Abstract
Progression through the cell cycle is largely dependent on waves of periodic gene expression, and the regulatory networks for these transcriptome dynamics have emerged as critical points of vulnerability in various aspects of tumor biology. Through RNA-sequencing of human cells during two continuous cell cycles (>2.3 billion paired reads), we identified over 1 000 mRNAs, non-coding RNAs and pseudogenes with periodic expression. Periodic transcripts are enriched in functions related to DNA metabolism, mitosis, and DNA damage response, indicating these genes likely represent putative cell cycle regulators. Using our set of periodic genes, we developed a new approach termed “mitotic trait” that can classify primary tumors and normal tissues by their transcriptome similarity to different cell cycle stages. By analyzing >4 000 tumor samples in The Cancer Genome Atlas (TCGA) and other expression data sets, we found that mitotic trait significantly correlates with genetic alterations, tumor subtype and, notably, patient survival. We further defined a core set of 67 genes with robust periodic expression in multiple cell types. Proteins encoded by these genes function as major hubs of protein-protein interaction and are mostly required for cell cycle progression. The core genes also have unique chromatin features including increased levels of CTCF/RAD21 binding and H3K36me3. Loss of these features in uterine and kidney cancers is associated with altered expression of the core 67 genes. Our study suggests new chromatin-associated mechanisms for periodic gene regulation and offers a predictor of cancer patient outcomes.
Similar content being viewed by others
Introduction
Eukaryotic cell division cycle requires periodic gene expression1,2. Previous studies in yeast, mouse and human cells have uncovered a periodic transcriptome comprised of classes of messenger RNAs (mRNAs) that oscillate during cell cycle and regulate cell cycle progression2,3,4,5,6,7. In addition to mRNAs, several RNAs of other types were also found to be periodic, including long non-coding RNAs (lncRNAs)8,9 and short non-coding RNAs8. However, the prevalence of these periodic non-coding RNAs remains mostly unknown, as previous studies surveyed cell cycle-dependent transcriptomes using microarrays that are unable to detect these non-canonical RNA species. Furthermore, cell type-specific periodic transcripts have been observed across different cell types and related species2,4, suggesting distinct regulatory mechanisms are in place to control periodicity in different cellular contexts. However, the commonly periodic (or cell type-independent) transcriptome is poorly defined, and the mechanisms that account for cell type differences remain unclear.
The misregulation of cell cycle is a hallmark of cancer10, however it remains unclear if specific patterns of periodic RNA expression across different tumor types bear any significance on patient outcomes or correlate with genetic alterations. In recent years, transcriptome profiling coupled with sophisticated statistical analysis has enabled molecular classification of tumors into subtypes that correlate with patient survival and/or drug responses11. Recently, The Cancer Genome Atlas (TCGA) has profiled nearly 5 000 samples from >20 tumor types for global alterations in DNA, RNA and protein12, making it possible to analyze differences and similarities across a wide spectrum of tumors. Curiously, such molecular classifications have revealed alterations in common biological pathways across multiple tumor types13. Unexpected regulators of both transcription and chromatin structure, including histone modifiers, ATP-dependent chromatin remodelers and regulators of three-dimensional chromatin structure, were found to be significantly mutated in a wide-variety of cancers14. Given that periodic gene expression drives cellular division and proliferation – major points of misregulation in almost all tumors – a thorough cell cycle-focused analysis of tumors should reveal additional similarities/differences across the pan-cancer landscape and potentially uncover specific vulnerabilities.
We report here deep sequencing of the human transcriptome through two continuous mitotic cycles followed by comprehensive analyses. We identified a large number of putative cell cycle regulators that have a unique gene structure and high capacity for protein-protein interactions (PPIs). By analyzing over 4 000 tumors, we provide a new molecular classification for tumors based on the cell cycle-dependent transcriptome. We demonstrate the significance of such classifications by showing that specific tumor classes differ in patient survival times irrespective of tissue origin. Analysis of transcription factors and histone modifications revealed novel regulatory mechanisms underlying periodic gene expression. Finally, we identify a core set of periodic genes that serve as protein interaction hubs for cell cycle control, and further reveal the functional consequence of misregulation of this core gene set across different tumor types.
Results
Transcriptome dynamics during cell cycle
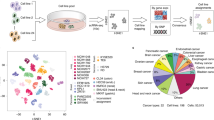
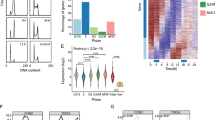
To obtain a high-resolution map of transcriptome dynamics during the mitotic cell cycle, we performed RNA-seq using synchronously dividing cells. Cells were synchronized by double thymidine block and collected every 1.5 h through two continuous divisions (∼32 h), in which the second cycle is considered to be chemically unperturbed. According to cellular DNA content measured at each time point, we selected 14 samples to cover matched stages between the two rounds of cell division (Supplementary information, Figure S1A). Cell synchrony was further confirmed by immunoblotting for eight well-known cell cycle markers (Figure 1A). From each sample we generated a cDNA library from poly-adenylated RNAs and performed RNA-seq with a depth of 100-250 million paired-end reads per sample (Figure 1B). In total, about 2.3 billion 100-nt reads were obtained (Supplementary information, Table S1) and mapped to the human genome (MapSplice, default parameters,15). We analyzed these data with Cufflinks to calculate transcript abundance for each gene16. Normalized FPKM values (fragments per kilobase of transcript per million mapped reads) were used to determine expression dynamics across the two cell cycles (see Materials and Methods section). We found that known periodic transcripts including CCNE, CCNB1, CCNB2, and CCNA2 exhibited the expected expression pattern (Figure 1C), providing a positive control and a starting point for the identification of additional genes with cyclical expression patterns.
Periodic mRNA expression during cell cycle. (A) HeLa cells were synchronized by double-thymidine-block, released from the block, and collected every 1.5 h for 36 h. 14 samples were selected so that the samples from the first cell cycle closely matched the samples of the second cycle. Cell synchronization was confirmed by quantification of DNA content (upper bar graph) and immunoblotting of cell cycle markers. (B) RNA-seq reads across 14 samples aligned by inferred cell cycle stages. (C) Relative expression of known cell cycle transcripts from RNA-seq data and heat map representation of ideal periodic seeds extracted from the known periodic transcripts. Periodic seeds are color-coded (left bar) and row-normalized. (D) Heat map representing 1 182 periodic transcripts. The color bar on the left represents seven periodic seed curves to which the identified periodic transcripts are matched (Euclidean distance ≤ 2.5), and the right colored bar represents the general peak expression stages. The periodic genes were further classified as G1-S if their temporal expression patterns are similar to the seed curves 1-2 or as G2-M if their temporal expression patterns are similar to the seed curves 3-7 in Supplementary information, Figure S1B. (E) Boxplot representation of gene length (top) and number of exons (bottom) for the periodic genes. (F) Periodic lncRNAs represented as a heat map (top panel for 37 lncRNAs with unknown function) or as normalized expression curves for two well-known lncRNAs (bottom panel, CCNB1 is shown for reference).
To identify periodically expressed transcripts we generated seven periodic seed curves with various peak expression timing modeled from the pattern of known cycling genes (Figure 1C; Supplementary information, Figure S1B). A simple distance metric was applied to interrogate all transcripts for their similarity to each periodic seed curve (see Materials and Methods section). Only transcripts with sufficient read coverage were considered for this analysis (see Materials and Methods section). In total, we identified 1 182 periodically expressed transcripts with peak expression times broadly distributed across all cell cycle phases (Figure 1D). Periodic genes were clustered according to two major peak expression phases (G1-S and G2-M). Consistent with prior reports3, the G2-M phase had a greater number of genes (304 in G1-S and 878 in G2-M). Interestingly, G2-M genes are significantly longer than those with expression peaks in the G1/S phase (1.8-fold, P = 7 × 10−11, Figure 1E). The increase in gene length tracked well with an increase in the total number of exons per transcript, with G2-M genes having about two more exons than the G1-S phase genes (Figure 1E). We also found that G2-M genes are significantly longer than all detected genes in this study (∼8 000 well-expressed genes), while the G1-S genes are generally shorter than all detected genes.
In addition to known protein coding genes, transcriptome profiling by RNA-seq enabled an unbiased detection of lncRNAs, which as a class, have emerged as key molecules that control cell proliferation and development9,17. To examine if the expression of lncRNAs is periodic, we mapped our reads to the annotated lncRNA database (http://www.broadinstitute.org/genome_bio/human_lincrnas/). Out of ∼400 lncRNAs with sufficient read coverage, 39 were identified as periodically expressed, including the known cell cycle-dependent lncRNA, MALAT118 (Figure 1F). The fact that we identified a previously reported periodic lncRNA confirms the sensitivity of our approach. This work represents the first comprehensive study of cell cycle-dependent expression of lncRNAs by sequencing and strongly suggests that these molecules may also play important roles in controlling cell cycle. Further analysis of the initial set of 1 182 genes also revealed that ∼200 of the genes identified in this study do not code for protein (i.e., representing ribosomal RNAs and pseudogenes). Taken together, we identified a large set of periodically expressed coding and non-coding transcripts, many of which have never been implicated in cell division or have unknown functions.
Functional classification of periodic genes
We next investigated the functional gene classes found within the periodic transcriptome. As expected, genes in both groups were enriched for cell cycle-related functional processes (Supplementary information, Figure S2A and Table S1). Specifically, genes in G1-S group were enriched for gene ontology (GO) terms related to “DNA metabolic process” (P = 6 × 10−13) and “DNA replication” (P = 6 × 10−14) while the G2-M group was highly enriched for the GO terms of “M phase” (P = 4 × 10−34) and “organelle fission” (P = 2 × 10−31). Further analysis of all of the identified GO terms revealed limited overlap in specific cell cycle functions between G2-M and G1-S genes. The shared category included the “DNA damage response” and “cellular response to stress” (P < 10−3 in both G1-S and G2-M genes), thus alluding to the requirement of cells to maintain genome fidelity during replication and mitosis.
Using PPIs as a measure of relatedness, we uncovered highly connected interaction maps in these periodic genes (see Materials and Methods section). The G1-S and G2-M genes were found in different interaction networks, indicating that these sets are functionally distinct with little co-expression overlap in dividing cells (Supplementary information, Figure S2B). Clustering of these interaction maps revealed highly connected subclasses with related functions (Supplementary information, Figure S2B). We next asked whether proteins corresponding to periodic genes have more interaction partners than a background set of proteins expressed at similar levels in our data set. Indeed, we found that periodic genes tend to have more interacting partners with the G2-M genes having the highest number of PPIs/gene (P = 2.2 × 10−16, Supplementary information, Figure S2C).
Systematic analysis of periodic gene promoters
Previous studies have indicated that transcriptional regulation is a highly conserved mechanism that controls periodic gene expression and cell division1,2. Classical regulators include the transcription factors E2F1 and FOXM1 that are required for the temporal control of gene expression in G1-S and G2-M stages of cell cycle, respectively4,19,20,21. Given that many of the 1 182 genes identified in this study have not been previously observed as being periodic, we sought to discover novel regulatory mechanisms underlying periodic gene expression. We first searched for motifs enriched at the promoters of periodic genes (see Materials and Methods section). We found that the promoters of the G1-S group harbored binding sites for cell cycle-related transcription factors such as E2F1, E2F4 and E2F6 motifs (Supplementary information, Figure S3A), while the top motifs for G2-M genes include binding sites for SP1, NFY, FOXO4, and MYC as well as unknown motifs (e.g., AACTT) that resemble known FOXM1-binding site22,23. Additional motif analysis revealed a modest overlap between the G1-S and G2-M groups, indicating differences in regulatory mechanisms controlling these groups (Supplementary information, Figure S3A). The list of novel motifs identified with possible cognate binding proteins is shown in Supplementary information, Table S2.
While motif enrichment analysis suggests possible interactions between transcription factors and DNA sequence, evidence of direct protein-DNA interactions may strengthen these findings. To this end, we analyzed 75 ChIP-seq (chromatin immunoprecipitaion followed by high-throughput sequencing) data sets performed in HeLa cells (ENCODE). We included transcriptional repressors, activators, and factors known to control cell cycle-dependent gene expression (see Supplementary information, Table S2). We examined ChIP signals in a 2kb region centered around the transcriptional start site (TSS), and compared promoters from G1-S or G2-M genes with a set of control genes with similar length and RNA abundance (Supplementary information, Figure S3B and S3C). Supporting the results of motif-based analysis, E2F1, E2F4 and E2F6 were significantly enriched in the G1-S genes, whereas FOXM1 and NFYB were enriched at the promoters of G2-M genes (Supplementary information, Figure S3A). We summarize the ChIP signals along with pairwise comparisons between the different expression phases and the associated P-values in Supplementary information, Table S2. This comprehensive analysis has uncovered both well-known and putative cell cycle transcriptional regulators of the newly identified periodic genes, including an emerging regulator of chromatin structure, CTCF, which we study further in mechanistic detail.
Analysis of regulators of chromatin structure on periodic genes
We identified enrichment of CTCF ChIP signals at the TSS of most genes, which has been observed as a general feature of many promoters24. CTCF was originally discovered as a transcriptional regulator but has also been shown to influence chromatin organization25,26. Intriguingly, we also identified CTCF enrichment at the transcription end site (TES) of the G1-S genes (but not the G2-M or control genes), and these G1-S gene regions were co-occupied by RAD21, a component of the cohesin complex that has been functionally associated with CTCF27 (Figure 2A). Recent evidence has also demonstrated that the transcriptional activator ZNF143 co-localizes with CTCF24,28. Indeed we observed ZNF143 co-localization with CTCF at G1-S genes with a modest, but significant, enrichment at the TES. Such chromatin architectural features are consistent with findings from Handoko et al.29, where enrichment of CTCF was shown to coalesce actively transcribed regions together into a chromatin loop. Our metagene analysis also shows that, in contrast to earlier works, CTCF was only present at the TES of a specific group of genes (Figure 2A and 2B and Supplementary information, Figure S3). Consistently, we also found a strong positive correlation between the presence of RAD21, CTCF and ZNF143 at the TES, indicating that these factors occupy the same regions (Figure 2C). These three proteins were again present at higher levels around the TES of G1-S genes compared with G2-M and background (Figure 2D). We also examined the TES of G1-S genes for enrichment of H3K4me1, a chromatin mark typically associated with enhancers30, and did not observe significant enrichment of H3K4me1 at the TES (data not shown). Based on these data we speculate that a unique distribution of chromatin architectural proteins CTCF, RAD21 and ZNF143 at the G1-S genes may enable gene loops that provide a potential mechanism for the rapid induction of these genes during cell cycle progression, a mechanism similar to the “transcriptional memory” or “bookmarking” for genes that undergo rapid cycles of activation and repression31. However, our results do not necessarily prove the existence of chromatin loops around G1-S genes, and further experiments are needed to reveal the detailed mechanism.
Novel chromatin features of periodic genes. (A) Metagene analysis of periodic genes. CTCF, RAD21, and ZNF143 ChIP-Seq signal was plotted across background (purple), G1-S (green), and G2-M (orange). ZNF143 inset represents an enlarged view of the transcription end site (TES). Read counts are normalized to per million mapped reads. (B) UCSC genome browser view of VPS18, a G1-S gene. CTCF, RAD21 and ZNF143 ChIP-seq data from ENCODE is shown at the VPS18 locus. (C) Comparison of CTCF, RAD21, and ZNF143 at TSS and TES of the G1-S genes. Respective ChIP marks were summed at either the TSS (black) or TES (green) and plotted against each other. Spearman's rank correlation is shown, and the significance of correlation coefficient (P-value) is calculated by t-test using non-correlation as null hypothesis. TSS was defined as the TSS and 1 kb upstream. TES was defined as the TES and 1 kb downstream. (D) Boxplot of TES ChIP signals. CTCF, RAD21, and ZNF143 ChIP signals were summed from TES to 1kb upstream and downstream in background (blue), G1-S (yellow), and G2-M (orange). (E) CTCF mutation frequency in and number of altered cases in tumors with gene expression data. UCEC: Uterine corpus endometrioid carcinoma; COAD: Colorectal adenocarcinoma; HNSC: Head and neck squamous cell carcinoma; BLCA: Bladder urothelial carcinoma; STAD: Stomach adenocarcinoma; MED: Medulloblastoma; BRCA: Breast invasive carcinoma. (F) Scatterplot representation of periodic gene expression levels in uterine corpus endometrioid carcinoma (UCEC) tumors with CTCF mutations. Data are drawn as a function of P-value. About 20% of periodic genes (249 genes) changed significantly in tumors (shown with a darker shade of red). (G) Boxplot representation of two G1-S genes, E2F1 and CCNE1, whose expression is lower in UCEC.
If CTCF is indeed critical for the activation of the G1-S expression wave, we would expect that this gene set would be deregulated in cells lacking CTCF. This hypothesis can be tested on a genomic scale using cancer genotyping data of uterine tumors (http://cbioportal.org), of which nearly 20% had mutations in CTCF (Figure 2E). We analyzed gene expression data from 240 primary uterine tumor samples, among which 44 samples harbor mutations in CTCF32. We found that nearly ∼20% of periodic genes were downregulated in tumors with mutant CTCF compared with tumors with wild type CTCF (Figure 2F and 2G), supporting the role of CTCF in mediating periodic gene expression. Altogether our analysis of the periodic transcriptome and the underlying transcription regulatory framework for the periodic genes reveals DNA cis-elements, transcriptional regulators and chromatin-associated proteins that may contribute to robust periodicity.
Mitotic trait classification of primary breast tissues
The features of the periodic transcriptome described thus far support the importance of cell cycle control in cancer. Since we were able to identify a transcriptional phenotype associated with CTCF loss in uterine cancer, we asked whether human tumors could be meaningfully classified based on their expression of periodic genes. We hypothesized that this periodic signature would be associated with the biology or clinical outcomes of human cancers. To test the feasibility of such analysis, we compared the expression of periodic genes between every pair of the 14 cell cycle samples (as shown in Figure 1) using Spearman's rank correlation, and found a high degree of positive correlation between samples in the same stages and robust negative correlation between different stages (Supplementary information, Figure S4A). These cell cycle stage-dependent correlations were only observed with the periodic transcripts, as a control set of 1 000 arbitrarily selected transcripts or all detected transcripts showed no significant correlation between samples representing the same cell cycle stage (Supplementary information, Figure S4A). This new analysis, referred to as “mitotic trait”, suggests that any gene expression data set can be compared with each of the 14 cell cycle samples as a measure of similarity to each stage. We tested mitotic trait analysis against two previous microarray-based transcriptome profiling data sets, and found a strong correlation of same cell cycle stages regardless of cell type used, confirming the reliability of this method (Supplementary information, Figure S4B). This approach is similar to gene expression signatures that have been identified to classify tumor types11,13. In this case, however, we used a newly generated cell cycle-dependent gene expression signature that measures the relative similarity of an expression data set to any cell cycle stage.
We then applied this approach to analyze data generated by TCGA12. We initially focused on breast cancers and their normal controls, since these tumors are known to be heterogeneous with well-annotated information on molecular subtypes and common mutations33. Mitotic trait analysis was carried out to compute the similarity of breast samples to the four major cell cycle stages, G1, S, G2 and M (Figure 3A). Strikingly, most normal breast tissues were very similar to G1 phase while tumor samples were heterogeneous in mitotic trait, consistent with a high degree of tumor-to-tumor variability. To further study this variability, we grouped breast samples according to molecular subtypes as defined by PAM50 analysis33 and found that different tumor subtypes correlated well with specific mitotic trait (Figure 3A). The Luminal A subtype was found to be more G1-like, which is consistent with the fact that most of these tumors have wild type TP5333, a key factor that drives cells into senescence and a G1/G0 state34. On the other hand, loss of TP53 in other subtypes correlates with an S-, G2-, or M- like mitotic trait. These results indicate that the classification based on mitotic trait can reliably cluster most, if not all, tumors and that this clustering correlates well specific genomic alterations. Interestingly, the published molecular subtype according to PAM50 has classified a small number of tumors as “normal-like”33. In our mitotic trait analysis, these “normal-like” tumors do not resemble the G1 stage like normal tissues (Figure 3A, shown by asterisk), suggesting that mitotic trait-based classification captures subtle differences in gene expression between normal and tumor-like states. Using the available TCGA BRCA survival data we detected a significant difference between G1 and non-G1 like tumors.
Classification of human tumors by mitotic trait analysis. (A) Mitotic trait analysis of human breast tumors using the RNA-seq data set from TCGA. Heat map representation of the similarity between normal (n = 79) and tumor (n = 739) breast and four representative cell cycle stages. Significant similarity correlations are indicated by a black line on the top (see Materials and Methods section). Breast tumors were grouped together according to PAM50 molecular subtype classification59. Common genetic mutations in each tumor sample are indicated. (B) Survival analysis of breast tumors the classified by mitotic trait. (C) Heat map representation of mitotic trait analysis of a distinct set of 1 809 human breast tumors clustered into two major classes, G1-like and non-G1 (left panel). This is from an old set of microarray analysis with matching survival results. Kaplan–Meier survival curves for the two major classes of breast tumors is shown (right panel). (D) Heat map representation of mitotic trait analysis of pan-cancer human tumors hierarchically clustered into two major classes (denoted by black or red bar). Tissue type is represented by color bar at bottom. BLCA, bladder urothelial carcinoma; BRCA, breast invasive carcinoma; COAD, colorectal adenocarcinoma; GBM, glioblastoma multiformae; HNSC, head and neck squamous cell carcinoma; KIRC, kidney renal clear cell carcinoma; LAML, acute myeloid leukemia; LUAD, lung adenocarcinoma; LUSC, lung squamous cell carcinoma; OV, ovarian serous cystadenocarcinoma; READ, rectum adenocarcinoma; UCEC, uterine corpus endometrioid carcinoma. G1-like and non-G1 (left panel). Kaplan–Meier survival curves for the three major classes of tumors are shown as G1, non-G1 and S + M types (right panel).
For a more rigorous assessment of our classifications and survival correlations we tested whether association with mitotic trait classes predicted variation in patient survival. We evaluated microarray-based expression data from 1 809 breast tumors with more extensive clinical survival data35. We first hierarchically classified tumors based on mitotic trait and into two major groups, which we term the G1-like and the non-G1, and plotted relapse-free survival (RFS) for each group (Figure 3C). Again, a striking survival difference was observed; non-G1 tumors showed significantly shorter median survival times compared with the G1-like tumors (Figure 3C). This analysis was performed independent of grade or PAM50 classification suggesting that classification according to mitotic trait may be capable of independently predicting tumor prognosis.
Pan-Cancer classification of tumors by mitotic trait
Our ability to classify breast tumors based on mitotic trait prompted us to determine if tumors arising from different tissue types are amenable to this analysis and whether these classifications would offer predictions of patient outcome. Expression data from 12 tumor types and matched normal tissues were subjected to mitotic trait analysis. Based on a similarity analysis of mitotic trait values, we found that all, but one, normal tissue types strongly resembled G1 phase (Supplementary information, Figure S4C). The outlier was normal head and neck tissue, which resembled the M-phase. In addition, we observed diversity in mitotic trait classification within most tumor types although some were relatively homogenous. For example, breast tumors (as shown above), lung adenocarcinomas, and uterine corpus endometrioid carcinoma were very heterogeneous, while kidney renal cell carcinoma was relatively homogenous across patients and most closely resembled G1. Such diversity may reflect meaningful tumor subtypes that are grouped together by histological or anatomic criteria (e.g., breast cancers).
Hierarchical clustering of the 12 tumor types revealed two major clusters of tumors: the G1-like and non-G1-like and a third cluster that was most similar to S and M phases (Figure 3D). Representative tumors from each of the tumor types were found in both clusters with the exception of clear cell renal cell carcinoma (ccRCC) that was predominantly G1-like, indicating that the two major clusters represent a mixture of distinct tissue types. Recent efforts to classify tumors by molecular characteristics (e.g., mutation types or RNA expression levels) rather than by tissue of origin have seen some success in identifying subclasses13 which may have implications for patient prognosis and drug responses. We asked whether survival differences could be observed in the two major classes uncovered by our mitotic trait analysis. Using the available survival data from 3 223 tumors across 12 types, we found that G1-like tumors showed a significantly longer median overall survival than the non-G1 cluster (Figure 3D). Interestingly, the third cluster had much lower survival times than both G1 and non-G1. This difference was observed in spite of the dramatic known differences in median survival for different tumor types (i.e., GBM vs. KIRC)36,37. Together these data indicate that mitotic trait classifications within single tumor types (Figure 3A and 3B) or across a diverse set of tumors (Figure 3D) correlate with patient outcome and mutation status, again demarcating the importance of cell cycle regulation in tumor biology.
Identification of highly functional core periodic genes
To address cell type-dependencies of periodic gene expression, we compared our data set with data from four other periodic gene expression studies in distinct cell lines, including foreskin fibroblasts, keratinocytes, osteosarcoma cells and HeLa cells. We noticed that the number of periodic genes shared by any two studies was modest, in some cases less than 30% (Supplementary information, Figure S5A-S5D). Possible explanations include differences in the method used to achieve cell synchrony, the type of analyses to identify periodic genes, the total numbers of genes initially detected, the platforms for expression analysis, as well as the inherent difference between cell types. Genes identified by other groups as periodic, but not in our analysis, exhibited some periodicity, albeit the periodicity failed to meet our threshold (Supplementary information, Figure S5A-S5D). The likelihood of a gene being identified across the different studies was most closely related to how robust the periodicity was for that specific gene (Supplementary information, Figure S5E), indicating conserved regulatory mechanisms for the highly periodic genes across cell- and tissue- types. Furthermore, by analyzing published results from genome-wide RNAi screens38,39,40, we found that a significant fraction (20-30%, P = 0.0003 by Fisher exact test) of periodic genes from most studies exhibit cell cycle-related defects after RNAi-mediated depletion (Supplementary information, Figure S5F), with the highly periodic genes being the most likely to display phenotypes upon depletion (Supplementary information, Figure S5G). These data indicate that there might be a “core” set of strongly periodic, cell type-independent genes that are functionally driving cell cycle processes.
We identified a core set of 67 genes that demonstrated periodicity across five different studies. We term these genes as “the core 67” (Figure 4A). The core 67 are significantly more periodic (i.e., lower periodic score, representing shorter distance from the seed curves) as compared with the total 1 182 periodic genes initially identified (P = 1.9 × 10−11 Supplementary information, Figure S5H). Interestingly, the average gene length of the core 67 is significantly shorter than the set of the 1 182 periodic genes or a control set of all genes, although the number of exons is slightly higher in the core 67 set (Figure 4B). Assuming constant transcription rate, this result suggests that the core 67 genes may be transcribed more rapidly, offering an explanation for their sharp increases in expression during cell cycle. About half of the core 67 genes lead to cell cycle defects when depleted by RNAi40, considerably more than what was observed for any of the periodic gene sets from five independent studies (Figure 4A right panel and Figure 4C). In addition, genes from the core 67 have more PPI partners than other periodic genes, forming a well-connected PPI network with two major clusters representing the G1-S and the G2-M (Figure 4D and 4E). The increased interactions seen for the core 67 (three fold greater PPI partners per gene than the total 1 182 periodic genes) is striking, given that these periodic genes already have a significantly higher number of PPIs than all detected genes (Figure 4D). These data suggest that the core 67 may function as periodic regulatory hubs that control cell cycle progression.
Features of the core cell cycle gene expression program. (A) Heat map showing the expression pattern of the core 67 during cell cycle (left) and cognate mitocheck.org phenotypes40 (right). (B) Average gene length (left) and number of exons (right) of a control set, all 1 182 periodic genes and the core 67 periodic genes. (C) Fraction of all periodic genes or the core 67 genes with known cell cycle defects from three independent genome-wide RNAi studies. (D) Boxplot showing the average number of interaction partners for the core 67 periodic genes. (E) Protein-protein interactions identified within the core 67 genes grouped into the G1-S and G2-M phases. Only interactions supported by experimental evidence were used for this analysis.
Given the unique features of the core 67, we asked whether these genes were misregulated in human cancers. To this end, we used microarray data from human breast tumors that have well-annotated clinical information. We found that 86% (58 out of the 67) of genes in the core 67 set were associated with shorter survival when expressed at high levels in breast tumors (Figure 5A-5C). Similar results were observed in ovarian and lung cancers (not shown). Given the possible number of total genes associated with the reduced survival, this association is significantly more than expected by chance (P < 4 × 10−43, hypergeometric test). Together these data indicate that the core 67 genes exhibit robust regulatory capacity across multiple cell lines and are misregulated in tumor samples.
High expression of core periodic genes correlates with poor survival. (A, B) Two representative Kaplan–Meier plots shown for two of the core 67 core genes. (C) Bar graph of the log-rank P-values determined for survival differences based on high vs. low expression of each of the 67 core periodic genes. As shown most core periodic genes (58/67) lead to decreased survival when over-expressed.
The core periodic genes have unique chromatin features that are misregulated in cancer
To identify chromatin features common to the core 67 genes, we analyzed the available ChIP-seq data sets for transcription factors, variant histones and histone modifications, and chromatin accessibility (ENCODE data sets as in Figure 2 and Supplementary information, Figure S3). We found a significant enrichment of histone modifications and transcription factors associated with active transcription around the TSS and the gene body of the core 67. For example, E2F1, E2F4, RNAPII, H3K4me3, H3K79me2, CTCF, RAD21 and ZNF143 were enriched at or around the TSS of these genes (Supplementary information, Figure S6). As expected, we also found a reduction in H3K27me3 and H3K9me3, modifications associated with repressed genes, at these loci (Supplementary information, Figure S6). Consistently, the transcription-associated histone modifications, H3K36me3 and H3K79me2, were preferentially enriched over the gene body (Figure 6A and 6B). Interestingly, we noticed that the absolute enrichment of H3K36me3 is higher in the core 67 as compared with either control, G1-S or G2-M genes. However, the increased signal was not associated with an increase in RNA levels (Figure 6A inset). This was unexpected since H3K36me3 levels generally correlate with RNA abundance.
SETD2 loss alters expression of the core cell cycle gene expression program. (A, B) Metagene plot of H3K36me3 and H3K79me2 ChIP-seq signals in a control set of genes, periodic genes and the core 67. Inset, average expression, FPKM, level of 67 genes compared with the background set of genes. (C) Scatterplot representation of the periodic gene expression levels in tumors with SETD2 mutations. Data are drawn as a function of P-value. (D) Boxplot representation of selected mis-regulated genes in tumors with SETD2 mutations. (E, F) Survival analysis of all kidney tumors based on the expression of CDCA3 and CDC20, members of the core 67 genes with mis-regulated expression in SETD2 mutant tumors.
Tri-methylation of H3K36 in human cells is catalyzed by the enzyme SETD2, which has emerged as a critical player in renal cancers37,41,42. We analyzed TCGA-derived RNA-seq data and found that in tumors with mutant SETD2, the expression levels of a subset of the core 67 genes were significantly higher than tumors with WT SETD2 (Figure 6C, two representative examples are shown in Figure 6D). This result is consistent with the observation that loss of SETD2 leads to aberrant transcription initiation43. To correlate increased expression of these genes with patient outcome, we associated CDC20 and CDCA3 expression levels with survival data and found that patients with higher CDC20 or CDCA3 expression had shorter event-free survival times (Figure 6E and 6F). In support of this observation, CDCA3 over-expression has been associated with poor prognosis in oral cancers by preventing G1 phase arrest44. Thus, the identification of a core gene set with periodic regulation across various tissue-types in multiple studies revealed a common epigenetic state in these genes, and uncovered an unexpected association of their expression levels with SETD2 mutation status in kidney tumors that was not reported in previous TCGA analysis.
Discussion
Through massive RNA sequencing of synchronous cells, we generated the first comprehensive transcriptome map of the cell cycle containing both coding and non-coding genes. We identified 1 182 periodic transcripts, most of which had not been observed in previous studies. Periodic genes encode proteins with higher capacity for PPIs and are unique in gene structure. An intriguing finding was the difference in gene length between G1-S and G2-M genes with the G1-S genes being significantly shorter. This difference may reflect the possibility that gene length may have evolved to accommodate the kinetic requirements for expression (i.e., rapid oscillations) during certain cell cycle stages. While oscillations in gene function may ultimately depend on the kinetics of protein synthesis, the unique structure of periodic genes likely contributes to temporal regulation of protein production. The high resolution mapping of transcriptome during cell cycle may also reveal additional mechanisms for temporal control of gene function at RNA level. In support of this notion, we discovered that more than a thousand of genes undergo periodic splicing during cell cycle, resulting in oscillations of splicing variants that play critical roles in cell cycle progression45.
We also uncovered 39 periodically expressed lncRNAs, including MALAT1 which was recently shown to oscillate during cell cycle in lung fibroblasts18. Interestingly, loss of MALAT1 leads to the activation of p53 and repression of the S-phase specific transcription factor MYBL2, providing a possible mechanism for cell cycle control. In addition, NEAT1, a structural component of para-speckles, also exhibited periodic expression, suggesting it may also have functions in cell cycle control. Non-coding RNAs arising from the promoter regions of 56 periodic genes were recently examined by custom microarrays8, which led to the identification of many putative non-coding RNAs expressed in a periodic fashion. However, we did not detect these non-coding RNAs probably due to their low expression levels. These findings lead us to speculate that lncRNAs play key roles in cell cycle control and like periodic mRNAs likely contribute to tumor progression. In addition to periodic changes in mRNA levels, we speculate that other gene-regulatory mechanisms including alternative splicing and alternative poly-adenylation will be periodic. Regulation of more than one of these gene regulatory steps could drastically influence protein levels during cell cycle.
To further understand the regulation of periodic genes we analyzed transcription factor binding data. As expected, E2F transcription factors were highly enriched at the promoters of the G1-S genes, while FOXM1 was enriched at the promoters of G2-M genes. STAT3, which is known to drive cell cycle in the context of immune cell activation or specific growth factor induction, was also observed at the G2-M genes and may help to drive the expression of these mRNAs46. Supporting this hypothesis is the data indicating that RNAi of STAT3 inhibits proliferation of certain tumor cells46. In addition to transcription factors, we also found CTCF and RAD21 at the transcription start and end sites of some periodic genes, as well as the enrichment of certain histone modifications at or near TSS. The insulator binding protein CTCF has been known to position nucleosomes containing histone H2A.Z47, a histone variant that is enriched around TSS during G1 phase of the cell cycle but is reduced at M phase48. Our results suggest that binding of CTCF happens at both TSS and transcription end site, and thus we speculate that CTCF enhances the periodicity of transcripts by altering chromosome architecture through DNA looping, a mechanism previously observed in various cell types29,49. Consistent with the novel role of CTCF in periodic gene expression, we were able to detect a significant decrease in the expression of 150 periodic genes in uterine corpus endometrioid carcinoma with CTCF mutations.
The high-throughput transcriptome map of cell cycle allowed us to develop a new bioinformatic analysis, “mitotic trait”, to quantify the association of any gene expression data set with specific cell cycle stages. Analysis of breast cancer data revealed that normal breast tissue resembles the G1 stage while breast tumors are varied. Our result reiterates the transcriptome variability among breast cancers and suggests, as expected, that cancers often have deregulated cell cycle. While Whitfield et al. reported misregulation of periodic genes in breast cancer, they proposed that rapidly dividing cells may have increased levels of certain periodic genes3. An alternative possibility, and one supported by our data, is that the transcriptome of a tumor could resemble a specific cell cycle stage. Histopathological studies have shown that increases in the number of mitotic figures or Ki67 staining within tumor samples correlate with a decrease in patient survival50,51. Here we found that the mitotic trait, which is based on gene expression at global scale, provides a quantitative classifier of tumors with different survival times. Tumors most similar to M-phase have the shortest survival, however it remains unclear if M-class tumors indeed have a higher number of cells in mitosis, as this information is not available. It is worth noting that mitotic figures estimation can be variable across studies52, making it difficult to interpret survival data. However, the RNA-seq analysis used in this study has been standardized and employed across thousands of samples. Mitotic trait classes were seen not only in breast, but also across 12 different tumor types. All except one normal tissue showed high similarity to G1 phase, while classification of 12 tumors revealed that some tumor types are heterogeneous in mitotic trait (e.g., breast) whereas others are very homogenous (e.g., kidney). Importantly, we performed survival analysis on a mixed population of tumors from different tissues separated only by mitotic trait and still observed differences in patient survival (Figure 3D). To date, most pan-cancer classifications have stratified tumors into the already known tissue types53, whereas mitotic trait can extract prognostic information in a tissues-independent fashion, providing a valuable new criterion for tumor classification. Because “mitotic trait” uses independent cell cycle genes that are expressed in most dividing cells, this predictive signature may suit most tumor types. Although a reliable validation of new cancer classifier requires clinical data that are independently collected without prior biases, combining “mitotic trait' with existing classifiers will likely help improve cancer classifications.
While nearly 4 000 genes have been labeled periodic in human cells by various groups, most of them are specific to certain cell types or experimental conditions. Our analysis reveals a core set of 67 genes across all studies using multiple cell types. The genes in this set have more PPIs and are shorter in length, are more likely to produce periodic proteins, and have severe cell cycle phenotypes when depleted. Further investigation of the epigenetic landscape revealed that H3K36me3 is enriched in the gene body of the core 67 genes. In addition, these genes were upregulated in kidney tumors that lack SETD2, the enzyme responsible for depositing this histone mark54. SETD2 has been shown to be mutated in KIRC, and recent reports suggest that kidney tumors with SETD2 mutations have poor prognosis42. What is less clear is the effect of SETD2 mutations (i.e., loss of H3K36me3) on cell proliferation and tumors. Here we show that H3K36me3 is highly enriched on a set of genes that promote cell proliferation (Figure 6). Consistently, these genes also correlated with patient outcomes in tumors across multiple tissues (e.g., 58/67 associated with shortened survival when up-regulated in breast tumors), reiterating their importance irrespective of cellular context. In addition, loss of SETD2 and H3K36me3 results in increased expression of these genes, suggesting that SETD2 mutations can affect tumor growth through mediating cell cycle regulation. Given the reported role of SETD2 in regulating splicing, we speculate that these transcripts are also mis-regulated at other gene regulatory steps. In sum this high-resolution map of RNA dynamics in cell cycle has uncovered novel features and potential regulatory mechanisms for periodic gene expression and elucidated critical aspects of tumor biology that correlate with patient survival.
Materials and Methods
Cell culture, synchronizations
HeLa cells were cultured in DMEM (Gibco) medium supplemented with 10% FBS (Gibco). Cell cycle synchronization was adapted from the protocol of Whitfield et al.3; ∼750 000 log phase HeLa cells were plated in 15 cm dishes in complete media and allowed to attach for 16 h, reaching < 30% confluence. Cells were subsequently treated with 2 mM thymidine (Sigma) for a total of 18 h, washed 2 times with 1× PBS, and supplemented with fresh complete media for 10 h. 2 mM thymidine was subsequently added for a second block of 18 h and washed as described previously.
RNA library preparations and sequencing
RNA from synchronized cells was extracted with TRIZOL (Invitrogen), treated with DNAse I (Qiagen), and purified on RNAeasy columns (Qiagen) according to the manufacturer's protocol. RNA-seq libraries were robotically prepared with Illumina TruSeq Total RNA Sample Prep kits according to the manufacturer's protocol and sequenced on one full flow cell of HiSeq 2000 (Illumina), using the pair-end protocol with 100 bp read length to obtain eight technical sequencing replicates per sample. The RNA-seq data reported in this paper are publically available through the NODE (National Omics Data Encyclopedia, http://www.biosino.org/node/) library with the access ID: ND00000019EP.
Correlation analysis for inference of cell cycle stage (Mitotic Trait)
All pan-cancer data were downloaded from the TCGA Network (https://tcga-data.nci.nih.gov/tcga/tcgaDownload.jsp). All tumor samples that satisfy TCGA quality control were used in analyses without further filtering. Microarray data were obtained from K-M plot.org35,55. The expression values for the 1 182 periodic genes were extracted from each data set to be analyzed. Before correlation analysis, gene expression values (RSEM values for TCGA data and Affymetrix gene expression values (gene-centric) for microarray) were normalized across all samples within the group. Normalization was performed in the same way as was done for the 14 cell cycle samples (Sample-min(Sample)/max(Sample)). Therefore, subsequent correlations reflect the relative similarity as compared with the rest of the samples within the normalization group. A pairwise sample to sample Spearman's Rank comparison was performed across all samples including the cell cycle stages. For simplicity, four samples representing G1 (sample 7), S (sample 9), G2 (sample 11) and M (sample 5) were shown in the analyses of cancer data. To assess the significance of the correlation, we carried out a permutation analysis by randomly shuffling the gene expression values for each tumor and calculating the correlation to each cell cycle stage. Permutations were carried out 1 000 times and the frequency in which a permutation had a better correlation score was calculated. We consider an FDR of 0.1 to meet the significance threshold. Tumors were then subjected to hierarchal clustering based on their similarity profile to each cell cycle stages and the major resulting clusters were subsequently separated for survival analysis.
Survival analysis
Breast cancer survival data were collected from K-Mplot.org35,55. Pan-cancer survival data were obtained from the Cancer Genome Database (https://tcga-data.nci.nih.gov/tcga/tcgaDownload.jsp)56. RFS was used for Kaplan–Meier plots for the breast cancer analysis. Time to death for Kaplan–Meier plots for the pan-cancer data set. All plots were drawn using the R version 3.1.1 survival function. For KIRC survival, K-M plots were generated and downloaded from cBioPortal.org. For breast cancer survival association with the core 67 genes (Figure 5), K-M were generated downloaded from K-M plotter.org35. To test the significance that 58/67 core genes correlate with patient survival. An assumption that 1 000 out of 8 835 (number of genes detected by this study) genes correlate with poor patient survival was made. A hypergeometric test was then performed to calculate the chance that if 67 genes are chosen at random from 8 835, 58 will correlate with poor patient survival (P = 4 × 10−43). In order for this find to not be statistically significant, nearly 7 000 genes would have to correlate with poor patient survival. Survival analysis of core 67 genes in kidney cancer was determined by cbioportals.org survival plot analysis of altered genes. Altered gene expression with default parameters set to expression being 2 standard deviations away from the mean was used.
Flow cytometry and cell cycle analysis
Cells were harvested with trypsin treatment, washed 2 times in cold 1xPBS and subsequently fixed in 80% ice-cold ethanol for at least 4 h. Cells were then washed twice with 1xPBS and suspended in propidium iodide/RNase staining buffer (BD pharmingen, cat # 550825). Cells were analyzed by flow cytometry to count 10 000 cells that satisfied gating criteria. Data collected were analyzed using ModFit software to discern 2N (G1), S-phase, and 4N (G2 and M) composition.
Mapping and filtering of RNA-seq data
The RNA-seq reads were mapped to the human genome (build hg19) using MapSplice informatics tool with default parameters15 and transcript abundance or FPKM was assessed by Cufflinks16. Eight technical replicates were sequenced and mapped independently for cell cycle data, and we required that the standard deviation for eight independently assigned FPKM values to be no greater than 0.5 fold of average FPKM (i.e., genes with larger variations between technical replicates were discarded). For each gene, at least 12 out of 14 cell cycle samples had to meet the FPKM variation criteria in order to be further analyzed. In the case where one or two samples did not satisfy these criteria, the final FPKM value for those outlier samples was adjusted to be the average of the sample before and the sample after (i.e., if gene expression of a gene in sample 5 did not meet these criteria then its gene expression value was adjusted to the mean of the gene expression of the same gene in sample 4 and 6). Finally, any gene with the average FPKM less than or equal to 0.5 across 14 samples was not further analyzed. In total 8 835 genes met our filtering criteria (i.e., reliably detected in HeLa cells with enough abundance for further statistic tests).
Identification of periodic RNAs
For each expressed gene, we normalized its gene expression as follows:

where i = 1 to 8 835 for all expressed genes; n = 1 to 14 for the14 samples; Emin is the minimum and Emax is the maximum gene expression value among the 14 samples.
To identify periodic genes, normalized gene expression values (i.e., normalized (en)) for the known periodic genes, CCNB1, CCNA2, CCNB2, and CENPE, were used as a starting point to subsequently add curves with broader or sharper peaks as well as shifted to left or right, resulting in 7 periodic expression curves that cover all the phases of cell cycle (Supplementary information, Figure S1B). These “ideal seed curves”, which capture intermittent peak times and phase shifts that were not well represented within the initial known periodic genes. To identify genes that have similar expression during the cell cycle time-series, we calculated the Euclidean Distance ED from each model seed curve to all detected genes as follows:

where m = 1 to 7 for all model seed curves, i = 1 to 8 835 for all expressed genes.
For each seed curve, we ranked the genes according to their ED and a cutoff of ED ≤ 2.5 was used in this study as a minimum requirement for periodicity (other cutoffs were also attempted, generating similar results). To reduce noise, we required a minimum of 1.5 fold change between the Eimin and Eimax for gene i to be considered periodic.
Gene ontology, protein interactions, heatmaps and promoter analysis
GENE-E (www.broadinstitute.org/cancer/software/GENE-E/) was used to generate heat maps and hierarchical clustering. Pearson correlations were calculated using R version 3.1.1. All heat maps shown are row-normalized for presentation purposes. Spearman's rank correlation with average linkage was used for clustering. GO terms shown are for biological process (GOTERM_BP_FAT) and were determined by DAVID (http://david.abcc.ncifcrf.gov/gene2gene.jsp). To identify enriched transcription factor motifs within the promoter of periodic genes, the WEB-based Gene SeT AnaLysis Toolkit bioinformatics application (http://bioinfo.vanderbilt.edu/webgestalt/) was employed using all human genes as background. Motifs were then converted into position weight matrix using TOMTOM57. Statistical significances were calculated with t-test, unless otherwise indicated.
ChIP-Seq analysis
ChIP-Seq data were downloaded from ENCODE with the exception of FOXM14. Comparisons of ChIP-Seq signal at transcriptional start site (TSS), gene body, and transcriptional end site (TES) was conducted using ngs.plot58. For comparative analysis in the scatter plot, promoter signal was defined between the TSS and 1 kb upstream of the TSS. For analysis of TES signal was summed for the region between the TES and 1kb downstream. The background ChIp-seq signals were calculated by using a randomly selected set of genes with matched expression levels to the periodic genes analyzed (shown in Supplementary information, Figure S3B and S3C). The significance between background and periodic genes was tested by two-tailed t-test, and similar results were observed with KS-test.
Author Contributions
DD and ZW conceived and designed the experiments. DD carried out the experiments. DD, Y-HT, NG, DKJ analyzed data. DD and ZW wrote the manuscript. Y-HT, NG, DKJ, ID provided manuscript suggestions.
Competing Financial Interests
The authors declare no competing financial interests.
References
Bertoli C, Skotheim JM, de Bruin RA . Control of cell cycle transcription during G1 and S phases. Nat Rev Mol Cell Biol 2013; 14:518–528.
Bahler J . Cell-cycle control of gene expression in budding and fission yeast. Ann Rev Genet 2005; 39:69–94.
Whitfield ML, Sherlock G, Saldanha AJ, et al. Identification of genes periodically expressed in the human cell cycle and their expression in tumors. Mol Biol Cell 2002; 13:1977–2000.
Grant GD, Brooks L 3rd, Zhang X, et al. Identification of cell cycle-regulated genes periodically expressed in U2OS cells and their regulation by FOXM1 and E2F transcription factors. Mol Biol Cell 2013; 24:3634–3650.
Cho RJ, Campbell MJ, Winzeler EA, et al. A genome-wide transcriptional analysis of the mitotic cell cycle. Mol Cell 1998; 2:65–73.
Bar-Joseph Z, Siegfried Z, Brandeis M, et al. Genome-wide transcriptional analysis of the human cell cycle identifies genes differentially regulated in normal and cancer cells. Proc Natl Acad Sci USA 2008; 105:955–960.
Spellman PT, Sherlock G, Zhang MQ, et al. Comprehensive identification of cell cycle-regulated genes of the yeast Saccharomyces cerevisiae by microarray hybridization. Mol Biol Cell 1998; 9:3273–3297.
Hung T, Wang Y, Lin MF, et al. Extensive and coordinated transcription of noncoding RNAs within cell-cycle promoters. Nat Genet 2011; 43:621–629.
Batista PJ, Chang HY . Long noncoding RNAs: cellular address codes in development and disease. Cell 2013; 152:1298–1307.
Hanahan D, Weinberg RA . Hallmarks of cancer: the next generation. Cell 2011; 144:646–674.
Chibon F . Cancer gene expression signatures - the rise and fall? Eur J Cancer 2013; 49:2000–2009.
Weinstein JN, Collisson EA, Mills GB, et al. The Cancer Genome Atlas Pan-Cancer analysis project. Nat Genet 2013; 45:1113–1120.
Ciriello G, Miller ML, Aksoy BA, Senbabaoglu Y, Schultz N, Sander C . Emerging landscape of oncogenic signatures across human cancers. Nat Genet 2013, 45:1127–1133.
Gonzalez-Perez A, Jene-Sanz A, Lopez-Bigas N . The mutational landscape of chromatin regulatory factors across 4,623 tumor samples. Genome Biol 2013; 14:r106.
Wang K, Singh D, Zeng Z, et al. MapSplice: accurate mapping of RNA-seq reads for splice junction discovery. Nucleic Acids Res 2010; 38:e178.
Trapnell C, Williams BA, Pertea G, et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 2010; 28:511–515.
Yoon JH, Abdelmohsen K, Gorospe M . Posttranscriptional gene regulation by long noncoding RNA. J Mol Biol 2013; 425:3723–3730.
Tripathi V, Shen Z, Chakraborty A, et al. Long noncoding RNA MALAT1 controls cell cycle progression by regulating the expression of oncogenic transcription factor B-MYB. PLoS Genet 2013; 9:e1003368.
Johnson DG, Schneider-Broussard R . Role of E2F in cell cycle control and cancer. Front Biosci 1998; 3:d447–d448.
Laoukili J, Kooistra MR, Bras A, et al. FoxM1 is required for execution of the mitotic programme and chromosome stability. Nat Cell Biol 2005; 7:126–136.
Grant GD, Gamsby J, Martyanov V, et al. Live-cell monitoring of periodic gene expression in synchronous human cells identifies Forkhead genes involved in cell cycle control. Mol Biol Cell 2012; 23:3079–3093.
Sanders DA, Ross-Innes CS, Beraldi D, Carroll JS, Balasubramanian S . Genome-wide mapping of FOXM1 binding reveals co-binding with estrogen receptor alpha in breast cancer cells. Genome Biol 2013; 14:R6.
Sadasivam S, Duan S, DeCaprio JA . The MuvB complex sequentially recruits B-Myb and FoxM1 to promote mitotic gene expression. Genes Dev 2012; 26:474–489.
Heidari N, Phanstiel DH, He C, et al. Genome-wide map of regulatory interactions in the human genome. Genome Res 2014; 24:1905–1917.
Ong CT, Corces VG . CTCF: an architectural protein bridging genome topology and function. Nat Rev Genet 2014; 15:234–246.
Holwerda SJ, de Laat W . CTCF: the protein, the binding partners, the binding sites and their chromatin loops. Philos Trans R Soc Lond B Biol Sci 2013; 368:20120369.
Rubio ED, Reiss DJ, Welcsh PL, et al. CTCF physically links cohesin to chromatin. Proc Natl Acad Sci USA 2008; 105:8309–8314.
Xie D, Boyle AP, Wu L, Zhai J, Kawli T, Snyder M . Dynamic trans-acting factor colocalization in human cells. Cell 2013; 155:713–724.
Handoko L, Xu H, Li G, et al. CTCF-mediated functional chromatin interactome in pluripotent cells. Nat Genet 2011; 43:630–638.
Smith E, Shilatifard A . Enhancer biology and enhanceropathies. Nat Struct Mol Biol 2014; 21:210–219.
Zaidi SK, Grandy RA, Lopez-Camacho C, et al. Bookmarking target genes in mitosis: a shared epigenetic trait of phenotypic transcription factors and oncogenes? Cancer Res 2014; 74:420–425.
Kandoth C, Schultz N, Cherniack AD, et al. Integrated genomic characterization of endometrial carcinoma. Nature 2013; 497:67–73.
Cancer Genome Atlas Network T. Comprehensive molecular portraits of human breast tumours. Nature 2012; 490:61–70.
Salama R, Sadaie M, Hoare M, Narita M . Cellular senescence and its effector programs. Genes Dev 2014; 28:99–114.
Gyorffy B, Lanczky A, Eklund AC, et al. An online survival analysis tool to rapidly assess the effect of 22,277 genes on breast cancer prognosis using microarray data of 1,809 patients. Breast Cancer Res Treat 2010; 123:725–731.
Thomas AA, Brennan CW, DeAngelis LM, Omuro AM . Emerging therapies for glioblastoma. JAMA Neurol 2014; 71:1437–1444.
Cancer Genome Atlas Network T. Comprehensive molecular characterization of clear cell renal cell carcinoma. Nature 2013; 499:43–49.
Kittler R, Pelletier L, Heninger AK, et al. Genome-scale RNAi profiling of cell division in human tissue culture cells. Nat Cell Biol 2007; 9:1401–1412.
Mukherji M, Bell R, Supekova L, et al. Genome-wide functional analysis of human cell-cycle regulators. Proc Natl Acad Sci USA 2006; 103:14819–14824.
Neumann B, Walter T, Heriche JK, et al. Phenotypic profiling of the human genome by time-lapse microscopy reveals cell division genes. Nature 2010; 464:721–727.
Simon JM, Hacker KE, Singh D, et al. Variation in chromatin accessibility in human kidney cancer links H3K36 methyltransferase loss with widespread RNA processing defects. Genome Res 2014; 24:241–250.
Hakimi AA, Ostrovnaya I, Reva B, et al. Adverse outcomes in clear cell renal cell carcinoma with mutations of 3p21 epigenetic regulators BAP1 and SETD2: a report by MSKCC and the KIRC TCGA research network. Clin Cancer Res 2013; 19:3259–3267.
Carvalho S, Raposo AC, Martins FB, et al. Histone methyltransferase SETD2 coordinates FACT recruitment with nucleosome dynamics during transcription. Nucleic Acids Res 2013; 41:2881–2893.
Uchida F, Uzawa K, Kasamatsu A, et al. Overexpression of cell cycle regulator CDCA3 promotes oral cancer progression by enhancing cell proliferation with prevention of G1 phase arrest. BMC cancer 2012; 12:321.
Dominguez D, Tsai YH, Weatheritt R, et al. An extensive program of periodic alternative splicing linked to cell cycle progression. Elife 2016; 5:e10288.
Inghirami G, Chiarle R, Simmons WJ, Piva R, Schlessinger K, Levy DE . New and old functions of STAT3: a pivotal target for individualized treatment of cancer. Cell Cycle 2005; 4:1131–1133.
Fu Y, Sinha M, Peterson CL, Weng Z . The insulator binding protein CTCF positions 20 nucleosomes around its binding sites across the human genome. PLoS Genet 2008; 4:e1000138.
Nekrasov M, Amrichova J, Parker BJ, et al. Histone H2A.Z inheritance during the cell cycle and its impact on promoter organization and dynamics. Nat Struct Mol Biol 2012; 19:1076–1083.
Rao SS, Huntley MH, Durand NC, et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 2014; 159:1665–1680.
Kontzoglou K, Palla V, Karaolanis G, et al. Correlation between Ki67 and breast cancer prognosis. Oncology 2013; 84:219–225.
Fitzgibbons PL, Page DL, Weaver D, et al. Prognostic factors in breast cancer. College of American Pathologists Consensus Statement 1999. Arch Pathol Lab Med 2000; 124:966–978.
Lehr HA, Rochat C, Schaper C, et al. Mitotic figure counts are significantly overestimated in resection specimens of invasive breast carcinomas. Mod Pathol 2013; 26:336–342.
Hoadley KA, Yau C, Wolf DM, et al. Multiplatform analysis of 12 cancer types reveals molecular classification within and across tissues of origin. Cell 2014; 158:929–944.
Edmunds JW, Mahadevan LC, Clayton AL . Dynamic histone H3 methylation during gene induction: HYPB/Setd2 mediates all H3K36 trimethylation. EMBO J 2008; 27:406–420.
Mihaly Z, Kormos M, Lanczky A, et al. A meta-analysis of gene expression-based biomarkers predicting outcome after tamoxifen treatment in breast cancer. Breast Cancer Res Treat 2013; 140:219–232.
Cline MS, Craft B, Swatloski T, et al. Exploring TCGA pan-cancer data at the UCSC cancer genomics browser. Sci Rep 2013; 3:2652.
Gupta S, Stamatoyannopoulos JA, Bailey TL, Noble WS . Quantifying similarity between motifs. Genome Biol 2007; 8:R24.
Shen L, Shao N, Liu X, Nestler E . ngs.plot: Quick mining and visualization of next-generation sequencing data by integrating genomic databases. BMC Genomics 2014; 15:284.
Parker JS, Mullins M, Cheang MC, et al. Supervised risk predictor of breast cancer based on intrinsic subtypes. J Clin Oncol 2009; 27:1160–1167.
Acknowledgements
We thank Dr Jan Prins and Darshan Singh for suggestions in analyzing RNA-seq data. We thank Drs William Marzluff, Jean Cook, Michael Emanuele and Mauro Calabrese for reading this manuscript and providing comments. We thank Peter Sudmant for his assistance and advice with permutation analysis and helpful comments on this manuscript. This work is supported by an NIH grant (R01CA158283) and Jefferson-Pilot fellowship to ZW.
Author information
Authors and Affiliations
Corresponding author
Additional information
( Supplementary information is linked to the online version of the paper on the Cell Research website.)
Supplementary information
Supplementary information, Figure S1
(A), Histograms of DNA content in synchronous HeLa cells upon G1/S release. (PDF 453 kb)
Supplementary information, Figure S2
(A), Heat map representation of gene ontology terms enriched in periodic genes grouped by peak stage (see methods). (PDF 618 kb)
Supplementary information, Figure S3
(A), Heat map representation of motifs discovered within the promoters of periodic genes (left). (PDF 850 kb)
Supplementary information, Figure S4
(A), Pairwise Spearman's rank correlation analysis on all possible pairs within the 14 synchronous samples based on the 1182 periodic transcripts, 1,000 arbitrarily selected non-periodic genes or all transcripts detected in Hela cells. (PDF 564 kb)
Supplementary information, Figure S5
(A-D), Venn diagrams showing the overlap in identified periodic genes in this study (red circle) and four other studies (black circles). (PDF 294 kb)
Supplementary information, Figure S6
ChIP-seq signals of selected histone modi_cations and proteins at the indicated genomic locations for the core 67, all periodic and background genes. (PDF 1170 kb)
Rights and permissions
About this article
Cite this article
Dominguez, D., Tsai, YH., Gomez, N. et al. A high-resolution transcriptome map of cell cycle reveals novel connections between periodic genes and cancer. Cell Res 26, 946–962 (2016). https://doi.org/10.1038/cr.2016.84
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2016.84
Keywords
This article is cited by
-
Single-cell transcriptomics identifies potential cells of origin of MYC rhabdoid tumors
Nature Communications (2022)
-
Dual RNA 3’-end processing of H2A.X messenger RNA maintains DNA damage repair throughout the cell cycle
Nature Communications (2021)
-
Joint changes in RNA, RNA polymerase II, and promoter activity through the cell cycle identify non-coding RNAs involved in proliferation
Scientific Reports (2021)
-
AKIRIN2 controls the nuclear import of proteasomes in vertebrates
Nature (2021)
-
Selective therapeutic strategy for p53-deficient cancer by targeting dysregulation in DNA repair
Communications Biology (2021)