Abstract
Human Y-chromosome haplogroup structure is largely circumscribed by continental boundaries. One notable exception to this general pattern is the young haplogroup R1a that exhibits post-Glacial coalescent times and relates the paternal ancestry of more than 10% of men in a wide geographic area extending from South Asia to Central East Europe and South Siberia. Its origin and dispersal patterns are poorly understood as no marker has yet been described that would distinguish European R1a chromosomes from Asian. Here we present frequency and haplotype diversity estimates for more than 2000 R1a chromosomes assessed for several newly discovered SNP markers that introduce the onset of informative R1a subdivisions by geography. Marker M434 has a low frequency and a late origin in West Asia bearing witness to recent gene flow over the Arabian Sea. Conversely, marker M458 has a significant frequency in Europe, exceeding 30% in its core area in Eastern Europe and comprising up to 70% of all M17 chromosomes present there. The diversity and frequency profiles of M458 suggest its origin during the early Holocene and a subsequent expansion likely related to a number of prehistoric cultural developments in the region. Its primary frequency and diversity distribution correlates well with some of the major Central and East European river basins where settled farming was established before its spread further eastward. Importantly, the virtual absence of M458 chromosomes outside Europe speaks against substantial patrilineal gene flow from East Europe to Asia, including to India, at least since the mid-Holocene.
Similar content being viewed by others
Introduction
Human populations across the world are characterized by generally low genetic differences as compared with their intrapopulation variation. These differences can be quantitative, pronounced in different frequencies of the same derived states of ancient polymorphic markers (eg, majority of the HapMap markers1), or qualitative, in which case younger derived variants are found restricted to a particular geographic region or population. The Y-chromosome haplogroup structure frequently shows a good qualitative correlation with continental boundaries, and the geographic specificity of the markers can most often be explained by their phylogenetic descent order rather than by drift alone.2, 3 Recently evolved polymorphisms unless amplified by selection or specific founder effects tend to have low frequencies in modern populations, characterized generally by increased effective population sizes in the Holocene period. One of the notable outliers to this rule, because of its high frequency and young age, is the transcontinental spread of haplogroup R1a.4, 5 Early observations have led to various interpretations associating R1a phylogeography with certain cultural developments of the past. Even though R1a occurs as the most frequent Y-chromosome haplogroup among populations representing a wide variety of language groups, such as Slavic, Indo-Iranian, Dravidian, Turkic and Finno-Ugric, many authors have been particularly interested in the link between R1a and the Indo-European language family. For example, R1a frequency patterns have been discussed6, 7 in the context of the purported link connecting Indo-European-speaking pastoralists and the archeological evidence on the distribution of the Kurgan culture in the Pontic steppe.8 A more precise interpretation of the underlying prehistoric and historic episodes of R1a chromosomes across this wide span of Eurasian geography remains largely unknown because of insufficient information on the phylogenetic subdivisions within haplogroup R1a. We address this shortcoming here by analyzing more than 11 000 DNA samples from across Eurasia, including more than 2000 from haplogroup R1a to ascertain the phylogenetic information of the newly discovered R1a-related SNPs. We also examine the STR diversity of the associated R1a subclades to better understand the demographic history and prehistoric cultural associations of one of the most widely spread and frequent Y-chromosome haplogroups in the world with post-Last Glacial Maximum origin.
Materials and methods
Twelve recently reported R1a markers ascertained in one R1a1 individual2, 9 across extensive but unspecified coverage and two new SNPs discovered in two R1a1 individuals during a scan of ∼44 kb10 were genotyped by denaturing high-performance liquid chromatography (DHPLC) and confirmed by direct sequencing in an initial screening of 18 DNA samples belonging to haplogroup R1a from different geographic regions spanning Scandinavia to India. Twelve of these markers were derived in all individuals carrying the M17 mutation, whereas one of the markers, Page68, exhibited an ancestral allele in all samples and was therefore not evaluated further. In addition, two new SNPs were discovered. One (M434) while surveying another SNP reported in the flanking sequence of DYS43811 by DHPLC in a globally representative collection of DNAs that included individuals from Pakistan, and another (M458) was discovered during the initial survey of the Hinds et al9 rs17250901 homopolymer variant. Markers M434 and M458 were variable in a subset of the 18 R1a screening samples and represent new informative subclades of R1a1. Another SNP (M334) was ascertained previously by DHPLC in one Estonian in a panel of 48 R1a1 samples. Marker M334 was not observed in an additional survey of 100 R1a1 Estonian samples and was not studied further. In the population surveys, the markers were genotyped either by DHPLC, RFLP or TaqMan (Applied Biosystems, Foster City, CA, USA) assays. Within specific haplogroups, median-joining networks were constructed. Specifications for the analyses are detailed in the relevant figure legends. The age of microsatellite variation within haplogroups was evaluated using the methodology described by Zhivotovsky et al12 as modified according to Sengupta et al13 using microsatellite evolutionary effective mutation rate of 6.9 × 10−4 per 25 years. Sample sizes and frequencies of the main R1a subclades are reported in Supplementary Tables 1–3. STR haplotype data are given in Supplementary Tables 4, 6 and 7. Supplementary Table 5 reports the primer sequences used in genotyping the informative SNPs.
Results and discussion
By using the new SNP markers, we were able to fractionate the R1a defining node into a nested series of branches that are reinforced by multiple phylogenetically equivalent mutations (inset, Figure 1). All chromosomes unresolved previously beyond the R1-M173* level14, 15, 35, 36 that were available to us are now attributed to either R1a*-M420 or R1b*-M343 haplogroups. Consequently, we revise the haplogroup nomenclature following the YCC guidelines.3, 5 Although the occurrences of the most basal haplogroup R1a*-M420(xSRY10831.2) and the intermediate haplogroup R1a1*-SRY10831.2(xM17) are rare (Supplementary Table S1), the descendent haplogroup R1a1a-M17 assemblage displays informative frequencies above a few percent in populations comprising a broad expanse of Eurasian geography ranging from Norway and Northeast Asia to south India, whereas frequencies above 10% occur in East Europe, West, South and central Asia (Supplementary Table S2, Figure 1). With the exception of a few localized low-frequency subhaplogroups,4, 14, 37 the majority of haplogroup R1a1a chromosomes have remained so far phylogenetically indistinct.
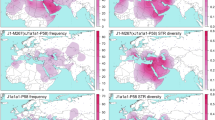
Geographic distribution of haplogroup R1a1a frequency. Spatial frequency map was obtained applying the frequencies from Supplementary Table S2 and for 8429 individuals representing 118 populations from literature.7, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34 Dots on the map indicate the approximate locations of the sampled populations. The frequency data were converted to isofrequency maps in Surfer software (version 7, Golden Software Inc., Golden, CO, USA) following the Kriging procedure. The inset map illustrates the available data (Supplementary Table S2) for the regional expansion times in KYA (thousands of years ago) of M17 Y-chromosomes. We note that especially in the latter case the density of the data points is too low for any viable geostatistical analyses. Phylogenetic tree relating SNP markers that define haplogroup R1a and its subgroups is shown in the inset. Previously described SNP markers3 are underlined. Markers M56, M157, M64.2, M87, M204, P98 and PK5 shown in gray font were not typed as they were previously detected at nonpolymorphic frequencies in other studies. PCR amplicons for 12 SNPs from Hinds et al9 (M420, M448, M449, M459, M511, M513, M516 and rs17250901) and 2 from Repping et al10 (Page07 and Page68) were designed and tested for male specificity using female control DNA. The phylogenetic relationships of these SNPs were evaluated in a geographically diverse panel of 18 R1a1 samples and 2 R1b* samples ranging from Northwest Europe to South Asia using DHPLC technology, and confirmed by direct sequencing of representative samples. Detailed specifications for these markers are given in Supplementary Table S5.
Recent Arabian Sea gene flow
The marker M434, defining the novel Y-chromosome haplogroup R1a1a6, was observed altogether in 14 individual samples in our screening of 691 R1a1a chromosomes (Supplementary Table S3). Given these data, the haplogroup R1a1a6 distribution seems to be restricted mainly to Pakistan whereas the Omani R1a1a6 samples, all three of which share the same STR haplotype, indicate recent gene flow across the Persian Gulf. The low STR haplotype diversity of R1a1a6 and its absence in 212 Indian R1a1a samples suggest that the M434 mutation may have arisen recently in Pakistan.
In situ diversification in Central Europe
In contrast to the restricted geographic pattern of M434, the R1a1a7 defining marker, M458, was found to be variable in a number of populations, and thus it provides the first significant geographic compartmentalization within the overarching haplogroup R1a distribution. The haplogroup R1a1a7 distribution is confined to Central and Eastern Europe and does not extend eastward beyond the Ural Mountains or southward beyond Turkey (Supplementary Table S2, Figure 2). Its spread in the Caucasus is specific: although absent in the Dagestanian group, it is present at low frequencies both in the northwestern and southern populations, and in particular in Karanogays, who only relatively recently were spread as pastoral nomadic people alongside the Ponto-Caspian steppe belt. The highest frequency of haplogroup R1a1a7 (over 30%) is observed in Central and Southern Poland. Frequencies higher than 10% occur among Western and Eastern Slavic populations whereas elsewhere in Europe, including Southern Slavic groups, the frequency of the derived M458G allele decreases rapidly away from its frequency peak that coincides broadly with the overall R1a1a frequency maximum in Poland (Figures 1 and 2). The R1a1a*(xM458) chromosomes on the other hand are less frequent in Poland and display frequency maximums in Belarus and southwest Russia (Supplementary Table S2).
Geographic distribution of haplogroup R1a1a7-M458 frequency. The spatial frequency map was obtained applying the frequencies from Supplementary Table S2 (dots on the map indicate the approximate locations of the sampled populations) to the Surfer software (version 7, Golden Software Inc., Golden, CO, USA) following the Inverse Distance to Power (Power 3.75; smoothness 0) procedure with added break lines indicated by dashed blue lines in the seas. Spatial distribution of the expansion times of the regional M458 derived Y-chromosomes is shown in the lower left inset map according to data in Supplementary Table S4. See text for discussion concerning the spread of M458 lineages with the major European river basins (shown in blue) and major Neolithic and Metal Age cultures.
Analysis of associated STR diversity profiles revealed that among the R1a1a*(xM458) chromosomes the highest diversity is observed among populations of the Indus Valley yielding coalescent times above 14 KYA (thousands of years ago), whereas the R1a1a* diversity declines toward Europe where its maximum diversity and coalescent times of 11.2 KYA are observed in Poland, Slovakia and Crete. As islands such as Crete have been subject to multiple episodes of colonization from different source regions, it is not inconsistent that R1a1a* Td predates the date of its first colonization by the first farmers approximately 9 KYA.38 Also noteworthy is the drop in R1a1a* diversity away from the Indus Valley toward central Asia (Kyrgyzstan 5.6 KYA) and the Altai region (8.1 KYA) that marks the eastern boundary of significant R1a1a* spread (Figure 1, Supplementary Table S4.). In Europe, Poland also has the highest R1a1a7-M458 diversity, corresponding to approximately an 11 KYA coalescent time (Supplementary Table S4). Other populations in Europe exhibit declining diversity when sampled at increasing distance away from Central Europe (Figure 2). Westward of the Rhine overall R1a1a frequency is low, signaling a genetic boundary with R1b varieties.39 However, the patterns of currently observed Y-chromosome diversity in East/Central Europe are unlikely to be explained solely by population movements of the last century.40
Although the median STR haplotype of the derived M458G allele differs from the median type of the ancestral M458A chromosomes at 3 of the 10 STR loci considered in our analyses, the STR data alone are not informative for unambiguous inference of whether an individual has the A or G allele (Supplementary Figures S1 and S2) underscoring the extent of STR saturation and the importance of SNP genotyping to assess phylogenetic ancestry even among closely related lineages.
Phylogeography
Haplogroup frequency, haplotype diversity and coalescent times are three parameters that can be considered as informative for making inferences about the origins and polarity of spread of alleles among populations. The most distantly related R1a chromosomes, that is, both R1a* and R1a1* (inset, Figure 1), have been detected at low frequency in Europe, Turkey, United Arab Emirates, Caucasus and Iran14, 41 (Supplementary Table S1). The highest STR diversity of R1a1a*(xM458) chromosomes are observed outside Europe, in particular in South Asia (Figure 1, Supplementary Table S4), but given the lack of informative SNP markers the ultimate source area of haplogroup R1a dispersals remains yet to be refined.
In Europe a large proportion of the R1a1a variation is represented by its presently identified subclade R1a1a7-M458 that is virtually absent in Asia. Its major frequency and relatively low diversity in Europe can be explained thus by a founder effect that according to our coalescent time estimation falls into the early Holocene period, 7.9±2.6 KYA (Supplementary Table S4). The highest regional date of 10.7±4.1 KYA among Polish R1a1a7 carriers falls into the period of recolonization of this region by Mesolithic (Swiderian and subsequent cultures) settlers.42, 43 The time window of 10−5 KYA BP is a culturally complex juncture period between the Mesolithic and early Neolithic in Europe, thus, not allowing us to relate founder effect with any particular culture specifically. Most broadly, the autochthonous European origin of haplogroup R1a1a7, its narrow spatial distribution and the inversely related decreasing expansion times with increased distance from its core frequency and diversity area are suggestive of a notably successful demic expansion starting from a small subset of radiating founder lineages during the early Holocene period. It should be noted, though, that the inevitably large error margins of our coalescent time estimates do not allow us to exclude its association with the establishment of the mainstream Neolithic cultures, including the Linearbandkeramik (LBK), that flourished ca. 7.5−6.5 KYA BP in the Middle Danube (Hungary) and was spread further along the Rhine, Elbe, Oder, Vistula river valleys and beyond the Carpathian Basin.44
Migratory and early agricultural zones
River valleys are migratory corridors for organisms including humans and such riparian habitats provide opportunities for the forager lifestyle, settled agriculture and establishment of trade networks. The Neolithic communities in Central Europe were primarily located on the margins of river valleys with fertile soils at elevations less than 500 m.45 Haplogroup R1a1a7-M458 diversity and frequency are highest in river basins known to be associated with several early and late Neolithic cultures (Figure 2, Supplementary Figure S3). Assuming the founder effect we detect originated in the sparse Mesolithic population of Central-North Europe, the genetic evidence suggests strong cultural interaction and admixture occurred between the pioneer horticultural groups and local foragers, which resulted in widespread adaptation of the Neolithic lifestyle by indigenous residents. This interpretation is consistent with computational models indicating that although the process of the expansion of farming communities throughout much of Europe would have been demic, even minute amounts of gene flow from foragers over a long time period would have lead to a predominantly Mesolithic contribution to their admixed offspring.46 Following this model, it would not be surprising to associate a localized Neolithic demic expansion with a genetic lineage absent in the Fertile Crescent where farming originated and where other Y-chromosome haplogroups, such as G and J, have been associated with the initial demic spread of farming toward Southeast Europe.38 However, it should be noted that ancient mtDNA evidence from the Central European Mesolithic and LBK sites shows a lack of substantial continuity between Mesolithic, Neolithic and presently living populations of the area.47, 48 Notably, mtDNA haplogroups R1a, U4, U5, HV3 and HV4, which have been inferred to have pre-Neolithic spread in East Europe, occur at marginally low frequencies in India.49
It is noteworthy that the LCT-13910T allele associated with lactase persistence and agricultural pastoralism overlaps broadly with the spatial distribution50 of the derived M458G allele. Direct ancient DNA evidence suggests that the lactase persistence allele would have reached high frequency in this area, likely due to strong positive selection, only after the LBK period.51 However, computer simulations have shown that its increased frequency particularly in North Europe does not necessarily imply stronger effect of positive selection there than in other parts of Europe.52 Ancient DNA evidence for the Y-chromosome M458G allele is still lacking and it is therefore possible only to speculate about its existence and prevalence in Neolithic Europe. Beyond its spread in the Central European river basins (Figure 2), the LBK extended around the northern Carpathians into the steppe zone of Ukraine and participated in the establishment of the Criş culture.53 Our data showing high frequency of R1a1a north of the Carpathians and its lower frequency to the South, in the Tisza river valley, are consistent with the genetic boundary previously reported for this region.16
Copper and Bronze age parallels
Figure 2 also shows a remarkable geographic concordance of the R1a1a7-M458 distribution with the Chalcolithic and Early Bronze Age Corded Ware (CW) cultures of Europe that prospered from ca. 5.5−4.5 KYA BP.54 Ancient DNA evidence from a 4600-year-old multiple burial unearthed near Eulau, Germany and attributed to the Central European CW culture, identified the remains of three males carrying the SRY10831.2 mutation and sharing the same YSTR haplotype, implying a single family lineage.55 Although haplogroup affiliation cannot be inferred with certainty from STR data alone, a composite 15-locus YSTR haplotype representing the ancient lineage suggests its potential R1a1a*(xM458) membership due to four alleles (DYS391=11, DYS439=10, DYS389B=17 and DYS458=15) shared with the median R1a1a*(xM458) haplotype (Supplementary Tables S4 and S7). Interestingly, from the list of regional median haplotypes, the ancient haplotype is most similar to the German R1a1a*(xM458) type.
Indo-Europeans
A final comment can be made concerning the relationship between R1a phylogeography and contested origin of Indo-Europeans that is generally, though not solely, attributed to either Anatolia, the South Caucasus or the North Pontic-Caspian regions (Gray and Atkinson56 and references therein). Haplogroup R1a1a occurs in all three of these areas and beyond at informative frequencies (Figure 1). Consistent with its wide geographic spread, the coalescent time estimates of R1a1a correlate with the timing of the recession of the Last Glacial Maximum and predate the upper bound of the age estimate of the Indo-European language tree. Although virtually absent among Romance, Celtic and Semitic speakers, the presence and overall frequency of haplogroup R1a does not distinguish Indo-Iranian, Finno-Ugric, Dravidian or Turkic speakers from each other. Some contrast, however, is unfolding in its subclade frequencies. Although the R1a1a* frequency and diversity is highest among Indo-Aryan and Dravidian speakers, the subhaplogroup R1a1a7-M458 frequency peaks among Slavic and Finno-Ugric peoples. Although this distinction by geography is not directly informative about the internal divisions of these separate language families, it might bear some significance for assessing dispersal models that have been proposed to explain the spread of Indo-Aryan languages in South Asia as it would exclude any significant patrilineal gene flow from East Europe to Asia, at least since the mid-Holocene period.
References
Frazer KA, Ballinger DG, Cox DR et al: A second generation human haplotype map of over 3.1 million SNPs. Nature 2007; 449: 851–861.
Underhill PA, Kivisild T : Use of Y chromosome and mitochondrial DNA population structure in tracing human migrations. Annu Rev Genet 2007; 41: 539–564.
Karafet TM, Mendez FL, Meilerman MB, Underhill PA, Zegura SL, Hammer MF : New binary polymorphisms reshape and increase resolution of the human Y chromosomal haplogroup tree. Genome Res 2008; 18: 830–838.
Underhill PA, Shen P, Lin AA et al: Y chromosome sequence variation and the history of human populations. Nat Genet 2000; 26: 358–361.
YCC: A nomenclature system for the tree of human Y-chromosomal binary haplogroups. Genome Res 2002; 12: 339–348.
Quintana-Murci L, Krausz C, Zerjal T et al: Y-chromosome lineages trace diffusion of people and languages in southwestern Asia. Am J Hum Genet 2001; 68: 537–542.
Wells RS, Yuldasheva N, Ruzibakiev R et al: The Eurasian heartland: a continental perspective on Y-chromosome diversity. Proc Natl Acad Sci USA 2001; 98: 10244–10249.
Gimbutas M : Proto-Indo-European culture. The Kurgan culture during the fifth, fourth, and third millenia B.C.; in Cardona G, Hoenigswald HM, Senn A (eds): Indo-European and Indo-Europeans. Philadelphia: University of Pennsylvania Press, 1970, pp 155–195.
Hinds DA, Stuve LL, Nilsen GB et al: Whole-genome patterns of common DNA variation in three human populations. Science 2005; 307: 1072–1079.
Repping S, van Daalen SK, Brown LG et al: High mutation rates have driven extensive structural polymorphism among human Y chromosomes. Nat Genet 2006; 38: 463–467.
Gusmão L, Alves C, Costa S et al: Point mutations in the flanking regions of the Y-chromosome specific STRs DYS391, DYS437 and DYS438. Int J Legal Med 2002; 116: 322–326.
Zhivotovsky LA, Underhill PA, Cinnioǧlu C et al: The effective mutation rate at Y chromosome short tandem repeats, with application to human population-divergence time. Am J Hum Genet 2004; 74: 50–61.
Sengupta S, Zhivotovsky LA, King R et al: Polarity and temporality of high-resolution Y-chromosome distributions in India identify both indigenous and exogenous expansions and reveal minor genetic influence of Central Asian pastoralists. Am J Hum Genet 2006; 78: 202–221.
Regueiro M, Cadenas AM, Gayden T, Underhill PA, Herrera RJ : Iran tricontinental nexus for Y-chromosome driven migration. Hum Hered 2006; 61: 132–143.
Cadenas AM, Zhivotovsky LA, Cavalli-Sforza LL, Underhill PA, Herrera RJ : Y-chromosome diversity characterizes the Gulf of Oman. Eur J Hum Genet 2008; 16: 374–386.
Stefan M, Stefanescu G, Gavrila L et al: Y chromosome analysis reveals a sharp genetic boundary in the Carpathian region. Eur J Hum Genet 2001; 9: 27–33.
Al-Zahery N, Semino O, Benuzzi G et al: Y-chromosome and mtDNA polymorphisms in Iraq, a crossroad of the early human dispersal and of post-Neolithic migrations. Mol Phylogenet Evol 2003; 28: 458–472.
Bosch E, Calafell F, Comas D, Oefner PJ, Underhill PA, Bertranpetit J : High-resolution analysis of human Y-chromosome variation shows a sharp discontinuity and limited gene flow between northwestern Africa and the Iberian Peninsula. Am J Hum Genet 2001; 68: 1019–1029.
Deng W, Shi B, He X et al: Evolution and migration history of the Chinese population inferred from Chinese Y-chromosome evidence. J Hum Genet 2004; 49: 339–348.
Di Giacomo F, Luca F, Anagnou N et al: Clinal patterns of human Y chromosomal diversity in continental Italy and Greece are dominated by drift and founder effects. Mol Phylogenet Evol 2003; 28: 387–395.
Karafet T, Xu L, Du R et al: Paternal population history of East Asia: sources, patterns, and microevolutionary processes. Am J Hum Genet 2001; 69: 615–628.
Karafet TM, Osipova LP, Gubina MA, Posukh OL, Zegura SL, Hammer MF : High levels of Y-chromosome differentiation among native Siberian populations and the genetic signature of a boreal hunter-gatherer way of life. Hum Biol 2002; 74: 761–789.
Karlsson AO, Wallerström T, Gotherström A, Holmlund G : Y-chromosome diversity in Sweden – a long-time perspective. Eur J Hum Genet 2006; 14: 963–970.
Katoh T, Munkhbat B, Tounai K et al: Genetic features of Mongolian ethnic groups revealed by Y-chromosomal analysis. Gene 2005; 346: 63–70.
Kharkov VN : Structure of Y-chromosomal Lineages in Siberian Populations. Tomsk: Siberian Division of Russian Academy of Medical Sciences, 2005.
Kivisild T, Rootsi S, Metspalu M et al: The genetic heritage of the earliest settlers persists both in Indian tribal and caste populations. Am J Hum Genet 2003; 72: 313–332.
Lappalainen T, Koivumaki S, Salmela E et al: Regional differences among the Finns: a Y-chromosomal perspective. Gene 2006; 376: 207–215.
Lappalainen T, Laitinen V, Salmela E et al: Migration waves to the Baltic Sea region. Ann Hum Genet 2008; 72: 337–348.
Qamar R, Ayub Q, Mohyuddin A et al: Y-chromosomal DNA variation in Pakistan. Am J Hum Genet 2002; 70: 1107–1124.
Sahoo S, Singh A, Himabindu G et al: A prehistory of Indian Y chromosomes: evaluating demic diffusion scenarios. Proc Natl Acad Sci USA 2006; 103: 843–848.
Semino O, Passarino G, Oefner PJ et al: The genetic legacy of Paleolithic Homo sapiens sapiens in extant Europeans: a Y chromosome perspective. Science 2000; 290: 1155–1159.
Semino O, Santachiara-Benerecetti AS, Falaschi F, Cavalli-Sforza LL, Underhill PA : Ethiopians and Khoisan share the deepest clades of the human Y-chromosome phylogeny. Am J Hum Genet 2002; 70: 265–268.
Zalloua PA, Xue Y, Khalife J et al: Y-chromosomal diversity in Lebanon is structured by recent historical events. Am J Hum Genet 2008; 82: 873–882.
Zerjal T, Wells R, Yuldasheva N, Ruzibakiev R, Tyler-Smith C : A genetic landscape reshaped by recent events: Y-chromosomal insights into Central Asia. Am J Hum Genet 2002; 71: 466–482.
Cinnioǧlu King R, Kivisild T et al: Excavating Y-chromosome haplotype strata in Anatolia. Hum Genet 2004; 114: 127–148.
Luis JR, Rowold DJ, Regueiro M et al: The Levant versus the Horn of Africa: evidence for bidirectional corridors of human migrations. Am J Hum Genet 2004; 74: 532–544.
Mohyuddin A, Ayub Q, Underhill PA, Tyler-Smith C, Mehdi SQ : Detection of novel Y SNPs provides further insights into Y chromosomal variation in Pakistan. J Hum Genet 2006; 51: 375–378.
King RJ, Ozcan SS, Carter T et al: Differential Y-chromosome Anatolian influences on the Greek and Cretan Neolithic. Ann Hum Genet 2008; 72: 205–214.
Kayser M, Lao O, Anslinger K et al: Significant genetic differentiation between Poland and Germany follows present-day political borders, as revealed by Y-chromosome analysis. Hum Genet 2005; 117: 428–443.
Woźniak M, Grzybowski T, Starzyñski J, Marciniak T : Continuity of Y chromosome haplotypes in the population of Southern Poland before and after the Second World War. Forensic Sci Int 2007; 1: 134–140.
Weale ME, Yepiskoposyan L, Jager RF et al: Armenian Y chromosome haplotypes reveal strong regional structure within a single ethno-national group. Hum Genet 2001; 109: 659–674.
Telegin D, Lillie M, Potekhina I, Kovaliukh M : Settlement and economy in Neolithic Ukraine: a new chronology. Antiquity 2003; 77: 456–470.
Zvelebil M : Innovating Hunter-Gathers. The Mesolithic in the Baltic; in Bailey G, Spikins P (eds): Mesolithic Europe. Cambridge: Cambridge University Press, 2008, pp 18–59.
Price T, Bentley R, Lüning J, Gronenborn D, Wahl J : Prehistoric human migration in the Linearbandkeramik of central Europe. Antiquity 2001; 75: 593–603.
Buchvaldek M : Corded pottery complex in central Europe. J Indo-European Studies 1980; 8: 393–406.
Currat M, Excoffier L : The effect of the Neolithic expansion on European molecular diversity. Proc Biol Sci 2005; 272: 679–688.
Bramanti B, Thomas MG, Haak W et al: Genetic discontinuity between local hunter-gatherers and central Europe's first farmers. Science 2009; 326: 137–140.
Haak W, Forster P, Bramanti B et al: Ancient DNA from the first European farmers in 7500-year-old Neolithic sites. Science 2005; 310: 1016–1018.
Malyarchuk B, Grzybowski T, Derenko M et al: Mitochondrial DNA phylogeny in Eastern and Western Slavs. Mol Biol Evol 2008; 25: 1651–1658.
Beja-Pereira A, Luikart G, England PR et al: Gene-culture coevolution between cattle milk protein genes and human lactase genes. Nat Genet 2003; 35: 311–313.
Burger J, Kirchner M, Bramanti B, Haak W, Thomas MG : Absence of the lactase-persistence-associated allele in early Neolithic Europeans. Proc Natl Acad Sci USA 2007; 104: 3736–3741.
Itan Y, Powell A, Beaumont MA, Burger J, Thomas MG : The origins of lactase persistence in Europe. PLoS Comput Biol 2009; 5: e1000491.
Anthony D : The Horse, the Wheel and Language. How Bronze-Age Riders from the Eurasian Steppes Shaped the Modern World. Princeton, NJ: Princeton University Press, 2007.
Sherratt A : The transformation of early agrarian Europe: the later Neolithic and Copper Ages 4500-2500 BC; in Cunliffe B (ed): Prehistoric Europe: An Illustrated History. Oxford: Oxford University Press, 1998, pp 167–201.
Haak W, Brandt G, de Jong HN et al: Ancient DNA, Strontium isotopes, and osteological analyses shed light on social and kinship organization of the Later Stone Age. Proc Natl Acad Sci USA 2008; 105: 18226–18231.
Gray RD, Atkinson QD : Language-tree divergence times support the Anatolian theory of Indo-European origin. Nature 2003; 426: 435–439.
Acknowledgements
We thank all the men who donated DNA samples used in this study. This research was supported by the European Union European Regional Development Fund through the Centre of Excellence in Genomics, Estonian Biocentre and Tartu University, EC Grant ECOGENE (205419) to Estonian Biocentre, Estonian Science Foundation Grant No. 7445 (to SR) and Estonian Basic Research Grant SF 0270177s08 (to RV), Croatian Ministry of Science, Education and Sports Grant 196-1962766-2751 (to PR), grants of the RAS Programs ‘Origin and Evolution of the Biosphere’ and ‘Molecular and Cell Biology’ to LZ, a grant to OS by V Compagnia di San Paolo and Progetti di Ricerca Interesse Nazionale 2007 (Italian Ministry of the University), and the grant of the Ministry of Education of the Slovak Republic and of Slovak Academy of Sciences – VEGA No. 1/3245/06 to MB. QA, AM and SQM were supported by The Wellcome Trust. We thank Scott R Woodward and the Sorenson Molecular Genealogy Foundation for providing support for AAL and PAU.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on European Journal of Human Genetics website (http://www.nature.com/ejhg)
Rights and permissions
About this article
Cite this article
Underhill, P., Myres, N., Rootsi, S. et al. Separating the post-Glacial coancestry of European and Asian Y chromosomes within haplogroup R1a. Eur J Hum Genet 18, 479–484 (2010). https://doi.org/10.1038/ejhg.2009.194
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2009.194
Keywords
This article is cited by
-
Contrasting maternal and paternal genetic histories among five ethnic groups from Khyber Pakhtunkhwa, Pakistan
Scientific Reports (2022)
-
The Y chromosome of autochthonous Basque populations and the Bronze Age replacement
Scientific Reports (2021)
-
Subdividing Y-chromosome haplogroup R1a1 reveals Norse Viking dispersal lineages in Britain
European Journal of Human Genetics (2021)
-
Sex-biased patterns shaped the genetic history of Roma
Scientific Reports (2020)
-
Middle eastern genetic legacy in the paternal and maternal gene pools of Chuetas
Scientific Reports (2020)