia. Schizophr Bull 2009; 35: 851–854.
Karayiorgou M, Morris MA, Morrow B et al: Schizophrenia susceptibility associated with interstitial deletions of chromosome 22q11. Proc Natl Acad Sci USA 1995; 92: 7612–7616.
Yan W, Jacobsen LK, Krasnewich DM et al: Chromosome 22q11.2 interstitial deletions among childhood-onset schizophrenics and ‘multidimensionally impaired’. Am J Med Genet 1998; 81: 41–43.
Ingason A, Rujescu D, Cichon S et al: Copy number variations of chromosome 16p13.1 region associated with schizophrenia. Mol Psychiatry 2009.
Rujescu D, Ingason A, Cichon S et al: Disruption of the neurexin 1 gene is associated with schizophrenia. Hum Mol Genet 2009; 18: 988–996.
Friedman JI, Vrijenhoek T, Markx S et al: CNTNAP2 gene dosage variation is associated with schizophrenia and epilepsy. Mol Psychiatry 2008; 13: 261–266.
Vrijenhoek T, Buizer-Voskamp JE, van der Stelt I et al: Recurrent CNVs disrupt three candidate genes in schizophrenia patients. Am J Hum Genet 2008; 83: 504–510.
Bassett AS, Marshall CR, Lionel AC, Chow EW, Scherer SW : Copy number variations and risk for schizophrenia in 22q11.2 deletion syndrome. Hum Mol Genet 2008; 17: 4045–4053.
Wilson GM, Flibotte S, Chopra V, Melnyk BL, Honer WG, Holt RA : DNA copy-number analysis in bipolar disorder and schizophrenia reveals aberrations in genes involved in glutamate signaling. Hum Mol Genet 2006; 15: 743–749.
Moon HJ, Yim SV, Lee WK et al: Identification of DNA copy-number aberrations by array-comparative genomic hybridization in patients with schizophrenia. Biochem Biophys Res Commun 2006; 344: 531–539.
de Kovel CG, Trucks H, Helbig I et al: Recurrent microdeletions at 15q11.2 and 16p13.11 predispose to idiopathic generalized epilepsies. Brain 2010; 133 (Part 1): 23–32.
Dibbens LM, Mullen S, Helbig I et al: Familial and sporadic 15q13.3 microdeletions in idiopathic generalized epilepsy: precedent for disorders with complex inheritance. Hum Mol Genet 2009; 18: 3626–3631.
Heinzen EL, Radtke RA, Urban TJ et al: Rare deletions at 16p13.11 predispose to a diverse spectrum of sporadic epilepsy syndromes. Am J Hum Genet 2010; 86: 707–718.
Helbig I, Mefford HC, Sharp AJ et al: 15q13.3 microdeletions increase risk of idiopathic generalized epilepsy. Nat Genet 2009; 41: 160–162.
Ching MS, Shen Y, Tan WH et al: Deletions of NRXN1 (neurexin-1) predispose to a wide spectrum of developmental disorders. Am J Med Genet B Neuropsychiatr Genet; 2010; 153B: 937–947.
Ben-Shachar S, Lanpher B, German JR et al: Microdeletion 15q13.3: a locus with incomplete penetrance for autism, mental retardation, and psychiatric disorders. J Med Genet 2009; 46: 382–388.
Elia J, Gai X, Xie HM et al: Rare structural variants found in attention-deficit hyperactivity disorder are preferentially associated with neurodevelopmental genes. Mol Psychiatry 2010; 15: 637–646.
Kumar RA, KaraMohamed S, Sudi J et al: Recurrent 16p11. microdeletions in autism. Hum Mol Genet 2008; 17: 628–638.
Marshall CR, Noor A, Vincent JB et al: Structural variation of chromosomes in autism spectrum disorder. Am J Hum Genet 2008; 82: 477–488.
Mefford HC, Sharp AJ, Baker C et al: Recurrent rearrangements of chromosome 1q21.1 and variable pediatric phenotypes. N Engl J Med 2008; 359: 1685–1699.
Pagnamenta AT, Wing K, Sadighi Akha E et al: A 15q13.3 microdeletion segregating with autism. Eur J Hum Genet 2009; 17: 687–692.
Sebat J, Lakshmi B, Malhotra D et al: Strong association of de novo copy number mutations with autism. Science 2007; 316: 445–449.
Sharp AJ, Mefford HC, Li K et al: A recurrent 15q13.3 microdeletion syndrome associated with mental retardation and seizures. Nat Genet 2008; 40: 322–328.
Sykes NH, Toma C, Wilson N et al: Copy number variation and association analysis of SHANK3 as a candidate gene for autism in the IMGSAC collection. Eur J Hum Genet 2009; 17: 1347–1353.
Szatmari P, Paterson AD, Zwaigenbaum L et al: Mapping autism risk loci using genetic linkage and chromosomal rearrangements. Nat Genet 2007; 39: 319–328.
Ullmann R, Turner G, Kirchhoff M et al: Array CGH identifies reciprocal 16p13.1 duplications and deletions that predispose to autism and/or mental retardation. Hum Mutat 2007; 28: 674–682.
Weiss LA, Shen Y, Korn JM et al: Association between microdeletion and microduplication at 16p11.2 and autism. N Engl J Med 2008; 358: 667–675.
Zhang D, Cheng L, Qian Y et al: Singleton deletions throughout the genome increase risk of bipolar disorder. Mol Psychiatry 2009; 14: 376–380.
Grozeva D, Kirov G, Ivanov D et al: Rare copy number variants: a point of rarity in genetic risk for bipolar disorder and schizophrenia. Arch Gen psychiatry 2010; 67: 318–327.
Lachman HM, Pedrosa E, Petruolo OA et al: Increase in GSK3beta gene copy number variation in bipolar disorder. Am J Med Genet B Neuropsychiatr Genet 2007; 144B: 259–265.
Craddock N, Hurles ME, Cardin N et al: Genome-wide association study of CNVs in 16 000 cases of eight common diseases and 3000 shared controls. Nature 2010; 464: 713–720.
Endicott J, Spitzer RL : A diagnostic interview: the schedule for affective disorders and schizophrenia. Arch Gen psychiatry 1978; 35: 837–844.
McGuffin P, Farmer A, Harvey I : A polydiagnostic application of operational criteria in studies of psychotic illness. Development and reliability of the OPCRIT system. Arch Gen psychiatry 1991; 48: 764–770.
Wang K, Li M, Hadley D et al: PennCNV: an integrated hidden Markov model designed for high-resolution copy number variation detection in whole-genome SNP genotyping data. Genome Res 2007; 17: 1665–1674.
Purcell S, Neale B, Todd-Brown K et al: PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 2007; 81: 559–575.
Jamain S, Betancur C, Quach H et al: Linkage and association of the glutamate receptor 6 gene with autism. Mol Psychiatry 2002; 7: 302–310.
Ahn Jo S, Ahn K, Kim JH et al: ApoE-epsilon 4-dependent association of the choline acetyltransferase gene polymorphisms (2384G>A and 1882G>A) with Alzheimer's disease. Clin Chim Acta 2006; 368: 179–182.
Kim KW, Suh YJ, Park WY et al: Choline acetyltransferase G +4 A polymorphism confers a risk for Alzheimer's disease in concert with Apolipoprotein E epsilon4. Neurosci Lett 2004; 366: 182–186.
Mubumbila V, Sutter A, Ptok U, Heun R, Quirin-Stricker C : Identification of a single nucleotide polymorphism in the choline acetyltransferase gene associated with Alzheimer's disease. Neurosci Lett 2002; 333: 9–12.
Hennah W, Tomppo L, Hiekkalinna T et al: Families with the risk allele of DISC1 reveal a link between schizophrenia and another component of the same molecular pathway, NDE1. Hum Mol Genet 2007; 16: 453–462.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
McQuillin, A., Bass, N., Anjorin, A. et al. Analysis of genetic deletions and duplications in the University College London bipolar disorder case control sample. Eur J Hum Genet 19, 588–592 (2011). https://doi.org/10.1038/ejhg.2010.221
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2010.221
Keywords
This article is cited by
-
Genomic Disorders in Psychiatry—What Does the Clinician Need to Know?
Current Psychiatry Reports (2017)
-
Copy number variation in bipolar disorder
Molecular Psychiatry (2016)
-
A pilot study on commonality and specificity of copy number variants in schizophrenia and bipolar disorder
Translational Psychiatry (2016)
-
Genomic structural variation in affective, anxiety, and stress-related disorders
Journal of Neural Transmission (2015)
-
Genomic insights into the overlap between psychiatric disorders: implications for research and clinical practice
Genome Medicine (2014)