Abstract
Craniofacial anomalies (CFAs) characterized by birth defects of skull and facial bones are the most frequent congenital disease. Genomic analysis has identified multiple genes responsible for CFAs; however, the underlying genetic mechanisms for the majority of CFAs remain largely unclear. Our previous study revealed that the Wwp2 E3 ubiquitin ligase facilitates craniofacial development in part through inducing monoubiquitination and activation of the paired-like homeobox transcription factor, Goosecoid (Gsc). Here we report that Gsc is also ubiquitinated and activated by the APCCdh1 E3 ubiquitin ligase, leading to transcriptional activation of various Gsc target genes crucial for craniofacial development. Consistenly, neural crest-specific Cdh1-knockout mice display similar bone malformation as Wwp2-deficient mice in the craniofacial region, characterized by a domed skull, a short snout and a twisted nasal bone. Mechanistically, like Wwp2-deficient mice, mice with Cdh1 deficiency in neural crest cells exhibit reduced Gsc/Sox6 transcriptional activities. Simultaneous deletion of Cdh1 and Wwp2 results in a more severe craniofacial defect compared with single gene deletion, suggesting a synergistic augmentation of Gsc activity by these two E3 ubiquitin ligases. Hence, our study reveals a novel role for Cdh1 in craniofacial development through promoting APC-dependent non-proteolytic ubiquitination and activation of Gsc.
Similar content being viewed by others
Introduction
Craniofacial anomalies (CFAs) are the most frequently occurring congenital diseases that affect the growth of the head and facial bones, accounting for about one-third of all human congenital defects1. CFAs seem to arise from a combination of genetic and environmental factors. Multiple genes mutated in children with CFAs have been identified, and most of them are transcription factors (e.g., Hox, Msx, Dlx and Gsc)2. However, the pathophysiological mechanisms governing craniofacial development are far from clear and the ∼100 gene defects accounting for the majority of CFAs have not been thoroughly investigated.
Wwp2 is a member of the NEDD4 family of E3 ligase proteins, containing an N-terminal membrane-targeting C2 domain, four internal double tryptophan (WW) domains and a C-terminal HECT domain that is responsible for its E3 ubiquitin ligase activity3. Our previous study found that Wwp2 plays a crucial role in craniofacial development in part by mono-ubiquitinating Goosecoid (Gsc), which amplifies the transcriptional activity of Gsc4. As such, mice deficient in Wwp2 exhibited craniofacial patterning defects that were largely due to a decreased expression of the key cartilaginous transcription factor Sox6, a well-characterized Gsc transcriptional target4. However, it remains undefined whether other upstream E3 ligases besides Wwp2 could also modulate the expression and/or activity of Gsc, and subsequently regulate craniofacial development.
Cdh1 is one of the two co-activators that associate with the anaphase-promoting complex/cyclosome (APC/C, also named APC) complex, which controls cell cycle progression mainly in the M and G1 phases through forming two distinct sub-complexes, APCCdh1 and APCCdc20, to promote the timely ubiquitination and subsequent degradation of mitotic cyclins and other cell cycle regulators5. Notably, Cdh1 recruits substrates to the APC core complex largely via the KEN Box or Destruction Box (D-box) motifs found in most APCCdh1 substrates6,7. Mice deficient in Cdh1 were embryonic lethal, while Cdh1 heterozygous mice displayed a decrease in survival and were more susceptible to developing epithelial tumors8, suggesting a possible tumor suppressor role for Cdh1. The tumor suppressor role of Cdh1 was further supported by recent studies revealing a decrease of Cdh1 expression in various human tumor tissues9,10,11. Besides its conventional roles in cell cycle regulation, Cdh1 has recently been found to play critical roles in a wide spectrum of cellular processes including DNA damage repair9,12, cellular metabolism13, cell migration14 and neuronal development15,16. Moreover, we recently identified a novel, APC-independent, role for Cdh1 in controlling osteoblast differentiation by disrupting the inter-molecular interaction of Smurf1 dimers, leading to the activation of Smurf1, another member of the NEDD4 family of E3 ligases17.
Here we reveal an important function of Cdh1 in positive regulation of craniofacial development through promoting APC-dependent monoubiquitination and activation of Gsc. Moreover, we show that APCCdh1 directly targets Gsc for monoubiquitination in a similar fashion as what we found for Wwp2, which activates the transcriptional activity of Gsc, therefore promoting the transcription of a cohort of downstream target genes to govern craniofacial development. To further investigate the functional consequence of APCCdh1-induced Gsc ubiquitination, we generated neural crest-specific Cdh1 knockout mice using Wnt1-Cre and found that Cdh1F/F;Wnt1-Cre (Cdh1Wnt1/+ for short) mice phenocopied Wwp2-deficient mice, showing reduced Gsc transcriptional activity and Sox6 expression. These results suggest that Cdh1 could regulate craniofacial development through modulating Gsc/Sox6 signaling via promoting non-proteolytic ubiquitination of Gsc in an APC-dependent manner.
Results
Cdh1 interacts with Gsc in a D-box-dependent manner
Most APCCdh1 substrates contain one or multiple D-box motifs (RxxLxxxN)5. Notably, we found that Gsc contains a canonical D-box motif, which is evolutionally conserved across multiple species (Figure 1A). We next examined whether Cdh1 interacts with Gsc. To this end, immunoprecipitation of Flag-tagged Gsc (or HA-tagged Cdh1 or Cdc20) followed by western blotting revealed that Cdh1, but not Cdc20, interacts with Gsc (Figure 1B). To further determine whether Gsc directly interacts with Cdh1 through the conserved D-box motif, we generated a Gsc D-box mutant by replacing two conserved residues within the D-box motif, arginine and leucine, with alanine (termed RLAA-Gsc thereafter; Figure 1A). As shown in Figure 1C, compared with WT Gsc, binding of Cdh1 with RLAA-Gsc was significantly reduced. The interaction between Cdh1 and Gsc was further confirmed at the endogenous level (Figure 1D). More importantly, the direct physical interaction between Gsc and Cdh1 was also established through in vitro GST pull-down experiments using purified recombinant proteins (Figure 1E). Consistently, an impaired interaction between RLAA-Gsc and Cdh1 was observed in the in vitro interaction analysis (Figure 1E). Additional analysis revealed that Cdh1 associates with Gsc through the N-terminal portion of Gsc that contains the conserved D-box motif (Figure 1F). On the other hand, Gsc mainly binds to the WD40 domain of Cdh1 (Figure 1G and 1H), which mediates the interaction of Cdh1 with most of its characterized substrates. These results together demonstrate that Cdh1 directly interacts with Gsc via the conserved D-box motif in Gsc, suggesting Gsc as a putative substrate for APCCdh1-mediated ubiquitination.
Cdh1 interacts with Gsc in a D-box-dependent manner. (A) Sequence alignment of the D-box-containing region among Gsc proteins from various species. Red and blue indicate WT and mutated amino acids within the Gsc D-box motif, respectively. (B) Immunoblot (IB) analysis of whole-cell lysates (WCL) and immunoprecipitate (IP) products derived from 293T cells transfected with HA-Cdh1 or HA-Cdc20 and Flag-Gsc constructs. Thirty-six hours post transfection, cells were pretreated with 10 μM MG132 for 10 h before harvesting. (C) IB analysis of WCL and IP products derived from 293T cells transfected with Flag-WT-Gsc or Flag-RLAA-Gsc and HA-Cdh1 constructs. Thirty-six hours post transfection, cells were pretreated with 10 μM MG132 for 10 h before harvesting. (D) IB analysis of WCL and anti-Cdh1 IP products derived from MCF-7 cells. Cells were pretreated with 10 μM MG132 for 10 h before harvesting. (E) Autoradiography of 35S-labelled Gsc bound to GST-Cdh1, but not the GST recombinant protein. (F) IB analysis of WCL and IP products derived from 293T cells transfected with HA-Cdh1 and the indicated Flag-Gsc constructs. Thirty-six hours post transfection, cells were pretreated with 10 μM MG132 for 10 h before harvesting. (G) IB analysis of WCL and IP products derived from 293T cells transfected with Flag-Gsc and the indicated Myc-Cdh1 constructs. Thirty-six hours post transfection, cells were pretreated with 10 μM MG132 for 10 h before harvesting. (H) A schematic illustration showing different fragments of Cdh1 (Myc-tag) and the binding with Gsc. “N” represents the N-terminal portion of Cdh1, which binds to the APC core complex; “F” represents the Fizzy domain of Cdh1, which also participates in the Cdh1-APC interaction; “WD40” represents the C-terminal part of Cdh1, which mediates the interaction between Cdh1 and its substrates; “+” represents definite interaction; “+/−” represents weak interaction; “−” represents no interaction.
APCCdh1 directly ubiquitinates Gsc in a D-box-dependent manner
The interaction between Cdh1 and Gsc prompted us to examine whether APCCdh1 could directly ubiquitinate Gsc. Notably, unlike most of the known Cdh1 substrates that are modified by APCCdh1-mediated polyubiquitination, Cdh1 promotes the monoubiquitination of Gsc (Figure 2A), which is similar to our previous finding that Wwp2 mainly triggers monoubiquitination and subsequent activation of Gsc (Figure 2B)4. In further support of this finding, we found that K0-ubiquitin, in which all seven lysine residues are mutated to arginine, could still be utilized by APCCdh1 to catalyze Gsc ubiquitination as efficiently as WT ubiquitin (Figure 2C). Since K0-ubiquitin is deficient in forming polyubiquitin chains, our results indicate that APCCdh1 mainly catalyzes monoubiquitination rather than polyubiquitination of Gsc. Furthermore, APCCdh1-mediated ubiquitination was largely attenuated for D-box-mutated RLAA-Gsc (Figure 2D). Considering that Gsc directly interacts with Cdh1 through its conserved D-box motif (Figure 1D), these findings suggest that APCCdh1 ubiquitinates Gsc through a direct association with this protein.
Cdh1 directly ubiquitinates Gsc in a D-box-dependent manner. (A) APCCdh1 targets Gsc for ubiquitination in HeLa cells. IB analysis of WCL and Ni-NTA pull-down products in 6 M guanidine-HCl-containing buffer derived from HeLa cells transfected with the indicated plasmids. Cells were pretreated with 10 μM MG132 for 10 h before harvesting. (B) APCCdh1 and Wwp2 target Gsc for ubiquitination in HeLa cells. IB analysis of WCL and Ni-NTA pull-down products in 6 M guanidine-HCl-containing buffer derived from HeLa cells transfected with the indicated plasmids. Cells were pretreated with 10 μM MG132 for 10 h before harvesting. (C) IB analysis of WCL and Ni-NTA pull-down products in 6 M guanidine-HCl-containing buffer derived from HeLa cells transfected with WT- or K0-His-Ub and the indicated plasmids. Cells were pretreated with 10 μM MG132 for 10 h before harvesting. (D) APCCdh1 failed to promote the ubiquitination of D-box-mutated RLAA-Gsc. IB analysis of WCL and Ni-NTA pull-down products in 6 M guanidine-HCl-containing buffer derived from HeLa cells transfected with the indicated plasmids. Cells were pretreated with 10 μM MG132 for 10 h before harvesting. (E) APCCdh1 targets Gsc for ubiquitination in vitro. Bacterially expressed and purified GST-tagged WT and RLAA Gsc proteins were incubated with the APCcdh1 complex purified from G1 phase HeLa cell extracts together with purified recombinant E1, E2 and ubiquitin as indicated at 30 °C for 60 min before being resolved by SDS-PAGE and probed with the anti-GST antibody. (F) APCCdh1 co-operated with Wwp2 to promote Gsc ubiquitination in cells. IB analysis of WCL and Ni-NTA pull-down products in 6 M guanidine-HCl-containing buffer derived from HeLa cells transfected with the indicated plasmids. Cells were pretreated with 10 μM MG132 for 10 h before harvesting.
In further support of Gsc as a bona fide substrate for APCCdh1-meditaed ubiquitination, an in vitro ubiquitination assay was performed using recombinant proteins, including WT or mutant Gsc, ubiqutin, ubiquitin-activating E1 enzyme UBE1 and ubiquitin-conjugating E2 enzyme UbcH10, together with the APCCdh1 complex purified from cell extracts. Notably, significant ubiquitination of Gsc was observed in the in vitro assay, while Gsc ubiquitination was largely reduced when RLAA-Gsc was used as a substrate (Figure 2E). These results support Gsc as a direct APCCdh1 substrate. Intriguingly, Cdh1 and Wwp2 co-operated to promote Gsc ubiquitination, suggesting their synergistic roles in controlling Gsc activity (Figure 2F).
Cdh1 augments the Gsc/Sox6 signaling pathway
We next sought to investigate whether Cdh1 could regulate Gsc transcriptional activity by utilizing a previously described reporter assay system encompassing a Gsc-Gal4 fusion protein and a Gal4-responsive luciferase reporter (Supplementary information, Figure S1A)4. Strikingly, we found that similar to Wwp2, ectopic expression of Cdh1 significantly enhanced Gsc transcriptional activity in a dose-dependent manner (Figure 3A). On the other hand, depletion of endogenous Cdh1 moderately suppressed Gsc activity (Figure 3B), indicating a positive regulation of Gsc transcriptional activity by Cdh1. Furthermore, we found that unlike Cdh1, Cdc20 or Cdh1 mutants, including ΔC-box-Cdh1 and ΔFizzy-Cdh1, which are deficient in promoting Gsc ubiquitination (Figure 3D and 3E) due to their impaired ability to interact with the APC core complex, failed to induce Gsc activity (Figure 3C). This result suggests that activation of Gsc by Cdh1 is largely dependent on the functional APCCdh1 E3 ligase holoenzyme.
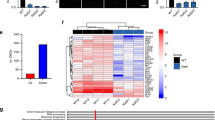
Cdh1 augments the activity of the Gsc/Sox6 signaling pathway. (A) Normalized luciferase reporter activities in 293T cells transfected with the Gal4-Gsc construct, Gal4-luciferase reporter and increased amounts of Cdh1. Fourty-eight hours post transfection, cells were harvested for luciferase reporter assays. Data represent relative luciferase activities shown as the mean ± SEM (n = 3, **P < 0.01, Student's t-test). (B) Normalized luciferase reporter activities in 293T cells transfected with the Gal4-Gsc construct, Gal4-luciferase reporter and the indicated pSuper-shRNA constructs. Fourty-eight hours post transfection, cells were harvested for luciferase reporter assays. Data represent relative luciferase activities shown as the mean ± SEM (n = 3, *P < 0.05, Student's t-test). (C) Normalized luciferase reporter activities in 293T cells transfected with the Gal4-Gsc construct, Gal4-luciferase reporter, the indicated WT or mutant Cdh1, and Cdc20 constructs. Fourty-eight hours post transfection, cells were harvested for luciferase reporter assays. Data represent relative luciferase activities shown as the mean ± SEM (n = 3, **P < 0.01, Student's t-test). (D) A schematic illustration of the domain structure of Cdh1. The location of C-Box, Fizzy Domain and WD40 Repeats were labelled. (E) IB analysis of WCL and Ni-NTA pull-down products in 6 M guanidine-HCl-containing buffer derived from HeLa cells transfected with Flag-Gsc and the indicated HA-Cdh1 plasmids. Cells were pretreated with 10 μM MG132 for 10 h before harvesting. (F) Normalized luciferase reporter activities in 293T cells transfected with Sox6 promoter luciferase reporter and the indicated constructs. Fourty-eight hours post transfection, cells were harvested for luciferase reporter assays. Data represent relative luciferase activities shown as the mean ± SEM (n = 3, *P < 0.05, **P < 0.01, Student's t-test). (G) Depletion of Cdh1 in ATDC5 chondrocytes led to a significant suppression in transcription of various downstream targets of the Gsc/Sox6 signaling pathway that governs the craniofacial development. ATDC5 cells were infected with the indicated lentiviral shRNA constructs. The infected cells were selected with 1 μg/ml hygromycin for 72 h to eliminate non-infected cells. The knockdown efficiency, as well as the expression of various characteristic craniofacial markers, was determined by real-time PCR analysis. Three sets of independent experiments were performed and data shown represent the mean ± SEM (n = 3, *P < 0.05, **P < 0.01, Student's t-test).
To further examine the mechanistic role of Cdh1 in controlling the Gsc/Sox6 signaling cascade, we measured the transcription activity of the Sox6 promoter using a luciferase reporter containing a truncated fragment of the Sox6 promoter described previously (Supplementary information, Figure S1B)4. The promoter reporter was verified to be responsive to Gsc expression (Supplementary information, Figures S1C). Interestingly, further introducing WT Cdh1, but not ΔC-box-Cdh1 or Cdc20, dramatically enhanced the Sox6 promoter activity in response to Gsc expression (Figure 3F). These results coherently support a model in which Cdh1 activates the Gsc/Sox6 signaling axis through directly influencing Gsc monoubiquitination.
The Gsc/Sox6 signaling cascade is essential for mounting the transcriptional activation of various target genes that are crucial in craniofacial development18. To further investigate the physiological role of APCCdh1 in this developmental process, we utilized the ATDC5 cell line derived from mouse teratocarcinoma cells, which is widely used as a model cell line for studying chondrogenesis4. Notably, consistent with Cdh1 being a positive regulator of the Gsc/Sox6 signaling pathway, depletion of Cdh1 in ATDC5 cells significantly decreased mRNA levels of various Gsc/Sox6 target genes including Col2a119, S100A1, S100B20 and Matn121 (Figures 3G). These results support a positive role of Cdh1 in regulating Gsc transcriptional activity.
Cdh1 conditional knockout mice displayed various craniofacial defects
We next examined whether deletion of Cdh1 would cause craniofacial defects. As germline Cdh1 knockout led to embryonic lethality8, we crossed Wnt1-Cre mice22 with Cdh1F/F mice8 to generate neural crest-specific Cdh1 knockout mice, Cdh1F/F;Wnt1-Cre mice (hereafter referred to as Cdh1Wnt1 mice; Supplementary information, Figure S2A). We harvested RNAs from different tissues and found reduced Cdh1 mRNA levels in dorsal root ganglion of Cdh1Wnt1 mice (Supplementary information, Figure S2B)23. Intriguingly, we found that Cdh1Wnt1 mice phenocopied Wwp2-deficient mice in displaying various craniofacial defects (Supplementary information, Figure S2C and S2D). Notably, compared with WT mice, Cdh1Wnt1 mice showed shorter stature and lighter weight (Figure 4A-4C). The major bone developmental defects were found in the craniofacial region (Figures 4D) rather than in limbs and ribs (Supplementary information, Figure S2E and S2F). Moreover, we observed abnormal craniofacial development in Cdh1Wnt1 mice as evidenced by the presence of a domed skull and a short snout (Figures 4E).
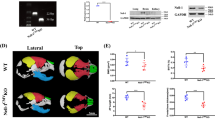
Cdh1 conditional knockout mice displayed various craniofacial defects. (A) Alcian blue/Alizarin red staining of 2-week-old WT, Cdh1Wnt1/+ and Cdh1Wnt1 mice. (B) Body weight of 2-week-old male WT and Cdh1Wnt1 mice. Values represent the means ± SD (n ≥ 5 for each genotype, **P < 0.001, Student's t-test). (C) Representative images of 5-week-old male WT and Cdh1Wnt1 mice. (D) Shorter snout (arrows) in Cdh1Wnt1 mice compared with WT littermates, as shown grossly (top) as well as by Alizarin red staining (bottom) of WT and Cdh1Wnt1 mouse skulls. (E) μCT analysis of skulls derived from WT (left) and Cdh1Wnt1 (middle and right) mice showing the short or twisted (arrows) nasal bone in Cdh1Wnt1 mice. (F) Quantitative analysis of the distances between the various landmarks labeled in E (red letters), in the skulls of 5-week-old WT and Cdh1Wnt1 mice. Values represent the means ± SD (n = 3 for each genotype; **P < 0.01, *P < 0.05, Student's t-test). (G) Relative mRNA levels of the indicated genes in primary calvarial cells from WT and Cdh1Wnt1 mice (n = 3 for each genotype; **P < 0.01, *P < 0.05, Student's t-test). (H) Relative mRNA levels of the indicated genes in immortalized Cdh1F/F calvarial cells treated with lentivirus expressing Cre (with GFP as the negative control). The infected cells were selected with 2 μg/ml puromycin for 72 h to eliminate non-infected cells. The knockdown efficiency and the expression of various characteristic craniofacial markers were determined by real-time PCR analysis. Three sets of independent experiments were performed to generate each data and data shown represent the mean ± SD (n = 3 for each genotype, **P < 0.01, *P < 0.05, Student's t-test).
To further quantify the craniofacial abnormalities, we used quantitative computed tomography (μ-QCT)4 to scan the skulls of the Cdh1Wnt1 mice and their WT littermates. Notably, we found that the nasal bones of Cdh1Wnt1 mice were decreased in length by ∼30% (Figure 4E and 4F). Although all Cdh1Wnt1 mice displayed craniofacial abnormalities, the severity of their phenotypes was variable due to unclear mechanism(s). Furthermore, a subset of Cdh1Wnt1 mice developed twisted nasal bones and abnormal teeth (Figure 4E), similar to Wwp2-deficient mice4. We next examined the expression of genes crucial for craniofacial development by qPCR and found that mRNA levels of Wwp2 and Gsc remained largely unchanged between WT and Cdh1Wnt1 mouse chondrocytes, which were isolated from the nasal bone of P4 mice (Figure 4G). On the other hand, in support of the craniofacial defects observed in Cdh1Wnt1 mice (Figure 4D-4F), mRNA levels of Sox6 and target genes of the Gsc/Sox6 signaling pathway including Col2a1, S100A1, S100B and Col9a2 were significantly reduced in Cdh1Wnt1 mice (Figure 4G). This result is consistent with the downregulated Gsc/Sox6 signaling upon Cdh1 knockdown in ATDC5 chondrocytes (Figure 3G).
To further confirm the regulatory roles of Cdh1 in the Gsc/Sox6 signaling pathway, we cultured nasal chondrocytes from Cdh1F/F mice and then infected these cells with lentivirus expressing GFP or Cre recombinase. As shown in Figure 4H, expression of Cre led to a markedly reduced Cdh1 expression accompanied by a significant decrease in expression levels of Sox6 and Gsc/Sox6 target genes. However, expression of Cre recombinase had no significant effects in WT cells (Supplementary information, Figure S3). Taken together, these data support that Cdh1 can regulate craniofacial development through activating the Gsc/Sox6 transcription network4.
Synergistic regulation of the Gsc/Sox6 signaling pathway by Wwp2 and Cdh1
Our results above demonstrate that both Wwp2 and APCCdh1 could activate the Gsc/Sox6 signaling pathway and regulate craniofacial development. Previous study from our group has identified that substitution of three lysine residues of Gsc (K219, K231 and K234) with arginine (Gsc-K3R) led to a dramatic decrease in the level of Wwp2-mediated Gsc ubiquitination4. We thus wonder whether these three lysines are also important for APCCdh1-mediated Gsc ubiquitination. As shown in Figure 5A, compared with WT Gsc, the Gsc-K3R mutant was largely resistant to APCCdh1-mediated monoubiquitination. We next examined whether Cdh1's effects on Gsc transcriptional activity could be impaired by the Gsc-K3R mutation. Indeed, luciferase reporter assay demonstrated that Cdh1 could further increase Gsc-induced transcription activity of the Sox6 promoter, however, Gsc-K3R could not activate the Sox6 promoter and Cdh1 failed to increase the transcriptional activity of Gsc-K3R (Figure 5B). We next sought to determine whether Wwp2 and Cdh1 could function in a synergistic manner. As shown in Figure 5C, co-transfection of Wwp2 and Cdh1 could synergistically increase Gsc transcriptional activity, indicating a cooperative effect of Cdh1 and Wwp2 in activation of Gsc in this experimental setting.
A synergistic regulation of the Gsc/Sox6 signaling pathway by Wwp2 and Cdh1. (A) IB analysis of WCL and Ni-NTA pull-down products in 6 M guanidine-HCl containing buffer derived from HeLa cells transfected with WT Gsc or K3R-Gsc mutant and the indicated plasmids. Cells were pretreated with 10 μM MG132 for 10 h before harvesting. (B) Normalized activities of Pax6 promoter luciferase reporter in 293T cells transfected with the indicated constructs. Fourty-eight hours post transfection, cells were harvested for luciferase reporter assays. Data shown represent the mean ± SD (n = 3, **P < 0.01, #P > 0.05, Student's t-test). (C) Gsc transcriptional reporter activity was evaluated in 293T cells transfected with Gal4-Gsc construct, Gal4-luciferase reporter and the indicated constructs. Fourty-eight hours post transfection, cells were harvested for luciferase reporter assays. Data shown represent the mean ± SD (n = 3, **P < 0.01, Student's t-test). (D) Depletion of Wwp2 in ATDC5 chondrocytes led to a significant suppression in transcription of various downstream targets of the Gsc/Sox6 signaling pathway that govern the craniofacial development. ATDC5 cells were infected with the indicated lentiviral shRNA constructs. The infected cells were selected with 2 μg/ml puromycin for 72 h to eliminate non-infected cells. The knockdown efficiency, as well as the expression of various characteristic craniofacial markers, was determined by real-time PCR analysis. Three sets of independent experiments were performed to generate each data and data shown represent the mean ± SD (n = 3, **P < 0.01, *P < 0.05, Student's t-test). (E) Depletion of Cdh1 or Wwp2 or simultaneous depletion of Cdh1 and Wwp2 in ATDC5 chondrocytes led to a significant suppression in transcription of various downstream targets of the Gsc/Sox6 signaling pathway that govern the craniofacial development. ATDC5 cells were infected with the indicated lentiviral shRNA constructs. The infected cells were selected with 2 μg/ml puromycin (for shWwp2) or with 1 μg/ml hygromycin (for shCdh1) for 72 h to eliminate non-infected cells. The Cdh1 and Wwp2 knockdown efficiency, as well as the expression of various characteristic craniofacial markers, was determined by real-time PCR analysis. Three sets of independent experiments were performed to generate each data and data shown represent the mean ± SD (n = 3, **P < 0.01, *P < 0.05, Student's t-test). (F) Immunoprecipitation experiments were conducted in nasal chondrocytes of Cdh1F/F cells. Endogenous Wwp2 and Gsc were pulled down by immunoprecipitation with anti-Cdh1 antibody.
To further determine the synergism between Wwp2 and Cdh1, we used ATDC5 cells and found that depletion of Wwp2 in ATDC5 cells significantly repressed the gene expression of Sox6 and Gsc/Sox6 target genes including Col2a1, Col9a2 and Matn1 (Figure 5D). Furthermore, co-depletion of Cdh1 and Wwp2 could further decrease the mRNA levels of Sox6 and various Gsc/Sox6 target genes (Figure 5E). Taken together, these data illustrate that Cdh1 and Wwp2 could synergistically promote the activation and transcriptional activity of Gsc. The synergism between Wwp2 and Cdh1 prompted us to examine whether Wwp2 and Cdh1 could interact with each other. We cultured nasal chondrocytes from WT mice and performed immunoprecipitation (IP) using anti-Cdh1 antibody. Co-IP experiments revealed the interaction between endogenous Cdh1 and Wwp2 (Figure 5F), further supporting that Cdh1/Wwp2 could regulate the Gsc/Sox6 signaling pathway in a synergistic manner.
CFAs in Cdh1/Wwp2 double heterozygous mice
As both Wwp2 and Cdh1 could induce Gsc ubiquitination and activate the Gsc/Sox6 signaling pathway, we sought to determine whether Wwp2 and Cdh1 could affect the craniofacial development in a synergistic manner. Cdh1Wnt1 (Figure 4) and Wwp2GT mice4 demonstrated similar craniofacial defects. Heterozygous Cdh1F/+;Wnt1-Cre (hereafter referred to as Cdh1Wnt1/+ mice) or Wwp2GT/+ mice exhibited no significant CFAs (Figure 6A). As Cdh1 and Wwp2 act synergistically, we hypothesized that double heterozygous mice may display CFAs. To this end, we mated Cdh1Wnt1/+ mice with Wwp2GT/+ mice, producing double heterozygous Cdh1Wnt1/+;Wwp2GT/+ mice. Notably, double heterozygous mice displayed shorter stature and lighter weight compared with WT and single heterozygous mice (Figure 6A-6B). Moreover, Cdh1/Wwp2 double heterozygous mice exhibited a domed skull and a shorter snout (Figures 6C). μ-QCT analysis of mouse skulls revealed that double heterozygous mice, but not single heterozygous mice, had severe CFAs (Figures 6D). Taken together, the CFAs in Cdh1Wnt1/+;Wwp2GT/+ double heterozygous mice support the synergistic effect between Cdh1 and Wwp2 in regulating craniofacial development.
Craniofacial anomalies in Wwp2/Cdh1 double heterozygous mice. (A) Representative images of 5-week-old male WT, Cdh1Wnt1/+, Wwp2GT/+ and Cdh1Wnt1/+;Wwp2GT/+ mice. (B) Body weight of 5-week-old male WT, Cdh1Wnt1/+, Wwp2GT/+ and Cdh1Wnt1/+;Wwp2GT/+ mice. Data shown represent the mean ± SD (n = 5 for each genotype, **P < 0.01, Student's t-test). (C) Alizarin red staining of skulls derived from 5-week-old male WT, Cdh1Wnt1/+, Wwp2GT/+ and Cdh1Wnt1/+;Wwp2GT/+ mice, showing the doomed skull, short and twisted nasal bone in Cdh1Wnt1/+;Wwp2GT/+ mice. (D) μCT analysis of skulls derived from 5-week-old male WT, Cdh1Wnt1/+, Wwp2GT/+ and Cdh1Wnt1/+;Wwp2GT/+ mice, showing the doomed skull, short and twisted nasal bone in Cdh1Wnt1/+;Wwp2GT/+ mice.
Discussion
Our previous study demonstrated that the E3 ligase Wwp2 could regulate craniofacial development through inducing a unique non-proteolytic ubiquitination of Gsc, which is required for the optimal activity of Gsc, a key transcription factor for craniofacial patterning4. Our current study extended this research and found that Gsc could be ubiquitinated and activated by another E3 ligase, APCCdh1. To assess how Cdh1 participates in craniofacial development, we generated Wnt1-Cre-mediated Cdh1 conditional knockout mice and found that Cdh1Wnt1 mice phenocopied Wwp2-deficient mice with shorter stature, lighter weight and abnormal craniofacial development (Figure 4). Consistently, the mRNA levels of various Gsc/Sox6 target genes were decreased in Cdh1Wnt1 mice, suggesting that similar to Wwp2, the APCCdh1 complex regulates craniofacial development through activating the Gsc/Sox6 signaling pathway. More strikingly, Cdh1/Wwp2 double heterozygous mice exhibited severe CFAs, although single heterozygous mice were normal (Figure 6), providing genetic evidence for the functional synergism between Cdh1 and Wwp2.
Cdh1 is a substrate recognition subunit of the APC complex, which can induce the timely ubiquitination and subsequent degradation of many cell cycle regulators24. The integrity of the whole APCCdh1 holoenzyme is important for the function of Cdh1 in cell cycle regulation. Consistent with this, Cdh1 associates with the APC core complex from early to mid G1 phase and regulates cell cycle progression primarily in controlling G1/S transition25,26,27. During the rest of the cell cycle, Cdh1 is largely APC free28. Our previous study has demonstrated that APC-free Cdh1 augments Smurf1 E3 ligase activity to govern osteoblast differentiation17. In the present study, we identified Gsc as a new substrate of APCCdh1 and further revealed a novel role for Cdh1 in craniofacial development through promoting APC-dependent, non-proteolytic ubiquitination and activation of Gsc.
It has been well characterized previously that APC promotes substrate ubiquitination in two distinct steps by associating with two different E2-conjugating enzymes, UBCH10 and Ube2S, which leads to initial monoubiquitination and subsequent K11-linkage polyubiquitination29. Previous studies have categorized APCCdh1 substrates into processive and distributive substrates30. For distributive substrates, instead of attaching a pre-formed polyubiquitin chain, APCCdh1 catalyzes the transfer of ubiquitin monomers, one at a time, to the substrate30,31. In further support of APCCdh1-mediated monoubiquitination of its substrates, a recent report revealed that cyclin B1 undergoes multiple monoubiquitinations at different lysine residues catalyzed by APC and UBCH10, which serve as destruction signals recognized by the 26S proteasome32. Taken together, our findings revealed that Gsc is a novel monoubiquitination substrate of APCCdh1. More importantly, unlike other known APCCdh1 substrates, monoubiquitination of Gsc mainly serves a non-proteolytic role, thus uncovering a new layer of Cdh1's biological functions.
Of note, we have recently found that Cdh1 suppresses Wwp2 E3 ligase activity in an APC-independent manner to modulate the PTEN/PI3K/Akt signaling pathway in cancer cells33. In the current study, however, we unveiled that in the chondrocyte setting, Cdh1 associates with the APC core complex and cooperates with Wwp2 to promote Gsc monoubiquitination. As most non-transformed somatic cells including chondrocytes are relatively low in Cdk kinase activity34, opposite to tumor cells, Cdh1 might be predominately in a hypo-phosphorylated form that favors its association with the APC core complex33. In support of this notion, we found that in ATDC5 cells, Cdh1 proteins existed mainly in the fractions with the molecular weight over 660 KDa, with little presence in the monomeric fractions33, suggesting that Cdh1 is predominantly associated with APC in ATDC5 cells. Consistently, we found that in ATDC5 cells, depletion of Cdh1 failed to alter the levels of Wwp2 or its downstream signaling factor PTEN (Supplementary information, Figure S4). The difference in the proportion of Cdh1 in association with the APC core complex, which is dictated by Cdk kinase activity, might explain the different functions of Cdh1 in tumor cells, suppressing Wwp2 activity in an APC-independent manner, and in chondrocytes, promoting the ubiquitination and activation of Gsc in an APC-dependent manner.
Genetic deletion of Gsc in mice results in perinatal lethality and skeletal patterning defects, including malformation of the nasal and frontal bones as well as abnormalities in the mandible and bones of the middle ear18,35. Gsc is known to contribute to craniofacial and skeletal development as well as early-stage organogenesis in various species, including Xenopus, Drosophila and human35,36,37,38,39,40. Gsc has also been revealed as a transcriptional suppressor for several genes and this suppressive activity is sensitive to SUMO modification36,38,40. Our previous work and the current study together demonstrate that Gsc could sct as a transcriptional activator and Gsc monoubiquitination is critical for its function as a transcriptional activator to drive the transcriptional program required for craniofacial development. In summary, our study reveals that a network of E3 ligases synergistically induce the non-proteolytic ubiquitination of Gsc, constituting a novel regulatory mechanism underlying Gsc activation, which is important for the regulation of craniofacial development.
Materials and Methods
Mouse models
Cdh1Wnt1 mice were generated by crossing Cdh1F/F mice with Wnt1-Cre mice22. The Cdh1F/F mice were from Cell Division and Cancer Group, Centro Nacional de Investigaciones Oncológicas (CNIO), Madrid, Spain. The Wnt1-Cre transgenic mice have been described previously22. Cdh1Wnt1/+;Wwp2GT/+ mice were generated by crossing Cdh1F/+;Wnt1-Cre mice with Wwp2GT/+ mice. Wwp2GT mice were described previously4. All Animals were maintained in pathogen-free conditions in accordance with the guidelines of the Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences.
Plasmids
pLKO-shRNA constructs against mouse Wwp2 and Cdh1 were purchased from Openbiosystems. His-Ubiquitin, HA-Cdh1, HA-Cdc20 and Myc-Cdh1 constructs were described previously17. Flag-Gsc, pRL-KT, Gal4-Gsc, (−238)Sox6pro-Luc and (−273) Sox6pro-Luc were described previously4. The targeting sequences of pLKO lentiviral expression vectors to knockdown mouse Cdh1 are: shCdh1-1 (sense: 5′-CGGCAGATCATCATCCAGAAT-3′), shCdh1-2 (sense: 5′-GATCTCTAAGATTCCCTTCAA-3′). The targeting sequences of pLKO lentiviral expression vectors to knockdown mouse Wwp2 are: shWwp2-1 (sense: 5′-CCGCTTTATCGGCAGATTCAT-3′), shWwp2-2 (sense: 5′-CTTCCGCTTTATCGGCAGATT-3′), shWwp2-3 (sense: 5′-GCAGCACTTCAGCCAAAGATT-3′). Site-directed mutagenesis to generate Flag-Gsc-RLAA was performed using the QuikChange XL Site-Directed Mutagenesis Kit (Stratagene) according to the manufacturer's instructions.
Antibodies
Anti-WWP2 antibody (sc-11897), anti-Gsc antibody (sc-22234), monoclonal anti-HA antibody (sc-7392), anti-Cyclin A antibody (sc-751) and Protein A/G PLUS-Agarose (sc-2003) were purchased from Santa Cruz Biotechnology. Anti-Cdh1 antibody (QB214618) was purchased from Thermo Scientific. Monoclonal anti-Flag antibody (F-3165), anti-Flag agarose beads (A-2220) and anti-HA agarose beads (A-2095) were purchased from Sigma. Anti-Myc antibody (AE010) was purchased from ABclonal.
Cell Culture, transfection and infection
Cell culture conditions including transfection have been described previously17. ATDC5 cell culture condition was described previously4. Lentiviral shRNA virus packaging and subsequent infection of various cell lines were performed according to the protocol described previously41.
Immunoblots and immunoprecipitation
Cells were lysed in the EBC buffer (50 mM Tris (pH 7.5), 120 mM NaCl and 0.5% NP-40) supplemented with protease inhibitors (Complete Mini, Roche) and phosphatase inhibitors (phosphatase inhibitor cocktail set I and II, Calbiochem). The protein concentrations of the lysates were measured using the Bio-Rad protein assay reagent on a Beckman Coulter DU-800 spectrophotometer. The lysates were then resolved by SDS-PAGE and immunoblotted with the indicated antibodies. For immunoprecipitation, 800 μg lysates were incubated with the appropriate antibodies (1-2 μg) for 3-4 h at 4 °C followed by 1-h incubation with Protein A/G plus agarose (Santa Cruz). Immunoprecipitated products were washed five times with the NETN buffer (20 mM Tris (pH 8.0), 100 mM NaCl, 1 mM EDTA and 0.5% NP-40) before being resolved by SDS-PAGE and immunoblotted with the indicated antibodies.
In vitro binding assays
Binding to immobilized GST proteins was performed as described previously12,42. Briefly, GST-Cdh1 proteins were expressed in the BL21 E. coli strain followed by purification using Glutathione Sepharose 4B (GE Healthcare #17-0756-01) according to the manufacturer's instruction. Purified GST and GST-Cdh1 proteins were incubated with 35S-labled in vitro transcribed and translated Gsc in the NETN buffer (20 mM Tris, pH 8.0, 100 mM NaCl, 1 mM EDTA and 0.5% NP-40) for 3-4 h at 4 °C followed by washing with the NETN buffer before being resolved by SDS-PAGE and immunoblotted with the indicated antibodies.
Real-time RT-PCR analysis
The cDNA synthesis was performed with the PrimeScript RT reagent Kit (Perfect Real Time; RR037A) and SYBR green qPCR components from Takara. The real-time reverse transcriptase (RT)-PCR reaction was performed with the Bio-Rad qRT system. The primer sets used were as follows: Cdh1 (general use), sense 5′-CGTGTACCTGTGGAGTGCAT-3′ and antisense 5′-ACGAAGCCCTTGTGTGTACC-3; Cdh1 (for knockout efficiency), sense 5′-GGCTCCTGCGGCAGATCA-3′ and antisense 5′-TGGGTGAAGACACTGGGGAG-3′; Wwp2, sense 5′-AAGTGGAGCGGAGTTAGGC-3′ and antisense 5′-AAGCTGGGACTTCTCAAAAGG-3′; Gsc, sense 5′-GCACCATCTTCACCGATGAG-3′ and antisense 5′-AGGAGGATCGCTTCTGTCGT-3′; Sox6, sense 5′-GACAGCGTTCTGTCATCTCAGCAA-3′ and antisense 5′-CGTTCCGGGGTTCCAAAAGTAACA-3′; Col2a1, sense 5′-CGGTCCTACGGTGTCAGG-3′ and antisense 5′-GCAGAGGACATTCCCAGTGT-3′; Col9a2, sense 5′-TATAAAGGCATGGTGGGCTC-3′ and antisense 5′-GTTCCCTTTGGGCCTGTTAT-3′; Col11a2, sense 5′-AGTCCCTTGCCATTCCTTG-3 and antisense 5′-GGGGGTCCCTCTACAAACAT-3′; Agc1, sense 5′-GCCAGGGGGAGTTGTATTCC-3′ and antisense 5′-TCACACTGCTCATAGCCTGC-3′; S100A1, sense 5′-GTGCCCTTCTGTCGAGAATC-3′ and antisense 5′-CACATTGATGAGGGTCTCCA-3′; S100B, sense 5′-TTACTCGGACACTGAAGCCA-3′ and antisense 5′-TTCAGCTTGTGCTTGTCACC-3′; Matn1, sense 5′-GTCCACGGGTACCATGACTG-3′ and antisense 5′-CTCCCATCAGTCACCACGAT-3′; Gapdh, sense 5′-TCGTCCCGTAGACAAAATGG-3′ and antisense 5′-TTGAGGTCAATGAAGGGGTC-3′.
Luciferase reporter assays
Cells were transfected with luciferase reporter plasmids and different combinations of expression constructs. Total amounts of transfected DNA were kept constant by supplementing with control empty expression vector plasmids as needed. All cells were co-transfected with pRL-TK (Promega) as a normalization control for transfection efficiency. Forty-eight hours after transfection, cells were collected and lysed in 1× passive lysis buffer (Promega). Luciferase assays were performed using the Dual-Luciferase Reporter Assay System (Promega).
Ubiquitination analysis in cells
APCCdh1-mediated Gsc ubiquitination was analyzed as described previously4,17. Briefly, 293T cells were transfected with Flag-Gsc, His-ubiquitin and HA-Cdh1. Thirty-six hours after the transfection, 10 μM MG132 was added to block proteasomal degradation, and cells were harvested in EBC buffer containing protease inhibitors. Whole-cell lysates (2 mg) were incubated with Ni-NTA beads for 4 h, followed by washing four times with NETN buffer. Then, the washed pellet was boiled in SDS-containing sample buffer and resolved by SDS-PAGE.
In vitro ubiquitination analysis
Gsc in vitro ubiquitination assays were performed as described previously17. The assay was performed in 20 μl ubiquitination assay buffer (50 mM Tris-HCl (pH 8.0), 50 mM NaCl, 1.5 mM dithiothreitol, 50 mM MgCl2 and 10 mM ATP), with 0.5 μg of E1, 1 μg of UbcH10 (E2) and 1.5 μg of HA-ubiquitin (Boston Biochem). 150 ng GST-Gsc proteins (WT or RLAA) together with 500 ng of APCCdh1 were added to the buffer to initiate the reaction. Samples were incubated at 30 °C for 15 min. The reactions were stopped by the addition of 2 × SDS-PAGE sample buffer, and the reaction products were resolved by SDS-PAGE and probed with the indicated antibodies.
μCT analysis of mouse skulls
Skulls isolated from age- and sex-matched 5-week-old mice were fixed in 70% ethanol and scanned using Quantum GX (PerkinElmer) with a spatial resolution of 24 μm. The scans were reconstructed into two-dimensional coronal slices. From these slices, we compared the distances among seven different landmarks individually as described previously4.
Statistical analysis
All quantitative data were presented as the mean ± SEM or the mean ± SD as indicated of at least three independent experiments. P < 0.05 was considered as statistically significant (Student's t-test).
Author Contributions
RS, JL and LW performed most of the experiments with assistance from GY, JZ, YH, JG, ZX, ZY, JL and MM. LW, WW and WZ designed the experiments and supervised the study. RS, LW and WZ wrote the manuscript with help from WW. All authors commented on the manuscript.
Competing Financial Interests
The authors declare no competing financial interests.
References
Alappat S, Zhang ZY, Chen YP . Msx homeobox gene family and craniofacial development. Cell Res 2003; 13:429–442.
Rice DP . Craniofacial anomalies: from development to molecular pathogenesis. Curr Mol Med 2005; 5:699–722.
Rotin D, Kumar S . Physiological functions of the HECT family of ubiquitin ligases. Nat Rev Mol Cell Biol 2009; 10:398–409.
Zou W, Chen X, Shim JH, et al. The E3 ubiquitin ligase Wwp2 regulates craniofacial development through mono-ubiquitylation of Goosecoid. Nat Cell Biol 2011; 13:59–65.
Peters JM . The anaphase promoting complex/cyclosome: a machine designed to destroy. Nat Rev Mol Cell Biol 2006; 7:644–656.
Burton JL, Tsakraklides V, Solomon MJ . Assembly of an APC-Cdh1-substrate complex is stimulated by engagement of a destruction box. Mol Cell 2005; 18:533–542.
Pfleger CM, Kirschner MW . The KEN box: an APC recognition signal distinct from the D box targeted by Cdh1. Genes Dev 2000; 14:655–665.
Garcia-Higuera I, Manchado E, Dubus P, et al. Genomic stability and tumour suppression by the APC/C cofactor Cdh1. Nat Cell Biol 2008; 10:802–811.
Bassermann F, Frescas D, Guardavaccaro D, Busino L, Peschiaroli A, Pagano M . The Cdc14B-Cdh1-Plk1 axis controls the G2 DNA-damage-response checkpoint. Cell 2008; 134:256–267.
Fujita T, Liu W, Doihara H, Date H, Wan Y . Dissection of the APCCdh1-Skp2 cascade in breast cancer. Clin Cancer Res 2008; 14:1966–1975.
Fujita T, Liu W, Doihara H, Wan Y . Regulation of Skp2-p27 axis by the Cdh1/anaphase-promoting complex pathway in colorectal tumorigenesis. Am J Pathol 2008; 173:217–228.
Gao D, Inuzuka H, Korenjak M, et al. Cdh1 regulates cell cycle through modulating the claspin/Chk1 and the Rb/E2F1 pathways. Mol Biol Cell 2009; 20:3305–3316.
Tudzarova S, Colombo SL, Stoeber K, Carcamo S, Williams GH, Moncada S . Two ubiquitin ligases, APC/C-Cdh1 and SKP1-CUL1-F (SCF)-beta-TrCP, sequentially regulate glycolysis during the cell cycle. Proc Natl Acad Sci USA 2011; 108:5278–5283.
Naoe H, Araki K, Nagano O, et al. The anaphase-promoting complex/cyclosome activator Cdh1 modulates Rho GTPase by targeting p190 RhoGAP for degradation. Mol Cell Biol 2010; 30:3994–4005.
Lasorella A, Stegmuller J, Guardavaccaro D, et al. Degradation of Id2 by the anaphase-promoting complex couples cell cycle exit and axonal growth. Nature 2006; 442:471–474.
Stegmuller J, Konishi Y, Huynh MA, Yuan Z, Dibacco S, Bonni A . Cell-intrinsic regulation of axonal morphogenesis by the Cdh1-APC target SnoN. Neuron 2006; 50:389–400.
Wan L, Zou W, Gao D, et al. Cdh1 regulates osteoblast function through an APC/C-independent modulation of Smurf1. Mol Cell 2011; 44:721–733.
Yamada G, Mansouri A, Torres M, et al. Targeted mutation of the murine goosecoid gene results in craniofacial defects and neonatal death. Development 1995; 121:2917–2922.
Lefebvre V, Li P, de Crombrugghe B . A new long form of Sox5 (L-Sox5), Sox6 and Sox9 are coexpressed in chondrogenesis and cooperatively activate the type II collagen gene. EMBO J 1998; 17:5718–5733.
Saito T, Ikeda T, Nakamura K, Chung UI, Kawaguchi H . S100A1 and S100B, transcriptional targets of SOX trio, inhibit terminal differentiation of chondrocytes. EMBO Rep 2007; 8:504–509.
Nagy A, Kenesi E, Rentsendorj O, et al. Evolutionarily conserved, growth plate zone-specific regulation of the matrilin-1 promoter: L-Sox5/Sox6 and Nfi factors bound near TATA finely tune activation by Sox9. Mol Cell Biol 2011; 31:686–699.
Danielian PS . Modification of gene activity in mouse embryos in uteroby a tamoxifen-inducible form of Cre recombinase. Curr Biol 1998; 8:1323–1326.
Surdenikova L, Ru F, Nassenstein C, Tatar M, Kollarik M . The neural crest- and placodes-derived afferent innervation of the mouse esophagus. Neurogastroenterol Motil 2012; 24:e517–525.
da Fonseca PC, Kong EH, Zhang Z, et al. Structures of APC/C(Cdh1) with substrates identify Cdh1 and Apc10 as the D-box co-receptor. Nature 2011; 470:274–278.
Geley S, Kramer E, Gieffers C, Gannon J, Peters JM, Hunt T . Anaphase-promoting complex/cyclosome-dependent proteolysis of human cyclin A starts at the beginning of mitosis and is not subject to the spindle assembly checkpoint. J Cell Biol 2001; 153:137–148.
Petersen BO, Wagener C, Marinoni F, et al. Cell cycle- and cell growth-regulated proteolysis of mammalian CDC6 is dependent on APC-CDH1. Genes Dev 2000; 14:2330–2343.
Zur A, Brandeis M . Securin degradation is mediated by fzy and fzr, and is required for complete chromatid separation but not for cytokinesis. EMBO J 2001; 20:792–801.
Hu D, Qiao X, Wu G, Wan Y . The emerging role of APC/CCdh1 in development. Semin Cell Dev Biol 2011; 22:579–585.
Meyer HJ, Rape M . Processive ubiquitin chain formation by the anaphase-promoting complex. Semin Cell Dev Biol 2011; 22:544–550.
Rape M, Reddy SK, Kirschner MW . The processivity of multiubiquitination by the APC determines the order of substrate degradation. Cell 2006; 124:89–103.
Wu T, Merbl Y, Huo Y, Gallop JL, Tzur A, Kirschner MW . UBE2S drives elongation of K11-linked ubiquitin chains by the anaphase-promoting complex. Proc Natl Acad Sci USA 2010; 107:1355–1360.
Dimova NV, Hathaway NA, Lee BH, et al. APC/C-mediated multiple monoubiquitylation provides an alternative degradation signal for cyclin B1. Nat Cell Biol 2012; 14:168–176.
Liu J, Wan L, Liu J, et al. Cdh1 inhibits WWP2-mediated ubiquitination of PTEN to suppress tumorigenesis in an APC-independent manner. Cell Discov 2016; 2:15044. doi:10.1038/celldisc.2015.44
Gao CY, Zelenka PS . Cyclins, cyclin-dependent kinases and differentiation. Bioessays 1997; 19:307–315.
Rivera-Perez JA, Mallo M, Gendron-Maguire M, Gridley T, Behringer RR . Goosecoid is not an essential component of the mouse gastrula organizer but is required for craniofacial and rib development. Development 1995; 121:3005–3012.
Boucher DM, Schaffer M, Deissler K, et al. goosecoid expression represses Brachyury in embryonic stem cells and affects craniofacial development in chimeric mice. Int J Dev Biol 2000; 44:279–288.
Hahn M, Jackle H . Drosophila goosecoid participates in neural development but not in body axis formation. EMBO J 1996; 15:3077–3084.
Latinkic BV, Smith JC . Goosecoid and mix.1 repress Brachyury expression and are required for head formation in Xenopus. Development 1999; 126:1769–1779.
Tassabehji M, Hammond P, Karmiloff-Smith A, et al. GTF2IRD1 in craniofacial development of humans and mice. Science 2005; 310:1184–1187.
Yao J, Kessler DS . Goosecoid promotes head organizer activity by direct repression of Xwnt8 in Spemann's organizer. Development 2001; 128:2975–2987.
Boehm JS, Hession MT, Bulmer SE, Hahn WC . Transformation of human and murine fibroblasts without viral oncoproteins. Mol Cell Biol 2005; 25:6464–6474.
Wei W, Ayad NG, Wan Y, Zhang GJ, Kirschner MW, Kaelin WG, Jr . Degradation of the SCF component Skp2 in cell-cycle phase G1 by the anaphase-promoting complex. Nature 2004; 428:194–198.
Acknowledgements
We thank Brian North, Shavali Shaik, Alan W. Lau, Matthew B. Greenblatt, Hiroyuki Inuzuka and Pengda Liu for critical reading of the manuscript, and members of the Zou and Wei labs for useful discussions. This work was supported by the Chinese Ministry of Science and Technology (MOST, 973 Program; 2014CB964704 and 2015CB964503), the National Natural Science Foundation of China (NSFC; 31371463) and NIH (GM089763, GM094777 and CA177910 to WW). WZ is a scholar of the 'National 1000 Young Talents Program of China' and 'the National Science Fund for Excellent Young Scholars' (NSFC; 81322027). WW is a Leukemia and Lymphoma Society Scholar and ACS Research Scholar.
Author information
Authors and Affiliations
Corresponding authors
Additional information
( Supplementary information is linked to the online version of the paper on the Cell Research website.)
Supplementary information
Supplementary information, Figure S1
Luciferase reporter constructs used in different luciferase assay. (PDF 410 kb)
Supplementary information, Figure S2
Cdh1 conditional knockout led to craniofacial development defects. (PDF 8349 kb)
Supplementary information, Figure S3
Cre virus did not induce Cdh1 knockdown in calvarial cells with WT-Cdh1 genetic background. (PDF 129 kb)
Supplementary information, Figure S4
Immunoblot analysis of whole cell lysates derived from ATDC5 cells infected with the indicated lentiviral constructs. (PDF 500 kb)
Rights and permissions
About this article
Cite this article
Shao, R., Liu, J., Yan, G. et al. Cdh1 regulates craniofacial development via APC-dependent ubiquitination and activation of Goosecoid. Cell Res 26, 699–712 (2016). https://doi.org/10.1038/cr.2016.51
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2016.51
Keywords
This article is cited by
-
The role of WWP1 and WWP2 in bone/cartilage development and diseases
Molecular and Cellular Biochemistry (2024)
-
The role of E3 ubiquitin ligase WWP2 and the regulation of PARP1 by ubiquitinated degradation in acute lymphoblastic leukemia
Cell Death Discovery (2022)
-
Gene regulatory network from cranial neural crest cells to osteoblast differentiation and calvarial bone development
Cellular and Molecular Life Sciences (2022)
-
Beraprost ameliorates postmenopausal osteoporosis by regulating Nedd4-induced Runx2 ubiquitination
Cell Death & Disease (2021)
-
Reply to ‘Dissecting the role of miR-140 and its host gene’
Nature Cell Biology (2018)