The 2019 novel coronavirus (2019-nCoV; severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)) has spread rapidly since its recent identification in patients with severe pneumonia in Wuhan, China. As of 10 February 2020, 2019-nCoV has been reported in 25 countries across 4 continents and >40,000 cases have been confirmed, with an estimated mortality risk of ~2%.
Unfortunately, no drug or vaccine has yet been approved to treat human coronaviruses. Several options can be envisaged to control or prevent emerging infections of 2019-nCoV, including vaccines, monoclonal antibodies, oligonucleotide-based therapies, peptides, interferon therapies and small-molecule drugs. However, new interventions are likely to require months to years to develop. Given the urgency of the 2019-nCoV outbreak, we focus here on the potential to repurpose existing antiviral agents approved or in development for treating infections caused by HIV, hepatitis B virus (HBV), hepatitis C virus (HCV) and influenza1, based on therapeutic experience with two other infections caused by human coronaviruses: severe acute respiratory syndrome (SARS) and Middle East respiratory syndrome (MERS).
Characteristics of 2019-nCoV
2019-nCoV is an enveloped, positive-sense, single-stranded RNA beta-coronavirus. Similar to SARS and MERS, the 2019-nCoV genome encodes non-structural proteins (such as 3-chymotrypsin-like protease, papain-like protease, helicase, and RNA-dependent RNA polymerase), structural proteins (such as spike glycoprotein) and accessory proteins (Online Fig. 1).
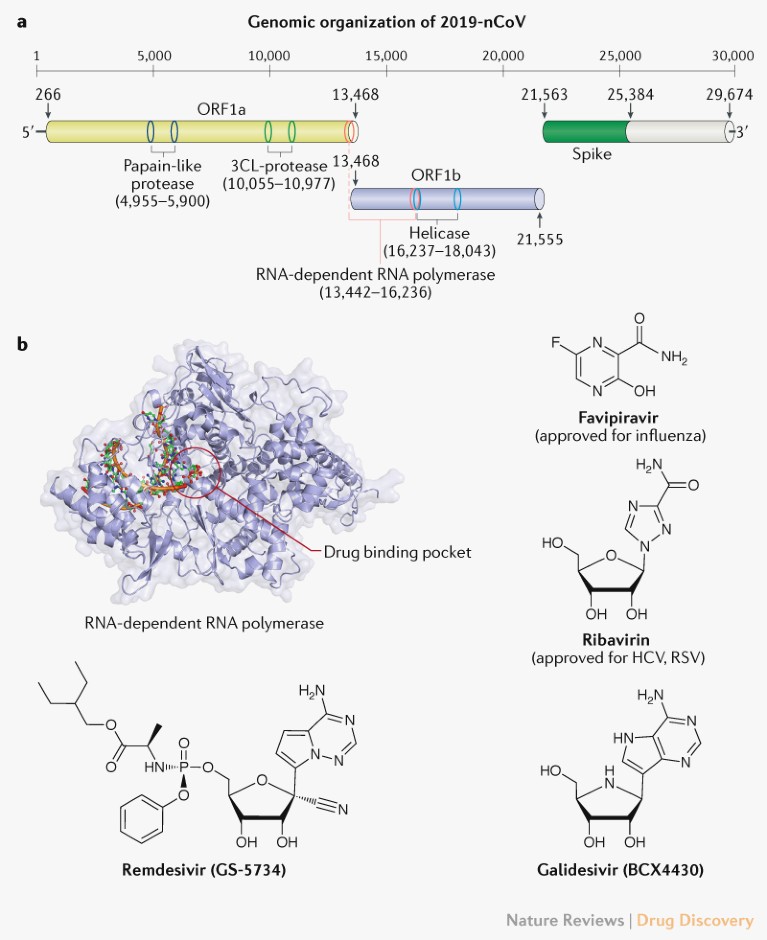
Fig. 1 | Potential drug targets for beta-coronaviruses. a | Genomic organization of 2019-nCoV (GenBank reference ID: MN908947.3), indicating the coding regions for proteins that are potential drug targets. b | A drug binding pocket is highlighted in the RNA-dependent RNA polymerase of SARS (PDB: 6NUR, 3H5Y), visualized using PyMOL V1.7 (https://pymol.org). Chemical structures of four potential inhibitors interfering with the RNA-dependent RNA polymerase of 2019-nCoV are also shown. 3CL, 3-chymotrypsin-like; HCV, hepatitis C virus; ORF, open reading frame; RSV, respiratory syncytial virus. Protein movies are available at www.virusface.com.
The four non-structural proteins mentioned above are key enzymes in the viral life cycle, and the spike glycoprotein is indispensable for virus–cell receptor interactions during viral entry2. These five proteins were therefore recognized as attractive targets to develop antiviral agents against SARS and MERS2.
Initial analyses of genomic sequences from 2019-nCoV indicate that the catalytic sites of the four 2019-nCoV enzymes that could represent antiviral targets are highly conserved, and share a high level of sequence similarity with the corresponding SARS and MERS enzymes3. Furthermore, protein structural analyses suggest that key drug-binding pockets in viral enzymes are probably conserved across 2019-nCoV, SARS and MERS3. It is, therefore, reasonable to consider repurposing existing MERS and SARS inhibitors for 2019-nCoV. Below, we discuss selected candidates with a focus on approved drugs or experimental agents that have been already tested in clinical trials for other diseases4. Supplementary Table 1 provides a longer list of anti-coronavirus agents, including preclinical compounds that could be considered for screening or starting points for optimizing antiviral agents against 2019-nCoV.
Potential repurposing candidates for 2019-nCoV
Virally targeted agents. Approved nucleoside analogues (favipiravir and ribavirin) and experimental nucleoside analogues (remdesivir and galidesivir) may have potential against 2019-nCoV. Nucleoside analogues in the form of adenine or guanine derivatives target the RNA-dependent RNA polymerase and block viral RNA synthesis in a broad spectrum of RNA viruses, including human coronaviruses4. Favipiravir (T-705), a guanine analogue approved for influenza treatment, can effectively inhibit the RNA-dependent RNA polymerase of RNA viruses such as influenza, Ebola, yellow fever, chikungunya, norovirus and enterovirus4, and a recent study reported its activity against 2019-nCoV (EC50 = 61.88 μM in Vero E6 cells)5. Patients with 2019-nCoV are being recruited in randomized trials to evaluate the efficacy of favipiravir plus interferon-α (ChiCTR2000029600) and favipiravir plus baloxavir marboxil (an approved influenza inhibitor targeting the cap-dependent endonuclease) (ChiCTR2000029544). Ribavirin is a guanine derivative approved for treating HCV and respiratory syncytial virus (RSV) that has been evaluated in patients with SARS and MERS, but its side effects such as anaemia may be severe at high doses2 and whether it offers sufficient potency against 2019-nCoV is uncertain. Remdesivir (GS-5734) is a phosphoramidate prodrug of an adenine derivative with a chemical structure similar to that of tenofovir alafenamide, an approved HIV reverse transcriptase inhibitor. Remdesivir has broad-spectrum activities against RNA viruses such as MERS and SARS in cell cultures and animal models, and has been tested in a clinical trial for Ebola. A recent study reported that remdesivir inhibited 2019-nCoV (EC50 = 0.77 μM in Vero E6 cells)5, and a US patient with 2019-nCoV recovered after receiving intravenous remdesivir in January6. Two phase III trials were initiated in early February to evaluate intravenous remdesivir (200 mg on day 1 and 100 mg once daily for 9 days) in patients with 2019-nCoV (NCT04252664 and NCT04257656), with estimated completion dates in April 2020. Galidesivir (BCX4430), an adenosine analogue that was originally developed for HCV, is currently in early-stage clinical studies evaluating its safety in healthy subjects and its efficacy against yellow fever, and has shown antiviral activities in preclinical studies against many RNA viruses, including SARS and MERS2.
Approved protease inhibitors including disulfiram, lopinavir and ritonavir have been reported to be active against SARS and MERS. Disulfiram, an approved drug to treat alcohol dependence, has been reported to inhibit the papain-like protease of MERS and SARS in cell cultures (Supplementary Table 1), but clinical evidence is lacking. Clinical trials (for example, ChiCTR2000029539) have been initiated to test HIV protease inhibitors such as lopinavir and ritonavir in patients infected with 2019-nCoV. Lopinavir and ritonavir were initially hypothesized to inhibit the 3-chymotrypsin-like protease of SARS and MERS, and appeared to be associated with improved clinical outcomes of patients with SARS in a non-randomized open-label trial2. However, it is debatable whether HIV protease inhibitors could effectively inhibit the 3-chymotrypsin-like and papain-like proteases of 2019-nCoV. HIV protease belongs to the aspartic protease family, whereas the two coronavirus proteases are from the cysteine protease family. Furthermore, HIV protease inhibitors were specifically optimized to fit the C2 symmetry in the catalytic site of the HIV protease dimer, but this C2-symmetric pocket is absent in coronavirus proteases. If HIV protease inhibitors alter host pathways to indirectly interfere with coronavirus infections, their potency remains a concern.
The spike glycoprotein is also a promising target. Griffithsin, a red-alga-derived lectin, binds to oligosaccharides on the surface of various viral glycoproteins, including HIV glycoprotein 120 and SARS-CoV spike glycoprotein2. Griffithsin has been tested in phase I studies as a gel or an enema for HIV prevention, but the potency and delivery systems of spike inhibitors should be re-evaluated for the treatment or prevention of 2019-nCoV.
Host-targeted agents. Pegylated interferon alfa-2a and -2b, approved for the treatment of HBV and HCV, could be used to stimulate innate antiviral responses in patients infected with 2019-nCoV, and trials involving interferons have been initiated, such as a trial testing the approved anti-HCV combination of a pegylated interferon plus ribavirin (ChiCTR2000029387). However, it is unclear whether a pegylated interferon and a nucleoside compound could act synergistically against 2019-nCoV. Owing to multiple adverse effects associated with subcutaneous interferon therapies, their evaluation should be closely monitored and dose reduction or discontinuation of therapy may be required.
Small-molecule agents approved for other human diseases may modulate the virus–host interactions of 2019-nCoV. An approved immune modulator, chloroquine, shows inhibitory effects against 2019-nCoV (EC50 = 1.13 μM in Vero E6 cells)5 and is being evaluated in an open-label trial (ChiCTR2000029609). Nitazoxanide, approved for diarrhea treatment, could also inhibit 2019-nCoV (EC50 = 2.12 μM in Vero E6 cells)5. The antiviral efficacy of such agents needs to be assessed in clinical studies. It is also worth mentioning that although many attempts have been made to develop host-targeted small molecules against viral infections in the past 50 years, only maraviroc has gained approval by the FDA, for HIV treatment1.
Outlook
The rapid identification of effective interventions against 2019-nCoV is a major challenge. Given the available knowledge on their safety profiles, and in some cases efficacy against closely related coronaviruses, repurposing existing antiviral agents is a potentially important near-term strategy to tackle 2019-nCoV. Phase III trials of remdesivir have been initiated, and many other trials are being established in China to test various treatment options such as umifenovir, oseltamivir and ASC09F (Supplementary Table 1). In addition, more than 50 existing MERS and/or SARS inhibitors, such as galidesivir, the protease inhibitors GC813 and compound 3k, the helicase inhibitor SSYA10‑001 and the nucleoside analogue pyrazofurin (Supplementary Table 1) could be screened against 2019-nCoV by facilities that have appropriate biocontainment capability. However, the reported EC50 and IC50 values of existing MERS and/or SARS inhibitors are mostly in the micromolar range, and further optimization of their activities against 2019-nCoV is probably needed before agents would be ready for clinical evaluation.
With the ongoing efforts to prevent the spread of 2019-nCoV worldwide, we hope that the outbreak may subside in a few months, as with SARS and MERS. Nevertheless, the outbreak has emphasized the urgent need for renewed efforts to develop broad-spectrum antiviral agents to combat coronaviruses.
