Abstract
Short-chain fatty acids and their corresponding acyl-CoAs sit at the crossroads of metabolic pathways and play important roles in diverse cellular processes. They are also precursors for protein post-translational lysine acylation modifications. A noteworthy example is the newly identified lysine 2-hydroxyisobutyrylation (Khib) that is derived from 2-hydroxyisobutyrate and 2-hydroxyisobutyryl-CoA. Histone Khib has been shown to be associated with active gene expression in spermatogenic cells. However, the key elements that regulate this post-translational lysine acylation pathway remain unknown. This has hindered characterization of the mechanisms by which this modification exerts its biological functions. Here we show that Esa1p in budding yeast and its homologue Tip60 in human could add Khib to substrate proteins both in vitro and in vivo. In addition, we have identified HDAC2 and HDAC3 as the major enzymes to remove Khib. Moreover, we report the first global profiling of Khib proteome in mammalian cells, identifying 6 548 Khib sites on 1 725 substrate proteins. Our study has thus discovered both the “writers” and “erasers” for histone Khib marks, and major Khib protein substrates. These results not only illustrate the landscape of this new lysine acylation pathway, but also open new avenues for studying diverse functions of cellular metabolites associated with this pathway.
Similar content being viewed by others
Introduction
About 17 000 cellular metabolites are now annotated in the Kyoto Encyclopedia of Genes and Genomes (KEGG) database1. Previous studies demonstrated a few major mechanisms by which cellular metabolites exert their functions, such as reversible binding to a protein to activate or inhibit its function, and serving as a precursor for protein post-translational modifications (PTMs)2,3,4. Nevertheless, the scope and broad functions of most cellular metabolites remain unknown, representing one of major black boxes in modern biology5. Short-chain fatty acids are a family of metabolites that are generated by either metabolism of host mammalian cells or microbial fermentation in the gut6. Emerging evidence suggests that the microbial-derived metabolites can be either harmful or beneficial to human health7,8. They have broad functions in signaling, host metabolism and immunity9. Despite much progress, the molecular mechanisms through which these molecules exert their function remain elusive. It was recently shown that short-chain fatty acids could serve as precursors for synthesis of their corresponding acyl-CoAs that in turn regulate histone modifications and gene expression10. This result leads to an intriguing hypothesis that metabolites from microbial fermentation can regulate epigenetic programs and control gene expression11.
2-Hydroxyisobutyrate, a cellular short-chain fatty acid, has been detected at high levels in the urine of obese people and is associated with the presence of some gut microbiota12,13. The dynamics of symbiotic gut microbiota-associated metabolites, including 2-hydroxyisobutyrate, have been associated with diverse host metabolic phenotypes13,14. Remarkably, 2-hydroxyisobutyrate is a precursor for the synthesis of 2-hydroxyisobutyryl-CoA and moreover, lysine 2-hydroxyisobutyrylation (Khib), a new type of histone PTM15. This histone mark poses unique features that differ from the widely studied histone lysine acetylation (Kac) and methylation (Kme) marks. It has a unique chemical structure, specific genomic distributions and exhibits varied dynamics among diverse model systems. ChIP-seq, gene expression analysis and immunodetection have indicated that in male germ cells H4K8hib is associated with regions of active gene transcription, in both meiotic and post-meiotic cells. These lines of evidence suggest that Khib is mechanistically and functionally different from histone Kac and Kme15. Importantly, we have identified a unique regulatory function of two cellular metabolites, 2-hydroxyisobutyrate and 2-hydroxyisobutyryl-CoA. Nevertheless, the key elements regulating this PTM pathway remain unknown, hindering functional studies of this modification in diverse biological systems and disease settings.
The history of PTM biology clearly shows that exploration of crucial regulatory proteins for a PTM pathway is key for studying its biology. In this respect, Kac provides an excellent example. Kac was originally discovered in core histones in the 1960s, findings that led to diverse correlative studies between histone Kac and chromatin structure and transcriptional activity16,17. The identification of histone acetyltransferases was a turning point for Kac biology18,19. Demonstration of the involvement of Kac in the regulation of p53 function inspired the research community to investigate the regulatory roles of Kac in transcription factors20. The identification of Kac substrates in cytosolic and mitochondrial proteins using proteomic approaches stimulated studies on the non-nuclear functions of this interesting PTM21,22,23,24. Likewise, discovery of the building blocks for the Khib pathway will undoubtedly lay a concrete foundation for studying its diverse cellular functions.
In this study, by screening a yeast knock-out (YKO) library and through mutational studies, we found that the Esa1-containing Saccharomyces cerevisiae piccolo NuA4 (picNuA4) histone acetyltransferase complex has Khib transferase activity. Esa1p's human homolog Tip60, a well-known MYST family acetyltransferase member25, is also identified as a “writer” able to catalyze Khib in mammalian cells both in vitro and in vivo. In addition, we demonstrated that histone deacetylases2 (HDAC2) and HDAC3 in mammalian cells can both serve as “erasers” to remove Khib in vitro and in vivo. Moreover, we report the first global identification of Khib substrates in human cells. Our screen identified 6 548 unique Khib sites across 1 725 proteins in HeLa cells. Analysis of the substrate proteins reveals that Khib is closely associated with the processes of transcription, translation, protein degradation and energy metabolism. Together, our study reveals key building blocks of the Khib pathway, and therefore offers a rich source for studying the role of Khib in diverse cellular process and disease development.
Results
Esa1p in yeast catalyzes the Khib modification in vivo and in vitro
Identification of the regulatory enzymes of histone marks is instrumental in studying their functions. Previous studies suggested that p300, a member of lysine acetyltransferases (KATs), can have enzymatic activity not only for lysine acetylation but also for lysine propionylation, butyrylation and crotonylation10,26. We therefore hypothesized that some KATs may also have activity toward Khib. To test this hypothesis, we took advantage of the haploid YKO collection and performed western blot analysis in each mutant strain in which a non-essential KAT was deleted. We found, unfortunately, that none of these enzymes was required for maintaining the H4K8hib level (Figure 1A).
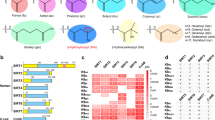
Esa1p in yeast catalyzes Khib reaction in vivo and in vitro. (A) Khib transferase activity assay of the non-essential KATs from haploid yeast knock-out collection. Cells of different deletion mutants at log phase were harvested and used to detect H4K8hib level. (B) Diagram illustrating the catalytic pocket of Esa1p bound with acetyl-CoA (left) and 2-hydroxyisobutyryl-CoA (right). Structures of acetyl-CoA-bound Esa1p (PDB: 5J9W) and 2-hydroxyisobutyryl-CoA (from PDB: 4R3U) were used for the modeling. (C) Comparing the levels of H4K8hib between an ESA1 temperature-sensitive mutant esa1-531 and its wild-type counterpart. esa1-531 decreased H4K8hib level at both permissive and non-permissive temperature. (D) Mutations in ESA1 (L264D within the ER motif) and EPL1 (F312D) decrease H4K8hib level. (E) Disruption of picNuA4 and NuA4 complexes decreased H4K8hib level. (F) PicNuA4 could catalyze the H4K8hib in vitro. NCP, nucleosome core particle; picNuA4, piccolo NuA4 complex.
In addition to these non-essential KATs, the yeast genome encodes another KAT, Esa1p, the catalytic subunit of the nucleosome acetyltransferase of the H4 (NuA4) complex, which is required for cell viability27. Esa1p associates with Yng2p, Eaf6p and Epl1p to form the core machinery termed picNuA4, which is joined by Eaf1p and other components to form the large complex28.
We first tested if Esa1p could bind 2-hydroxyisobutyryl-CoA using structural modeling. Recently, we solved the crystal structure of the NuA4 core complex, which revealed a space-sequence double recognition mechanism for targeting the N-terminal tail of histone H429. When we examined the structure of the catalytic pocket of Esa1p bound to acetyl-CoA (PDB 5J9W), we found that the catalytic pocket of Esa1p is large enough to accommodate other alkylated-CoAs with bigger acyl tails (Figure 1B). To test if 2-hydroxyisobutyryl-CoA would fit into the catalytic pocket of Esa1p, we modeled an Esa1p/2-hydroxyisobutyryl-CoA structure based on the structure of acetyl-CoA-bound Esa1p (PDB 5J9W) with the terminal acetyl group replaced with a 2-hydroxyisobutyryl group (from PDB 4R3U). As anticipated, the manner in which 2-hydroxyisobutyryl-CoA can bind to Esa1p is similar to that seen for the acetyl-CoA/Esa1p complex and allows a good fit of the 2-hydroxyisobutyryl group into the catalytic pocket (Figure 1B). Therefore, it is possible that multiple acylations, including Khib, could be catalyzed by the same enzyme.
We then carried out a series of experiments to test whether Esa1p could catalyze H4K8hib acylation in vivo. First, we compared the levels of H4K8hib between an ESA1 temperature-sensitive mutant esa1-531 and its wild-type counterpart. We found that H4K8hib persisted in the mutant at permissive temperature (25 °C), despite the reduced level of modification compared to the wild type. In contrast, at the non-permissive temperature (37 °C), the modification was completely abolished (Figure 1C). Second, we constructed a strain carrying an amino-acid substitution (L264D) within the ER motif to diminish the enzymatic activity of Esa1p29. As shown in Figure 1D, H4K8hib was dramatically decreased in this mutant. Therefore, a functional Esa1p is required for H4K8hib. Third, we asked if Esa1p-mediated H4K8hib requires the intact picNuA4 complex, as known for acetylation. We then performed western blot analysis in two strains: one carrying a point mutation in Epl1p, F312D, which disrupts its interaction with Esa1p; and the other with a deletion of YNG2. We found the amount of H4K8hib was drastically decreased in both strains (Figure 1D and 1E). Finally, we tested if the NuA4 complex is also required for H4K8hib by knocking out EAF1, which maintains the picNuA4 complex but disrupts NuA4 complex30, or EAF3, another component of the NuA4 complex. The EAF1 deletion strain showed moderate effect on H4K8hib whereas the EAF3 deletion strain did not (Figure 1E). Taken together, these results indicate that the picNuA4 complex is responsible for histone H4K8hib in S. cerevisiae.
In order to demonstrate that Esa1p catalyzes H4K8hib directly, we performed an in vitro assay using the purified picNuA4 complex and nucleosome core particles (NCPs)29. The 2-hydroxyisobutyryl-CoA was synthesized chemically (Supplementary information, Figure S1) and supplied as the donor. The reaction was carried out at room temperature and the production of H4K8hib was visualized by western blot. We found that picNuA4 can efficiently catalyze 2-hydroxyisobutyrylation on H4K8 in vitro (Figure 1F). Together, the results of these experiments carried out both in vivo and in vitro demonstrate that Esa1p is the “writer” of H4K8hib in yeast. Given the broad acetylation substrate spectrum of Esa1p31, H4K8 may serve one of the histone Khib substrate sites for Esa1p.
Tip60 has Khib transferase activity in vitro and in vivo
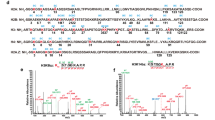
Given the Khib transferase activity of Esa1p in yeast, we next hypothesized that Tip60, the human homolog of Esa1p, might also be able to catalyze the Khib reaction. To test this hypothesis, we first carried out Khib reactions in vitro using recombinant histone H4 protein and 2-hydroxyisobutyryl-CoA as substrate and cofactor, respectively. Acetyl-CoA was used as a positive control for the reactions. Western blot results showed that Tip60 could increase the global level of the histone H4Khib modification (Figure 2A), although it had a lower intrinsic preference toward the 2-hydroxyisobutyryl-CoA (Supplementary information, Figure S2). Mass spectrometry analysis identified H4K8, H4K12, H4K16 and H4K31 as Khib substrates of Tip60 (Supplementary information, Figure S3), indicating that Tip60 is a potential mammalian Khib transferase in vitro. To confirm this result, a Tip60-specific inhibitor, TH183432, was used to determine whether it could prevent H4 from undergoing 2-hydroxyisobutyrylation. Upon treatment of TH1834, the Khib level was significantly decreased in a similar fashion as Kac (Figure 2B), further suggesting that Tip60 has Khib transferase activity in vitro.
Tip60 has lysine 2-hydroxyisobutyryltrasferase activity in vitro and in vivo. (A) In vitro assay showing Khib transferase activity of Tip60. Khib or Kac activities of recombinant Tip60 were assayed using recombinant H4 as substrate. Reaction products were detected by western blot with indicated antibodies. (B) TH1834, a Tip60 inhibitor, inhibits Tip60 activity in vitro. Khib or Kac activities of recombinant Tip60 were assayed using recombinant H4 as substrate. TH1834 at 200 μM, 500 μM and 1 mM was added to the assay. Reaction products were detected by western blot with indicated antibodies. (C) Western blot analysis showing that Tip60 knockdown impairs histone Khib in vivo. HeLa cells were treated with siRNAs against Tip60 for 48 h before being subjected to western blot analysis. The efficiency of siRNA knockdown was verified by western blot. (D) TH1834 inhibits Tip60 activity in vivo. HeLa cells were treated with 200 μM of TH1834 for 1 h or 2 h. Levels of H4K8ac and H4K8hib were detected by western blot and H4 served as loading control. (E) Western blot analysis showing that overexpression of Tip60 increases histone Khib in vivo. Flag-Tip60 was transfected into HEK293 cells for 48 h before being subjected to western blot analysis.
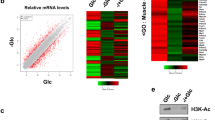
We next examined whether Tip60 regulates Khib in vivo. Knockdown of Tip60 by short interfering RNA reduced the global levels of Khib and Kac on histones, while the Khib level on a specific residue, H4K8, was decreased slightly (Figure 2C). In support of this observation, we found that treatment of cells with the Tip60 inhibitor, TH1834, clearly decreased H4K8hib signals (Figure 2D). By contrast, overexpression of full-length Tip60 by transient transfection increased both global histone Khib and H4K8hib levels and enhanced the Kac positive control as expected (Figure 2E). Quantitative mass spectrometry showed that the levels of histone H4K8hib, H4K12hib and H4K16hib were increased 35%-46% in response to overexpression of Tip60, indicating that these sites are Tip60-targeted Khib substrates (Supplementary information, Figure S4).
Taken together, these results demonstrate that Tip60 can regulate Khib both in vitro and in vivo.
HDAC2 and HDAC3 are Khib deacylases in vitro and in vivo
HDACs, which are also called lysine deacetylases (KDACs), are a family of enzymes that can remove acetyl groups from the amine at the epsilon position of lysine side chain. Given the fact that some HDACs, such as Sirt5 and Sirt6, can also catalyze removal of other forms of lysine acylation33,34,35, we hypothesized that some HDAC member might also have enzymatic activity toward Khib. To test this possibility, we first carried out an in vitro screen of the 11 class I, II and IV deacetylases (HDAC1-HDAC11) using core histones as the substrates (Figure 3A). This revealed that HDAC2 and HDAC3 have the highest activity for de-2-hydroxyisobutyrylation, whereas HDAC1 only showed marginal activity in vitro. The in vitro de-2-hydroxyisobutyrylation activity of HDAC3 could be further confirmed by the inhibition of NaBu and TSA; two inhibitors for class I and II HDACs, but not NAM, an inhibitor for class III HDAC15. Together, these data suggested that HDAC2 and HDAC3 could remove Khib in vitro.
HDAC2 and HDAC3 are Khib deacylases in vitro and in vivo. (A) In vitro screen of HDACs' Khib deacylase activities. In each reaction, 2 μg of core histones were incubated with 11 class I, II and IV deacetylases (HDAC1-HDAC11) at 37 °C for 12 h. Reaction products were detected by western blot with indicated antibodies. (B) HDAC1 knockdown slightly affects global histone Khib level in vivo. HEK293T cells were treated with siRNAs against HDAC1 for 48 h before being subjected to western blot analysis. The efficiency of siRNA knockdown was verified by western blot. (C) Western blot analysis showing that HDAC1 overexpression slightly affects global histone Khib level in HEK293T cells. An empty vector PCDNA3.1 and a vector encoding PCDNA3.1-HA-HDAC1 were transfected into HEK293T cells for 48 h before being subjected to western blot analysis. (D) HDAC3 knockdown increases histone Khib in vivo. U2OS cells were transfected twice with ON-TARGETplus SMARTpool siRNA for human HDAC3 (DHARMACON) for 24 h. (E) HDAC2 knockout and HDAC2/HDAC3 double depletion increases histone Khib in vivo. HEK293T cells were transfected with an empty vector PX458 and an HDAC2 knockout vector PX458-sgRNA (HDAC2) for 48 h, respectively. Depletion of HDAC2 and HDAC3 was further achieved by using HDAC2 knockout HEK293T cell line transfected with siRNA mix against HDAC3. (F) Western blot analysis showing that HDAC2 and/or HDAC3 overexpression decrease histone Khib in HEK293T cells. An empty vector PCDNA3.1, a vector encoding PCDNA3.1-HA-HDAC2, a vector encoding PCDNA3.1-HA-HDAC3 and mixed vectors encoding PCDNA3.1-HA-HDAC2 and PCDNA3.1-HA-HDAC3 were transfected into HEK293T cells for 48 h before being subjected to western blot analysis. Coomassie staining of total histone was used as loading control.
Next, we sought to determine whether HDACs1-3 could remove Khib in vivo. Consistent with the in vitro assay, knockdown and overexpression of HDAC1 slightly affected the global level of histone Khib (Figure 3B and 3C), while knockdown of HDAC3 or knockout of HDAC2 increased the global level of histone Khib more obviously (Figure 3D and 3E). Compared with HDAC2 knockout, the double depletion of HDAC2 and HDAC3 led to a similar increase of the global histone Khib level. In contrast, overexpression of HDAC2 or HDAC3 reduced the global level of histone Khib (Figure 3F). SILAC quantification of the histone Khib dynamics indicated that the levels of Khib decreased by 30% or more after HDAC3 overexpression (Supplementary information, Table S1). Moreover, overexpression of both HDAC2 and HDAC3 enhanced such a decrease.
Together, these data support the conclusion that HDAC2 and HDAC3 are Khib deacylases both in vitro and in vivo.
Proteomics screening of Khib peptides in HeLa cells
To globally identify Khib substrate proteins and their modification sites, we carried out proteomic screening in HeLa cells involving peptide fractionation, affinity enrichment of Khib-peptides and HPLC-MS/MS analysis (Figure 4A). The specificity of the pan anti-Khib antibody used for immunoaffinity purification was verified with a dot blot assay; the antibody could only detect the peptide library bearing a fixed 2-hydroxyisobutyrylated lysine but not the peptide library bearing a fixed unmodified lysine, acetylation lysine, butyrylated lysine, crotonylated lysine or β-hydroxybutyrylated lysine (Supplementary information, Figure S5). The acquired raw MS data were analyzed by Maxquant software with a false discovery rate of < 1% at protein, peptide and site levels. We further removed those hits with Andromeda scores lower than 40. The experiments were performed as biological triplicates. Using this procedure, we identified 7 937 Khib sites on 1 901 proteins. To improve the reliability of the identified peptides, we eliminated 1 213 Khib sites with localization probability scores < 0.75, and eliminated redundant sites. Using these criteria, we identified 6 548 unique Khib sites on 1 725 proteins in triplicate analysis (Supplementary information, Data S1A). Of the 6 548 sites, 74% (4 867) were identified in at least two biological replicates (Figure 4B; Supplementary information, Data S1B), demonstrating the high reproducibility of our procedure. These Khib sites were used as high-confidence data set in subsequent analysis.
Systematic profiling of lysine 2-hydroxyisobutyrylation in HeLa cells. (A) Schematic overview of experimental workflow for the identification of Khib in HeLa cells. (B) Pie chart shows experimental reproducibility of three biological replicates. (C) Distribution of the number of Khib sites per protein.
Sequence preference and subcellular localization of the Khib proteome
The identified Khib substrates have varied numbers of modification sites. We found that 617 proteins were modified at only one Khib site, while 80 proteins had more than 10 Khib sites, including Plectin (PLEC) with 58 sites and Myosin-9 (MYH9) with 44 sites (Figure 4C; Supplementary information, Data S1B). To determine whether there are common sequence motifs in Khib peptides, we aligned the amino-acid sequences surrounding Khib sites against all human background sequences. We found that the negatively-charged amino acids (aspartic acid and glutamic acid) were enriched at both −1 and +1 positions, whereas the positively-charged amino acid lysine was enriched at −6, −5, +5 and +6 positions (Figure 5A). In addition, arginine and lysine residues were under-represented at the −1 and +1 positions, respectively (Figure 5A). Interestingly, proline was largely depleted at most of the positions, making the sites distinct from the reported flanking sequence preference studies of Kac, lysine malonylation (Kmal) and lysine succinylation (Ksucc) (Figure 5A)36,37,38,39.
Characterization of lysine 2-hydroxyisobutyrylation proteome in HeLa cells. (A) Consensus sequence logo shows a representative sequence for all Khib sites. (B) Venn diagram shows cellular compartment distribution of 2-hydroxyisobutyrylated proteins. (C) Distribution of the number of Khib sites on diverse enzymes. The enzymes with < 10 Khib sites are not shown in this bar graph. (D) Three-dimensional structure of alpha-enolase (ENO1, PDB entry 2PSN) shown with Khib sites. The important residues in the substrate binding pocked are shown in detail. (E) Table of the Khib sites on key residues involving substrate or cofactor binding.
To explore the subcellular distribution of Khib substrates in cells, we performed a cellular compartment analysis of the Khib proteome. Recently, some PTMs, such as Kmal and Ksucc, were reported being significantly enriched in mitochondria37,40. In contrast, only ∼15% of Khib proteins were annotated in mitochondria, whereas 61% and 86% of Khib proteins localized exclusively or partially in nucleus and cytosol, respectively (Figure 5B). This suggests that the Khib modification has a very different regulatory mechanism from Kmal and Ksucc. In contrast to Kmal and Ksucc, the subcellular distribution of Kac substrates is similar to that of Khib substrates. The majority of Kac substrates often reside in either the cytoplasm or the nucleus, while mitochondria account for ∼5% or less of lysine-acetylated proteins22. Given that cytoplasmic, mitochondrial and nuclear Kac can regulate various cellular processes, the Khib pathway's major functions are likely widespread in diverse subcellular compartments.
Functional annotation of the Khib proteome
In order to explore the possible pathways affected by Khib, we performed a KEGG pathway enrichment analysis of the Khib proteins1. Ribosome (adjusted P = 5.65 × 10−35), spliceosome (adjusted P = 6.60 × 10−25) and proteasome (adjusted P = 4.94 × 10−13) pathways were most significantly enriched (Supplementary information, Data S2A). Notably, 66%, 48% and 57% of proteins in these pathways were 2-hydroxyisobutyrylated, respectively (Supplementary information, Data S2A). In addition, macromolecule transport-related pathways, such as RNA transport (adjusted P = 2.65 × 10−10) and protein export (adjusted P = 2.28 × 10−6) pathways, were also enriched (Supplementary information, Data S2A). Moreover, our data showed that Khib proteins were enriched in energy metabolic networks, such as the citric cycle (adjusted P = 2.28 × 10−11), fatty acid metabolism (adjusted P = 1.24 × 10−5) and pyruvate metabolism (adjusted P = 2.08 × 10−5) (Supplementary information, Data S2A). Similar to the pathway analysis, unbiased gene ontology biological process and UniProt keywords annotation of the Khib proteome showed that Khib proteins were associated with metabolic process and RNA processing (Supplementary information, Figure S6).
We also performed a protein complex enrichment analysis of the Khib proteome with a manually curated CORUM database41. In accord with the pathway analysis, the top enriched protein complexes are associated with ribosome, spliceosome and proteasome (Supplementary information, Data S2B). In addition to these complexes, we identified significant enrichment of Khib in the CCT micro-complex (adjusted P = 6.09 × 10−10), the H2AX complex (adjusted P = 2.29 × 10−8), the TNF-α/NF-κB signal transduction pathway (adjusted P = 5.95 × 10−8) and the DNA-PK-Ku-eIF2-NF90-NF45 complex (adjusted P = 6.44 × 10−8) (Supplementary information, Data S2B). The TNF-α/NF-κB signal transduction pathway plays a pivotal role in various biological processes, and dysregulation of this pathway is associated with many diseases42. Our data showed that 9 out of 14 subunits in this complex were 2-hydroxyisobutyrylated (Supplementary information, Data S2B). The DNA-PK-Ku-eIF2-NF90-NF45 complex is involved in DNA double-strand break repair43. We found that 87.5% of proteins (7 out of 8) in this complex were 2-hydroxyisobutyrylated (Supplementary information, Data S2B). These results suggest that Khib can be involved in diverse cellular functions and networks that include protein synthesis and degradation, cellular signaling and DNA repair.
Khib in enzymes
Of the 346 2-hydroxyisobutyrylated enzymes, 219 enzymes have multiple Khib sites. Of these, 21 enzymes were heavily modified and had more than 10 Khib sites (Figure 5C; Supplementary information, Data S3). The DNA-dependent protein kinase catalytic subunit (PRKDC), the core component of the DNA-PK-Ku-eIF2-NF90-NF45 complex, has 37 Khib sites (Figure 5C). Glycolysis is a catabolic process that converts glucose into pyruvate via 10 enzymatic steps44. Strikingly, 5 out of the 10 key enzymes required for glycolysis were heavily modified, including phosphoglycerate kinase 1 (PGK1), alpha-enolase (ENO1), pyruvate kinase isoform M2 (PKM2), glyceraldehyde-3-phosphate dehydrogenase (GAPDH) and fructose-bisphosphate aldolase A (ALDOA) (Figure 5C). ENO1 is a highly-conserved enzyme that coverts 2-phosphoglycerate to the high-energy intermediate phosphoenolpyruvate. Remarkably, its activity center residue, K343, was 2-hydroxyisobutyrylated (Figure 5D). Previous experiments showed that mutations on this site (Lys to Ala or Met) abolished enzyme activity45. In addition, the crystal structure of ENO1 shows that two magnesium ions are located in the active pocked within 4Å to the epsilon nitrogen atom of K343 (Figure 5D), and both magnesium ions are thought to participate in the catalytic reaction45. Therefore, the K343hib could not only inactivate the epsilon nitrogen atom, but is also likely to occupy the space of magnesium ions, thus disrupting the interaction network of the active-site residues and hampering enzymatic activity.
In addition to these enzyme active sites, we also found that many substrates or cofactor binding sites were 2-hydroxyisobutyrylated (Figure 5E). For example, K186 of adenosylhomocysteinase; K147 of ALDOA; K171 of glucose-6-phosphate 1-dehydrogenase; and K147 of glutamate dehydrogenase 1 are known to be important for substrate binding (www.uniprot.com). The K745 residue of epidermal growth factor receptor (EGFR), K112 of heat shock protein HSP 90-alpha A2, K168 of endoplasmin and K220 of PGK1 are critical residues for ATP binding (www.uniprot.com). Khib on these positions is most likely to affect the protein functions by disrupting the binding interaction.
Discussion
Emerging evidence indicates that diverse newly-discovered lysine acylations are associated with normal cellular physiology and disease. Histone butyrylation, crotonylation and β-hydroxybutyrylation are associated with gene expression in diverse cellular systems10,46,47,48. These modifications can also have specific “readers” to mediate their functions49,50,51,52. Lysine propionylation, butyrylation, malonylation, succinylation, glutarylation all contribute to multiple heritable genetic diseases36,40,53,54,55. These lines of evidence suggest that lysine acylations are a family of PTMs that are physiologically relevant.
Lysine acetylation not only occurs on core histones, but also on diverse other proteins in nuclei, cytosol and mitochondria, many of which are metabolic enzymes. Previous studies have suggested that the lysine acetylation pathway has diverse functions. Given the significant structural changes associated with Khib and also its widespread abundance, its close association with gene expression, it seems that the Khib pathway is also highly likely to function similar to lysine acetylation and to have diverse regulatory roles. However, study of the Khib pathway is hindered by a lack of knowledge of its regulatory enzymes and key substrates, the major focus of this work.
Our findings on the Khib activities of Esa1p/TIP60, HDAC2 and HDAC3 broaden the enzymatic activities of the previously annotated HATs and HDACs. TIP60 is a cellular lysine acetyltransferase that is involved in the regulation of gene expression and DNA damage response. It exerts cellular functions principally through acetylation of histones and critical cellular proteins like p53, p21 and ATM kinase56. To our knowledge, TIP60 has not yet been found to have acyltransferase activity other than acetylation. Thus, many of the annotated acetylation-regulatory enzymes have an expanded repertoire of acyl-transferase activities, and our result provides additional complexity of enzymatic activities of these enzymes.
Our study of the comprehensive global lysine 2-hydroxyisobutyrylome has provided a data set of 6 548 reliable Khib sites on 1 725 proteins in HeLa cells. This data set represents the first Khib proteome in mammalian cell to date, and illustrates the broad landscape of the Khib pathway. Notably, 14 Khib sites were located at substrate or cofactor-binding positions, suggesting Khib on these positions may negatively regulate protein functions. Second, this study provides a valuable resource for understanding the molecular mechanism whereby Khib is associated with enzyme functions and human diseases. We detected 1 252 Khib sites on 346 enzymes in the HeLa Khib proteome (Supplementary information, Data S3) and many of these enzymes are heavily 2-hydroxyisobutyrylated, suggesting important roles of Khib in diverse cellular processes and development of disease. Third, our further analysis of the Khib proteome has shed additional light on the crosstalk between Khib and phosphorylation. We identified 115 Khib sites in the HeLa Khib proteome located within five residues of reported phosphorylation sites (Supplementary information, Data S4) by mapping the Khib proteome against the phosphorylome in UniProt database. Given that phosphorylation can serve as a functional on/off switch, these data suggest potential roles for Khib in tuning protein functions through interplay with nearby phosphorylation sites.
Our study also sheds new light on the ways that microbial metabolites might affect human health and disease. The human gut harbors trillions of microorganisms that produce multiple metabolites, which have been demonstrated to play important roles in the development of diverse diseases8. For example, high levels of 2-hydroxyisobutyrate in the urine of obese people have been found associated with reduced bacterial diversity in “obese” gut microbiota12. However, the underlying mechanisms of how the microbial metabolites affect human health and disease are poorly understood. Emerging evidence has demonstrated that fluctuations in the availability of short-chain fatty acids could directly impinge on levels of their corresponding PTMs10,53,57. In this context, given the potentially important roles of Khib in various cellular processes revealed in this study, it is highly likely that microbial metabolites, including 2-hydroxyisotutarate, play a regulatory role at the level of PTMs such as Khib in a variety of host tissues and can ultimately affect host protein functions and the overall phenotype.
Identification of key regulatory elements for the Khib pathway, including enzymes and substrate proteins, will mark a key step forward toward description of this pathway. It is expected that histone Khib pathway will have unique binding modules in a similar fashion to histone lysine acetylation and crotonylation49,50. It seems highly likely that the relative abundance of histone Kac and Khib in a given substrate protein such as a histone protein is regulated by metabolic status and concentrations of acyl-CoAs in a similar fashion as histone lysine crotonylation10. We now require functional studies of these enzymes and substrates, together with the discovery and characterization of their binding proteins (or “readers”), and investigations of the governing principles that acyl-CoA metabolism might have on these proteins. Such future studies will greatly enhance our understanding of the Khib pathway and molecular regulation of cellular functions mediated by its cognate short-chain fatty acid.
Materials and Methods
Plasmids, siRNA, antibodies and cell lines
Tip60 was cloned into pcDNA3.0 with a Flag tag. The construction was verified by DNA sequencing. siRNA was purchased from Dharmacon (L-006301-00-0005). The antibodies used here were anti-Pan Kac (PTM Biolabs, PTM-101), anti-Pan Khib (PTM Biolabs, PTM-801), anti-H4K8ac (PTM Biolabs, PTM-120), anti-H4K8hib (PTM Biolabs, PTM-805), anti-H4 (Abcam, ab31830), anti-Flag (Sigma-Aldrich, F7425), anti-Tubulin (Abcam, ab6160) and anti-Tip60 (Abcam, 23886). Cell lines were purchased from ATCC (www.atcc.org) and used without further authentication, including HEK293 (ATCC CRL-1573), HEK293T (ATCC CRL-3216), HeLa (ATCC CCL-2) and U2OS (ATCC HTB-96). No mycoplasma contamination was detected using a MycoAlert Mycoplasma Detection Kit (Lonza, LT07-118).
Stable isotope labeling of cells and transfections
HEK293T cells were grown in lysine-free DMEM supplemented with 10% dialyzed FBS, and either light (12C614N2-L-Lysine) or heavy (13C614N2-L-Lysine) lysine (100 mg/L). Cells were grown for more than seven generations to achieve more than 98% labeling efficiency. For the transfections, HeLa and HEK293 cells were cultured in DMEM medium supplemented with 10% FBS, 50 U/mL penicillin and 50 mg/mL streptomycin in a 5% CO2 atmosphere at 37 °C. Transfection to achieve transient overexpression was performed with Lipofectamine 2000 (Invitrogen) according to the manufacturer's instructions. siRNA transfection was performed with Lipofectamine RNAiMAX (Invitrogen) according to manufacturer's instructions. For HDAC1, HDAC2 and HDAC3 overexpression, five pools of HEK293T cells were transfected with an empty vector PCDNA3.1, a vector encoding PCDNA3.1-HA-HDAC1, a vector encoding PCDNA3.1-HA-HDAC2, a vector encoding PCDNA3.1-HA-HDAC3 and mixed vectors encoding PCDNA3.1-HA-HDAC2 and PCDNA3.1-HA-HDAC3 for 48 h.
Depletion of HDAC1, HDAC2 and HDAC3
Two pools of HEK293T cells were transfected twice with HDAC1 siRNA (Genepharma, siHDAC1: (#1) 5′-GCUCCUCUGACAAACGAAUTT-3′; (#2) 5′-CCGGUCAUGUCCAAAGUAATT-3′; (#3) 5′-GCUGUACAUUGACAUUGAUTT-3′) and a negative control siRNA (Genepharma, 5′-UUCUCCGAACGUGUCACGUTT-3′), respectively. About 48 h after the second transfection, the cell lysates were probed with anti-Kac and anti-Khib antibodies.
HEK293T cells were transfected with an empty vector PX458 and an HDAC2 knockout vector PX458-sgRNA (HDAC2), respectively, to get two stable cell lines. Depletion of HDAC2 and HDAC3 was further achieved by using the HDAC2 knockout HEK293T cell line transfected twice with a siRNA mix against HDAC3 (Genepharma, siHDAC3: (#1) 5′-CCGCCAGACAAUCUUUGAATT-3′; (#2) 5′-GAGCUUCCCUAUAGUGAAUTT-3′; (#3) 5′-GGGAAUGCGUUGAAUAUGUTT-3′). The HDAC2 knockout HEK293T cells transfected twice with siRNA (Genepharma, 5′-UUCUCCGAACGUGUCACGUTT-3′) were used as negative control. About 48 h after the last transfection, the Khib and Kac levels were assessed by western blot using extractions of whole-cell lysates.
Preparation of cell lysate
Cells were sonicated for 3 min on ice using a high-intensity ultrasonic processor (Scientz) in lysis buffer (8 M urea, 2 mM EDTA, 3 μM TSA, 50 mM NAM, 5 mM DTT and 1% Protease Inhibitor Cocktail III). The remaining debris was removed by centrifugation at 18 000× g at 4 oC for 3 min. The protein concentration was determined using a 2-D Quant kit according to the manufacturer's instructions.
Trypsin digestion of cell lysate
Proteins in the cell lysate were reduced with 10 mM DTT for 1 h at 37 oC, alkylated with 20 mM iodoacetamide for 45 min at room temperature in darkness, and the excess iodoacetamide was blocked by 20 mM cysteine. Then the protein sample was diluted by adding 100 mM NH4HCO3 to reduce the urea concentration to < 2 M. Trypsin was added at 1:50 trypsin-to-protein mass ratio for the first digestion overnight and 1:100 trypsin-to-protein mass ratio for a second 4 h-digestion. Finally, 18 mg of proteins from the HeLa cells lysate was digested for subsequent experiments.
Peptide fractionation with high-pH reverse-phase chromatography
The peptides from HeLa cells lysate were then divided into three equal parts, and each of them (6 mg) was fractionated into six fractions by high pH reverse-phase HPLC using an Agilent 300 Extend C18 column (5 μm particles, 4.6 mm ID, 250 mm length). Briefly, peptides were first separated using a gradient of 2%-60% acetonitrile in 10 mM ammonium bicarbonate at pH 10 over 90 min into 90 fractions. The peptides were then combined into six fractions and dried by vacuum centrifugation.
Immunoaffinity enrichment
To enrich Khib peptides, total peptides dissolved in NETN buffer (100 mM NaCl, 1 mM EDTA, 50 mM Tris-HCl, 0.5% NP-40, pH 8.0) were incubated with pre-washed pan anti-Khib beads (PTM Biolabs Inc., Chicago, IL) at 4 oC overnight with gentle shaking. The beads were washed four times with NETN buffer and twice with ddH2O. The bound peptides were eluted from the beads with 0.1% trifluoroacetic acid, and the eluted fractions were combined and vacuum-dried.
Mass spectrometry
Samples were dissolved in 0.1% formic acid and loaded onto a reversed-phase pre-column (Acclaim PepMap 100, Thermo Scientific). Peptide separation was performed using a reversed-phase analytical column (Acclaim PepMap RSLC, Thermo Scientific) with a gradient of 5-80% HPLC buffer B (0.1% formic acid in 90% acetonitrile, v/v) in buffer A (0.1% formic acid in water, v/v) at a flow rate of 300 nl/min over 60 min on an EASY-nLC 1000 UPLC system. The samples were analyzed by Q Exactive Plus Mass Spectrometers (ThermoFisher Scientific). A data-dependent procedure that alternated between one full mass scan followed by the top 20 most intense precursor ions was applied with 30 s dynamic exclusion. Intact peptides were detected with a resolution of 70 000 at 200 m/z, and ion fragments were detected with a resolution of 17 500 at 28% normalized collision energy.
Database search and data filter criteria
The resulting MS/MS data were processed using MaxQuant with integrated Andromeda search engine (v.1.3.0.5)58. Tandem mass spectra were searched against the UniProt Human protein database (88 277 entries, http://www.uniprot.org) concatenated with reverse decoy database. Trypsin/P was specified as cleavage enzyme allowing up to two missing cleavages and five modifications per peptide. Carbamidomethylation on cysteine was specified as fixed modification. Oxidation on methionine, 2-hydroxyisobutyrylation on lysine, acetylation on lysine and acetylation on protein N-terminus were specified as variable modifications. False discovery rate thresholds for protein, peptide and modification site were specified at 1%. Minimum peptide length was set at 7. All the other parameters in MaxQuant were set to default values. Khib identified on peptides from reverse or contaminant protein sequences, peptides with an Andromeda score below 40, site localization probability below 0.75 and redundant Khib sites were removed. In addition, Khib sites on the peptide C-terminus were eliminated, unless the peptide C-terminus was also the corresponding protein C-terminus. For the SILAC sample data, Khib site ratios were normalized by the quantified protein expression levels.
In vitro acetylation and 2-hydroxyisobutyrylation assay
Recombinant human Tip60 and Histone H4 proteins were purchased from Cayman (10783) and NEB (M2504S), respectively. For each reaction, 250 ng of Tip60 protein, 2.5 μg of H4 protein and 10 μM of CoA were added in reaction buffer (50 mM Tris-CI, pH 8.0, 10% glycerol, 10 mM butyric acid, 0.1 mM EDTA, 1 mM DTT and 1 mM PMSF). The reaction mixtures were incubated at 30 oC for 1 h, followed by addition of SDS sample buffer. The levels of acetylation and 2-hydroxyisobutyrylation were assessed by western blot.
PicNuA4 in vitro assay
The NCP with 147 bp 601 “wisdom” sequence was prepared using luger's protocol. PicNuA4 complex was prepared using an unpublished method. The reaction system was 1 μM NCP, 0.01 μM picNuA4, 50 μM 2-hib-CoA, 50 mM NaCl, 10 mM HEPES (pH 7.0), 0.1 mg/mL BSA. The assay was performed at room temperature, utilizing picNuA4 to start the reactions with a time-based gradient. SDS-loading buffer was used to stop the reactions with incubation at 100 °C for 1 min. The signal was detected by western blot.
Protein function annotation analysis
Enrichment analysis for KEGG pathway and PFAM domain was performed using a hypergeometric test in GOstats package in R59. Protein complexes were enriched basing on that manually curated CORUM protein complex database41 for all mammals using a hypergeometric test. Unbiased gene ontology terms and UniProt keywords enrichment analyses were performed using a web tool (https://agotool.sund.ku.dk)60.
Protein-protein interaction network analysis
The STRING database (v10, http://www.string-db.org/) was used for analyzing the protein-protein interaction network of the Tip60-regulated Khib proteome. Interactions with the highest confidence score (above 0.9) were selected and the network was visualized in Cytoscape (v.3.2.1).
Correlation between Khib and mutation/phosphorylation
In-house developed scripts were applied to map Khib to known mutations and phosphorylation sites. Disease-related mutations were extracted from UniProt (http://www.uniprot.org) and COSMIC (Catalogue of Somatic Mutations in Cancer, http://cancer.sanger.ac.uk/cosmic) databases. Protein dysfunction-related mutations, substrate/cofactor-binding sites and reported phosphorylation sites were extracted from UniProt database.
Chemical synthesis of 2-hydroxyisobutyryl-CoA
The method used to synthesize 2-hydroxyisobutyryl-CoA was modified from a previous study61. About 0.88 g (10 mmol) 2-hydroxyisobutyric acid and 1 mL thiophenol were dissolved in 50 mL pre-cooled dimethylformamide (DMF), to which 2.52 g (12.2 mmol) dicyclohexylcarbodiimide in 50 mL DMF was added drop by drop and the mixture was stirred for 3 h on ice bath. About 40 mL of cold water was then added and the solution was filtered. The filtrate was extracted with 100 mL of ether and the organic layer was washed three times with saturated NaCl. The ether extract was dried using anhydrous sodium sulfate and ether evaporated. The residue was then purified by silica gel column chromatography with an elution solvent consisting of ethylacetate:hexane (20:1). The roughly purified product was further purified by silica thin-layer chromatography with an elution solvent consisting of ethylacetate:hexane (1:4). The target band was collected and the purified S-phenyl 2-hydroxy-2-methylpropanethioate was used as the reactant for the next step. 16 mg of S-phenyl 2-hydroxy-2-methylpropanethioate was dissolved in 500 μL 0.1 M NaHCO3 (pH 8.0) and 200 μL dioxane mixture was added to a solution of 10 mg sodium salt of CoA dissolved in 1 mL NaHCO3 (pH 8.0) at 0 °C. The mixture was reacted overnight, and then was neutralized by adding 1 N HCl to pH 7.0 to stop the reaction. About 8 mL ether was used to extract the unreacted reactant and this step was repeated five times. Then 8 mL of ethylacetate was used to extract the water phase and this step was repeated eight times. The final product remained in the water phase. Water was allowed to evaporate at 30 °C in order to achieve the solid end product.
Data availability
The mass spectrometry proteomics data have been deposited in the ProteomeXchange Consortium via the PRIDE62 partner repository with the data set identifier PXD005414.
Author Contributions
YZ and JD designed and coordinated the whole project. HH performed the proteomics experiments, data analysis and bioinformatics analysis, and helped coordinate the project. SQ performed the immunoprecipitation and Tip60-related experiments. JH and ZL were involved in yeast acyltransferases assay and verification experiments. XW, LG, WZ, LD and WG were involved in the HDAC assay and verification experiments. PX and ZC performed the modeling experiments. FL and JW synthesized 2-hydroxyisobutyryl-CoA. HH, YZ and JD wrote the manuscript. All authors discussed the results and commented on the manuscript.
Competing Financial Interests
YZ is on the science advisory board of PTM Biolabs.
References
Kanehisa M, Goto S . KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res 2000; 28:27–30.
Fierz B, Muir TW . Chromatin as an expansive canvas for chemical biology. Nat Chem Biol 2012; 8:417–427.
Walsh CT, Garneau-Tsodikova S . Gatto GJ Jr . Protein posttranslational modifications: the chemistry of proteome diversifications. Angew Chem Int Ed Engl 2005; 44:7342–7372.
DeBerardinis RJ, Thompson CB . Cellular metabolism and disease: what do metabolic outliers teach us? Cell 2012; 148:1132–1144.
Moellering RE, Cravatt BF . Functional lysine modification by an intrinsically reactive primary glycolytic metabolite. Science 2013; 341:549–553.
Koh A, De Vadder F, Kovatcheva-Datchary P, Backhed F . From dietary fiber to host physiology: short-chain fatty acids as key bacterial metabolites. Cell 2016; 165:1332–1345.
Russell WR, Hoyles L, Flint HJ, Dumas ME . Colonic bacterial metabolites and human health. Curr Opin Microbiol 2013; 16:246–254.
Lee WJ, Hase K . Gut microbiota-generated metabolites in animal health and disease. Nat Chem Biol 2014; 10:416–424.
den Besten G, van Eunen K, Groen AK, Venema K, Reijngoud DJ, Bakker BM . The role of short-chain fatty acids in the interplay between diet, gut microbiota, and host energy metabolism. J Lipid Res 2013; 54:2325–2340.
Sabari BR, Tang Z, Huang H, et al. Intracellular crotonyl-CoA stimulates transcription through p300-catalyzed histone crotonylation. Mol Cell 2015; 58:203–215.
Sabari BR, Zhang D, Allis CD, Zhao YM . Metabolic regulation of gene expression through histone acylations. Nat Rev Mol Cell Biol 2017; 18:90–101.
Calvani R, Miccheli A, Capuani G, et al. Gut microbiome-derived metabolites characterize a peculiar obese urinary metabotype. Int J Obes 2010; 34:1095–1098.
Li M, Wang B, Zhang M, et al. Symbiotic gut microbes modulate human metabolic phenotypes. Proc Natl Acad Sci USA 2008; 105:2117–2122.
Koppel N, Balskus EP . Exploring and understanding the biochemical diversity of the human microbiota. Cell Chem Biol 2016; 23:18–30.
Dai L, Peng C, Montellier E, et al. Lysine 2-hydroxyisobutyrylation is a widely distributed active histone mark. Nat Chem Biol 2014; 10:365–370.
Allfrey VG, Faulkner R, Mirsky AE . Acetylation and methylation of histones and their possible role in the regulation of RNA synthesis. Proc Natl Acad Sci USA 1964; 51:786–794.
Allis CD, Glover CV, Bowen JK, Gorovsky MA . Histone variants specific to the transcriptionally active, amitotically dividing macronucleus of the unicellular eucaryote, Tetrahymena thermophila. Cell 1980; 20:609–617.
Brownell JE, Zhou J, Ranalli T, et al. Tetrahymena histone acetyltransferase A: a homolog to yeast Gcn5p linking histone acetylation to gene activation. Cell 1996; 84:843–851.
Brownell JE, Allis CD . An activity gel assay detects a single, catalytically active histone acetyltransferase subunit in Tetrahymena macronuclei. Proc Natl Acad Sci USA 1995; 92:6364–6368.
Gu W, Shi XL, Roeder RG . Synergistic activation of transcription by CBP and p53. Nature 1997; 387:819–823.
Kim SC, Sprung R, Chen Y, et al. Substrate and functional diversity of lysine acetylation revealed by a proteomics survey. Mol Cell 2006; 23:607–618.
Choudhary C, Kumar C, Gnad F, et al. Lysine acetylation targets protein complexes and co-regulates major cellular functions. Science 2009; 325:834–840.
Zhao S, Xu W, Jiang W, et al. Regulation of cellular metabolism by protein lysine acetylation. Science 2010; 327:1000–1004.
Chen Y, Zhao W, Yang JS, et al. Quantitative acetylome analysis reveals the roles of SIRT1 in regulating diverse substrates and cellular pathways. Mol Cell Proteomics 2012; 11:1048–1062.
Sapountzi V, Logan IR, Robson CN . Cellular functions of TIP60. Int J Biochem Cell Biol 2006; 38:1496–1509.
Chen Y, Sprung R, Tang Y, et al. Lysine propionylation and butyrylation are novel post-translational modifications in histones. Mol Cell Proteomics 2007; 6:812–819.
Allard S, Utley RT, Savard J, et al. NuA4, an essential transcription adaptor/histone H4 acetyltransferase complex containing Esa1p and the ATM-related cofactor Tra1p. EMBO J 1999; 18:5108–5119.
Boudreault AA, Cronier D, Selleck W, et al. Yeast enhancer of polycomb defines global Esa1-dependent acetylation of chromatin. Genes Dev 2003; 17:1415–1428.
Xu P, Li CM, Chen ZH, et al. The NuA4 core complex acetylates nucleosomal histone H4 through a double recognition mechanism. Mol Cell 2016; 63:965–975.
Mitchell L, Lambert JP, Gerdes M, et al. Functional dissection of the NuA4 histone acetyltransferase reveals its role as a genetic hub and that Eaf1 is essential for complex integrity. Mol Cell Biol 2008; 28:2244–2256.
Smith ER, Eisen A, Gu WG, et al. ESA1 is a histone acetyltransferase that is essential for growth in yeast. Proc Natl Acad Sci USA 1998; 95:3561–3565.
Gao C, Bourke E, Scobie M, et al. Rational design and validation of a Tip60 histone acetyltransferase inhibitor. Sci Rep 2014; 4:5372.
Peng C, Lu Z, Xie Z, et al. The first identification of lysine malonylation substrates and its regulatory enzyme. Mol Cell Proteomics 2011; 10:M111.012658.
Du J, Zhou Y, Su X, et al. Sirt5 is a NAD-dependent protein lysine demalonylase and desuccinylase. Science 2011; 334:806–809.
Jiang H, Khan S, Wang Y, et al. SIRT6 regulates TNF-alpha secretion through hydrolysis of long-chain fatty acyl lysine. Nature 2013; 496:110–113.
Park J, Chen Y, Tishkoff DX, et al. SIRT5-mediated lysine desuccinylation impacts diverse metabolic pathways. Mol Cell 2013; 50:919–930.
Rardin MJ, He W, Nishida Y, et al. SIRT5 regulates the mitochondrial lysine succinylome and metabolic networks. Cell Metab 2013; 18:920–933.
Nishida Y, Rardin MJ, Carrico C, et al. SIRT5 regulates both cytosolic and mitochondrial protein malonylation with glycolysis as a major target. Mol Cell 2015; 59:321–332.
Svinkina T, Gu HB, Silva JC, et al. Deep, quantitative coverage of the lysine acetylome using novel anti-acetyl-lysine antibodies and an optimized proteomic workflow. Mol Cell Proteomics 2015; 14:2429–2440.
Colak G, Pougovkina O, Dai L, et al. Proteomic and biochemical studies of lysine malonylation suggest its malonic aciduria-associated regulatory role in mitochondrial function and fatty acid oxidation. Mol Cell Proteomics 2015; 14:3056–3071.
Ruepp A, Waegele B, Lechner M, et al. CORUM: the comprehensive resource of mammalian protein complexes--2009. Nucleic Acids Res 2010; 38:D497–501.
Bouwmeester T, Bauch A, Ruffner H, et al. A physical and functional map of the human TNF-alpha/NF-kappa B signal transduction pathway. Nat Cell Biol 2004; 6:97–105.
Ting NS, Kao PN, Chan DW, Lintott LG, Lees-Miller SP . DNA-dependent protein kinase interacts with antigen receptor response element binding proteins NF90 and NF45. J Biol Chem 1998; 273:2136–2145.
Lunt SY, Vander Heiden MG . Aerobic glycolysis: meeting the metabolic requirements of cell proliferation. Annu Rev Cell Dev Biol 2011; 27:441–464.
Pancholi V . Multifunctional alpha-enolase: its role in diseases. Cell Mol Life Sci 2001; 58:902–920.
Goudarzi A, Zhang D, Huang H, et al. Dynamic competing histone H4 K5K8 acetylation and butyrylation are hallmarks of highly active gene promoters. Mol Cell 2016; 62:169–180.
Xie Z, Zhang D, Chung D, et al. Metabolic regulation of gene expression by histone lysine beta-hydroxybutyrylation. Mol Cell 2016; 62:194–206.
Tan M, Luo H, Lee S, et al. Identification of 67 histone marks and histone lysine crotonylation as a new type of histone modification. Cell 2011; 146:1016–1028.
Li YY, Sabari BR, Panchenko T, et al. Molecular coupling of histone crotonylation and active transcription by AF9 YEATS domain. Mol Cell 2016; 62:181–193.
Zhao D, Guan HP, Zhao S, et al. YEATS2 is a selective histone crotonylation reader. Cell Res 2016; 26:629–632.
Andrews FH, Shinsky SA, Shanle EK, et al. The Taf14 YEATS domain is a reader of histone crotonylation. Nat Chem Biol 2016; 12:396–398.
Flynn EM, Huang OW, Poy F, et al. A subset of human bromodomains recognizes butyryllysine and crotonyllysine histone peptide modifications. Structure 2015; 23:1801–1814.
Hirschey MD, Zhao YM . Metabolic regulation by lysine malonylation, succinylation, and glutarylation. Mol Cell Proteomics 2015; 14:2308–2315.
Tan M, Peng C, Anderson KA, et al. Lysine glutarylation is a protein posttranslational modification regulated by SIRT5. Cell Metab 2014; 19:605–617.
Pougovkina O, Te Brinke H, Wanders RJ, Houten SM, de Boer VC . Aberrant protein acylation is a common observation in inborn errors of acyl-CoA metabolism. J Inherit Metab Dis 2014; 37:709–714.
Squatrito M, Gorrini C, Amati B . Tip60 in DNA damage response and growth control: many tricks in one HAT. Trends Cell Biol 2006; 16:433–442.
Sabari BR, Zhang D, Allis CD, Zhao Y . Metabolic regulation of gene express through histone acylations. Nat Rev Mol Cell Biol 2017; 18:90–101.
Cox J, Mann M . MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat Biotechnol 2008; 26:1367–1372.
Falcon S, Gentleman R . Using GOstats to test gene lists for GO term association. Bioinformatics 2007; 23:257–258.
Scholz C, Lyon D, Refsgaard JC, Jensen LJ, Choudhary C, Weinert BT . Avoiding abundance bias in the functional annotation of post-translationally modified proteins. Nat Methods 2015; 12:1003–1004.
Padmakumar R, Gantla S, Banerjee R . A rapid method for the synthesis of methylmalonyl-coenzyme A and other CoA-esters. Anal Biochem 1993; 214:318–320.
Vizcaino JA, Csordas A, del-Toro N, et al. 2016 update of the PRIDE database and its related tools. Nucleic Acids Res 2016; 44:D447–D456.
Acknowledgements
Our work was supported by the National Institutes of Health (NIH) award GM105933, DK107868 and GM115961 (YZ). This work was also supported by the National Key Research and Development Program of China (2017YFA0505103 to JD), Research Fund for the Doctoral Program of Higher Education of China (20120002110022 to JD) and the National Natural Science Foundation of China (81570060 to LD).
Author information
Authors and Affiliations
Corresponding authors
Additional information
( Supplementary information is linked to the online version of the paper on the Cell Research website.)
Supplementary information
Supplementary information, Figure S1
Synthesis of 2-hydroxyisobutyryl-CoA. (PDF 502 kb)
Supplementary information, Figure S2
Competition assay of Tip60's acyltransferase activity towards Acetyl-CoA and 2-Hydroxyisobutyryl-CoA. (PDF 242 kb)
Supplementary information, Figure S3
Spectra of the H4Khib peptides identified in the in vitro assay. (PDF 599 kb)
Supplementary information, Figure S4
The ratios of quantifiable histone H4 N-terminal Khib sites between Tip60 overexpressed (OE) 293T cells and wildtype (WT) 293T cells. (PDF 225 kb)
Supplementary information, Figure S5
Specificity of the pan anti-Khib antibody. (PDF 273 kb)
Supplementary information, Figure S6
Gene Ontology (GO) terms and UniProt keywords enrichment analysis on the HeLa Khib proteome. (PDF 193 kb)
Supplementary information, Table S1
The ratios of quantifiable histone Khib sites between HDAC3 overexpressed 293T cells and control 293T cells. (PDF 170 kb)
Supplementary information, Data S1
List of Khib Sites Identified in HeLa cells, Related to Figure 1. (XLSX 2051 kb)
Supplementary information, Data S2
KEGG Pathway and Protein Complex Analysis of the Khib Proteins Identified in HeLa cells. (XLSX 27 kb)
Supplementary information, Data S3
Khib Enzymes Identified in HeLa cells, Related to Figure 2. (XLSX 89 kb)
Supplementary information, Data S4
Potential Crosstalk between Reported Phosphorylation Sites and Khib Sites. (XLSX 20 kb)
Rights and permissions
About this article
Cite this article
Huang, H., Luo, Z., Qi, S. et al. Landscape of the regulatory elements for lysine 2-hydroxyisobutyrylation pathway. Cell Res 28, 111–125 (2018). https://doi.org/10.1038/cr.2017.149
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2017.149
Keywords
This article is cited by
-
A glimpse into novel acylations and their emerging role in regulating cancer metastasis
Cellular and Molecular Life Sciences (2024)
-
Roles of protein post-translational modifications in glucose and lipid metabolism: mechanisms and perspectives
Molecular Medicine (2023)
-
Proteomic analysis of protein lysine 2-hydroxyisobutyrylation (Khib) in soybean leaves
BMC Plant Biology (2023)
-
Lysine 2-hydroxyisobutyrylation of NAT10 promotes cancer metastasis in an ac4C-dependent manner
Cell Research (2023)
-
Computational investigation on the effect of the lysine 2-hydroxyisobutyrylation on argininosuccinate synthetase 1 conformational dynamics in Botrytis cinerea
Journal of Molecular Modeling (2023)