Abstract
Ryanodine receptors (RyRs) are a class of giant ion channels with molecular mass over 2.2 mega-Daltons. These channels mediate calcium signaling in a variety of cells. Since more than 80% of the RyR protein is folded into the cytoplasmic assembly and the remaining residues form the transmembrane domain, it has been hypothesized that the activation and regulation of RyR channels occur through an as yet uncharacterized long-range allosteric mechanism. Here we report the characterization of a Ca2+-activated open-state RyR1 structure by cryo-electron microscopy. The structure has an overall resolution of 4.9 Å and a resolution of 4.2 Å for the core region. In comparison with the previously determined apo/closed-state structure, we observed long-range allosteric gating of the channel upon Ca2+ activation. In-depth structural analyses elucidated a novel channel-gating mechanism and a novel ion selectivity mechanism of RyR1. Our work not only provides structural insights into the molecular mechanisms of channel gating and regulation of RyRs, but also sheds light on structural basis for channel-gating and ion selectivity mechanisms for the six-transmembrane-helix cation channel family.
Similar content being viewed by others
Introduction
Ryanodine receptors (RyRs) mediate calcium signaling in a variety of cells, including muscle and neuron cells. In muscle cells, RyRs are responsible for the rapid release of Ca2+ ions from sarcoplasmic/endoplasmic reticulum (SR/ER) into the cytoplasm, a key event that triggers muscle contraction1,2,3,4. Three mammalian isoforms, RyR1, RyR2, and RyR3, share about 70% sequence identity. RyR1 and RyR2 are primarily expressed in skeletal and cardiac muscles, respectively. RyR3 was originally found in the brain, but it was later found to be expressed in a variety of tissues5,6,7. RyRs also share considerable sequence similarity with the inositol-1,4,5-triphosphate (Ins(1,4,5)P3 or IP3) receptor (IP3R), which is another major sarcoplasmic/endoplasmic reticulum Ca2+ release channel8,9,10. RyRs are the largest known ion channels. They have a molecular mass of more than 2.2 mega-Daltons (MDa) and consist of four identical subunits, each containing over 5 000 residues, more than 80% of which are folded into the cytoplasmic assembly; the remaining residues form the transmembrane region5,11. Four identical transmembrane segments enclose a central pore that functions to conduct ions, whereas the cytoplasmic assembly interacts with diverse ligands ranging from ions such as Ca2+ to proteins such as the voltage-gated Ca2+ channel/dihydropyridine receptor (DHPR, also known as Cav1.1). RyRs can therefore respond to a large number of stimuli with complex regulatory mechanisms11,12,13,14.
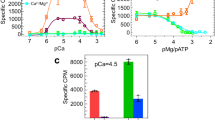
Ca2+ ions are a primary regulator of RyRs3,4,11,12,13,14,15,16,17,18,19. Single-channel recordings of RyR1 revealed that the probability plot of channel opening versus Ca2+ concentration is bell-shaped4,19. Specifically, when the Ca2+ concentration is lower than the nano-molar level, RyR1 is in a resting, closed state. When the Ca2+ concentration is between the nano-molar and micro-molar level, RyR1 begins to be activated, and its probability of opening increases as the Ca2+ concentration is elevated and reaches a maximum at 10∼100 μM Ca2+. With a Ca2+ concentration higher than 100 μM, RyR1 begins to be inactivated; it completely closes at Ca2+ concentrations ≥1 mM. In cardiac myocytes, voltage-gated opening of the Ca2+ channel Cav1.2 causes an influx of Ca2+ ions, and this subsequently activates RyR2. In skeletal myocytes, action potential-induced conformational changes of the Ca2+ channel Cav1.1 are physically coupled to channel opening of RyR1. In essence, Cav1.1 serves as the voltage sensor for RyR120,21,22. A variety of other chemicals (such as ATP/Mg2+, caffeine, ryanodine, PCB-95) and proteins (such as FK506-binding protein (FKBP), calmodulin, calsequestrin, triadin, junctin) modulate RyRs through independent or synergistic mechanisms (for review, Van Petegem14). As RyRs have an essential role in muscle contraction, their functional mutations are associated with several debilitating diseases. Aberrant function of RyR1 contributes to malignant hyperthermia and central core disease. Mutations in RyR2 may lead to heart disorders. In total, more than 500 disease-derived mutations have been mapped to the primary sequences of RyRs, with several hot spots of particular interest14.
Previously, we and two other groups independently determined the structure of a closed-state RyR123,24,25. We built an atomic structure model of RyR1. On the basis of this structural model, we discovered that (1) RyRs belong to the six-transmembrane-helix cation channel superfamily; (2) RyRs have a hierarchical organization, and are composed of the channel region, the central region, the peripheral region, and the N-terminal domain (NTD), with each region containing several subdomains; (3) all domains are interconnected in space23. The structure model served as a framework for exploring the regulation of the RyR channel by ligands.
The regulation of the RyR1 channel is known to be complex; previous structural and functional studies have demonstrated that the opening of the RyR1 channel can be regulated by effectors at every tier of the RyR1 structure — i.e., from the top, from the middle, from the bottom, and from the side14 — indicating that the regulation of RyR channel occurs through a long-range allosteric mechanism. However, the structural mechanism through which such long-range allostery occurs remains unknown. Here we utilized a cryo-electron microscopy (cryo-EM) approach to determine an open-state structure of RyR1 that is activated by Ca2+ and further blocked/stabilized by ruthenium red (RR), an open channel blocker26,27. Structural analyses and comparisons with the apo/closed-state RyR1 provide novel insights into the long-sought structural mechanisms of long-range allosteric Ca2+ activation and channel gating, and the structural basis for broad-spectrum ion conductivity.
Results
Global structural changes of RyR1 upon activation by Ca2+
To understand the structural mechanisms underlying long-range allosteric Ca2+ activation and channel gating, we activated the RyR1 channel with 100 μM Ca2+, a concentration at which the RyR1 channel has been demonstrated to be in a highly activated state (open probability > 0.85)4. This activated state was further blocked and stabilized by the addition of RR, which is known to lock the activated RyR1 channel in an open state26,27. We captured this Ca2+-activated and RR-blocked open-state (hereafter referred to as “open-state”) of RyR1 by fast freezing, and determined its structure with cryo-EM and a local-optimization refinement algorithm (Supplementary information, Figures S1-S5, and Tables S1-S3). The EM map of this open-state is of high resolution (Supplementary information, Figure S1A), which enabled us to trace the individual domains/motifs (Supplementary information, Figure S1B and S1C) and amino acid residues in the EM map (Supplementary information, Figures S4 and S5), and subsequently built up an atomic structural model (Supplementary information, Figure S1D).
The resolution of the whole molecule of the open-state RyR1 as determined by this approach is 4.9 Å; the resolution of the core region, which describes the central region of the molecule that contains the EF-hand motif, the U-motif, the voltage-sensor like domain (VSL), the C-terminal domain (CTD), and the transmembrane region is 4.2 Å (Supplementary information, Figure S2A). The local resolutions in the core region are between 3.8–4.6 Å (Supplementary information, Figure S2B). At this resolution, the densities of amino acid side chains of the core region are resolved and can be traced in the EM map (Supplementary information, Figures S4 and S5). We built up a three-dimensional (3D) structure model of the open-state RyR1 (Supplementary information, Figure S1D, Table S2 and Movie S1) based on the model of the previously reported apo/closed RyR1 structure23. It should be noted that the extra density observed in ion conduction pore of the open-state RyR1, which may correspond to RR, is absent in the closed-state EM maps, confirming that the current RyR1 conformation is in an RR-blocked open state (Supplementary information, Figure S6 and Data S1).
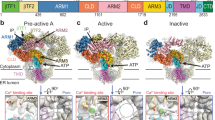
Structural analyses revealed that RyR1 undergoes pronounced global conformational changes when activated by Ca2+ (Figure 1 and Supplementary information, Table S3). The whole molecule displays a remarkable “breathing” motion (Figures 1 and 2A, and Supplementary information, Movie S2). In the transition from the apo/closed-state (hereafter, referred to as “closed-state”) to the open-state, the helical domain 2 (HD2) moves both outward and downward as viewed from the side (Figure 2A and Supplementary information, Movie S2); the helical domain 1 (HD1) and NTD each move upward and outward as viewed from the side (Figure 2A and Supplementary information, Movie S2). These movements together increase the width (from 225.7 to 237.9 Å; Figure 1A) and height (from 86.8 to 91.4 Å; Figure 1B) of the cytoplasmic assembly. The transmembrane region, composed of the pore-forming helix pair S5-S6, pore helices 1 and 2, the outer helix bundle of S1-S3, helix S4, the linker helix of S4-S5, and the extension helix S1′ (Figure 2A)23, displays a lateral dilation, shortening its height (from 40.1 to 37.8 Å; Figure 1B) and increasing its width (from 117.2 to 129.2 Å; Figure 1C).
Structural comparison between apo/closed and activated/open RyR1s. Structure models (colored cylinders and lines) of closed-state RyR1 and open-state RyR1 are superimposed on the cryo-EM maps (semi-transparent grey) in top (A), side (B), and bottom (C) views. In the top views, the side and diagonal dimensions of the cytoplasmic assembly of each state are labeled. In the side views, the heights of the cytoplasmic assembly and the transmembrane region of each state are marked. In the bottom views, only the transmembrane region, including helices S1-S6, the linker helix S4-S5, and extension helix S1′, is displayed with the diameter labeled.
Conformational changes of RyR1 upon Ca2+ activation and the initiation of conformational changes at the EF-hand motif. (A) Overall conformational comparison between the apo/closed and the activated/open forms of RyR1s. The structure models are displayed with individual domains/motifs depicted in different colors. For nomenclature and abbreviations of the domains/regions, see Supplementary information, Table S3. To facilitate visualization, only two opposing subunits are displayed here. (B) Structural transitions of the EF-hand module, central domain, U-motif, and CTD. The conformations of the EF-hand module in the closed and open states are denoted as the angles between E and F helices for the two E/F pairs (also Supplementary information, Table S4). In the open state, the conformational change of EF-hand 1 is depicted by a curved arrow at the bottom and the outward movement of whole EF-hand module is indicated by an arrow on the top, and the Ca2+ binding site is denoted as *Ca2+. (C) Structural transitions of the U-motif, CTD, VSL, and transmembrane region. In the open state, the outward movement of the U-motif is indicated by an arrow on the top. In all figures, the movements of individual domains/motifs are depicted as arrows with the same color-coding as the individual domains/motifs. In B and C, the left panels show the conformational changes and the right panels show the structural details in the closed and open states. See also Supplementary information, Figure S5.
Initiation of global conformational changes by the EF-hand motif
An overall comparison of the open-state structure with the closed-state structure reveals global structural differences. The open-state structure is “expanded” throughout the whole RyR1 molecule (Figures 1 and 2A), indicating that conformational changes propagate over the whole molecule. To reveal the structural mechanism underlying this conformational propagation, we conducted a detailed analysis of the open-state structure in comparison with the closed-state structure.
Sequence analysis has predicted that RyR1 possesses several EF-hand-type domains/motifs5,28,29. Our structural analyses of both the closed-state and the open-state RyR1 reveal that amino acids (aa) 4 071-4 132 in the central domain of RyR1 constitute the EF-hand motif (Supplementary information, Figure S1B). This motif contains two EF hands, namely EF-hand 1 and EF-hand 2, and each EF hand is composed of a helix pair (helix E1/F1 and helix E2/F2; Supplementary information, Figure S5A). Consistent with our structural analyses, the aa 4 081-4 127 region has been predicted to harbor a EF-hand motif containing two putative Ca2+-binding sites29, and mutational and functional studies have shown that EF-hand 1 is likely respnsible for Ca2+ activation of RyR129. Although several other regions have also been predicted to contain EF hands, our structural results reveal that these regions either do not form an EF-hand-like structure (aa 594-656), or display disordered conformation (aa 4 253-4 499) and thus are unlikely to form stable EF-hand-like structures that would be able to bind Ca2+ and regulate RyR1. Indeed, mutations in three predicted EF-hand motifs in the aa 4 253-4 499 region, either singly or in combination, do not affect the function of RyR129.
Comparison of the structures of the EF-hand motif in the open- and closed-states reveals significant difference (Figure 2B and Supplementary information, Figure S7B). It is known that when the EF hand binds Ca2+, the angle between the E and F helices changes30. To evaluate whether the two EF hands of the open-state RyR1 are occupied by Ca2+, we compared the angles between the E and F helices of EF-hand 1 and EF-hand 2 in the open-state structure with those in the closed-state structure. The angle between the E and F helices in EF-hand 1 changes from 71° in the closed-state to 38° in the open-state, and the E1 helix undergoes an anti-clockwise rotation (42.3°) and an outward movement (2.4 Å) during the transition. The angle between the E and F helices in EF-hand 2, however, exhibits only a slight shift from 87° (closed-state) to 76° (open-state; Figure 2B and Supplementary information, Figure S7B, Table S4 and Movie S2). These structural features indicate that EF-hand 1 is most likely the site occupied by Ca2+ whereas EF-hand 2 likely remains void. In combination with previous functional results which showed that only mutations of the putative EF-hand motif in the aa 4 081-4 127 region affect the function of RyR129, our structural data suggest that EF-hand 1 in the central region of RyR1 binds Ca2+ and is most likely responsible for the initiation of Ca2+ activation.
Structural transitions of the central domain of RyR1
We next investigated how the initial conformational changes that occur at the EF-hand motif are propagated to other domains. To be clear, in the following descriptions of the results of our structural analyses, we define the movement toward the central channel pore as “inward movement”, the movement away from the central channel pore as “outward movement”, the movement toward the cytosol as “upward movement”, and the movement toward the lumen as “downward movement”.
The EF-hand motif is directly linked to the rest of the central domain via the link between helices E1 and helix 18 and the link between helices F2 and 19 (Figure 2B and Supplementary information, Figure S5B). In addition to these direct linkages, helix E2 has hydrophobic interactions with helices 18 and 19. Upon Ca2+ binding, the EF-hand motif retains its basic fold but nevertheless undergoes pronounced conformational changes. The center of mass of the EF-hand motif moves upward (1.8 Å) and outward (2.6 Å). Due to the direct linkages and the hydrophobic interactions of the EF-hand motif with the rest of the central domain, the center of mass of the central domain itself moves outward (2.3 Å) (Figure 2B).
The U-motif (also referred to as the “thumb and forefingers” motif24) is an extension motif from the C-terminal end of the central domain and is composed of helices 20-22 and two anti-parallel β-sheets23. The movement of the central domain brings about a coordinated outward movement of the U-motif (Figure 2B). The CTD is clamped by the U-motif via hydrophobic interactions of clustered hydrophobic residues (Figure 2C and Supplementary information, Figure S5C). The outward movement of the U-motif is relayed to the CTD and leads to a corresponding outward movement of the CTD (Figure 2C and Supplementary information, Movie S2). The U-motif is also coupled to the VSL through hydrophobic interactions between helices 22 and S2′ (Figure 2C and Supplementary information, Figure S5C). The outward movement of the U-motif pushes the VSL outward; the center of mass of VSL moves outward by 2.3 Å.
Structural transitions of the RyR1 transmembrane domain
Since the CTD is an extension motif of the inner helix S6, and the VSL is directly linked to the transmembrane helices S2-S3 that are coupled with inner helix S6 via helices S4 and S5, and the S4-S5 linker (Figure 3A and Supplementary information, Figure S4A), the movements of both CTD and VSL propagate to inner helix S6, thus causing the channel to open.
Structural transitions of the transmembrane region. (A) The coordination of the movements of the U-motif, CTD, VSL, and transmembrane helices. (B) Interactions of the inner S6 helices of the neighboring subunits, the U-motif, and the CTD in the closed-state and open-state structures of RyR1. NS6 denotes the inner helix S6 from the neighboring subunit. (C-E) Structural transitions of the transmembrane helices from side (C), cytoplasmic (D), and luminal (E) views. The movements of individual structural elements are depicted as arrows with the same color-coding. See also Supplementary information, Figure S5.
The CTD is connected to inner helix S6 via a loop between the helix H1 of the CTD and the N-terminal end of inner helix S6, and this loop has hydrophobic interactions with the U-motif (Figure 3B and Supplementary information, Figure S5D). Upon Ca2+ activation, the outward movements of the U-motif and CTD trigger an outward movement of inner helix S6. The hydrogen bond (E4948-Q4949) between the S6 helices in the neighboring subunits that exists in the closed-state and restrains the movement of S6, thus “locking” the channel in the closed-state, is broken by the movement of helix S6 (Figure 3B).
VSL (also referred to as the S2-S3 domain24) is an insertion domain between transmembrane helices S2 and S3. In the open-state, VSL moves outward (Figure 3A), and this movement triggers an outward movement of the helix bundle S1-S3. This helix bundle is held via hydrophobic interactions among helices S1, S2, and S3, and is coupled to inner helix S6 via helices S4 and S5, and the S4-S5 linker (Figure 3C and Supplementary information, Figure S5E and S5F). Upon Ca2+ activation, the transmembrane helix bundle S1-S3 undergoes a 3.2 Å outward movement. Helix S4 has a downward movement that drives the linker helix S4-S5 to move downward (Figure 3C). The movement of the linker helix S4-S5 pulls the outer helix S5, moving it outward.
In contrast to most transmembrane helices whose lengths only just span the lipid bilayer, inner helix S6 of RyR1 is a long helix that is double the length of a typical transmembrane helix; half of this helix is in the lipid bilayer and the other half extends into the cytoplasm. S6 is inclined toward the membrane and has a kink at residue Gly4934 that is located in the midpoint of helix S6; this kink acts as a “hinge”. Since the outer helix S5 is tightly packed with the transmembrane region of inner helix S6 via hydrophobic interactions (Supplementary information, Figure S5F), the outward movement of the outer helix S5 pulls the transmembrane region of the inner helix S6 outward around the hinge of Gly4934 (Figure 3C). The cytoplasmic region of inner helix S6 is coupled with the CTD (Figure 3B and Supplementary information, Figure S5D) and the CTD moves outward upon Ca2+ activation (Figure 3A); the movement of the CTD causes the cytoplasmic region of inner helix S6 to move outward around the hinge of Gly4934 (Figure 3B). Thus, the outward movements of VSL and CTD induce an overall lateral dilation of the transmembrane domain (Figure 3C-3E) that results in the coordinated outward movement of inner helix S6.
Ca2+-activated opening of the RyR1 channel
The outward movement of S6 leads to an increase in the distance between the S6 helices of opposing subunits (from 24.3 to 28.4 Å; Figure 4A); this breaks the “seal” formed by the gate residues Ile4937 of the four subunits23 and expands the ion-conduction pathway of RyR1 (Figure 4B), thus opening the channel (Figure 4C, 4D and Supplementary information, Movie S2).
Structural transitions underlying channel gating. (A) The different arrangements of the inner helix S6 in the closed-state and open-state structures of RyR1. The distance between the top ends of two S6 helices in opposing subunits, which represents the diameter of the cytoplasmic entrance of the ion-conduction pathway, is marked, and the kink angle between the upper half and the lower half of the inner helix S6 is indicated. The positions of the gate residue Ile4937 and the kink residue Gly4934 in inner helix S6 are indicated. (B) The channel passage, calculated by MOLE 2.073, is depicted by colored volume in the open-state (right) and closed-state (left). (C) Size change of the pore at the gate that is formed by the gate residues Ile4937 of the four subunits, as viewed from the side. (D) Size change of the pore at the gate from a top view. Density maps of the gate in the closed-state and open-state are shown with atomic models fitted. The gate residues Ile4937 of the four subunits, which form the hydrophobic seal of the gate, are highlighted with side chains as red sticks. (E) Superimposition of the inner pore region of the closed- and open-state RyR1 structures. In A and C, only the structural elements of two opposing subunits are displayed for clarity; in B, D, and E, the structural elements of all four subunits are shown. See also Supplementary information, Figure S5.
In the closed-state, the pore diameter conferred by the gate residues Ile4937 of the four subunits is 4.9 Å, blocking the passage of a hydrated Ca2+ ion (diameter ∼8.2 Å31). In the open-state, the pore diameter increases to 9.8 Å, a diameter sufficient to allow the passage of a hydrated Ca2+ ion (Figure 4C-4E).
Structural transitions of the selectivity filter of RyR1
The selectivity filter of an ion channel is typically formed by residues from all of the individual subunits of a multi-subunit channel protein; these subunits are often arranged symmetrically around the ion-conduction pathway of ion channel. The selectivity filter of RyR1 is formed by components from all four subunits, each of which contributes a pore helix (pore helix 1, P1), a short helix (pore helix 2, P2), and a loop between P1 and P2 (P1-P2 loop). The residues putatively conferring ion selectivity (G4894GGIGDE4900) are located in the P1-P2 loop23,32,33.
The selectivity filter of RyR1 is not stationary; its conformation changes during the state transition. P1 has an outward (1.3 Å) and an anti-clockwise rotational movement (9.5°) as viewed from the luminal side; P2 has an outward (1.8 Å) and an anti-clockwise rotational movement (7.4°; Figure 5A). Owing to these structural changes, the aperture of the selectivity filter expands. The narrowest pore diameter of the filter conferred by residues Gly4894 of the four subunits expands from 8.8 to 16.7 Å upon Ca2+ activation (Figure 5B and Supplementary information, Movie S2).
Structural transitions of the selectivity filter. (A) Superimposition of the selectivity filter region of the closed- and open-state RyR1 structures in the luminal (left) and side (right) views. The movements of P1 and P2 in the open-state, relative to the closed-state, are indicated with arrows. Substantial expansion of the filter aperture in the open-state structure, relative to the closed-state structure, is noted. (B) Comparisons of the selectivity filter regions in the closed-state and open-state of RyR1. Density maps of the selectivity filter are shown with atomic models fitted. The aperture diameters conferred by residues Gly4894 of the four subunits are labeled to indicate the expansion of the filter upon Ca2+ activation. (C) Comparisons of the selectivity filter regions from a side view. The selectivity filter-conferring sequence, GGGIGDE, is colored in red. Note that the filter sequence is not aligned in parallel with the ion conduction pathway but is rather oriented in a zig-zag pattern.
Surprisingly, the selectivity filter sequence GGGIGDE (Figure 5B and 5C, highlighted in red) in RyR1 is not aligned in parallel with the ion-conduction pathway. Rather, it is arranged in a zig-zag fashion (Figure 5C). Given that the selectivity filter is not aligned in parallel with the ion conduction pathway and is flexible, and that the aperture of the selectivity filter expands upon activation, the ion selectivity of RyR1 appears to be controlled by an electric field produced by the negatively charged residues located in the selectivity filter (Asp4899 and Glu4900), rather than by ion coordination or hydrogen bonding of hydrated water with the oxygen atoms of these residues, a scenario that would require the oxygen atoms from the four subunits to form a stationary aperture with a specific size and geometry in order to form coordination with the ion to be conducted. Actually, it was demonstrated that replacement of these two acidic residues in RyR1 with asparagine and glutamine profoundly affected ion permeation and selectivity33,34. The negatively charged residues and the mobile nature of the selectivity filter are responsible for making RyR1 a broad-spectrum cation-conduction channel. Indeed, it has been demonstrated that that RyR1 is a not just a Ca2+ channel, it is also a cation channel that can conduct most of the IA/mono- and IIA/divalent cations of the periodic table of elements35,36.
Structural transitions of the peripheral region of RyR1
The central domain acts as a “relay station” for communication between different regions of RyR1. The structural change in the central domain is not only transmitted downwards to the channel region, but is also transmitted upwards to the peripheral region.
The central domain is coupled with the handle domain via an interaction between helices 1 and 14 (Figure 6A). The structural change in the central domain is communicated to the handle domain and induces pronounced structural changes; in the transition from the closed-state to the open-state, helix 14 undergoes a 3.1 Å outward movement and the whole handle domain helix bundle is pushed by the central domain and moves outward (Figure 6A). Some peripheral helices display even larger outward movements. Helices 2, 3, and 7, for example, move 5.7, 7.1, and 4.5 Å, respectively, during the transition from the closed-state to the open-state (Figure 6A).
Structural transitions of the peripheral region. (A-C) Coordinated structural transitions of the handle domain (A), the helical domain (B), and the NTD (C) with the central domain. (D) Detailed structural changes of the NTD. All the nomenclatures of the domains and secondary structure elements are based on Yan et al.23.
The helical domain is coupled to the handle domain through interactions between helices 6 and 1a (Figure 6B). Consequently, the structural changes of the handle domain are relayed to the helical domain and induce the largest structural changes observed in RyR1. From the closed-state to the open-state, the whole helical domain is pushed by the handle domain and has a profound outward movement (Figure 6B). The helices in HD1 undergo considerable upward movement; helix 5a′, for example, moves 4.3 Å upwards. The helices in HD2 also undergo profound outward and/or downward movements. Helices 13a and 15a, for example, move 6.4 and 7.4 Å outward, and helix 17a moves downward 2.3 Å during the transition from the closed-state to the open-state (Figure 6B). These movements make RyR1 expand and thus account for the observed increase of the cytoplasmic assembly in both the height and width (Figure 1).
The profound structural changes of the helical domain indicate that it is the most dynamic domain of RyR1. The dynamic nature of the helical domain suggests that it might be sensitive to stimuli and may thus act as a platform for receiving stimulation signals such as those from the DHPR, which has been shown to be located just above HD1 and has been proposed to be physically coupled to RyR1 during excitation-contraction coupling21.
Structural transitions of the NTD of RyR1
The RyR1 NTD is composed of three subdomains: NTD-A, NTD-B, and NTD-C. The NTD is coupled to the handle domain through interactions between helices 2 and 3 of the handle domain and helices 11 and 12 of the NTD-C subdomain (Figure 6C). Structural changes in the handle domain are propagated to the NTD and lead to the corresponding conformational changes of the NTD. In the transition from the closed-state to the open-state, the whole NTD-C subdomain undergoes a large upward and outward movement (Figure 6C) during which helices 10, 11, and 12, for example, move outward 6.3, 5.3, and 6.1 Å, respectively.
The structural transition of the NTD affects the interactions among the subdomains of the NTD and also alters the interactions with the NTDs of the neighboring subunits. In the transition from the closed-state to the open-state, the distances between the centers of mass of subdomains A, B, and C within the NTD increase considerably (Figure 6D, top panel). The A-B, B-C, and C-A distances increase from 30.9, 36.7, and 38.9 to 32.3, 37.8, and 40.4 Å, respectively, indicating that the couplings between subdomains A, B, and C are weakened in this transition. Meanwhile, the distances between the centers of mass of the NTDs of neighboring subunits undergo profound increases (from 70.4 to 76.4 Å; Figure 6D, middle panel). The distance between subdomains A and B of neighboring NTDs increases from 35.3 to 38.6 Å (Figure 6D, bottom panel), indicating that the couplings between neighboring NTDs are substantially weakened. These structural data suggest that both the interactions between subdomains A, B, and C of the NTD and the interactions between neighboring NTDs likely play important roles in Ca2+-induced channel gating.
A model for long-range allosteric channel gating
Ca2+ is the primary regulator of RyRs; micromolar concentrations of Ca2+ activate RyR1 channel, while millimolar concentrations of Ca2+ actually inactivates it3,4,11,12,13,14,15,16,17,18,19. On the basis of structural model presented in the current study, we propose a model for the Ca2+-activated long-range allosteric channel gating of RyR1 (Figure 7A and 7B).
Schematic model of RyR1 channel activation by Ca2+ and allosteric communication pathway among domains. (A) Mechanism of channel activation by Ca2+, which results in the consecutive outward movements of domains/motifs/structural elements. The movements of domains are indicated as arrows. Only two opposing subunits are depicted for clarity. (B) Long-range allosteric communication transition pathway of RyR1 underlying Ca2+ activation. Ca2+ binding to the EF-hand in the central domain initiates a cascade of conformational transmissions via the U-motif, downwards to the VSL, the CTD and S6, inducing S6 to change conformation thus opening the channel. Meanwhile, the conformational changes of the central domain also transmit upwards to the peripheral cytoplasmic domains (NTD, handle, HD1 and HD2), causing global conformational changes of RyR1.
In the apo/closed state, the RyR1 channel is blocked by the hydrophobic gate residues Ile4937 of the four subunits23. In the Ca2+-activated open state, Ca2+ binding to the EF-hand motif in the central region induces a conformational change. This structural change is first communicated to the central domain and the central domain moves outward, away from the central channel pore. The conformational change in the central domain is then transmitted, via the U-motif, to the channel domain through the CTD, which is directly connected to the C-terminal end of inner helix S6, and induces an outward movement of inner helix S6. Meanwhile, the conformational change in the EF-hand motif is propagated, also via the U-motif, to the VSL. The outward movement of the VSL causes the transmembrane helix bundle S1-S3 to move outward and helix S4 to undergo a downward movement. This causes the linker helix S4-S5 to move downward. The movement of the S4-S5 linker drives outer helix S5 and inner helix S6 to move outward. The outward movement of inner helix S6 breaks the hydrogen bond between the S6 helices in the neighboring subunits. The outward movement of inner helix S6 breaks the hydrophobic interactions between the gate residues Ile4937 of the four subunits and moves these residues outwards, away from the central pore, thus opening the channel.
The movement of the central domain is also propagated to the NTD and the handle domain, and subsequently to the helical domain, thereby inducing significant conformational changes in the peripheral regions of RyR1 (Figure 7B). The detailed roles and transitions of each functional domain have been summarized in Supplementary information, Table S3.
Discussion
The present study greatly advances our understanding of the structural mechanisms of RyR1 channel gating and ion selectivity in four aspects.
First, we demonstrate that the RyR1 channel can be regulated by a regulator (in this case, Ca2+) from a remote site located far away from the channel domain; this regulation is achieved via a cascade of allosteric conformational transmissions. Our structural data revealed that RyR1 is essentially a giant molecular “machine” composed of multiple domains/motifs that act as “component parts”. The domains/motifs and the transmembrane helices are coupled, domain-to-domain or helix-to-helix, through hydrophobic or electrostatic interactions. These domain-domain and helix-helix interactions play a fundamental role in RyR1 long-range allosteric conformational transmission. As all domains/motifs are coupled through domain-domain/helix-helix interactions, RyR1 undergoes global conformational changes through allosteric conformational transmissions upon being activated. These structural characteristics suggest that the RyR1 channel can be regulated from certain domains via long-range conformational transmission. This explains structural basis of the previously reported observations that the RyR1 channel can be regulated by effectors at every tier of the RyR1 structure — i.e., from the top, the middle, the bottom, and the side14. The structural mechanisms of allosteric conformational transmission and channel gating can explain the previously reported observations that mutations of some critical residues in the interfaces between domains and between helices can cause debilitating diseases such as central core disease, congenital fiber type disproportion, and malignant hyperthermia23 (Supplementary information, Table S5), as these residues are essential in mediating domain-domain and/or helix-helix interactions that are essential for conformational transmission across domains and helices. Mutations in these residues would presumably impair the interactions between domains and/or between helices and disrupt the allosteric conformational transmissions that control channel gating, thereby leading to dysfunction of the RyR1 channel.
The structural mechanisms that we demonstrate here are consistent with our previous proposal that the RyR1 domains/motifs are interconnected in space23 and are consistent with the hypothesis that Ca2+ binding to the EF-hand motif located in the central domain activates the RyR1 channel via allosteric conformational transmissions as proposed by Marks and colleagues and ourselves23,24. Our structural data are also consistent with previous low-resolution structural studies suggesting that RyR1 exhibits global structural changes and expansion when activated by Ca2+37,38. The structural mechanisms reported here also fully elucidate the molecular interactions underlying the previously reported mutagenesis- and electrophysiological-study-based observations that the S4-S5 linker, Gly4934, and the CTD are all critical for RyR1 channel gating39,40,41,42,43,44,45.
Second, we here elucidate a novel channel gating mechanism for a six-transmembrane-helix cation channel family protein. The RyR1 channel is not only gated by the action of the VSL domain that is coupled to the pore-forming helix S6 through the transmembrane voltage-sensor domain in a way analogous to the voltage-gated Na+, K+ and Ca2+ channels46, but also by CTD that is directly coupled to the cytoplasmic region of the pore-forming helix S6. These enable the RyR1 channel to be gated not only from the transmembrane region but also from the cytosolic domain. In fact, the RyR CTD is a zinc-finger-like domain23 and Zn2+ can indeed modulate RyR gating from the cytosol47.
Third, our structural work reveals a novel ion selectivity mechanism of an ion channel. The ion selectivity filter of RyR1 is mobile and hence the ion selectivity is controlled by an electric field produced by the negatively charged residues in the selectivity filter. The negatively charged residues and the mobile nature of the selectivity filter would allow all cations smaller than the flexible aperture of the selectivity filter to pass through upon activation, hence making the RyR1 a broad-spectrum cation-conduction channel. In contrast, the ion selectivity filters of K+ and Na+ channels are typically stationary and the ion selectivity of these channels is often conferred by coordination of oxygen atoms from the residues located in stationary aperture of the selectivity filter with the ion to be conducted48,49. The coordination and the stationary nature of the selectivity filter make the K+ and Na+ channels highly selective ion channels.
In addition, it has long been known that RyR1 is activated by DHPR/Cav1.1, which is essential for skeletal muscle excitation-contraction coupling, and DHPR/Cav1.1 has been proposed to be physically coupled with RyR1 at a position corresponding to the helical domain20,21,22. On the basis of the structural mechanism of allosteric conformational transmission and channel gating that we deduce here, we propose that, upon depolarization of the surface membrane of skeletal muscle cells, the depolarization-induced action potential will induce a conformational change in DHPR/Cav1.1. This conformational change in DHPR/Cav1.1 will first be transmitted to the helical domain of RyR1 and subsequently initiate a cascade of conformational changes in RyR1 that propagate across the handle domain and NTD, to the central domain and the U-motif, and finally, to the channel domain, leading to the channel opening.
The structural mechanism of allosteric conformational transmission and channel gating of RyR1 that we characterized in the present study provides a framework for further exploration of the molecular mechanism of skeletal muscle excitation-contraction coupling and its regulation by RyR1 ligands. Future work will be conducted to decipher the structural mechanism of DHPR-RyR1 coupling and RyR1 regulation by modulatory proteins such as FKBP, calmodulin, calsequestrin, triadin, junctin, among others14,50,51, with the goal of understanding the molecular mechanism of excitation-contraction coupling and RyR1 regulation at the atomic level.
Materials and Methods
Materials
All chemicals were from Sigma-Aldrich, unless otherwise specified.
Preparation of skeletal heavy SR vesicles
Skeletal heavy SR vesicles were prepared from New Zealand white rabbit skeletal muscle according to a published method2.
RyR1 purification
RyR1 was purified from CHAPS-solubilized skeletal heavy SR essentially as described previously, with slight modifications3,52. Briefly, skeletal heavy SR vesicles were suspended in buffer A (1 M NaCl, 20 mM Na-HEPES, pH 7.4, 2 mM DTT, 2 mM PMSF, 1:1 000 diluted protease inhibitor cocktail (Sigma-Aldrich, P8340) and 1.2% CHAPS (AMRESCO)/0.6% soybean lecithin (Sigma-Aldrich, 11145); the CHAPS/protein ratio was 12 (wt/wt)). The sample was centrifuged for 1 h at 110 000× g after incubation on ice for 30 min with shaking. The supernatant was loaded onto a 5-ml hydroxyapatite ceramic (Bio-Rad) column equilibrated with buffer B (200 mM NaCl, 10 mM Na-HEPES, pH 7.4, 0.5% CHAPS/0.25% soybean lecithin, 2 mM DTT, 2 mM PMSF, 1:1 000 diluted protease inhibitors cocktail). The column was washed with buffer B containing 10 mM K2HPO4, followed washing with buffer B containing 50 mM K2HPO4. Proteins were then eluted with 15 ml of buffer B containing 200 mM K2HPO4. The eluate was collected and concentrated by centrifugation at 1 000× g in a 100-kDa cut-off Amicon centrifugal filter (Millipore) and loaded on the top of a 5% - 20% (w/v) linear sucrose gradient in buffer B. After centrifugation for 16 h in a Beckman SW28 rotor at 26 000 rpm, the gradient was fractionated into 1.5-ml fractions. After checking by SDS-PAGE, the RyR1-enriched fractions were collected and concentrated, rapidly frozen in liquid nitrogen, and stored in small aliquots at −80 °C.
Sample preparation for cryo-EM
It is known that the RyR1 particles prepared in buffer containing the detergent CHAPS have preferred orientations when they are frozen in vitreous ice; this hinders structural determination of RyR1 in high-resolution cryo-EM and single-particle analysis53. To overcome this difficulty, we replaced CHAPS with amphipol A8-35, which has been used successfully in the structural determination of transient receptor potential cation channel subfamily V member 1 (TRPV1), the first ion channel characterized by single-particle cryo-EM to near-atomic resolution54. It has been shown that RyR1 retains structural and functional integrity after replacement of CHAPS with amphipol A8-3553. We therefore replaced CHAPS with amphipol A8-35 and performed cryo-EM in buffer containing amphipol A8-35. Briefly, the purified RyR1 in buffer B was mixed with amphipol A8-35 (Anatrace) at 1:1 (wt/wt) to a final concentration of 5 mg/ml (∼2.5 μM). After incubation at 4 °C for 4 h, 200 mg of SM-2 bio-beads (Bio-Rad) was added and incubated overnight. The bio-beads were then removed over a spin column (Pierce), and RyR1 was further separated using a PD-10 column (GE) in buffer composed of 20 mM HEPES, pH 7.4, 300 mM NaCl, 2 mM DTT, 2 mM PMSF, and 1:1 000 diluted protease inhibitor cocktail. RyR1 (∼2.5 μM) was incubated with 100 μM Ca2+ for 10 min to activate the channel, and then mixed with 10 μM RR shortly before cryo-EM grid preparation to lock the channel in an open state. Ruthenium red is known to block the open RyR1 channel26,27 and lock the channel in an open state (Supplementary information, Data S1).
Cryo-EM
Aliquots of 3 μl of purified RyR1 (∼5 mg/ml) were placed on glow-discharged 400-mesh R2.0/2.0 Quantifoil holy carbon grids (Quantifoil Micro Tools GmbH). Grids were blotted for 2 seconds and flash-frozen in liquid ethane using an FEI Mark IV Vitrobot plunger. Grids were transferred to an FEI Titan Krios electron microscope that was operating at 300 kV. Images were collected automatically using SerialEM55 and recorded in video mode (17 frames/s) using an FEI Falcon-II detector at a nominal magnification of 59 000× g and a pixel size of 1.396 Å. A dose rate of 24 electrons per Å2 per second 24e−/(sec·Å2) and an exposure time of 2 s were used.
Image processing
The 31 frames of each video were processed into 10 images by merging 3 adjacent frames; these were then subjected to motion correction using the dosefgpu_driftcorr program56. A sum of frames in each image stack were used for further processing. We used the Gautomatch program (developed by Zhang K, MRC Laboratory of Molecular Biology, Cambridge, UK) for automated selection of 376 537 particles from 4 931 micrographs. Contrast transfer function parameters were estimated using Gctf57. All two- (2D) and three-dimensional (3D) classifications and refinements were performed using the RELION1.4 software58. We used initial analyses of 2D classification to discard bad particles, and selected 94 788 particles for an initial 3D classification. A 60-Å low-pass filtered cryo-EM reconstruction of RyR1 (Electron Microscopy Data Bank (EMDB) accession number 1606)38 was used as an initial model. 3D classification was performed with an angular sampling of 7.5 degrees with C4 symmetry imposed and further refined with local angular searches. The particles were therefore classified into four classes. Among these 4 classes, the three reasonable classes yielded a subset of 72 039 particles, which was subjected to 3D auto-refinement, particle-based motion correction, and radiation-damage weighting. The resulting 'polished particles' were used for an additional 3D classification analysis with 2 classes, and C4 symmetry was performed. Class 1 (41 743 particles) and class 2 (30 296 particles) yielded reconstructions at 4.9 and 7.7 Å, respectively, using 3D auto-refinement with C4 symmetry imposed. Subsequent focused refinement used a local-optimization refinement algorithm59,60 to subtract the signal of the peripheral region from the whole density map; this yielded maps of the core region (including the central region containing the EF-hand motif, the U-motif, VSL, CTD, and the transmembrane region) of class 1 and class 2 at 4.2 and 4.9 Å, respectively. Detailed structural analyses revealed that the gate diameter of the class 1 structure is 9.8 Å, sufficient to allow the hydrated Ca2+ (diameter 8.2 Å31) to pass through; we therefore designated the class 1 structure as “open-state”. The gate diameter of the class 2 structure is 7.1 Å, which is larger than that in the apo/closed-state structure (diameter 4.9 Å23) but is insufficiently wide to allow the hydrated Ca2+ to pass through; the class 2 structure therefore represents a “non-conducting state”. In this paper, we focus on the open-state; the non-conducting state will be described in a separate manuscript.
Reported resolutions are based on the gold-standard Fourier shell correlation (FSC) 0.143 criterion, and FSC curves were corrected for the effects of a soft mask on the FSC curve using high-resolution noise substitution61. Prior to visualization, all density maps were corrected for the modulation transfer function of the detector; these were then sharpened by applying a negative B-factor that was estimated using automated procedures62. Local resolution variations were estimated using ResMap63.
The complete process of cryo-EM data collection, image processing, three-dimensional reconstruction, structure validation, and resolution assessment is depicted in Supplementary information, Figures S2 and S3, and the cryo-EM data collection statistics is summarized in Supplementary information, Table S1.
Tilt pair validation
Tilt pairs were used to validate the cryo-EM maps of Ca2+-activated RyR1. Electron micrographs were taken at a 0° and 15° tilt angles (1-s exposure time was used for both angles) with an FEI Titan Kiros transmission electron microscope operating at 300 kV with an FEI Falcon II detector, a pixel size of 1.396 Å, and a dose rate of 24e−/(sec·Å2). The images were recorded at approximately 3-μm defocus. Movies were recorded at 16 frames per second and corrected for beam-induced drift by aligning the individual frames of each movie using whole-image motion correction56. CTF parameters for drift-corrected micrographs were estimated using Gctf57. A total of 936 particle pairs were picked manually using EMAN264 from 45 pairs of micrographs. After 2D and 3D classification, approximately 60% of particles were discarded. The screened particle pairs were then combined with the particles that were used for the final refinement of the corresponding structure. After another round of 3D refinement, the angular parameters of the particles pairs were extracted and analyzed using TiltDiffRelion65,66.
Model building
Local resolution assessment indicated that the resolutions of the core region of the final EM-map of the Ca2+-activated, open-state RyR1 are between 3.8–4.6 Å (Supplementary information, Figure S2B). At these resolutions, the densities of the side chains of amino acid residues are resolved and could be traced in the EM-map (Supplementary information, Figures S4 and S5).
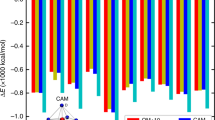
We built up an atomic structure model of the Ca2+-activated, open-state RyR1 via an integrated approach combining molecular dynamics and crystallographic methods. Briefly, the atomic model of the closed-state RyR1 (PDB code: 3J8H) was first split into four protomers and these were separately fitted into the overall open-state cryo-EM density map using “Fit in Map” in Chimera67. The density of one protomer was extracted using “Surface Zone”. The atomic model of one protomer was then used as an initial model for molecular dynamics flexible fitting (MDFF). All MDFF and atomistic MD simulations were performed using NAMD 2.968 and the CHARMM22 force field69, with CAMP corrections70. First, MDFF was run for 1 ns with a scaling factor 0.3 kcal·mol−1, yielding converged structures. Second, energy minimization was performed with 2 ps and a scaling factor of 10 kcal/mol.
After MDFF, the model was adjusted in Coot71. Some regions not seen in the closed-state RyR1 model were built de novo according to the map of the core region. In this way, most of residues in the core region were built (Supplementary information, Figures S4 and S5). The model of the core region was then adjusted in Coot and was refined by REFMAC5 in CCP4–7.072. The resolution limit for refinement was set at 4.2 Å. The statistics of model refinement is listed in Supplementary information, Table S2.
Finally, the atomic models of the tetramer for the whole RyR1 molecule were generated by imposing four-fold rotation symmetry to the atomic model of the protomer in accordance with the EM maps of the tetramer, as described previously23.
All structural figures here were generated with UCSF Chimera67 and PyMOL (www.pymol.org). The size of the Ca2+ channel of RyR1 was quantitatively analyzed by using MOLE 2.073.
Accession codes
The overall cryo-EM map of Ca2+-activated RyR and the cryo-EM map of its core region have been deposited in EMDB with the accession codes EMD-8073 and EMD-8074, respectively. The refined coordinates of Ca2+-activated RyR against the overall cryo-EM map have been deposited in the Protein Data Bank (PDB) with the accession code 5J8V.
Author Contributions
CCY and FS designed research; RW performed biochemical work and collected EM data; XW collected EM data, performed image processing and 3D reconstruction; YZ performed MDFF; LZ collected EM data and performed image processing; SM and AJW conducted single channel recording experiments; QC, XH, SJ, CL, SL, GW, YX and SZ performed biochemical work; RW, WX, FS and CCY analyzed data; CCY and FS wrote the paper with the input from other authors.
Competing Financial Interests
The authors declare no competing financial interests.
References
Pessah IN, Waterhouse AL, Casida JE . The calcium-ryanodine receptor complex of skeletal and cardiac muscle. Biochem Biophys Res Commun 1985; 128:449–456.
Inui M, Saito A, Fleischer S . Purification of the ryanodine receptor and identity with feet structures of junctional terminal cisternae of sarcoplasmic reticulum from fast skeletal muscle. J Biol Chem 1987; 262:1740–1747.
Lai FA, Erickson HP, Rousseau E, Liu QY, Meissner G . Purification and reconstitution of the calcium release channel from skeletal muscle. Nature 1988; 331:315–319.
Smith JS, Imagawa T, Ma J, Fill M, Campbell KP, Coronado R . Purified ryanodine receptor from rabbit skeletal muscle is the calcium-release channel of sarcoplasmic reticulum. J Gen Physiol 1988; 92:1–26.
Takeshima H, Nishimura S, Matsumoto T, et al. Primary structure and expression from complementary DNA of skeletal muscle ryanodine receptor. Nature 1989; 339:439–445.
Otsu K, Willard HF, Khanna VK, Zorzato F, Green NM, MacLennan DH . Molecular cloning of cDNA encoding the Ca2+ release channel (ryanodine receptor) of rabbit cardiac muscle sarcoplasmic reticulum. J Biol Chem 1990; 265:13472–13483.
Hakamata Y, Nakai J, Takeshima H, Imoto K . Primary structure and distribution of a novel ryanodine receptor/calcium release channel from rabbit brain. FEBS Lett 1992; 312:229–235.
Mignery GA, Sudhof TC, Takei K, De Camilli P . Putative receptor for inositol 1,4,5-trisphosphate similar to ryanodine receptor. Nature 1989; 342:192–195.
Seo MD, Velamakanni S, Ishiyama N, et al. Structural and functional conservation of key domains in InsP3 and ryanodine receptors. Nature 2012; 483:108–112.
Fan G, Baker ML, Wang Z, et al. Gating machinery of InsP3R channels revealed by electron cryomicroscopy. Nature 2015; 527:336–341.
Van Petegem F . Ryanodine receptors: structure and function. J Biol Chem 2012; 287:31624–31632.
Meissner G . Ryanodine receptor/Ca2+ release channels and their regulation by endogenous effectors. Annu Rev Physiol 1994; 56:485–508.
Lanner JT, Georgiou DK, Joshi AD, Hamilton SL . Ryanodine receptors: structure, expression, molecular details, and function in calcium release. Cold Spring Harb Perspect Biol 2010; 2:a003996.
Van Petegem F . Ryanodine receptors: allosteric ion channel giants. J Mol Biol 2015; 427:31–53.
Endo M, Tanaka M, Ogawa Y . Calcium induced release of calcium from the sarcoplasmic reticulum of skinned skeletal muscle fibres. Nature 1970; 228:34–36.
Smith JS, Coronado R, Meissner G . Single channel measurements of the calcium release channel from skeletal muscle sarcoplasmic reticulum. Activation by Ca2+ and ATP and modulation by Mg2+. J Gen Physiol 1986; 88:573–588.
Hymel L, Inui M, Fleischer S, Schindler H . Purified ryanodine receptor of skeletal muscle sarcoplasmic reticulum forms Ca2+-activated oligomeric Ca2+ channels in planar bilayers. Proc Natl Acad Sci USA 1988; 85:441–445.
Ma J, Fill M, Knudson CM, Campbell KP, Coronado R . Ryanodine receptor of skeletal muscle is a gap junction-type channel. Science 1988; 242:99–102.
Bezprozvanny I, Watras J, Ehrlich BE . Bell-shaped calcium-response curves of Ins(1,4,5)P3- and calcium-gated channels from endoplasmic reticulum of cerebellum. Nature 1991; 351:751–754.
Rios E, Brum G . Involvement of dihydropyridine receptors in excitation-contraction coupling in skeletal muscle. Nature 1987; 325:717–720.
Paolini C, Fessenden JD, Pessah IN, Franzini-Armstrong C . Evidence for conformational coupling between two calcium channels. Proc Natl Acad Sci USA 2004; 101:12748–12752.
Paolini C, Protasi F, Franzini-Armstrong C . The relative position of RyR feet and DHPR tetrads in skeletal muscle. J Mol Biol 2004; 342:145–153.
Yan Z, Bai XC, Yan C, et al. Structure of the rabbit ryanodine receptor RyR1 at near-atomic resolution. Nature 2015; 517:50–55.
Zalk R, Clarke OB, des Georges A, et al. Structure of a mammalian ryanodine receptor. Nature 2015; 517:44–49.
Efremov RG, Leitner A, Aebersold R, Raunser S . Architecture and conformational switch mechanism of the ryanodine receptor. Nature 2015; 517:39–43.
Ma J . Block by ruthenium red of the ryanodine-activated calcium release channel of skeletal muscle. J Gen Physiol 1993; 102:1031–1056.
Xu L, Tripathy A, Pasek DA, Meissner G . Ruthenium red modifies the cardiac and skeletal muscle Ca(2+) release channels (ryanodine receptors) by multiple mechanisms. J Biol Chem 1999; 274:32680–32691.
Xiong H, Feng X, Gao L, et al. of a two EF-hand Ca2+ binding domain in lobster skeletal muscle ryanodine receptor/Ca2+ release channel. Biochemistry 1998; 37:4804–4814.
Fessenden JD, Feng W, Pessah IN, Allen PD . Mutational analysis of putative calcium binding motifs within the skeletal ryanodine receptor isoform, RyR1. J Biol Chem 2004; 279:53028–53035.
Xie X, Dwyer MD, Swenson L, Parker MH, Botfield MC . Crystal structure of calcium-free human sorcin: a member of the penta-EF-hand protein family. Protein Sci 2001; 10: 2419–2425.
Conway BE . Ionic hydration in chemistry and biophysics. Amsterdam; New York: Elsevier Scientific Pub. Co., 1983.
Balshaw D, Gao L, Meissner G . Luminal loop of the ryanodine receptor: a pore-forming segment? Proc Natl Acad Sci USA 1999; 96:3345–3347.
Gillespie D, Xu L, Wang Y, Meissner G . (De)constructing the ryanodine receptor: modeling ion permeation and selectivity of the calcium release channel. J Phys Chem B 2005; 109:15598–15610.
Wang Y, Xu L, Pasek DA, Gillespie D, Meissner G . Probing the role of negatively charged amino acid residues in ion permeation of skeletal muscle ryanodine receptor. Biophys J 2005; 89:256–265.
Chen DP, Xu L, Tripathy A, Meissner G, Eisenberg B . Selectivity and permeation in calcium release channel of cardiac muscle: alkali metal ions. Biophys J 1999; 76:1346–1366.
Carney J, Mason SA, Viero C, Williams AJ . The ryanodine receptor pore: is there a consensus view? Curr Top Membr 2010; 66:49–67.
Serysheva II, Schatz M, van Heel M, Chiu W, Hamilton SL . Structure of the skeletal muscle calcium release channel activated with Ca2+ and AMP-PCP. Biophys J 1999; 77:1936–1944.
Samso M, Feng W, Pessah IN, Allen P . D Coordinated movement of cytoplasmic and transmembrane domains of RyR1 upon gating. PLoS Biol 2009; 7:e85.
Gao L, Tripathy A, Lu X, Meissner G . Evidence for a role of C-terminal amino acid residues in skeletal muscle Ca2+ release channel (ryanodine receptor) function. FEBS Lett 1997; 412:223–226.
Stewart R, Zissimopoulos S, Lai FA . Oligomerization of the cardiac ryanodine receptor C-terminal tail. Biochem J 2003; 376:795–799.
Pouliquin P, Pace SM, Curtis SM, et al. Effects of an alpha-helical ryanodine receptor C-terminal tail peptide on ryanodine receptor activity: modulation by Homer. Int J Biochem Cell Biol 2006; 38:1700–1715.
Murayama T, Kurebayashi N, Oba T, et al. Role of amino-terminal half of the S4-S5 linker in type 1 ryanodine receptor (RyR1) channel gating. J Biol Chem 2011; 286:35571–35577.
Ramachandran S, Chakraborty A, Xu L, et al. Structural determinants of skeletal muscle ryanodine receptor gating. J Biol Chem 2013; 288:6154–6165.
Euden J, Mason SA, Viero C, Thomas NL, Williams AJ . Investigations of the contribution of a putative glycine hinge to ryanodine receptor channel gating. J Biol Chem 2013; 288:16671–16679.
Mei Y, Xu L, Mowrey DD, et al. Channel gating dependence on pore lining helix glycine residues in skeletal muscle ryanodine receptor. J Biol Chem 2015; 290:17535–17545.
Tombola F, Pathak MM, Isacoff EY . How does voltage open an ion channel? Annu Rev Cell Dev Biol 2006; 22:23–52.
Woodier J, Rainbow RD, Stewart AJ, Pitt SJ . Intracellular zinc modulates cardiac ryanodine receptor-mediated calcium release. J Biol Chem 2015; 290:17599–17610.
Bagnéris C, Naylor CE, McCusker EC, Wallace BA . Structural model of the open-closed-inactivated cycle of prokaryotic voltage-gated sodium channels. J Gen Physiol 2015; 145:5–16.
Kuang Q, Purhonen P, Hebert H . Structure of potassium channels. Cell Mol Life Sci 2015; 72:3677–3693.
Rebbeck RT, Karunasekara Y, Board PG, et al. Skeletal muscle excitation-contraction coupling: who are the dancing partners? Int J Biochem Cell Biol 2014; 48:28–38.
Yuchi Z, Van Petegem F . Ryanodine receptors under the magnifying lens: Insights and limitations of cryo-electronmicroscopy and X-ray crystallography studies. Cell Calcium 2016; 59:209–227.
Yin CC, Lai FA . Intrinsic lattice formation by the ryanodine receptor calcium-release channel. Nat Cell Biol 2000; 2:669–671.
Baker MR, Fan G, Serysheva II . Single-particle cryo-EM of the ryanodine receptor channel in an aqueous environment. Eur J Transl Myol 2015; 25:35–48.
Liao M, Cao E, Julius D, Cheng Y . Structure of the TRPV1 ion channel determined by electron cryo-microscopy. Nature 2013; 504:107–112.
Mastronarde DN . Automated electron microscope tomography using robust prediction of specimen movements. J Struct Biol 2005; 152:36–51.
Li X, Mooney P, Zheng S, et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nat Methods 2013; 10:584–590.
Zhang K . Gctf: Real-time CTF determination and correction. J Struct Biol 2016; 193:1–12.
Scheres SH . RELION: implementation of a Bayesian approach to cryo-EM structure determination. J Struct Biol 2012; 180:519–530.
Bai XC, Rajendra E, Yang G, Shi Y, Scheres SH . Sampling the conformational space of the catalytic subunit of human gamma-secretase. Elife 2015; 4:e11182.
Shan H, Wang Z, Zhang F, Xiong Y, Yin CC, Sun F . A local-optimization refinement algorithm in single particle analysis for macromolecular complex with multiple rigid modules. Protein Cell 2016; 7:46–62.
Scheres SH, Chen S . Prevention of overfitting in cryo-EM structure determination. Nat Methods 2012; 9:853–854.
Rosenthal PB, Henderson R . Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J Mol Biol 2003; 333:721–745.
Kucukelbir A, Sigworth FJ, Tagare HD . Quantifying the local resolution of cryo-EM density maps. Nat Methods 2014; 11:63–65.
Tang G, Peng L, Baldwin PR, et al. EMAN2: an extensible image processing suite for electron microscopy. J Struct Biol 2007; 157:38–46.
Russo CJ, Passmore LA . Robust evaluation of 3D electron cryomicroscopy data using tilt-pairs. J Struct Biol 2014; 187:112–118.
Wasilewski S, Rosenthal PB . Web server for tilt-pair validation of single particle maps from electron cryomicroscopy. J Struct Biol 2014; 186:122–131.
Pettersen EF, Goddard TD, Huang CC, et al. UCSF Chimera--a visualization system for exploratory research and analysis. J Comput Chem 2004; 25:1605–1612.
Phillips JC, Braun R, Wang W, et al. Scalable molecular dynamics with NAMD. J Comput Chem 2005; 26:1781–1802.
MacKerell AD, Bashford D, Bellott M, et al. All-atom empirical potential for molecular modeling and dynamics studies of proteins. J Phys Chem B 1998; 102:3586–3616.
Mackerell AD Jr, Feig M, Brooks CL 3rd . Extending the treatment of backbone energetics in protein force fields: limitations of gas-phase quantum mechanics in reproducing protein conformational distributions in molecular dynamics simulations. J Comput Chem 2004; 25:1400–1415.
Emsley P, Lohkamp B, Scott WG, Cowtan K . Features and development of Coot. Acta Crystallogr D Biol Crystallogr 2010; 66:486–501.
Murshudov GN, Skubak P, Lebedev AA, et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr D Biol Crystallogr 2011; 67:355–367.
Sehnal D, Svobodová Vařeková R, Berka K, et al. MOLE 2.0: advanced approach for analysis of biomacromolecular channels. J Cheminform 2013; 5:39.
Acknowledgements
We thank Dr Anthony Lai (Wales Heart Research Institute, Cardiff University School of Medicine, UK) for advice and suggestions during the course of this research and critically reading this manuscript. This research was supported by grants from the Strategic Priority Research Program of Chinese Academy of Sciences (XDB08030202 to FS), the National Basic Research Program (973 Program), the Ministry of Science & Technology of China (Programs 2012CB917200 to CCY and 2014CB910700 to FS), the National Natural Science Foundation of China (31270768 to CCY) and the Ministry of Education of China (111 Program China to CCY). All the cryo-EM work and all the intensive computations were performed at Centre for Biological Imaging, Institute of Biophysics, Chinese Academy of Sciences (CBI, http://cbi.ibp.ac.cn), with the help from Drs Xiaojun Huang and Lunjiang Lin.
Author information
Authors and Affiliations
Corresponding authors
Additional information
( Supplementary information is linked to the online version of the paper on the Cell Research website.)
Supplementary information
Supplementary information, Figure S1
The structure of Ca2+-activated open-state RyR1. (PDF 405 kb)
Supplementary information, Figure S2
RyR1 cryo-EM map resolution and quality assessment. (PDF 355 kb)
Supplementary information, Figure S3
Single particle cryo-EM analysis of RyR1. (PDF 393 kb)
Supplementary information, Figure S4
Cryo-EM map quality of different domains ofthe open-state RyR1. (PDF 455 kb)
Supplementary information, Figure S5
Cryo-EM map of important structural elements in the open-state. (PDF 604 kb)
Supplementary information, Figure S6
Occlusion of open RyR1 channel by ruthenium red. (PDF 217 kb)
Supplementary information, Figure S7
Structural comparison with the published Ca2+-modified structure (Efremov et al., 2015). (PDF 474 kb)
Supplementary information, Table S1
Summary of cryo-EM data collection (PDF 145 kb)
Supplementary information, Table S2
Statistics for the model refinement of the core region (PDF 143 kb)
Supplementary information, Table S3
Summary of functions of the major RyR1 domains/motifs upon Ca2+ activation (PDF 264 kb)
Supplementary information, Table S4
Angles between E and F helices in different states (PDF 137 kb)
Supplementary information, Table S5
Summary of mutations located in the domain/domain and helix/helix interfaces and related diseases of RyR1 (PDF 168 kb)
Supplementary information, Movie S1
Cryo-EM map quality of the open-state RyR1. This movie displays the cryo-EM density map of RyR1 with an atomic structure model fitted in the open-state and its slices in the horizontal and vertical orientations. (MOV 5154 kb)
Supplementary information, Movie S2
Structural transitions of RyR1 upon Ca2+ activation. This movie shows the conformational changes of the whole molecule of RyR1 and the core region (including EF-hand motif, U-motif, VSL, CTD, and the transmembrane region), as manifest in an atomic structure model, from the apo/closed-state to the Ca2+-activated, open-state, with highlights for the conformational changes of the EF-hand module/the central domain, the gate, and the selectivity filter. (MOV 9835 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 Unported License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Wei, R., Wang, X., Zhang, Y. et al. Structural insights into Ca2+-activated long-range allosteric channel gating of RyR1. Cell Res 26, 977–994 (2016). https://doi.org/10.1038/cr.2016.99
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2016.99
Keywords
This article is cited by
-
Molecular basis for gating of cardiac ryanodine receptor explains the mechanisms for gain- and loss-of function mutations
Nature Communications (2022)
-
Regulatory mechanisms of ryanodine receptor/Ca2+ release channel revealed by recent advancements in structural studies
Journal of Muscle Research and Cell Motility (2021)
-
Retrieving functional pathways of biomolecules from single-particle snapshots
Nature Communications (2020)
-
Intracellular calcium leak in heart failure and atrial fibrillation: a unifying mechanism and therapeutic target
Nature Reviews Cardiology (2020)
-
Smart polymer-based calcium-ion self-regulated nanochannels by mimicking the biological Ca2+-induced Ca2+ release process
NPG Asia Materials (2019)