Abstract
Crocodilians are diving reptiles that can hold their breath under water for long periods of time and are crepuscular animals with excellent sensory abilities. They comprise a sister lineage of birds and have no sex chromosome. Here we report the genome sequence of the endangered Chinese alligator (Alligator sinensis) and describe its unique features. The next-generation sequencing generated 314 Gb of raw sequence, yielding a genome size of 2.3 Gb. A total of 22 200 genes were predicted in Alligator sinensis using a de novo, homology- and RNA-based combined model. The genetic basis of long-diving behavior includes duplication of the bicarbonate-binding hemoglobin gene, co-functioning of routine phosphate-binding and special bicarbonate-binding oxygen transport, and positively selected energy metabolism, ammonium bicarbonate excretion and cardiac muscle contraction. Further, we elucidated the robust Alligator sinensis sensory system, including a significantly expanded olfactory receptor repertoire, rapidly evolving nerve-related cellular components and visual perception, and positive selection of the night vision-related opsin and sound detection-associated otopetrin. We also discovered a well-developed immune system with a considerable number of lineage-specific antigen-presentation genes for adaptive immunity as well as expansion of the tripartite motif-containing C-type lectin and butyrophilin genes for innate immunity and expression of antibacterial peptides. Multifluorescence in situ hybridization showed that alligator chromosome 3, which encodes DMRT1, exhibits significant synteny with chicken chromosome Z. Finally, population history analysis indicated population admixture 0.60-1.05 million years ago, when the Qinghai-Tibetan Plateau was uplifted.
Similar content being viewed by others
Introduction
The amniotes diverged into mammals and reptiles ∼320 million years ago (Mya)1. The early reptiles evolved into a diverse fauna, including lizards, snakes, turtles, crocodilians, and birds. The body plan of modern crocodilians has remained somewhat unchanged since the earliest crocodilians appeared 240 Mya2,3, with their adaptations to aquatic environments and a diving lifestyle. An impressive feature of crocodilians is their ability to remain submerged for long periods of time. Their diving behaviors are derived from a number of traits. First, the crocodilians immerse themselves in the water to ambush prey, which they kill by drowning4. Crocodilians never stray far from the water and usually enter the water to escape predation4. They are ectotherms that behaviorally regulate their body temperature by shuttling between terrestrial and aquatic environments and controlling their diving depth5. Crocodilians also submerge themselves during social interactions, such as mating6.
Monitoring of free dives7 has shown that the crocodilians exhibit two types of submergence behavior: long (∼12 min) resting dives, the duration of which depends on body mass, and short (∼1 min) active dives, which vary in length based on the animal's activities. Previous data suggest that the external environment is the most important determinant in prolonged aerobic dives7, which might be extended to 1-2 h8. Crocodilians have evolved a series of physical adaptations for diving, including development of a palatal valve at the back of the throat to prevent entry of water into the throat, esophagus, and trachea when the animal is submerged9; ear-flaps and eye-lids to protect the inner ear and cornea, respectively9; thickening of the lung wall to resist underwater pressure9; and increased lung capacity8. Aerobic dive duration is achieved when oxygen deprivation prompts the crocodilian heart to stop blood flow to the muscles, ensuring aerobic energy expenditure in the brain9. Switching of cardiac performance is controlled by pulmonary oxygen tension10.
Crocodilians are mostly active at dawn and dusk9,11 and require a powerful sensory system to detect predators and prey, sense environmental changes, and engage in social interactions. They have a collection of highly adapted morphological characteristics for their crepuscular lifestyle9, including two large olfactory lobes in the brain, nerve ending-enriched sensory pits on the jaws to sense extremely small vibrations, a reflecting layer in the eye to improve night vision, and 2-4-fold more auditory fibers than birds and mammals12, contributing to their remarkable hearing under water13,14. Crocodilians also have a minimal body profile, allowing their sensory organs to break the water surface while the remainder of their body is hidden from view; this is considered one of their most significant adaptations.
Crocodilians live in marshes, lakes, and rivers, and often suffer from serious injuries — males during fights for mates and females during battles for nests9; however, they appear to recover quickly from open wounds in water and are thought to have a robust immune system that resists microbial infections15. Merchant et al.16 revealed that crocodilians generate antimicrobial peptides in the blood and possess a powerful first-line defense against pathogens in aquatic environments. In addition to their physical and physiological adaptations, we expect whole-genome analysis to reveal molecular evidence of diving, sensory, and immune adaptations in crocodilians.
Amniotes have evolved diverse sex-determining mechanisms: mammals exhibit XY-type genetic sex determination (GSD); birds exhibit ZW-type GSD; and non-avian reptiles exhibit XY-GSD, ZW-GSD, and temperature-dependent sex determination (TSD)17. Although complete genomes are available for 1 lizard1, 3 birds18,19,20, and various mammals, all of these species have sex chromosomes. In contrast, crocodilians exhibit TSD and do not possess sex chromosomes21. Therefore, genome sequencing of a crocodilian may provide novel insights into sex chromosome evolution.
There are 23 species of crocodilians, divided into three groups: Alligatoridae, Crocodylidae, and Gavialidae22. The Chinese alligator (Alligator sinensis), a freshwater crocodilian endemic to China, is one of the most endangered crocodilian species23. Currently, there are ∼100 Chinese alligators in the wild and ∼10 000 captive individuals in Zhejiang and Anhui Provinces24. We chose the Chinese alligator for genome sequencing with the hope of providing information that could help design scientific captive-breeding programs for population recovery project of this endangered species.
Results
Assembly and annotation
We collected a male Chinese alligator from Changxing Yinjiabian Chinese Alligator Nature Reserve (Zhejiang Province, China) and sequenced its genome using a whole-genome shotgun strategy. We obtained 314.03 Gb of raw sequence on a next-generation sequencing platform (IlluminaHiseq 2000). SOAPdenovo25 was used to assemble the genomic sequence, resulting in a 2.3-Gb assembly with contig and scaffold N50 values of 23.4 kb and 2.2 Mb (Table 1). We assessed the assembly quality using bacterial artificial chromosome (BAC) clones and found that the scaffolds were reliably assembled with the exclusion of the GC-rich and repeat-rich regions, which were filled by gaps (Supplementary information, Figure S1).
A total of 22 200 genes were predicted in the alligator with integration of de novo prediction, homolog-based prediction, and RNA-Seq data (Supplementary information, Table S1); 79.35% of these were functionally annotated (Supplementary information, Table S2). We then annotated interspersed repeats and found that about 37.93% of the alligator genome consists of DNA transposons, long interspersed nuclear elements (LINEs), long terminal repeats, and short interspersed nuclear elements (Supplementary information, Table S3); the LINEs were most abundant, comprising about 29.13% of the genome.
Genome landscape
GC content
We calculated the GC content of the Chinese alligator genome and detected a clear preference for GC-rich regions, averaging 44.5%; Chinese alligator showed the highest GC level among the studied organisms (Supplementary information, Figure S2). The GC patterns of Chinese alligator differed from those of most representative animals, but are similar to that of humans (Homo sapiens) (Supplementary information, Figure S2), with a wider GC range and a lower GC-content peak. In contrast, the green anole lizard (Anolis carolinensis) and clawed frog (Xenopus tropicalis) yielded narrow elevated curves. Previous studies demonstrated that the green anole lizard and clawed frog possess an unusually homogenous GC distribution, while humans have a heterogeneous GC curve1,26. Thus, the Chinese alligator may have a heterogeneous GC distribution. We compared GC content in different regions of the alligator genome and found that GC frequencies were highest in gene regions (Supplementary information, Table S4). GC-rich regions often trigger gene recombination18, suggesting that the Chinese alligator might undergo more gene conversion events than other amniotes.
Repeat elements
We compared transposable elements (TEs) of four reptiles — all with abundant LINEs, with the alligator showing relatively long LINE members (Supplementary information, Table S5). Then, we analyzed the accumulation of different TEs based on divergence from the consensus sequence, which resembles the motifs of ancestral repeats27, and found different patterns in the Chinese alligator, green anole lizard, chicken (Gallus gallus), and zebra finch (Taeniopygia guttata) (Supplementary information, Figure S3). The alligator genome has accumulated the most highly divergent (old) TEs, showing 1 obvious LINE peak at high divergence rates of 0.10-0.30. The lizard presented a slight fluctuation of TE coverage across different divergence rates, suggesting the loss of some old TEs and acquisition of relatively young TEs in comparison to Chinese alligator. The TE coverage of the chicken and zebra finch genomes exhibited half of the TE abundance found in the lizard and one-third of the alligator TE abundance (Supplementary information, Figure S3). As a result, the high genome coverage of old LINEs might account for the larger genome of Chinese alligator in comparison to the other three animals, especially birds.
Segmental duplication (SD) analysis
Genomic sequences of the Chinese alligator, green anole lizard, chicken, and zebra finch were subjected to whole-genome alignment and SD (length > 1 kb and similarity > 90%) analyses to assess genomic features. A total of 35.90, 201.21, 68.85, and 122.32 Mb non-redundant SD blocks were identified in the alligator, lizard, chicken, and finch genomes, occupying 2.05%, 11.18%, 6.21%, and 9.92% of the genome, respectively. Short read-based assembled Chinese alligator genome revealed fewer SD blocks than the other three BAC-based assembled genomes, possibly due to susceptibility of the short read sequencing to lost assembly of recently duplicated genome regions28. The four species were uniformly biased toward smaller SDs (Supplementary information, Figure S4A-S4C), suggesting that the ancestral reptilian genome might have been characterized by frequent duplication of short segments. The identities of the SD blocks were evenly divergent SDs in the lizard, more similar in the birds, and more divergent in the alligator (Supplementary information, Figure S4D); thus, like the TEs (Supplementary information, Figure S3), older blocks were retained in Chinese alligator and more blocks were newly duplicated in chicken and zebra finch.
Genomic alignments between the alligator and chicken showed better synteny than the pairing of alligator and lizard (Supplementary information, Figure S5), supporting the notion that Chinese alligator is a close relative of birds. The blocks that showed poor synteny between chicken and alligator were largely found in regions containing the most dense SDs, TEs, and small gaps (Supplementary information, Figure S5A), suggesting that the large-scale syntenic breaks between alligator and lizard might be due to the relative abundance of SDs and TEs in the lizard (Supplementary information, Figure S5B). The chicken presented many SD and TE islands with heterogeneous distributions (Supplementary information, Figure S5A) in contrast to the relatively homogenous SD and TE curves in the lizard and alligator (Supplementary information, Figure S5B). Consistent isochores between TEs and SDs were seen in these species, especially in the chicken genome (Supplementary information, Figure S5), indicating that the occurrence of SDs was related to that of TEs. In view of the most abundant LINEs in the reptiles (Supplementary information, Table S5), we further examined the relationship between the SD and LINE distributions and found a significant positive correlation (r = 0.89; P = 0.001), suggesting that the SD events might be triggered by LINEs, which has also been demonstrated in mammals29.
The genomes of chicken and green anole lizard have been assembled at the chromosomal level (Ensembl release 63). Some concordant gaps were seen in the distributions of SDs and TEs; these coincided precisely with the highest densities of Ns (nucleotides) in the gaps (Supplementary information, Figure S5). Furthermore, each chromosome of the chicken and green anole lizard had only a single SD-TE gap. Centromeres typically contain numerous repeats30, leading to unclosed gaps in the assemblies of centromeric DNA regions, as seen in the human genome assembly (NCBI Build 37.3). Thus, the coincident gaps among the SDs, TEs, and N-filled regions of these genomes might represent centromeres.
Genome adaptive divergence
We resolved features of genome adaptive divergence from an increase in gene copy number, an increase in nonsynonymous (dN) over synonymous (dS) substitutions, and lineage-specific genes. These genetic signatures of the Chinese alligator genome were then matched to the biological traits of crocodilians.
Expansion of gene families
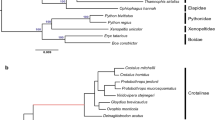
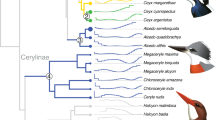
We employed TreeFam to deduce gene clusters from two non-avian reptiles (Chinese alligator and green anole), three avian reptiles (birds: chicken, turkey, and zebra finch), and one mammal (human; outgroup species). We found that among the five reptiles, the Chinese alligator had developed more unique paralogs and unclustered genes (Figure 1A), suggesting that the alligator has more lineage-specific genomic features. From the unique paralogs, we identified 413 alligator-specific multi-copy gene families (Supplementary information, Table S6). Based on the single-copy gene families of these six species, we constructed a phylogenetic tree and calculated divergence time using fossil records. Our results revealed that the crocodilian lineage split from the common ancestor of birds ∼241 Mya (Figure 1B). According to divergence times and phylogenetic relationships, we adopted CAFÉ to assess clustering relationship and discriminated 363 gene families that were significantly expanded in the Chinese alligator (P < 0.05) (Figure 1C). This result highlights the presence of distinct components in the alligator genome.
Comparisons of orthologous and paralogous genes in the genomes of different species. (A) TreeFam-based clustering of gene families. (B) Divergence time of six species. (C) Expansion and contraction of CAFÉ-based gene families. (D) Gene family of functional olfactory receptors (ORs). aMann-Whitney U test, P < 0.01; bMann-Whitney U test was not performed because the number of α-ORs was less than 30.
Rapid evolution analysis
In addition to an expansion in paralogs, adaptation features of genome divergence usually induce an excess of nonsynonymous over synonymous substitutions (dN > dS) at orthologous genes31, which can be identified as positively selected genes (PSGs) by the likelihood ratio test and lineage-specific accelerated evolving gene ontology (GO) categories by the binominal test. We achieved 7 337 strictly filtered 1:1:1:1:1 orthologous genes in the Chinese alligator, green anole lizard, zebra finch, chicken, and turkey (Meleagris gallopavo) genomes and identified 219 PSGs (Supplementary information, Table S7) and 86 rapidly evolving GO classes (Supplementary information, Table S8).
Lineage-specific gene analysis
From the perspective of combined paralogous and orthologous genes, adaptive differences could be determined by analyzing lineage-specific gene pools, as shown in the potato32. Comparative genomic analysis of the alligator and 18 representative species of fish (Tetraodon nigroviridis, Gasterosteus aculeatus, Oryzias latipes, Takifugu rubripes, and Danio rerio), amphibians (X. tropicalis), mammals (Dasypus novemcinctus, Bos taurus, Canis familiaris, Loxodonta africana, H. sapiens, Monodelphis domestica, Ornithorhynchus anatinus, Rattus norvegicus, and Choloepus hoffmanni), and reptiles (Anolis carolinensis, T. guttata, and G. gallus) revealed 6 122 alligator lineage-specific genes (Supplementary information, Table S9), of which 1 543 (25.20%) are functionally annotated.
Diving behavior adaptation
Crocodilians inherited their terrestrial style from the ancestral amniotes33, greatly challenging their diving ability. Thus, we examined their genetic signatures from multiple angles, including oxygen transport, energy supply, urinary excretion, and cardiac muscle contraction systems.
Oxygen transport
Hemoglobin (Hb) is responsible for oxygen (O2) transport and is composed of two α and two β subunits. The phylogenetic tree of reptilian Hbs shows that the Chinese alligator has four crocodilian-specific Hb genes: 1 α (HBA1) and 3 β (HBB2, HBB4, and HBB5) (Supplementary information, Figure S6). The transcriptome and proteome data demonstrate that these genes are all highly expressed in the blood (Figure 2A). Previous studies demonstrated that crocodilian hemoglobin had been mutated from a routine bisphosphoglycerate-binding type to a special bicarbonate (HCO3−)-binding form to increase O2 release34. The Hb sequence alignment shows that the alligator HBA1, HBB2, and HBB4 are HCO3−-binding subunits but HBB5 is a routine β subunit (Figure 2A). Thus, our results reveal that in the Chinese alligator: (1) the HCO3−-binding β gene has been duplicated once; (2) it possesses routine phosphate-binding and special HCO3−-binding double Hb effectors; and (3) the only HBA1 subunit simultaneously participates in the assembly of two types of Hb molecules (Figure 2A). We examined PSGs and found that anion exchanger 1, which facilitates O2 unloading in the standard erythrocyte pathway35,36, has undergone positive selection (Supplementary information, Table S7), presenting evidence of active routine O2 transport. Therefore, our study has identified multiple unique O2 transport pathways in the Chinese alligator (Figure 2B).
Diving adaptations in the Chinese alligator genome. (A) Alignment of hemoglobin genes. Asterisk represents HCO3−-binding sites34. CA, AA, NC, and SC represent the Chinese alligator, American alligator, Nile crocodile, and Spectacled caiman. (B) Standard and HCO3−-binding O2 transportation pathways36. (C) Positively selected genes (PSGs) have been mapped to metabolic pathways by iPath37. (D) The pathways directly and indirectly participate in oxidative phosphorylation (according to map 01100, map 00190, and iPath). The color-shaded pathways correspond to those in C. (E) NH4HCO3 excretory pathways35 and maps 00460, 00480, and 00910. (F) Cardiac muscle contraction pathways (maps 05410 and 04260). The bicarbonate feature is highlighted in green, and the PSGs are shown in red. Bold lines depict the major pathways underwater, and the dotted lines indicate multiple linked steps in the pathway.
Energy supply system
We enriched KEGG pathways for PSGs and found that alligator PSGs are mostly metabolism-related (Supplementary information, Table S10). In particular, the oxidative phosphorylation (OXPHOS) in charge of energy production is overrepresented in the Chinese alligator (Supplementary information, Table S10). We then used iPath37 to visualize the mutual relationship of PSGs in metabolic pathways and found that the PSGs were obviously focused on the glycan, fatty acid, terpenoids, and OXPHOS metabolic pathways (Figure 2C). Of the 28 iPath-mapped PSGs, 24 are directly and indirectly part of OXPHOS metabolism at the mitochondrial inner membrane (Figure 2D). The most remarkable result is that two ATP synthases (ATPeF0B and ATPeVAC39) have undergone positive selection in Chinese alligator (Figure 2D). The rapidly evolving GO results indicate that the “ATP catabolic process”, “phosphorylation”, “glucose homeostasis”, “mitochondrial inner membrane”, “mitochondrion”, “ATPase activity”, and “heme binding” GO categories have experienced strong selective pressure (Figure 3 and Supplementary information, Table S8). Consequently, positive selection in the energy metabolism-related genes and GO classes suggests special energy demand in alligators during diving.
Rapidly evolving GO categories of the Chinese alligator. (A) GO supergenes containing ≥ 20 orthologous genes and fast evolving biological process (B), cellular component (C) and molecular function (D) classes. The full list of GO categories is provided in Supplementary information, Table S8.
Urinary excretion
Amniotes remove carbon dioxide (CO2) by lung ventilation38. As no gas exchange is possible during a dive, crocodilians secrete ammonium bicarbonate (NH4HCO3) in the urine as the major excretory route9. We surveyed the genetics of the excretory system and discovered that the ammonium (NH4+) transporter (PF00909) and HCO3− transporter (PF00955) families are overrepresented in the PSGs (Supplementary information, Table S11). Furthermore, the PSG-enriched pathways, cyanoamino acid metabolism (map 00460) and glutathione metabolism (map 00480) (Supplementary information, Table S10), generate formamide and glutamate, which are the precursors of ammonia (NH3) (map 00910; Figure 2E). CA4 (carbonic anhydrase IV), which binds the plasma membrane of the tubular lumen39 and controls urine HCO3− concentrations by catalyzing the reversible dehydration of carbonic acid (Figure 2E), and RHCG (Rhesus blood group, C glycoprotein), which excretes NH4+ in kidney tubules and regulates body acid-base balance40, have both undergone positive selection in Chinese alligator (Supplementary information, Table S7). These results provide evidence for the positive selection of NH4HCO3 secretion through the urinary system in the alligator.
Cardiac muscle contraction
Once oxygen depletion occurs during submergence, the crocodilians slow their heart rate and supply oxygenated blood only to the brain9. The alligator lineage-specific rapidly evolving GO results demonstrate that the diving hypoxia adaptation-related categories, including the “response to hypoxia”, “heart development”, “response to stress”, and “heat shock protein binding” (Figure 3B and 3D), have undergone fast evolution in the Chinese alligator. We further examined the PSG evidence for the crocodilian cardiovascular system and found strengthened striated muscles, showing four PSGs related to the assembly of sarcomeres (Figure 2F). Mapping of PSGs to the cardiac muscle contraction pathway (map 04260) indicates that the Na+/K+-ATPase β subunit (ATP1B) is a PSG (Figure 2F and Supplementary information, Table S7) that controls the repolarization/relaxation of the cardiac muscle41. Furthermore, two other PSGs, SCN4B and KCNJ8 (Supplementary information, Table S7), have also been associated with heart repolarization42,43. Thus, these positive selection (dN/dS test) results suggest that a longer resting state and slower heartbeat of robust cardiac muscle may meet the special demand of minimizing metabolic rate during alligator diving.
Sensory system signatures
We extracted molecular evidence of an excellent olfactory ability in the Chinese alligator genome. First, the alligator lineage-specific rapidly evolving GO results show that the “receptor activity”, “transporter activity”, and “ion channel activity” have experienced rapid evolution (Figure 3D). We then enriched the GO domains of the CAFÉ-based gene families and found that the “receptor activity”, “transmembrane signaling receptor activity”, and “olfactory receptor (OR) activity” genes are overrepresented (all P = 0.0000) in the molecular function category of the Chinese alligator (Supplementary information, Table S12). We thus re-annotated the OR families for the Chinese alligator, green anole lizard, zebra finch, chicken, turkey, and human and found that the Chinese alligator developed the most ORs (Supplementary information, Table S13). We performed a CAFÉ-based analysis of these OR gene families and found that the Chinese alligator expanded the most gene families and contracted the fewest ORs (Supplementary information, Figure S7). We then built a phylogenetic tree for the ORs of six species and found that these ORs are grouped into a single-copy θ basal branch and two multiple-copy α and γ clusters (Supplementary information, Figure S8). This clustering relationship indicated that the alligator possessed the most abundant functional γ- and α-type ORs relative to other reptiles and a similar number of ORs to human (Figure 1D). The γ- and α-type ORs bind airborne and water-soluble odorant molecules44, respectively, and the α-type ORs in the human are putative relics of ancestral tetrapods45. Thus, we calculated the dN/dS values of the alligator γ- and α-ORs relative to its θ gene and found that Chinese alligator presented a significantly higher dN/dS ratio in the γ set than α (Figure 1D; Mann-Whitney U test, P = 0.0082). Hence, the large quantity and high selective pressure (dN/dS) of γ ORs suggest that the Chinese alligator relies heavily on airborne odorant detection.
In addition, we identified genetic signatures from the alligator nervous system. Two synaptic genes (SYT11 and NLGN3)46,47 have been positively selected (Supplementary information, Table S7) and neurological system process (GO_0050877) was the most overrepresented biological process (BP) among the alligator-specific genes (Supplementary information, Table S14). Finally, four nerve-related cellular components (CCs) have rapidly evolved, including the “neuronal cell body”, “dendrite”, “synapse”, and “synaptosome” (Figure 3C).
We also obtained evidence of enhanced auditory and ocular systems in the Chinese alligator. We found that visual perception has rapidly evolved in the alligator (Figure 3B), and opsin (OPN3) and otopetrin (OTOP1) have been positively selected (Supplementary information, Table S7). Opsin senses light48 and is associated with dark adaptation49, whereas the otopetrin triggers development of the otolith in fish50, which is important for detecting sounds under water51.
Features of the immune system
The single-copy genes in the “immune response” GO category have undergone fast evolution in the Chinese alligator (Figure 3B). The alligator lineage-specific genes indicate that the BP GO classes of “antigen processing and presentation” (0019882), “defense response” (0006952), “immune response” (0006955), and “immune system process” (0002376) are overrepresented (Supplementary information, Table S14), suggesting that the Chinese alligator genome features antigen-triggered adaptive immunity. The major histocompatibility complex (MHC) is responsible for antigen presentation and is divided into class I and class II molecules52. The adaptive immune system of the Chinese alligator was characterized by the lineage-specific class I MHC genes, as shown by enriched CC categories of GO_0042612 (MHC class I protein complex) and GO_0042611 (MHC protein complex) (Supplementary information, Table S14).
The alligator-specific gene families show that the BP categories of “immune response” (GO_0006955), “immune system process” (GO_0002376), and “innate immune response” (GO_0045087) are overrepresented (Supplementary information, Table S15), reflecting strong innate immunity in the Chinese alligator. We then carried out Ensembl-based classification of immune gene families from different animals and found that 43 had expanded in the alligator relative to the other reptiles (Figure 4). The first-, second-, and third-rank gene families are the tripartite motif (TRIM)-containing, C-type lectin (CLEC), and butyrophilin, respectively (Figure 4). The TRIM superfamily is a versatile effector in innate immunity and participates in resistance to different pathogens, especially lentiviruses such as HIV53. The CLEC is a major receptor on the natural killer cells that regulate innate immunity54, while butyrophilin is a regulator responding to inflammation55. Thus, the obvious expansions of the TRIM, CLEC, and butyrophilin gene families in Chinese alligator suggest its strong innate immunity function.
Crocodilian blood kills bacteria in vitro and it is thought to possess antimicrobial activity in vivo56,57. We scanned for genes that intersected in the blood transcriptome and proteome and found that cathelicidin (PF00666), a major class of antimicrobial peptides58, is overrepresented in the blood (Supplementary information, Table S16). Thus, the results suggest that alligator blood carries an effective system with non-specific defense against microbial infection.
Our study derived three significant findings from the Chinese alligator immune system, including evolution of alligator-specific genes related to adaptive immunity, expansion of genes related to innate immunity, and expression of antibacterial peptides in the blood, which indicate that Chinese alligator possesses a well-developed immune defense system.
Sex chromosome evolution and DMRT1 alternative splicing analyses
Sex chromosome evolution
Reptiles show XY-type GSD, ZW-type GSD, and TSD sex determinant mechanisms17. We sequenced genital gland transcriptomes and found 8 743 differentially expressed genes (DEGs) between the ovary and testis of the Chinese alligator (Supplementary information, Figure S9). We extracted the top 20% DEGs in the Chinese alligator gonads and performed chromosomal assignment of the orthologs in humans (Supplementary information, Figure S10A) and chickens (Supplementary information, Figure S10B). Apart from the genes located on the autosomal chromosomes, other orthologs were uniformly allocated to the human X and chicken Z chromosomes (Supplementary information, Figure S10). We then adopted the orthologous genes to all of the DEGs (Figure 5A) to assess their expression profiles in the ovary and testis of the alligator, human, and chicken (Figure 5B and 5C). We found that the testis produced a slight difference (Mann-Whitney U test, P = 0.5988) between the alligator and chicken expression profiles, while all other pairwise comparisons revealed significant differences (Mann-Whitney U test, all P < 0.05), suggesting that cognate genes in the Chinese alligator may resemble the ZW system of the chicken.
Sex chromosome evolutionary features and DMRT1 splice variants. (A) Human X and chicken Z chromosome assignments of genes orthologous to the Chinese alligator gonadal differentially expressed genes (DEGs). (B) Expression profiles of DEGs orthologous to X chromosome-located genes. (C) Expression profiles of DEGs orthologous to Z chromosome-located genes. (D) Multi-color fluorescence in situ hybridization (M-FISH) analysis of DMRT1 and other representative genes in males. (E) M-FISH analysis in females. (F) Syntenic relationship between alligator chromosome 3 and chicken chromosome Z. Sex-related genes were selected from synteny comparison results (Supplementary information, Figure S14). (G) Genomic structure of the Chinese alligator DMRT1 splice variants and their expression profiles in male and female gonads.
The sex determination system of the chicken is ZZ for males and ZW for females; the expression dosage of the DMRT1 (doublesex and mab-3 related transcription factor 1) gene located on the Z chromosome controls testis development and the W-located genes influence ovary development59,60. Two W-located genes, ASW (avian sex-specific W-linked) and FET1 (female expressed transcript 1), are expressed specifically in the embryonic gonads of female chickens59. The ASW was also called HINTW (W-linked histidine triad nucleotide binding protein) due to the presence of its homologous copy (HINTZ) on the Z chromosome61. We annotated 1 DMRT1, 1 HINT, and 130 FET1 genes from the alligator genome, and then constructed phylogenetic trees for the HINT and FET1 genes for classification. The results revealed that the alligator HINT homolog should be HINTZ, as it clustered with the chicken HINTZ gene (Supplementary information, Figure S11). Alligator FET1 genes also differed from chicken FET1 genes because they grouped with the non-ovary-specific FET1 genes of the chicken (Supplementary information, Figure S12). We scanned the BAC library for the single-copy DMRT1 and HINTZ and obtained two BACs, 316D7 (∼120 kb) for DMRT1 and 324H6 (∼80 kb) for HINTZ, for fluorescence in situ hybridization (FISH). The FISH results show that the DMRT1 and HINTZ genes are located on the p-arm of chromosome 3 of the Chinese alligator (Supplementary information, Figure S13). We performed synteny analysis between the alligator scaffolds and the chicken Z chromosome and chose the three largest syntenic blocks from scaffolds 560_1, 573_1, and 240_1 (Supplementary information, Figure S14), where we again selected the three most significant DEGs of genital glands (farnesyltransferase, FNTA; glutathione peroxidase, GPx; terminal uridylyltransferase, TUT) for BAC library scanning. We finally obtained 877H6 (∼70 kb) for the FNTA, 785B7 (∼125 kb) for the GPx, and 1638C10 (∼120 kb) for the TUT, and subjected them to multi-color FISH (M-FISH), together with the 316D7 of DMRT1 and the 324H6 of HINTZ to examine inter-chromosome synteny. The male and female M-FISHs present identical assignments of the five BACs to chromosome 3 (Figure 5D and 5E) and a perfect synteny to the chicken Z chromosome (Figure 5F). Therefore, alligator chromosome 3 and chicken chromosome Z shared an ancestral chromosome.
DMRT1 splice variant
The key sex determination gene, DMRT1, exhibits sex-specific alternative splicing62,63. In this study, we observed 10 alternatively spliced DMRT1 variants in the genital transcriptome sequences of the Chinese alligator due to exon skipping and intron inclusion, six of which were specific to the testis (Figure 5G). Comparisons of the DMRT1 genes of the Chinese alligator, chicken62, and Mugger crocodile (Crocodylus palustris)63 revealed large discrepancies in the lengths of the DMRT1 genomic and coding sequences; the Chinese alligator DMRT1 is composed of five exons and occupies a 100-kb genomic fragment, presenting a sharp contrast to the Mugger crocodile DMRT1, which consisted of three exons spanning only 4 kb (Supplementary information, Figure S15A). This extreme diversification of the DMRT1 structure was accompanied by various alternative splicing options such as the inclusion of introns, exon skipping, the occurrence of pre-stop codons, and frame-shift mutations (Supplementary information, Figure S15B-S15D). The DMRT1 gene is well-known for its DM (d oublesex and m ab-3 related) domain, which is a cysteine-rich DNA-binding motif first recognized in proteins encoded by the Drosophila sex determination gene, doublesex (DSX)64. The DSX gene undergoes sex-specific alternative splicing, and the resultant male- and female-specific isoforms direct male and female development in the fruit fly64. In addition to the DM domain, the DMRT1 gene usually contains another DMRT1 domain (NCBI CDD pfam12374)62. Alignment of the alternatively spliced variants of DMRT1 showed that all isoforms harbored the DM domain, and allowed us to identify alternative splicing hotspots at the ends of the DM and DMRT1 domains across different reptiles (Supplementary information, Figure S16). Of the six testis-specific isoforms (e-j) of the Chinese alligator DMRT1, four isoforms (g-j) contain the DM domain but not the DMRT1 domain (Supplementary information, Figure S16). Furthermore, no DM-only DMRT1 isoforms were expressed in the ovary (Figure 5G), suggesting a DM-biased genital expression profile in Chinese alligator.
Previous studies have shown that the DM-only genes act as independent sex-determining factors in some species, such as the W-linked female-specific DMW in the African clawed frog X. laevis65, and the Y-linked male-specific DMY/DMRT1Y in the medaka Oryzias latipes66. Comparisons of DMRT1 genes between two TSD crocodilians revealed that a nearly identical DMRT1 isoform was shared by the Chinese alligator and the Mugger crocodile (Alligator sinensis DMRT1g and C. palustris DMRT1b) (Supplementary information, Figure S16). Interestingly, DMRT1b is the only isoform in the Mugger crocodile that contains the DM domain alone (Supplementary information, Figure S16). Furthermore, an alternative splicing study identified chicken DM-only DMRT1-V4 and found that it was the only isoform specific to male embryonic gonads62. Thus, we named the alligator DMRT1g isoform “DMZ” due to its inclusion of only the DM domain and its high similarity to the chicken Z-linked DMRT1. As the W-located DMW in frogs and the Y-located DMY in fishes trigger sex differentiation, the expression bias and splicing site similarities between Chinese alligator DMZ, Mugger crocodile DMRT1b, and chicken DMRT1-V4 suggest that DMZ may play an important role in the sex determination of the Chinese alligator.
Single nucleotide polymorphism (SNP) and population history analyses
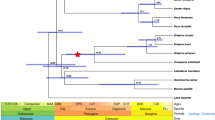
The Chinese alligator is an endangered species, making its population history another issue of interest in conservation biology. Moreover, the sequenced individual was collected from the severely bottlenecked Changxing Chinese alligator population, which developed from 11 founders in 197967. This severe population bottleneck was examined in this study. We aligned clean reads to the genome sequence and identified 318 283 SNPs. The heterozygosity rate was 0.15 × 10−3, which is much lower than those of the green anole, chicken, and human (Figure 6A). The SNP heterozygosities of the coding sequences (CDS) and intronic regions were similar in the Chinese alligator, whereas the ratios were all approximately 0.5 in the lizard, chicken, and human (Figure 6A), suggesting a rapid loss of genetic variation in the non-coding sequences of the Chinese alligator. Furthermore, the SNP curve of the Chinese alligator depicted a continuously elevated number of homozygous SNPs while others entered the descending phase (Supplementary information, Figure S17). These results provide evidence for the ongoing bottleneck in the Chinese alligator.
Single nucleotide polymorphism (SNP) (A) and population history (B) analyses. The shaded area represents the time span of the Qinghai-Tibetan Plateau uplift. *The effective population sizes of the Chinese alligator, green anole and human are indicated on the left; the chicken is on the right (Supplementary information, Figure S18).
Based on the SNP data, we estimated the population history of Chinese alligator (and that of the chicken, green anole, and human) by using the pairwise sequentially Markovian coalescent (PSMC) model, which has been used to deduce human population history68. The alligator PSMC curve depicts a unique increase in its effective population size (Ne) between 0.60 and 1.05 Mya, when other tested species present a consistent decline in population size (Supplementary information, Figure S18). In the human PSMC curve, the younger peak may correspond to an increase in Ne induced by population separation and subsequent admixture68. The Chinese alligator lives in the Yangtze River, the third longest river in the world. The source of the Yangtze River lies in the Tanggula Mountains of the Qinghai-Tibetan Plateau, which experienced widespread and rapid uplifting between 0.6 and 1.1 Mya69. The fossil record indicates that the Chinese alligator once resided in Xinjiang Province of the Qinghai-Tibetan region11. Thus, the concordance between the Ne increase and the Qinghai-Tibetan Plateau uplift (Figure 6B) suggests that Chinese alligators living in the upper Yangtze River would have been forced to swim toward the middle-lower Yangtze. The resulting gene exchange between the upper and lower stream alligators would explain the enhanced Ne in the range of 0.60-1.05 Mya.
Discussion
The amniotes diverged from the tetrapods ∼340 Mya70 and attained the ability to inhabit terrestrial environments. Many mammals and reptiles then returned to the water and regained adaptations to aquatic life. Crocodilians are semi-aquatic reptiles with unique diving, sensory, and immune adaptations. The Chinese alligator genome sequence has unraveled the genetic basis of secondary aquatic adaptations in the circulatory, metabolic, excretory, cardiac, olfactory, nervous, ocular, auditory, and the innate and adaptive immune systems, presenting evidence for co-evolution of multiple systems specific to the back-to-the water transition. Thus, this study provides a good example of how terrestrial-style reptiles adapt to aquatic environments.
Hypoxia usually refers to passive low O2 at high altitude31,71 or in aquatic conditions72. Aerobic diving is a distinctive form of voluntarily tolerant hypoxia. Crocodilians have perfect diving ability, which can help them to resist atmospheric hypoxia9. Consequently, the unique molecular signatures of the alligator diving adaptation provide a new perspective into hypoxia resistance.
The Chinese alligator genome is the first complete crocodilian genome to become available, and this will provide a comparative genomic target for deducing the characteristics of the ancestral reptilian genome and rooting the complicated phylogeny of birds. The alligator is also the first TSD species whose genome has been sequenced; therefore, it fills an important gap in resolving sex chromosome evolution.
In addition, this single Chinese alligator genome provides a snapshot of the 1 million-year-old population demography of the Chinese alligator species and effectively captures molecular evidence of past geophysical events and historical gene flow. Genome sequencing of the Chinese alligator provides a valuable resource for future efforts in designing better strategies to help protect this endangered reptile.
Materials and Methods
All samples for genome and transcriptome sequencing were provided by the Changxing Yinjiabian Chinese Alligator Nature Reserve. Illumina sequencing and the SOAPdenovo algorithm were used to assemble the Chinese alligator genome. Sequence similarity at the nucleotide and protein levels was applied to identify repeat sequences in Chinese alligator. Genes were annotated based on the repeat-masked genome sequence using ab initio, homology- and RNA-based gene prediction models. Genome adaptive features were extracted from gene families, dN/dS tests, and lineage-specific genes. Sex chromosome evolution was analyzed by synteny comparison and M-FISH. Population history was reconstructed from SNP data.
Full Materials and Methods are provided in Supplementary information, Data S1.
References
Alfoldi J, Di Palma F, Grabherr M, et al. The genome of the green anole lizard and a comparative analysis with birds and mammals. Nature 2011; 477:587–591.
Brown B . An ancestral crocodile. Novitates: American Museum of Natural History, 1933; 638:1–4.
Colbert EH, Mook CC . The ancestral crocodilian Protosuchus. Bull Am Mus Nat Hist 1951; 97:143–182.
Andersen HT . Physiological adjustments to prolonged diving in the American alligator, Alligator mississippiensis. Acta Physiol Scand 1961; 53:23–45.
Fish FE, Cosgrove LA . Behavioral thermoregulation of small American alligators in water: postural changes in relation to the thermal environment. Copeia 1987; 1987:804–807.
Martin S . Global diversity of crocodiles (Crocodilia, Reptilia) in freshwater. Hydrobiologia 2008; 595:587–591.
Campbell HA, Sullivan S, Read MA, Gordos MA, Franklin CE . Ecological and physiological determinants of dive duration in the freshwater crocodile. Funct Ecol 2010; 24:103–111.
Wright JC, Kirshner DS . Allometry of lung-volume during voluntary submergence in the saltwater crocodile Crocodylus porosus. J Exp Biol 1987; 130:433–436.
Huchzermeyer FW, eds. Crocodiles: Biology, Husbandry and Diseases. Cambridge: CABI Publishing, 2003.
Wright JC . Energy metabolism during unrestrained submergence in the saltwater crocodile Crocodylus porosus. Physiol Zool 1987; 60:515–523.
Thorbjarnarson J, Wang X, eds. The Chinese Alligator: Ecology, Behavior, Conservation and Culture. Baltimore: The Johns Hopkins Universiity Press, 2010.
Gleich O . Auditory primary afferents in the starling: Correlation of function and morphology. Hear Res 1989; 37:255–267.
Wever EG . Hearing in the crocodilia. Proc Natl Acad Sci USA 1971; 68:1498–1500.
Higgs D, Brittan-Powell E, Soares D, et al. Amphibious auditory responses of the American alligator (Alligator mississipiensis). J Comp Physiol A 2002; 188:217–223.
Merchant ME, Mills K, Leger N, Jerkins E, Vliet KA, McDaniel N . Comparisons of innate immune activity of all known living crocodylian species. Comp Biochem Physiol B Biochem Mol Biol 2006; 143:133–137.
Merchant ME, Leger N, Jerkins E, et al. Broad spectrum antimicrobial activity of leukocyte extracts from the American alligator (Alligator mississippiensis). Vet Immunol Immunopathol 2006; 110:221–228.
Janzen FJ, Paukstis GL . Environmental sex determination in reptiles: ecology, evolution, and experimental design. Q Rev Biol 1991; 66:149–179.
Hillier LW, Miller W, Birney E, et al. Sequence and comparative analysis of the chicken genome provide unique perspectives on vertebrate evolution. Nature 2004; 432:695–716.
Dalloul RA, Long JA, Zimin AV, et al. Multi-platform next-generation sequencing of the domestic Turkey (Meleagris gallopavo): genome assembly and analysis. PLoS Biol 2010; 8:e1000475.
Warren WC, Clayton DF, Ellegren H, et al. The genome of a songbird. Nature 2010; 464:757–762.
Cohen MM, Gans C . The chromosomes of the order Crocodilia. Cytogenetics 1970; 9:81–105.
Willis RE, McAliley LR, Neeley ED, Densmore LD . Evidence for placing the false gharial (Tomistoma schlegelii) into the family Gavialidae: inferences from nuclear gene sequences. Mol Phylogenet Evol 2007; 43:787–794.
Thorbjarnarson J, Wang X . The conservation status of the Chinese alligator. Oryx 1999; 33:152–159.
Wang ZH, Yao H, Ding YZ, John T, Wang XM . Testing reintroduction as a conservation strategy for the critically endangered Chinese alligator: movements and home range of released captive individuals. Chin Sci Bull 2011; 56:2586–2593.
Li R, Zhu H, Ruan J, et al. De novo assembly of human genomes with massively parallel short read sequencing. Genome Res 2010; 20:265–272.
Costantini M, Cammarano R, Bernardi G . The evolution of isochore patterns in vertebrate genomes. BMC Genomics 2009; 10:146.
Gilbert N, Labuda D . CORE-SINEs: Eukaryotic short interspersed retroposing elements with common sequence motifs. Proc Natl Acad Sci USA 1999; 96:2869–2874.
Alkan C, Sajjadian S, Eichler EE . Limitations of next-generation genome sequence assembly. Nat Methods 2011; 8:61–65.
She X, Cheng Z, Zollner S, Church DM, Eichler EE . Mouse segmental duplication and copy number variation. Nat Genet 2008; 40:909–914.
Wong LH, Choo KHA . Evolutionary dynamics of transposable elements at the centromere. Trends Genet 2004; 20:611–616.
Qiu Q, Zhang G, Ma T, et al. The yak genome and adaptation to life at high altitude. Nat Genet 2012; 44:946–949.
Xu X, Pan S, Cheng S, et al. Genome sequence and analysis of the tuber crop potato. Nature 2011; 475:189–195.
Farmer CG, Carrier DR . Ventilation and gas exchange during treadmill locomotion in the American alligator (Alligator mississippiensis). J Exp Biol 2000; 203:1671–1678.
Komiyama NH, Miyazaki G, Tame J, Nagai K . Transplanting a unique allosteric effect from crocodile into human haemoglobin. Nature 1995; 373:244–246.
Romero MF, Fulton CM, Boron WF . The SLC4 family of HCO3− transporters. Pflügers Archiv 2004; 447:495–509.
Jensen FB, Wang T, Jones DR, Brahm J . Carbon dioxide transport in alligator blood and its erythrocyte permeability to anions and water. Am J Physiol 1998; 274:R661–R671.
Letunic I, Yamada T, Kanehisa M, Bork P . iPath: interactive exploration of biochemical pathways and networks. Trends Biochem Sci 2008; 33:101–103.
Carvalho O, Gonçalves C, Gómez-Soler M, et al. Comparative physiology of the respiratory system in the animal kingdom. Open Biol J 2011; 4:35–46.
Brown D, Zhu XL, Sly WS . Localization of membrane-associated carbonic anhydrase type IV in kidney epithelial cells. Proc Natl Acad Sci USA 1990; 87:7457–7461.
Bakouh N, Benjelloun F, Hulin P, et al. NH3 is involved in the NH4+ transport induced by the functional expression of the human Rh C glycoprotein. J Biol Chem 2004; 279:15975–15983.
Joubert PH, Grossmann M . Local and systemic effects of Na+/K+ATPase inhibition. Eur J Clin Invest 2001; 31:1–4.
Medeiros-Domingo A, Kaku T, Tester DJ, et al. SCN4B-encoded sodium channel beta4 subunit in congenital long-QT syndrome. Circulation 2007; 116:134–142.
Delaney JT, Muhammad R, Blair MA, et al. A KCNJ8 mutation associated with early repolarization and atrial fibrillation. Europace 2012; 14:1428–1432.
Liu AH, Zhang X, Stolovitzky GA, Califano A, Firestein SJ . Motif-based construction of a functional map for mammalian olfactory receptors. Genomics 2003; 81:443–456.
Niimura Y, Nei M . Evolution of olfactory receptor genes in the human genome. Proc Natl Acad Sci USA 2003; 100:12235–12240.
Inoue S, Imamura A, Okazaki Y, et al. Synaptotagmin XI as a candidate gene for susceptibility to schizophrenia. Am J Med Genet B Neuropsychiatr Genet 2007; 144B:332–340.
Yu J, He X, Yao D, Li Z, Li H, Zhao Z . A sex-specific association of common variants of neuroligin genes (NLGN3 and NLGN4X) with autism spectrum disorders in a Chinese Han cohort. Behav Brain Funct 2011; 7:13.
Terakita A . The opsins. Genome Biol 2005; 6:213.
Kefalov VJ, Crouch RK, Cornwall MC . Role of noncovalent binding of 11-cis-retinal to opsin in dark adaptation of rod and cone photoreceptors. Neuron 2001; 29:749–755.
Hughes I, Blasiole B, Huss D, et al. Otopetrin 1 is required for otolith formation in the zebrafish Danio rerio. Dev Biol 2004; 276:391–402.
Popper AN, Ramcharitar J, Campana SE . Why otoliths? Insights from inner ear physiology and fisheries biology. Mar Freshw Res 2005; 56:497–504.
Mantegazza AR, Magalhaes JG, Amigorena S, Marks MS . Presentation of phagocytosed antigens by MHC class I and II. Traffic 2013; 14:135–152.
Ozato K, Shin D-M, Chang T-H, Morse HC, III . TRIM family proteins and their emerging roles in innate immunity. Nat Rev Immunol 2008; 8:849–860.
Brown MG, Scalzo AA . NK gene complex dynamics and selection for NK cell receptors. Semin Immunol 2008; 20:361–368.
Abeler-Dörner L, Swamy M, Williams G, Hayday AC, Bas A . Butyrophilins: an emerging family of immune regulators. Trends Immunol 2012; 33:34–41.
Leelawongtawon R, Siruntawineti J, Chaeychomsri W, Sattaponpan C . Antibacterial and antifungal activities from Siamese crocodile blood. J Med Assoc Thai 2010; 93:S58–S64.
Preecharram S, Daduang S, Bunyatratchata W, Araki T, Thammasirirak S . Antibacterial activity from Siamese crocodile (Crocodylus siamensis) serum. Afr J Biotechnol 2008; 7:3121–3128.
Cederlund A, Gudmundsson GH, Agerberth B . Antimicrobial peptides important in innate immunity. FEBS J 2011; 278:3942–3951.
Smith CA, Sinclair AH . Sex determination: insights from the chicken. Bioessays 2004; 26:120–132.
McQueen HA, Clinton M . Avian sex chromosomes: dosage compensation matters. Chromosome Res 2009; 17:687–697.
Backstrom N, Ceplitis H, Berlin S, Ellegren H . Gene conversion drives the evolution of HINTW, an ampliconic gene on the female-specific avian W chromosome. Mol Biol Evol 2005; 22:1992–1999.
Zhao Y, Lu H, Yu HS, Cheng HH, Zhou RJ . Multiple alternative splicing in gonads of chicken DMRT1. Dev Genes Evol 2007; 217:119–126.
Anand A, Patel M, Lalremruata A, et al. Multiple alternative splicing of DMRT1 during gonadogenesis in Indian mugger, a species exhibiting temperature-dependent sex determination. Gene 2008; 425:56–63.
Manolakou P, Lavranos G, Angelopoulou R . Molecular patterns of sex determination in the animal kingdom: a comparative study of the biology of reproduction. Reprod Biol Endocrinol 2006; 4:59.
Yoshimoto S, Okada E, Umemoto H, et al. A W-linked DM-domain gene, DM-W, participates in primary ovary development in Xenopus laevis. Proc Natl Acad Sci USA 2008; 105:2469–2474.
Matsuda M, Nagahama Y, Shinomiya A, et al. DMY is a Y-specific DM-domain gene required for male development in the medaka fish. Nature 2002; 417:559–563.
SFA. Status Quo and Future of Conservation for Chinese Alligator and Crocodiles in the World. International Workshop on Conservation and Reintroduction of Chinese Alligator. Beijing: Chinese Forestry Publishing House, 2002.
Li H, Durbin R . Inference of human population history from individual whole-genome sequences. Nature 2011; 475:493–496.
Zhou SZ, Wang XL, Wang J, Xu LB . A preliminary study on timing of the oldest Pleistocene glaciation in Qinghai-Tibetan Plateau. Quat Int 2006; 154:44–51.
Hedges SB, Kumar S . Precision of molecular time estimates. Trends Genet 2004; 20:242–247.
Beall CM . Two routes to functional adaptation: Tibetan and Andean high-altitude natives. Proc Natl Acad Sci USA 2007; 104:8655–8660.
Urbina MA, Forster ME, Glover CN . Leap of faith: voluntary emersion behaviour and physiological adaptations to aerial exposure in a non-aestivating freshwater fish in response to aquatic hypoxia. Physiol Behav 2011; 103:240–247.
Acknowledgements
We are grateful to the Changxing Yinjiabian Chinese Alligator Nature Reserve for providing samples. We especially thank Drs Zeng XM, Han FP, and Chen ZK for their help in performing the FISH experiments. This work was supported by the Major Research Plan of the National Natural Science Foundation of China (30730019 and 31230010), the National Basic Research Program of China (the 973 program; 2007CB411600), the State Forestry Administration, and the Fundamental Research Funds for the Central Universities of China.
Author information
Authors and Affiliations
Corresponding author
Additional information
( Supplementary information is linked to the online version of the paper on the Cell Research website.)
Supplementary information
Supplementary information, Figure S1
Quality assessment of de novo genome assembles using the full sequences of 5 repeat-rich and GC-rich bacterial artificial chromosomes (BACs) as references. (PDF 330 kb)
Supplementary information, Figure S2
Comparisons of GC content in the Chinese alligator, green anole, chicken, zebra finch, human, and clawed frog. (PDF 72 kb)
Supplementary information, Figure S3
Divergence comparisons of transposable elements from the Chinese alligator, green anole, chicken, and zebra finch. (PDF 322 kb)
Supplementary information, Figure S4
Comparisons of the distribution of segmental duplications among the Chinese alligator, green anole, chicken, and zebra finch. (PDF 370 kb)
Supplementary information, Figure S5
Circos ideograms depicting genomic synteny between the Chinese alligator and chicken (A) and between the Chinese alligator and green anole (B). (PDF 681 kb)
Supplementary information, Figure S6
The maximum likelihood phylogenetic trees of hemoglobin (Hb) alpha (A) and beta (B) genes from the alligator and other reptiles. (PDF 217 kb)
Supplementary information, Figure S7
Expansion and contraction of CAFÉ-based olfactory receptor gene families. (PDF 113 kb)
Supplementary information, Figure S8
Neighbor-joining phylogenetic analysis of olfactory receptors from 6 species. (PDF 434 kb)
Supplementary information, Figure S9
Scatter plots of differentially expressed genes in the testis and ovary glands of the Chinese alligator. (PDF 124 kb)
Supplementary information, Figure S10
Chromosomal assignments of orthologs in the human (A) and chicken (B) to genes found to be differentially expressed (log16 RPKM (testis/ovary) ≥1; ∼ top 20% of differentially expressed genes) between the ovary and testis of the Chinese alligator. (PDF 412 kb)
Supplementary information, Figure S11
The maximum likelihood phylogenetic tree of the HINTZ and HINTW genes from 5 reptiles. (PDF 162 kb)
Supplementary information, Figure S12
The maximum likelihood phylogenetic trees of the FET1 genes. (PDF 177 kb)
Supplementary information, Figure S13
Fluorescence in situ hybridization of DRMT1 (A) and HINTZ (B) in the alligator. (PDF 61 kb)
Supplementary information, Figure S14
Genomic synteny between the alligator scaffolds and chicken Z chromosome (A) and the differentially expressed genes in the large syntenic blocks in panel A (B). (PDF 236 kb)
Supplementary information, Figure S15
Comparison of DMRT1 genes in the alligator, chicken, and crocodile. (PDF 212 kb)
Supplementary information, Figure S16
Alignment of amino acid sequences deduced from the DMRT1 splice variants of the alligator, crocodile, and chicken. (PDF 707 kb)
Supplementary information, Figure S17
Distributions of single nucleotide polymorphisms from the Chinese alligator (A), chicken (B), green anole (C), and human (D). (PDF 143 kb)
Supplementary information, Figure S18
Pairwise sequentially Markovian coalescent estimates for the population histories of the Chinese alligator (A), chicken (B), green anole (C), and human (D). (PDF 270 kb)
Supplementary information, Table S1
Predicted protein-coding genes of the Chinese alligator (PDF 14 kb)
Supplementary information, Table S2
Functional annotation of the Chinese alligator (PDF 8 kb)
Supplementary information, Table S3
Transposable elements (TEs) in the Chinese alligator. (PDF 13 kb)
Supplementary information, Table S4
GC content in different genomic regions (PDF 7 kb)
Supplementary information, Table S5
Transposable elements (TEs) in the Chinese alligator, green anole, chicken, and zebra finch (PDF 14 kb)
Supplementary information, Table S6
Gene families deduced by TreeFam (PDF 12 kb)
Supplementary information, Table S7
Positively selected genes (PSGs) identified by comparison with orthologous genes in the green anole lizard, zebra finch, chicken, and turkey (PDF 102 kb)
Supplementary information, Table S8
Rapidly evolving GO categories in the alligator lineage (PDF 76 kb)
Supplementary information, Table S9
Functional annotation of alligator lineage-specific genes in the alligator genome (PDF 9 kb)
Supplementary information, Table S10
KEGG pathway enrichment of positively selected genes (PSGs) in the Chinese alligator genome (PDF 12 kb)
Supplementary information, Table S11
Enriched Pfam domains of positively selected genes (PSGs) in the alligator genome (PDF 43 kb)
Supplementary information, Table S12
GO enrichment of significantly expanded gene families in the alligator; CAFÉ analysis of 4 species (PDF 37 kb)
Supplementary information, Table S13
Olfactory receptors (ORs) re-annotated for 6 species (PDF 26 kb)
Supplementary information, Table S14
GO enrichment of alligator lineage-specific genes (PDF 32 kb)
Supplementary information, Table S15
GO enrichment of alligator-specific gene families derived from TreeFam analysis of 6 species (PDF 32 kb)
Supplementary information, Table S16
Enriched Pfam domains of the genes identified in the blood transcriptome and proteome (PDF 29 kb)
Supplementary information, Data S1
Materials and Methods (PDF 381 kb)
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0
About this article
Cite this article
Wan, QH., Pan, SK., Hu, L. et al. Genome analysis and signature discovery for diving and sensory properties of the endangered Chinese alligator. Cell Res 23, 1091–1105 (2013). https://doi.org/10.1038/cr.2013.104
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2013.104
Keywords
This article is cited by
-
Evidence of venom factor-like in crocodilians
Organisms Diversity & Evolution (2023)
-
Comparative transcriptomic analysis of the brain in Takifugu rubripes shows its tolerance to acute hypoxia
Fish Physiology and Biochemistry (2021)
-
Multi-omics analysis reveals that natural hibernation is crucial for oocyte maturation in the female Chinese alligator
BMC Genomics (2020)
-
Molecular analysis and bioinformatic characterization of cooper, zinc-superoxide dismutase (Cu/Zn-sod) gene of Caiman latirostris
Molecular Biology Reports (2020)
-
Evidence that DNA repair genes, a family of tumor suppressor genes, are associated with evolution rate and size of genomes
Human Genomics (2019)