Abstract
The sterol regulatory element-binding protein (SREBP) and SREBP cleavage-activating protein (SCAP) are central players in the SREBP pathway, which control the cellular lipid homeostasis. SCAP binds to SREBP through their carboxyl (C) domains and escorts SREBP from the endoplasmic reticulum to the Golgi upon sterol depletion. A conserved pathway, with the homologues of SREBP and SCAP being Sre1 and Scp1, was identified in fission yeast Schizosaccharomyces pombe. Here we report the in vitro reconstitution of the complex between the C domains of Sre1 and Scp1 as well as the crystal structure of the WD40 domain of Scp1 at 2.1 Å resolution. The structure reveals an eight-bladed β-propeller that exhibits several distinctive features from a canonical WD40 repeat domain. Structural and biochemical characterization led to the identification of two Scp1 elements that are involved in Sre1 recognition, an Arg/Lys-enriched surface patch on the top face of the WD40 propeller and a 30-residue C-terminal tail. The structural and biochemical findings were corroborated by in vivo examinations. These studies serve as a framework for the mechanistic understanding and further functional characterization of the SREBP and SCAP proteins in fission yeast and higher organisms.
Similar content being viewed by others
Introduction
The sterol regulatory element-binding proteins (SREBPs) play a pivotal role in lipid homeostasis by controlling the synthesis of cholesterol, fatty acids, and triglycerides1,2. In addition, mounting evidence suggests critical roles for SREBPs in diabetes, immune responses, and cancer3. SREBPs represent a unique family of basic helix-loop-helix leucine zipper transcription factors that are synthesized as endoplasmic reticulum (ER) membrane-anchored precursors4. The SREBP cleavage-activating protein (SCAP) forms a complex with the SREBP precursor through their C-terminal domains. Upon sterol depletion, SCAP facilitates the translocation of SREBPs from the ER to the Golgi5, where SREBPs are first cleaved by the Site-1 protease (S1P) in the luminal loop, producing an intermediate in which the N-terminal transcription factor remains attached to the membrane and a C-terminal fragment attached to SCAP. The N-terminal membrane-attached intermediate is further cleaved by Site-2 protease (S2P), liberating the transcription factor from the membrane. SCAP recycles from the Golgi to the ER6,7. C-terminal SREBP fragments are believed to be removed from the bound SCAP through an unknown mechanism. Then SCAP binds to newly-synthesized SREBP precursor for additional rounds of SREBP transport and cleavage.
The SREBP pathway is also present in fungi exemplified by fission yeast Schizosaccharomyces pombe (S. pombe)8,9, where low oxygen activates SREBP to promote hypoxic adaptation10,11. Sre1 and Scp1 are the yeast orthologues of mammalian SREBP and SCAP, respectively11. Despite low sequence identities, Sre1 and Scp1 were predicted to share identical topologies with their respective mammalian homologues (Figure 1A). S1P and S2P are absent from fission yeast8. Rather, Sre1 cleavage occurs through a different mechanism that requires the Golgi Dsc E3 ubiquitin ligase complex12. Despite the variation for Sre1 proteolytic cleavage, the mechanism of sterol-regulated ER-to-Golgi transport of SCAP11 and SREBP cleavage-triggered Golgi-to-ER recycling of SCAP6 is conserved.
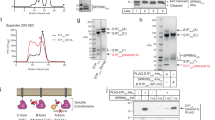
In vitro reconstitution of the Sre1-Scp1 cytosolic complex. (A) Schematic illustration of the domain organizations of Scp1 and Sre1. Transmembrane helices are colored red and the carboxyl terminal tail of Scp1 is colored green. “SSD” stands for sterol sensing domain. The eight WD40 repeats of Scp1 are numbered 1-8. (B) Purified recombinant proteins of the C-terminal domains of Scp1 and Sre1 form complex in vitro. The MBP-fused regulatory domain of Sre1 (residues 628-900, and named MBP-Sre1) was immobilized on amylose resin to pull down Scp1-WD40 (residues 567-1 085). MBP was tested as negative control. Details of the experiments can be found in Materials and Methods.
Both mammalian and fungal SCAP proteins consist of an amino terminal domain of eight transmembrane helices (TM) and a carboxyl (C) terminal WD40 domain11,13, TMs 2-6 of SCAP constitute a sterol-sensing domain14, which is also found in several membrane proteins involved in sterol homeostasis and cell signaling such as HMG-CoA reductase15, Niemann-Pick C116,17, patched18 and dispatched19. The cytosolic WD40 domain of SCAP interacts with the C-terminal regulatory domain of SREBPs to achieve the escort function20. Notably, the WD40 domains of mammalian SCAPs and fungal Scp1 consist of more than 500 amino acids, considerably larger than canonical WD40 domains that typically contain 300-400 residues21.
Despite the essential function of SREBP and SCAP in the regulation of lipid homeostasis22,23, structural information of these proteins is lacking due to technical challenges concerning the manipulation of recombinant proteins. For the same reason, the complex between SREBP and SCAP has never been reconstituted in vitro using purified recombinant proteins. In this study, we recapitulated the complex formation between the C-terminal cytoplasmic domains of Scp1 and Sre1 using recombinant proteins purified to homogeneity. We also report the high-resolution crystal structure of the WD40 domain of S. pombe Scp1. Structure-guided in vitro and in vivo mutational analyses allowed identification of Scp1 residues that are essential for the interaction with Sre1.
Results
Reconstitution of the Sre1-Scp1 complex in vitro
To obtain soluble and well-behaved recombinant proteins of the C-terminal cytoplasmic domains of SREBP and SCAP for biochemical and structural analysis, we screened homologues from more than 10 species ranging from yeast to human. We also exploited multiple expression systems including E. coli, yeasts, and baculovirus-infected insect cells. After extensive effort, only the C-terminal segments of Sre1 and Scp1 from S. pombe yielded soluble recombinant proteins. The Scp1-WD40 domain expressed in Sf9 insect cells yielded well-behaved proteins, whereas soluble C-terminal domain of Sre1 could be obtained from E. coli. Importantly, the purified recombinant proteins formed a complex in vitro, supporting a direct interaction of the C-terminal domains between Sre1 and Scp1 free of any other co-factors (Figure 1B).
Structure determination of the Scp1-WD40 domain
We then sought to resolve the structure of the Sre1-Scp1 complex, as well as the individual components. Despite rigorous trials, these proteins defied our crystallization efforts. To solve this problem, we launched a systematic protein engineering strategy and finally determined the high-resolution crystal structure of Scp1-WD40.
Briefly, limited proteolysis and sequence alignment analysis of Scp1 suggested that the less conserved fragment (residues 962-985) could be flexible (Supplementary information, Figure S1). Supporting this notion, co-expression of the two fragments containing residues 567-961 and 986-1 085 in E. coli yielded well-behaved proteins that retained interaction with Sre1. Using this construct (hereafter named Scp1-Δ24), further C-terminal truncation and substitution of cysteine residues were screened. Eventually, the engineered Scp1-WD40 protein (residues 567-961 and 986-1 054, C618S/C671S/C680S/C756S/C873S/C901S/C920S/C941S/C1010S; Figure 2A) was crystallized and the crystals diffracted X-rays beyond 2 Å resolution. The structure of Scp1-WD40 was solved by single anomalous diffraction of selenium (Se-SAD) and the final atomic model was refined to 2.1 Å resolution (Figure 2B, Supplementary information, Figure S2 and Table S1). In the final structural model, the visible segments contain residues 567-941 and 992-1 054.
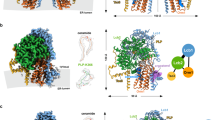
Crystal structure of the Scp1 WD40 domain. (A) Protein purification of the Scp1-WD40 protein (residues 567-961 and 986-1 054, C618S/C671S/C680S/C756S/C873S/C901S/C920S/C941S/C1010S) used for crystallization studies. Two fragments (residues 567-961 and 986-1 054) were co-expressed in E.coli. A representative chromatogram of size exclusion chromatography of the recombinant Scp1-WD40 is shown. Peak fractions were analyzed by SDS-PAGE and visualized by Coomassie blue staining. (B) Overall structure of the Scp1 WD40 domain. Two perpendicular views are shown. The structure is shown in rainbow color, with the N- and C-termini colored blue and red, respectively. The eight blades are numbered 1 to 8 and the four strands in each blade are labeled A to D from the center to the outer ring. All structure figures were prepared with PyMol31.
Structural analysis of Scp1-WD40
In contrast to the sequence-based prediction of five or seven WD40 repeats in mammalian SCAPs and yeast Scp111,13,24, the crystal structure of Scp1-WD40 revealed an eight-bladed β-propeller. Following the convention for illustrating WD40 propellers, these blades are numbered 1 to 8, and the four strands within each blade are labeled A, B, C, and D from the center to the outer ring of the propeller (Figure 2B, Supplementary information, Figure S1). The top and bottom surfaces of the propeller are formed by BC/DA loops and AB/CD loops, respectively. Similar to the canonical WD40 propeller21, strand D of blade 8 (strand 8D) is formed by sequences preceding strand A of blade 1 (strand 1A; Figure 2B). Although Scp1-WD40 consists of more residues than most eight-bladed WD40 domains, the core structure is similar to other eight-bladed WD40 domains such as that of Sif225 (Supplementary information, Figure S3). The number of residues in each blade of the Scp1-WD40 propeller varies considerably due to the differences in the length of strands and their intervening loops (Supplementary information, Figure S1). Consequently, the structure of Scp1-WD40 has a number of unique features compared to canonical WD40 propellers.
In the structure of Scp1-WD40, an extended N-terminal segment (residues 570-573) preceding strand 8D forms an antiparallel strand with strand 7D (Figure 2B). Blades 1/2/4/5/8 of Scp1-WD40 are similar to each other, with root-mean-square deviation values of ∼0.6-0.8 Å when superimposed pairwise (Figure 3A). Blades 3 and 7 have longer B/C/D strands protruding from the bottom and top faces, respectively (Figure 3B). Blade 6 is most divergent among the eight blades in that it has exceptionally elongated AB and CD loops at the bottom face (Figure 3C). The CD loop of blade 6 (6CD loop), consisting of 39 amino acids (residues 870-908), adopts a twisted conformation in a shape similar to “8”. The electron density of this loop is well resolved, indicating a relatively stable conformation. Indeed, 6CD loop is stabilized through extensive interactions with the 6AB loop. These unusual features of blades 3/6/7 result in the overall asymmetric contour of the WD40 propeller (Figure 2B).
Structural comparison of the eight blades in the Scp1-WD40 β-propeller. (A) Structural superimposition of the blades 1/2/4/5/8. These five blades share similar repeat structures. (B) Blades 3 and 7 have elongated B/C/D strands. Shown here are structural superimpositions of blade 1 with blades 3 (left) and 7 (right). (C) Blade 6 has elongated AB and CD loops extending out of the bottom face. Left: structural comparison between blades 1 and 6. The 6AB and 6CD loops are highlighted in cyan and purple, respectively. Right: The 2Fo-Fc omit electron density map of 6CD loop (residues 870-908), contoured at 1σ, is shown as blue mesh.
Identification of the Scp1 residues for Sre1 binding
The successful in vitro reconstitution of the Sre1-Scp1 complex, as well as structure determination of Scp1-WD40 provided an opportunity to examine the recognition mechanism between Sre1 and Scp1. We had two serendipitous discoveries during protein engineering. First, truncation of the 30 C-terminal residues of Scp1 led to reduced binding with Sre1 (Figure 4C, Scp1-M6, Lane 15). Second, Scp1 variant (C618S/C680S) appeared to have decreased binding affinity with Sre1 (Figure 4C, Scp1-M4, Lane 13). As the C-terminal fragment is missing, Ser618 and Ser680 can be accurately mapped on the structure. These two residues are located on the top face of blades 1 and 2 of the propeller and distanced by ∼25 Å (Figure 4A). Interestingly, electrostatic surface potential analysis reveals that the two residues are intervened by a continuous patch of positive potentials that are constituted by a cluster of Arg/Lys residues. We name this surface region the “RK patch” (Figure 4A).
Identification of Sre1-binding surface based on the Scp1-WD40 structure. (A) The top surface of the Scp1-WD40 propeller. The left and right panels depict the electrostatic surface potential and the corresponding cartoon representations, respectively. Six basic residues from blades 1 and 2 constitute a positively charged surface patch, the “RK patch”. (B) A close-up view on the residues involved in Sre1 binding. Six basic residues, one acidic residue and two Ser residues are colored blue, red and yellow, respectively. (C) Interaction between Scp1-WD40 variants and Sre1 regulatory domain assessed by MBP-mediated pull-down assay. The WT (residues 567-1 085) and Scp1-Δ24 were included as control. Scp1-M1/M2/M3 almost completely lost association with Sre1 (Lanes 10-12, boxed in red). Scp1-M4/M5/M6 showed decreased binding with Sre1 (Lanes 13-15, boxed in green). The experiments were performed with the same protocol as that for Figure 1B.
To test whether the RK patch is involved in Sre1 recognition, we generated three Scp1-WD40 variants (Scp1-M1: R617E; Scp1-M2: K635E/R640E/K643E; Scp1-M3: K659E/K685E) on the basis of Scp1-Δ24. The three mutant proteins, which exhibited similar yield and solution behavior to the wild-type (WT) protein (Supplementary information, Figure S4), were purified to homogeneity. All three variants almost completely lost binding with Sre1 when tested in the pull-down assay (Figure 4C, Lanes 10-12). These observations support the notion that the RK patch is essential for Sre1 recognition. Interestingly, substitution of a negatively charged residue Asp1023 on blade 8 (Scp1-M5: D1023K) also led to decreased complex formation (Figure 4B and 4C, Lane 14), implying an extended Sre1-binding surface on the top side of Scp1-WD40 propeller.
To look for additional surface elements that may contribute to Scp1-Sre1 complex formation, we generated more surface mutations (Supplementary information, Figure S5). Mutation of the charged residues in the vicinity of Asp1023, including E862K (Supplementary information, Figure S5C, Scp1-M7) and D916K/E919K (Supplementary information, Figure S5C, Scp1-M8) had no detectable impact on Sre1 binding, suggesting that Asp1023 may demarcate the Sre1-binding surface of Scp1 on one side (Supplementary information, Figure S5A). Substitutions of two Glu residues on the top surface of blade 3, E759K/E762K (Supplementary information, Figure S5C, Scp1-M9), also showed no effect on complex formation. Scp1 variants with residue substitutions at the bottom face, including D697K/D698K/E699K (Supplementary information, Figure S5C, Scp1-M10) and E722K/E724K (Supplementary information, Figure S5C, Scp1-M11), showed no difference in Sre1 association either. These results further support the specific recognition of Sre1 by Scp1 through the RK patch.
Functional analysis of Scp1 mutants in vivo
To characterize the Sre1-Scp1 interaction in a cellular context, we generated fission yeast strains expressing Myc epitope-tagged scp1 WT or M1-M6 mutants from the endogenous scp1+ promoter. Our previous studies demonstrated that Scp1 was required to stabilize Sre1 precursor and activate Sre1 processing. scp1Δ cells failed to grow in cobalt chloride-containing medium and anaerobic Sre1N activation was abolished in scp1Δ cells11. In addition, in scp1Δ cells, Sre1 precursor was rapidly degraded in the ER and the degradation required Hrd1, an E3 ubiquitin ligase26. Based on these findings, we expected that Scp1 mutants defective for Sre1 binding would (i) show reduced Sre1 precursor due to Hrd1-dependent degradation, (ii) fail to accumulate Sre1N under low oxygen, and (iii) fail to grow in the presence of cobalt chloride due to reduced activation of Sre1.
We first assayed Sre1 precursor levels. Consistent with the in vitro analysis (Supplementary information, Figure S4), the expression level of scp1 mutants was similar to that of the WT (Figure 5A, Myc blot), supporting the notion that the introduction of mutations did not result in mis-folded products. As expected, scp1 mutants M1, M2, and M3 that failed to bind Sre1 showed reduced level of Sre1 precursor (Figure 5A, lanes 2-4). Interestingly, while Scp1 variants M4-M6 showed decreased binding to Sre1 in the in vitro test, Sre1 precursor level remained almost unchanged in Scp1 mutants M4-M6 (Figure 5A, lanes 5-7). Deletion of hrd1+ restored Sre1 precursor level in mutants M1, M2 and M3 (Figure 5B, lanes 3-8), suggesting that the reduction of Sre1 precursor resulted from Hrd1-dependent degradation in the ER. Consistent with these data, mutants M1-M3 failed to grow in medium containing cobalt (Figure 5C) or activate Sre1 under low oxygen (Figure 5D, lanes 3-8), while M4 and M5 behaved like WT cells in these assays. Intriguingly, although mutant M6 showed normal levels of Sre1 precursor (Figure 5A, lane 7), this mutant failed to accumulate Sre1 under low oxygen (Figure 5D, lanes 13-14) and grow on cobalt medium (Figure 5C), indicating that the C-terminal tail of Scp1 (residues 1 055-1 085) is required for Sre1 activation. These cellular data demonstrate that the RK patch of Scp1 is essential for Sre1 binding and subsequent activation, corroborating the in vitro binding studies.
Functional analysis of Scp1 mutants in vivo. (A) Western blot of membrane fractions from WT and the indicated scp1 mutant S. pombe strains probed with anti-Sre1 serum, anti-Myc IgG or anti-Dsc5 serum. (B) Western blot of whole-cell lysates from WT and the indicated scp1 mutant S. pombe strains in the presence or absence of hrd1+ probed with anti-Sre1 serum or anti-Dsc5 serum. P denotes Sre1 precursor form. Asterisks indicate cross-reacting protein. (C) WT and mutant yeast (5 000, 1 000, 200, 40, and 8 cells) were grown on rich medium (3 days) or rich medium containing cobalt chloride (1.6 mM; 6 days). (D) Western blot of whole-cell lysates from WT and the indicated scp1 mutant S. pombe strains grown in the presence or absence of oxygen for 3 h probed with anti-Sre1 serum or anti-Dsc5 serum. P and N denote Sre1 precursor and nuclear forms, respectively. Asterisks indicate cross-reacting proteins. (E) Two potential models for Sre1-Scp1 complex formation.
Discussion
WD40 domains are widely-distributed scaffold modules that participate in a variety of biological processes27. They provide distinct surface features, including the top, the bottom, and the side surfaces, for interaction with diverse ligands involving proteins, peptides, and nucleic acids21. In this study, we determined the high-resolution crystal structure of the eight-bladed WD40 domain of Scp1 and identified two Sre1-binding elements that were verified by in vitro and in vivo examinations. An Arg/Lys-enriched surface patch (the RK patch) that is located on the top side of the propeller is essential for Sre1 recognition. The C-terminal tail (residues 1 055-1 085) also appears to participate in Sre1 binding (Figure 4C).
In the crystal structure of Scp1-WD40, the last visible residue is Phe1054, which is placed at the outer edge of the bottom face. There is one possibility that the C-tail of Scp1 can extend to the top surface and stay in the vicinity of the RK patch to collectively constitute one single Sre1-binding surface (Figure 5E, left). Alternatively, the C-terminal tail may represent an independent binding element that recognizes a separate Sre1 surface (Figure 5E, right). The detailed recognition mechanism awaits the determination of the complex structure between Sre1 and Scp1.
Like mammals, S. pombe Scp1 was required to stabilize Sre1 precursor in the ER and activate Sre1 processing. The RK patch scp1 mutants defective for Sre1 binding in the pull-down assay (M1, M2, and M3) phenocopied scp1Δ, demonstrating that the RK patch in Scp1 WD40 domain plays an essential role in Scp1-Sre1 interaction. The mutants M4 and M5 showed compromised Sre1 binding in the pull-down assay, but no effect in the in vivo examination, suggesting that slight changes in the binding affinity between Scp1 and Sre1 may not affect their functions under physiological condition. scp1 mutant M6 was particularly interesting. The fact that Scp1-M6 stabilized Sre1 precursor in the ER but failed to activate Sre1 processing suggests that the C-terminal 30 amino acids are critical for Sre1 processing. Sre1 activation requires the Golgi-localized Dsc E3 ligase12. Given that Scp1 is not required for recognition of Sre1 by the Dsc E3 ligase12, the Scp1 30-amino acid tail may function upstream of ER exit. One possibility is that the Scp1 tail is the site for COPII binding. Indeed, the 30-amino acid Scp1 tail contains di-acidic and di-hydrophobic motifs that are common ER-exit signals found in the cytosolic C-terminal tails of membrane proteins (Supplementary information, Figure S1)28. Alternatively, the deletion of the C-terminal tail may prevent the COPII binding to other ER-exit signals, such as the MELADL-like motif in the Loop 6 region29,30.
Due to low sequence similarities between the WD40 domain of mammalian SCAP and yeast Scp1, it is impractical to build a reliable structural model of the WD40 domain of the mammalian SCAPs on the basis of the Scp1-WD40 structure. Nevertheless, considering the similar length, we speculate that the mammalian SCAP-WD40 may also be an eight-bladed β-propeller. Notably, the corresponding sequence for the RK patch of Scp1-WD40 cannot be identified in the mammalian SCAP-WD40. Co-evolution may have occurred to SCAP and SREBP in other organisms to preserve the complex formation, which allows the functional conservation of SCAP. The structural, biochemical, and cell-based studies reported here have laid a foundation for further mechanistic characterizations of the SREBP and SCAP proteins in fission yeast, as well as higher organisms.
Materials and Methods
Protein expression and purification
The complementary DNA of the Scp1-WD40 (residues 567-1 085) was subcloned into pFastBac1 vector (Invitrogen) with N-terminal 10× His tag. The recombinant protein was then expressed in Sf9 insect cells (Invitrogen) using the pFastBac baculovirus system (Invitrogen). After infection for 48 h, the Sf9 cells were harvested and the protein was purified to homogeneity as described below. The two fragments of Scp1-WD40 (residues 567-961 and residues 986-1 085 or 986-1 054) were co-expressed in E.coli BL21 (DE3). Briefly, the cDNAs of the two fragments were subcloned into pETDuet-1 vector (Novagen) with 6× His tag at the N-terminus of the small fragment. Co-expression was induced by 0.2 mM isopropyl--D-thiogalactoside at an OD600nm of 1.2. After growing at 20 °C for 16 h, the cells were collected and homogenized in a buffer containing 25 mM Tris-HCl, pH 8.0, and 150 mM NaCl. After sonication and centrifugation, the supernatant was applied to Ni2+ affinity resin (Ni-NTA; Qiagen) and further purified by ion-exchange chromatography (Source 15Q; GE Healthcare) and size-exclusion chromatography (Superdex 200; GE Healthcare). The peak fraction was collected and concentrated to ∼10 mg/ml for crystallization. Selenomethionine (SeMet)-substituted Scp1-WD40 was prepared in E.coli following the same protocol. All the mutants were generated using two-step PCR or standard PCR-based cloning strategy and subcloned, expressed and purified in the same way. The complementary DNA of the regulatory domain of Sre1 (residues 628-900) was cloned into pET15b vector (Novagen) with N-terminal MBP fusion. Similar protocol was applied to the overexpression and purification of the regulatory domain of Sre1.
Crystallization, data collection, and structure determination
Crystals of an engineered Scp1 WD40 domain (residues 567-961 and 986-1 054, C618S/C671S/C680S/C756S/C873S/C901S/C920S/ C941S/C1010S) were grown at 18 °C by the hanging drop vapor diffusion method by mixing 1 μl protein with 1 μl reservoir solution containing 20% (v/v) PEG3350, and 0.2 M Na3Citrate (pH 8.3). Crystals appeared in 1-2 days and grew to full size in 1 week. SeMet-labeled protein was crystallized in the same way. Both native and Se-Met crystals were soaked in mother liquor plus 20% ethylene glycol before flash-freezing in a cold nitrogen stream at 100 K.
The native data sets were collected at SSRF beamline BL17U or Spring-8 beamline BL41XU, and the SeMet-derived data were collected at SSRF beamline BL17U. All data sets were integrated and scaled using HKL200032. Further processing was carried out using programs from CCP4 suite33. Data collection statistics are summarized in Supplementary information, Table S1. To begin with, molecular replacement methods were tried by using structures composed of WD40 repeats from PDB. However, no suitable models were able to provide the correct phase. To solve the phase problem, SeMet-derived crystals were generated and a set of SAD data was collected. The selenium positions were identified by SHELXD34, and initial phases were generated by SHELXE. A starting model was traced automatically using the program BUCCANEER35. The initial model was then manually rebuilt in COOT36 and refined with PHENIX37. The resulting map was in good quality for unambiguous sequence assignment with the aid of selenium sites in the anomalous difference Fourier map. Using this as initial model for native data sets, a higher resolution structure was solved by molecular replacement in PHASER38. The final model was refined with COOT and PHENIX. The atomic coordinates of Scp1-WD40 have been deposited in the Protein Data Bank with the accession code 4YHC.
MBP-mediated pull-down assay
Approximately 240 μg MBP-Sre1 protein was incubated with 50 μl amylose resin (GE Healthcare). Approximately 300 μg of WT or Scp1-WD40 variants were incubated with the Sre1-bound resin at room temperature for 30 min. The resin was then extensively rinsed with the buffer containing 25 mM Tris, pH 8.0, and 150 mM NaCl to remove unbound proteins. The resin was resuspended in 500 l buffer containing 25 mM Tris, pH 8.0, and 150 mM NaCl. Then 7.5 μl suspension was directly applied to SDS-PAGE followed by Coomassie blue staining.
Circular dichroism (CD)
CD scanning spectra were collected between 190 and 260 nm at 25 °C using a quartz cuvette with 1 mm path length on Chirascan-plus CD Spectrometer (Applied Photophysics). Measurements were performed with protein samples at concentrations of ∼5 μM in 50 mM Na2HPO4/NaH2PO4 (pH 7.5). Spectra were collected at 1.0 nm interval at a scan rate of 120 nm/min and subtracted for buffer signal.
Reagents
We obtained yeast extract, peptone, agar from BD Biosciences; glass beads (G8772, for yeast cell lysis) from Sigma; Geneticin from GIBCO for the kanMX marker selection (final 100 μg/l); clonNAT from Werner BioAgents for the natMX marker selection (final 100 μg/l); Protein G magnetic beads (#S1430S) from NEB; EDTA-free protease inhibitor cocktail tablets (#11873580001), and Alkaline phosphatase (10713023001) from Roche Diagnostics.
S. pombe strains
We obtained WT haploid S. pombe KGY425 and KGY461 from ATCC. Strains scp1Δ and hrd1 have been described previously11,26. To generate scp1+ WT and mutants (M1-M6) with C-terminal Myc epitope tagging, scp1+ integrating plasmids were linearized, transformed into scp1Δ strain PEY1748 and selected on medium containing 100 μg/l Geneticin. scp1Δ strain PEY1748 marked with natMX marker was generated by transforming linearized pFA6a-NatMX39 into PEY554 and selecting on medium containing 100 μg/l clonNAT. To construct scp1+ integrating plasmids for strain generation, pFA6a-13XMyc-kanMX640 was digested by Bgl II and Nco I to remove TEF promoter upstream of kanMX, and a fragment containing 500 bp ura4+ promoter flanked by Bgl II and Nco I restriction sites was inserted. We then inserted a 2 443 bp fragment starting 619 bp upstream of the scp1+ coding sequence to the sequence coding for Scp1 proline 589 into the Sma I site by Gibson Cloning to generate the intermediate plasmid pWS569. Fragments coding for WT or mutated scp1 were inserted between sequences coding for P589 and Myc epitope tag of the plasmid pWS569 to generate scp1+ integrating plasmids, respectively. To further delete hrd1+ from scp1+ WT and mutants (M1-M3), hrd1Δ strain PEY1756 was mated to scp1+ WT and mutants (M1-M3), respectively, and selected on medium containing both clonNAT (100 μg/l) and Geneticin (100 μg/l). hrd1Δ strain PEY1756 marked with natMX marker was generated by transforming linearized pFA6a-NatMX into PEY893 and mating with KGY461 to switch the mating type. Supplementary information, Table S2 contains a complete list of strains used in this study.
S. pombe cell culture
All strains were grown to log phase at 30 °C in YES medium (5 g/l yeast extract plus 30 g/l glucose and supplements, 225 mg/l each of uracil, adenine, leucine, histidine, and lysine). On day 0, yeast cells were inoculated from saturated cultures into 10 ml fresh YES medium and grown overnight at 30 °C. On day 1, exponential phase cells were inoculated into 10 ml fresh YES medium under aerobic or anaerobic conditions for 3 h. For hypoxic treatment, cells were grown in YES medium, collected by centrifugation, and oxygenated medium was removed by aspiration. Cells were resuspended in deoxygenated YES medium inside an in vivo 400 hypoxic workstation (Biotrace). Anaerobic conditions were maintained in the workstation using 10% hydrogen gas with balanced nitrogen in the presence of palladium catalyst. Deoxygenated medium was prepared by preincubation for > 24 h in the workstation. After resuspension, cultures were agitated at 30 °C for the indicated time, and 2 × 107 cells (whole-cell lysates) or 5 × 107 cells (microsomes) of exponential phase culture were harvested for protein extraction and immunoblotting.
Protein extraction and immunoblotting
Whole-cell extracts were prepared and treated with phosphatase as described previously11. Microsome extracts were prepared as described previously6. Protein concentration was measured using the BCA Kit (Pierce), and the samples were mixed with 5× SDS loading buffer (150 mM Tris-HCl, pH 6.8, 15% SDS, 25% glycerol, 0.2% Bromophenol Blue, and 12.5% β-mercaptoethanol). After heating at 37 °C for 30 min, proteins were subjected to SDS-PAGE and transferred to nitrocellulose membranes (Amersham). The filters were incubated with the antibodies described in the figure legends. Bound antibodies were visualized with IRDye-conjugated goat anti-mouse or anti-rabbit IgG using the LI-COR Odyssey CLx Infrared Imaging System. Gels were calibrated with pre-stained molecular mass markers (Bio-Rad). Working concentrations of antibodies were: 0.2 μg/ml mouse monoclonal anti-Myc IgG 9E10 (Santa Cruz Biotechnology), 1:2 500 diluted rabbit anti-Sre1 anti-sera11, 1:20 000 diluted rabbit polyclonal anti-Dsc5 anti-sera41, 1:20 000 diluted IRDye-680/800 conjugated goat anti-mouse/rabbit secondary IgGs (LI-COR Biosciences).
Accession codes
References
Horton JD, Goldstein JL, Brown MS . SREBPs: activators of the complete program of cholesterol and fatty acid synthesis in the liver. J Clin Invest 2002; 109:1125–1131.
Brown MS, Goldstein JL . The SREBP pathway: regulation of cholesterol metabolism by proteolysis of a membrane-bound transcription factor. Cell 1997; 89:331–340.
Shao W, Espenshade PJ . Expanding roles for SREBP in metabolism. Cell Metab 2012; 16:414–419.
Espenshade PJ, Hughes AL . Regulation of sterol synthesis in eukaryotes. Annu Rev Genet 2007; 41:401–427.
Goldstein JL, DeBose-Boyd RA, Brown MS . Protein sensors for membrane sterols. Cell 2006; 124:35–46.
Shao W, Espenshade PJ . Sterol regulatory element-binding protein (SREBP) cleavage regulates Golgi-to-endoplasmic reticulum recycling of SREBP cleavage-activating protein (SCAP). J Biol Chem 2014; 289:7547–7557.
Nohturfft A, DeBose-Boyd RA, Scheek S, Goldstein JL, Brown MS . Sterols regulate cycling of SREBP cleavage-activating protein (SCAP) between endoplasmic reticulum and Golgi. Proc Natl Acad Sci USA 1999; 96:11235–11240.
Bien CM, Espenshade PJ . Sterol regulatory element binding proteins in fungi: hypoxic transcription factors linked to pathogenesis. Eukaryot Cell 2010; 9:352–359.
Osborne TF, Espenshade PJ . Evolutionary conservation and adaptation in the mechanism that regulates SREBP action: what a long, strange tRIP it's been. Genes Dev 2009; 23:2578–2591.
Todd BL, Stewart EV, Burg JS, Hughes AL, Espenshade PJ . Sterol regulatory element binding protein is a principal regulator of anaerobic gene expression in fission yeast. Mol Cell Biol 2006; 26:2817–2831.
Hughes AL, Todd BL, Espenshade PJ . SREBP pathway responds to sterols and functions as an oxygen sensor in fission yeast. Cell 2005; 120:831–842.
Stewart EV, Nwosu CC, Tong Z, et al. Yeast SREBP cleavage activation requires the Golgi Dsc E3 ligase complex. Mol Cell 2011; 42:160–171.
Nohturfft A, Brown MS, Goldstein JL . Topology of SREBP cleavage-activating protein, a polytopic membrane protein with a sterol-sensing domain. J Biol Chem 1998; 273:17243–17250.
Kuwabara PE, Labouesse M . The sterol-sensing domain: multiple families, a unique role? Trends Genet 2002; 18:193–201.
Luskey KL, Stevens B . Human 3-Hydroxy-3-Methylglutaryl coenzyme-a reductase - conserved domains responsible for catalytic activity and sterol-regulated degradation. J Biol Chem 1985; 260:271–277.
Carstea ED, Morris JA, Coleman KG, et al. Niemann-Pick C1 disease gene: homology to mediators of cholesterol homeostasis. Science 1997; 277:228–231.
Loftus SK, Morris JA, Carstea ED, et al. Murine model of Niemann-Pick C disease: mutation in a cholesterol homeostasis gene. Science 1997; 277:232–235.
Ingham PW, Taylor AM, Nakano Y . Role of the Drosophila patched gene in positional signalling. Nature 1991; 353:184–187.
Burke R, Nellen D, Bellotto M, et al. Dispatched, a novel sterol-sensing domain protein dedicated to the release of cholesterol-modified hedgehog from signaling cells. Cell 1999; 99:803–815.
Sakai J, Nohturfft A, Cheng D, Ho YK, Brown MS, Goldstein JL . Identification of complexes between the COOH-terminal domains of sterol regulatory element-binding proteins (SREBPs) and SREBP cleavage-activating protein. J Biol Chem 1997; 272:20213–20221.
Stirnimann CU, Petsalaki E, Russell RB, Muller CW . WD40 proteins propel cellular networks. Trends Biochem Sci 2010; 35:565–574.
Matsuda M, Korn BS, Hammer RE, et al. SREBP cleavage-activating protein (SCAP) is required for increased lipid synthesis in liver induced by cholesterol deprivation and insulin elevation. Genes Dev 2001; 15:1206–1216.
Moon YA, Liang G, Xie X, et al. The Scap/SREBP pathway is essential for developing diabetic fatty liver and carbohydrate-induced hypertriglyceridemia in animals. Cell Metab 2012; 15:240–246.
Hua X, Nohturfft A, Goldstein JL, Brown MS . Sterol resistance in CHO cells traced to point mutation in SREBP cleavage-activating protein. Cell 1996; 87:415–426.
Cerna D, Wilson DK . The structure of Sif2p, a WD repeat protein functioning in the SET3 corepressor complex. J Mol Biol 2005; 351:923–935.
Hughes BT, Nwosu CC, Espenshade PJ . Degradation of sterol regulatory element-binding protein precursor requires the endoplasmic reticulum-associated degradation components Ubc7 and Hrd1 in fission yeast. J Biol Chem 2009; 284:20512–20521.
Yu L, Gaitatzes C, Neer E, Smith TF . Thirty-plus functional families from a single motif. Protein Sci 2000; 9:2470–2476.
Barlowe C . Signals for COPII-dependent export from the ER: what's the ticket out? Trends Cell Biol 2003; 13:295–300.
Sun LP, Li L, Goldstein JL, Brown MS . Insig required for sterol-mediated inhibition of Scap/SREBP binding to COPII proteins in vitro. J Biol Chem 2005; 280:26483–26490.
Hughes AL, Stewart EV, Espenshade PJ . Identification of twenty-three mutations in fission yeast Scap that constitutively activate SREBP. J Lipid Res 2008; 49:2001–2012.
DeLano WL . The PyMOL Molecular Graphics System on World Wide Web http://www.pymol.org. 2002.
Otwinowski Z, Minor W . Processing of X-ray diffraction data collected in oscillation mode. Method Enzymol 1997; 276:307–326.
Collaborative Computational Project, Number 4. The Ccp4 suite: programs for protein crystallography. Acta Crystallogr D Biol Crystallogr 1994; 50(Pt 5):760–763.
Schneider TR, Sheldrick GM . Substructure solution with SHELXD. Acta Crystallogr D Biol Crystallogr 2002; 58(Pt 10 Pt 2):1772–1779.
Cowtan K . The Buccaneer software for automated model building. 1. Tracing protein chains. Acta Crystallogr D Biol Crystallogr 2006; 62(Pt 9):1002–1011.
Emsley P, Cowtan K . Coot: model-building tools for molecular graphics. Acta Crystallogr D Biol Crystallogr 2004; 60(Pt 12 Pt 1):2126–2132.
Adams PD, Grosse-Kunstleve RW, Hung LW, et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr D Biol Crystallogr 2002; 58(Pt 11):1948–1954.
Mccoy AJ, Grosse-Kunstleve RW, Adams PD, Winn MD, Storoni LC, Read RJ . Phaser crystallographic software. J Appl Crystallogr 2007; 40:658–674.
Hentges P, Van Driessche B, Tafforeau L, Vandenhaute J, Carr AM . Three novel antibiotic marker cassettes for gene disruption and marker switching in Schizosaccharomyces pombe. Yeast 2005; 22:1013–1019.
Bahler J, Wu JQ, Longtine MS, et al. Heterologous modules for efficient and versatile PCR-based gene targeting in Schizosaccharomyces pombe. Yeast 1998; 14:943–951.
Stewart EV, Lloyd SJ, Burg JS, et al. Yeast sterol regulatory element-binding protein (SREBP) cleavage requires Cdc48 and Dsc5, a ubiquitin regulatory X domain-containing subunit of the Golgi Dsc E3 ligase. J Biol Chem 2012; 287:672–681.
Acknowledgements
This work was supported by funds from the Ministry of Science and Technology of China (2015CB910101, 2011CB910501 and 2009CB918802), the National Natural Science Foundation of China (31321062 and 31125009), and the National Institutes of Health (HL077558 to PE). The research of NY was supported in part by an International Early Career Scientist grant from the Howard Hughes Medical Institute and an endowed professorship from Bayer Healthcare.
Author information
Authors and Affiliations
Corresponding authors
Additional information
( Supplementary information is linked to the online version of the paper on the Cell Research website.)
Supplementary information
Supplementary information, Figure S1
Sequence alignment of the Scp1 WD40 domain from eight closely related species. (PDF 928 kb)
Supplementary information, Figure S2
Structural determination of the Scp1 WD40 domain. (PDF 745 kb)
Supplementary information, Figure S3
Structural comparison of the WD40 domains of Scp1 and Sif2. (PDF 286 kb)
Supplementary information, Figure S4
The Scp1-WD40 RK patch mutants are well-folded. (PDF 186 kb)
Supplementary information, Figure S5
The examined surface residues of Scp1-WD40 that may not be involved in Sre1 binding. (PDF 275 kb)
Supplementary information, Table S1
Crystallographic data collection and refinement statistics for Scp1-WD40 (PDF 170 kb)
Supplementary information, Table S2
S. pombe strain list (PDF 153 kb)
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0
About this article
Cite this article
Gong, X., Li, J., Shao, W. et al. Structure of the WD40 domain of SCAP from fission yeast reveals the molecular basis for SREBP recognition. Cell Res 25, 401–411 (2015). https://doi.org/10.1038/cr.2015.32
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2015.32
Keywords
This article is cited by
-
SREBP-regulated lipid metabolism: convergent physiology — divergent pathophysiology
Nature Reviews Endocrinology (2017)
-
Complex structure of the fission yeast SREBP-SCAP binding domains reveals an oligomeric organization
Cell Research (2016)