Abstract
Single-nucleotide polymorphisms (SNPs) in the 3′ region of myosin IXB (MYO9B) gene have recently been reported to associate with different inflammatory or autoimmune diseases. We monitored for the association of MYO9B variants to multiple sclerosis (MS) in four Northern European populations. First, 18 SNPs including 6 SNPs with previous evidence for association to immune disorders, were tested in 730 Finnish MS families, but no linkage or family-based association was observed. To ensure the power to detect variants with a modest effect size, we further analyzed 10 variants in 899 Finnish cases and 1325 controls, and in a total of 1521 cases and 1476 controls from Denmark, Norway and Sweden, but found no association. Our results thereby do not support a major function of the tested MYO9B variants in MS.
Similar content being viewed by others
Introduction
Multiple sclerosis (MS) is a chronic inflammatory disease of the central nervous system, which leads to demyelination and neurodegeneration. Although the exact etiology of MS is unknown, it is considered a putative autoimmune disease (AID) caused by a combination of multiple genetic and environmental factors.1 Like many AIDs, MS has consistently shown strong linkage and association to the human leukocyte antigen (HLA) locus.
In addition to the HLA, also other loci such as PTPN22 and CTLA4, have shown linkage or association with more than one AID, indicating that AIDs may share some common genetic background.2 More recently, allelic variants in the gene encoding for myosin IXB (MYO9B) have been reported to associate to different AIDs or inflammatory disorders (ulcerative colitis, rheumatoid arthritis, systemic lupus erythematosus and type 1 diabetes) following the initial report by Monsuur et al3 in which association to celiac disease (CD) was found with six MYO9B 3′ region single-nucleotide polymorphisms (SNPs) in the Dutch population.3, 4, 5, 6, 7, 8 To further investigate the relevance of MYO9B in different immune disorders, we have tested the association of its genetic variants to MS in four Northern European populations.
Materials and methods
Subjects and genotyping
The Finnish MS family sample consisted of 730 families (340 full trios). Nineteen families were informative for linkage containing 2–4 affected individuals and available healthy family members. In the other families, one affected and at least one healthy parent or sibling were genotyped. The family cases were also included in a case–control analysis with 169 additional Finnish MS patients, 1325 unrelated Finnish population controls and 1521 MS cases and 1476 population controls from Denmark, Norway and Sweden. All cases had been diagnosed according to Poser's criteria and informed consent was obtained from all subjects. The study was approved by the ethics committees of all institutions involved.
Eighteen tagging SNPs selected to represent all major haploblocks in MYO9B were genotyped in 730 Finnish MS families in two multiplexes (Figure 1). All genotyping was conducted with Sequenom iPLEX platform (Sequenom Inc., San Diego, CA, USA, detailed protocols and primer sequences available on request). One of the multiplex assays (10 SNPs) was then genotyped in cases and controls. These 18 and 10 SNPs, respectively, capture 56 and 48% of the variation in MYO9B in HapMap CEPH samples with pairwise r2=0.7 (phase II data).9 Overall genotyping success rate per SNP was 97–100% in all populations.
Haploblock structure of MYO9B based on HapMap CEPH genotype data. The coloring reflects level of linkage disequilibrium between single-nucleotide polymorphisms (SNPs) in HapMap CEPH data, darker shades representing higher pairwise r2. The 10 SNPs genotyped in all samples are indicated with black arrows. SNPs genotyped in Finnish MS families only are indicated with gray arrows. SNPs previously showing association to other disorders are marked with asterisks.
Statistical analysis
Power calculations were performed with the Genetic Power Calculator.10 Haploview and PedCheck were used for data quality checks (Hardy–Weinberg equilibrium P-value ≥0.01 in healthy family founders, cases and controls in all populations, at most one Mendelian error/SNP).11, 12 Family-based analyses were performed with Merlin (nonparametric linkage test using Kong and Cox LOD score),13 transmission disequilibrium test (TDT; Haploview) and Gamete competition test (in Mendel software package version 7.0), which uses families with missing genotype data in addition to full trios.14, 15 Case–control association test was conducted as a two-tailed χ2-test in Haploview.
Results and discussion
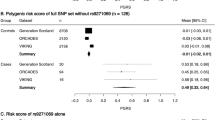
We first genotyped 18 SNPs including 6 SNPs with previous association to other AIDs in 730 Finnish MS families. The power to detect association at rs2305764 (P=0.05) in TDT was 0.96 assuming risk allele frequency of 35.7% and odds ratios 1.66 and 2.27 for heterozygotes and homozygotes, respectively, as reported by Monsuur et al3 for CD. We did not observe significant evidence of association in single SNP TDT or gamete competition tests (Table 1), or in haplotype TDT. Furthermore, no evidence of linkage was observed (Merlin nonparametric single point LOD scores 0.59–0.72).
We acknowledge, however, that with lower effect sizes our family sample may have been underpowered to detect association. We therefore genotyped a set of Northern European MS cases and controls for a single multiplex of 10 SNPs. This included rs1457092, which efficiently tags the CD risk haplotype identified by Monsuur et al,3 and has been among the most strongly and consistently associated SNPs in other studies. Concordantly with our results from the family-based analysis, we failed to see evidence for association in any single population or in combined analysis (Cochran–Mantel–Haenszel test; Table 2). Furthermore, no association was observed after stratifying the cases for gender or HLA-DRB1*15 (results not shown).
Although association of MYO9B has been reported to more than one AID or inflammatory disease, its role remains controversial because of failed replication efforts.16, 17, 18, 19, 20, 21 In this study we tested the association of MYO9B variants to yet another inflammatory disorder, MS, but failed to find association. This is concordant with the lack of association with MYO9B in the recent international genomewide association study for MS,22 which, however, contained no SNPs in the previously associated 3′ haploblock. Considering the reasonable power of our sample, the previously AID-associated MYO9B variants are unlikely to be involved in MS.
References
Ebers GC : Environmental factors and multiple sclerosis. Lancet Neurol 2008; 7: 268–277.
Serrano NC, Millan P, Páez MC : Non-HLA associations with autoimmune diseases. Autoimmun Rev 2006; 5: 209–214.
Monsuur AJ, de Bakker PI, Alizadeh BZ et al: Myosin IXB variant increases the risk of celiac disease and points toward a primary intestinal barrier defect. Nat Genet 2005; 37: 1341–1344.
van Bodegraven AA, Curley CR, Hunt KA et al: Genetic variation in myosin IXB is associated with ulcerative colitis. Gastroenterology 2006; 131: 1768–1774.
Núñez C, Oliver J, Mendoza JL et al: MYO9B polymorphisms in patients with inflammatory bowel disease. Gut 2007; 56: 1321–1322.
Latiano A, Palmieri O, Valvano MR et al: The association of MYO9B gene in Italian patients with inflammatory bowel diseases. Aliment Pharmacol Ther 2008; 27: 241–248.
Sánchez E, Alizadeh BZ, Valdigem G et al: MYO9B gene polymorphisms are associated with autoimmune diseases in Spanish population. Hum Immunol 2007; 68: 610–615.
Santiago JL, Martínez A, Núñez C et al: Association of MYO9B haplotype with type 1 diabetes. Hum Immunol 2008; 69: 112–115.
The International HapMap Consortium: A second generation human haplotype map of over 3.1 million SNPs. Nature 2007; 449: 851–861.
Purcell S, Cherny SS, Sham PC : Genetic Power Calculator: design of linkage and association genetic mapping studies of complex traits. Bioinformatics 2003; 19: 149–150. URL: http://pngu.mgh.harvard.edu/~purcell/gpc/.
Barrett JC, Fry B, Maller J, Daly MJ : Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 2005; 21: 263–265.
O'Connell JR, Weeks DE : PedCheck: a program for identifying genotype incompatibilities in linkage analysis. Am J Hum Genet 1998; 63: 259–266.
Abecasis GR, Cherny SS, Cookson WO, Cardon LR : Merlin-rapid analysis of dense genetic maps using sparse gene flow trees. Nat Genet 2002; 30: 97–101.
Sinsheimer JS, Blangero J, Lange K : Gamete-competition models. Am J Hum Genet 2000; 66: 1168–1172.
Lange K, Cantor R, Horvath S et al: MENDEL version 4.0: a complete package for the exact genetic analysis of discrete traits in pedigree and population data sets. Am J Hum Genet 2001; 69 (Suppl): 504.
Amundsen SS, Monsuur AJ, Wapenaar MC et al: Association analysis of MYO9B gene polymorphisms with celiac disease in a Swedish/Norwegian cohort. Hum Immunol 2006; 67: 341–345.
Hunt KA, Monsuur AJ, McArdle WL et al: Lack of association of MYO9B genetic variants with coeliac disease in a British cohort. Gut 2006; 55: 969–972.
Núñez C, Márquez A, Varadé J et al: No evidence of association of the MYO9B polymorphisms with celiac disease in the Spanish population. Tissue Antigens 2006; 68: 489–492.
Cirillo G, Di Domenico MR, Corsi I et al: Do MYO9B genetic variants predispose to coeliac disease? An association study in a cohort of South Italian children. Dig Liver Dis 2007; 39: 228–231.
Latiano A, Mora B, Bonamico M et al: Analysis of candidate genes on chromosomes 5q and 19p in celiac disease. J Pediatr Gastroenterol Nutr 2007; 45: 180–186.
Giordano M, Marano C, Mellai M et al: A family-based study does not confirm the association of MYO9B with celiac disease in the Italian population. Genes Immun 2006; 7: 606–608.
International Multiple Sclerosis Genetics Consortium Hafler DA, Compston A, Sawcer S et al: Risk alleles for multiple sclerosis identified by a genomewide study. N Engl J Med 2007; 357: 851–862.
Acknowledgements
We are indebted to the patients and their families for participation to the study. This work was financially supported by Neuropromise EU grant LSHM-CT-2005-018637, NIH grant NS 43559, The Center of Excellence for Disease Genetics of the Academy of Finland, the Sigrid Juselius Foundation, Helsinki University Central Hospital Research Foundation and the Helsinki Biomedical Graduate School (AK). Large scale DNA extraction and storage facility of the National Public Health Institute is acknowledged for handling of Finnish samples. The Norwegian Bone Marrow Donor Registry is acknowledged for collaboration in establishment of the Norwegian control material.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kemppinen, A., Suvela, M., Tienari, P. et al. MYO9B polymorphisms in multiple sclerosis. Eur J Hum Genet 17, 840–843 (2009). https://doi.org/10.1038/ejhg.2008.251
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2008.251