Abstract
The Hedgehog (Hh) family of secreted proteins is essential for development in both vertebrates and invertebrates. As one of main morphogens during metazoan development, the graded Hh signal is transduced across the plasma membrane by Smoothened (Smo) through the differential phosphorylation of its cytoplasmic tail, leading to pathway activation and the differential expression of target genes. However, how Smo transduces the graded Hh signal via the Costal2 (Cos2)/Fused (Fu) complex remains poorly understood. Here we present a model of the cell response to a Hh gradient by translating Smo phosphorylation information to Fu dimerization and Cubitus interruptus (Ci) nuclear localization information. Our findings suggest that the phosphorylated C-terminus of Smo recruits the Cos2/Fu complex to the membrane through the interaction between Smo and Cos2, which further induces Fu dimerization. Dimerized Fu is phosphorylated and transduces the Hh signal by phosphorylating Cos2 and Suppressor of Fu (Su(fu)). We further show that this process promotes the dissociation of the full-length Ci (Ci155) and Cos2 or Su(fu), and results in the translocation of Ci155 into the nucleus, activating the expression of target genes.
Similar content being viewed by others
Introduction
One of the central questions in the developmental biology is how diverse cell types are generated and organized to form appropriate body patterns from a single cell type. A theme that has emerged from the study of model organisms is that cells choose fates and form patterns based on the positional information they have received, which is often provided by the evolutionarily conserved families of secreted signaling molecules, morphogens. The Hedgehog (Hh) family of secreted proteins represents one of the major families of these signaling molecules. Members of the Hh family govern cell proliferation, differentiation and patterning in a range of tissues during development in both vertebrates and invertebrates 1. Malfunction of Hh signaling causes congenital disorders and cancers, such as birth defects, basal cell carcinoma and obesity in humans 2, 3, 4. Thus, the study of Hh signaling transduction is of great importance in understanding the mechanism of oncogenesis and obesity, and how different thresholds of a morphogen can generate distinct development outputs.
The Hh pathway, which was first elucidated in Drosophila, is composed of a series of repressive reactions 3, 5. In the absence of Hh, downstream gene transcription is blocked by the Hh receptor, the 12-transmembrane protein Patched (Ptc), which inhibits the function of the 12-transmembrane GPCR-like protein Smoothened (Smo) 3, 6. Under this condition, a large protein complex, which contains the kinesin-related protein Costal2 (Cos2) and the Ser/Thr kinase Fused (Fu), associates with the transcription factor Cubitus interruptus (Ci) and other kinases including cAMP-dependent protein kinase (PKA), GSK3 and CKI. The full-length Ci (Ci155) is phosphorylated and processed to generate a truncated form of Ci, Ci75, which functions as a Hh pathway repressor to block downstream gene expression. In the presence of Hh, however, the binding of Hh to Ptc allows Smo to be activated, resulting in the transduction of the Hh signal through the Cos2/Fu complex. In this case, Ci155 is stabilized and translocates to the nucleus, where it functions as a transcriptional activator, turning on downstream gene expression 3. Therefore, a graded Hh signal could be translated into different transcriptional responses via Hh signaling components. However, more questions remain regarding Hh signaling cascade. For example, it is not known how the gradient information of a Hh signal is transduced through Smo and the Cos2/Fu complex to distinct downstream target genes. In addition, although many of the canonical Hh signal components are phosphorylated in Drosophila, including Smo, Cos2, Fu and Su(fu) 7, the specific role for these phosphorylation events in the regulation of Hh signal transduction remains poorly understood. For instance, Smo is phosphorylated by PKA and CKI in the presence of Hh 8, 9, 10 and transduces the Hh signal by interacting with Cos2/Fu complex 11, 12, 13, 14, 15, 16. However, it is poorly understood how the kinase activity of Fu is regulated and how activated Fu functions in this signaling cascade.
In this study, we found that Smo activation, different levels of which are induced by different Smo mutants, recruits the Cos2/Fu complex to the membrane through the interaction between Smo and Cos2, which further induces Fu dimerization. Dimerized Fu is auto-phosphorylated and transduces the Hh signal by phosphorylating Cos2 and Su(fu). This process promotes the dissociation of the full-length Ci and Cos2 or Su(fu), resulting in the translocation of Ci155 into the nucleus, which activates the expression of target genes. Our findings indicate that cells respond to Hh signaling through the regulation of Fu dimerization and its kinase activity.
Results
The recruitment of the Cos2/Fu complex by Smo to the plasma membrane is regulated by the Hh gradient
Smo could potentially transduce Hh signaling by physically interacting with the Cos2/Fu complex 11, 12, 13, 14. To investigate how the interaction between Smo and the Cos2/Fu complex is regulated by Hh, we first examined the co-localization of Smo and the Cos2/Fu complex in the presence or absence of Hh signaling. By co-transfecting Smo (tagged by CFP at its C terminus, Smo-CFPC), Cos2 (tagged by YFP at its C terminus, Cos2-YFPC) and Fu (tagged by Myc at its N terminus, Myc-Fu) in S2 cells, we found that Cos2 co-localized with Fu independent of Hh signaling. In the absence of Hh, however, the Cos2/Fu complex only partially co-localized with Smo in the cytoplasm. In the presence of Hh, the Cos2/Fu complex was highly co-localized with Smo at the cell membrane (Figure 1A). To investigate whether the accumulation of the Cos2/Fu complex at the plasma membrane, induced by Hh signaling, were dependent on Smo activity, we examined the co-localization of Cos2/Fu with two Smo mutants: SmoSA, a phosphorylation-defective Smo that fails to respond to Hh signaling, and SmoSD, a constitutively active Smo irrespective of Hh signaling 9, 17. If the plasma membrane accumulation of the Cos2/Fu complex, induced by Hh, was independent of Smo activity, the Cos2/Fu complex co-transfected with SmoSA would be located at the cell membrane following Hh treatment. However, if the accumulation was dependent on Smo activity, the Cos2/Fu complex would be located at the cell membrane following co-transfection with SmoSD in the absence of Hh. We found that, the Cos2/Fu complex remained in the cytoplasm when co-transfected with SmoSA-CFPC, even after Hh treatment. In contrast, when SmoSD-CFPC was co-transfected, the Cos2/Fu complex was recruited to the cell membrane even without Hh treatment (Figure 1A). These results suggest that the cellular membrane accumulation of the Cos2/Fu complex is dependent on the activation of Smo. We next investigated how the Hh signal controls the interaction between Smo and Cos2/Fu. We used SmoSA-CFPC, SmoWT-CFPC, SmoSD1-CFPC, SmoSD12-CFPC and SmoSD-CFPC to examine the effects of different Hh concentration inputs because the three Smo mutants (SmoSD1, SmoSD12 and SmoSD), which contain Ser to Asp substitution in one, two and three phosphorylation clusters, respectively, and exhibit progressively higher levels of constitutive signaling can mimic low to high activity, respectively 9. We found that an increasing amount of Smo co-localized with the Cos2/Fu complex when Smo activation was increased, indicating that the interaction between Smo and Cos2/Fu might be regulated by the Hh gradient (Figure 1B).
Hh induces the recruitment of Cos2/Fu to the membrane through the interaction between Smo and the Cos2/Fu complex. (A) S2 cells were co-transfected with Smo-CFPC, SmoSA-CFPC, SmoSD-CFPC(green) and Cos2-YFPC (red), and Myc-tagged Fu, treated with or without Hh, and immunostained with anti-Myc antibody (blue). In the absence of Hh, only a few Cos2/Fu complexes co-localized with wild-type Smo. In the presence of Hh, the majority of Co2/Fu complexes co-localized with wild-type Smo and was recruited to the cell plasma membrane. When SmoSA was co-transfected, Cos2/Fu complexes were still localized in the cytoplasm following Hh treatment. When SmoSD was co-transfected, Cos2/Fu complexes were localized on the cell plasma membrane even without Hh treatment. (B) Quantification of Smo co-localized with Cos2 by using Smo mutants to mimic Hh gradient. (C, E) FRET efficiency from the indicated constructs in S2 cells treated with or without Hh-condition medium (mean ± sd, n ≥ 15). (D, F) FRET efficiency between Smo-CFPC/Cos2-YFPC(D), or Cos2-CFPC/Cos2-YFPC(F), expressed in wing discs (mean ± sd, n ≥ 8). A, A-compartment cells away from the A/P boundary; P, P-compartment cells; A/P, A-compartment cells adjacent to the A/P boundary.
Fluorescence resonance energy transfer (FRET) has been broadly employed in the detection of protein-protein interaction in living cells 18. To investigate how Smo interacts with the Cos2/Fu complex, we measured FRET between the Smo C-terminal tail and Cos2 or Fu in both S2 cells and wing imaginal discs. The last 59 amino acids of the Smo C-terminal tail could interact with Fu directly 19. However, no FRET activity was detected between Smo and Fu by using either Smo-CFPC/Fu-YFPN (Fu tagged by YFP at N-terminus) or Smo-CFPC/Fu-YFPC (Fu tagged by YFP at C-terminus) in our experiments (data not shown). The last 59 amino acids of the Smo C-terminal tail interacts directly with Fu 19, but the CFP tag on the C-terminus of Smo might be too far away from the N- or C-terminus of Fu. This may be the reason why we could not observe a FRET signal between Fu and Smo. We then found that FRET between Cos2-YFPC and Smo-CFPC was increased in response to Hh or following the co-transfection of Smo phosphor-mimetic mutants. Moreover, increased FRET was also well correlated with Smo activation (Figure 1C). In addition, we employed CFP and YFP as a control. In both the absence and presence of Hh, CFP and YFP showed a very low FRET signal (Figure 1C). Because Hh can stabilize Smo and increase Smo protein levels, increased FRET between Smo-CFPC and Cos2-YFPC might simply be due to the increase of FRET donor level. To test this possibility, we switched the donor to Cos2-CFPC and the receptor to Smo-YFPC. The results suggested that FRET between Smo and Cos2 increased dramatically with Hh treatment (Figure 1C). A similar change in FRET was found when Smo-CFPC and Cos2-YFPC were expressed in fly wing imaginal discs (Figure 1D and Supplementary information, Figure S1). Collectively, our data indicate that, in response to a Hh gradient and Smo activity, the Cos2/Fu complex is recruited to the cell membrane through an interaction between Smo and Cos2.
Cos2 is a kinesin-related protein that may form a dimer 20, 21. We measured FRET changes between Cos2-CFPC (Cos2 tagged CFP at its C terminus) and Cos2-YFPC (Cos2 tagged YFP at its C terminus) to investigate the dimerization of Cos2. We found that Cos2 formed dimers in both S2 cells and wing imaginal discs that were not dependent on the presence of Hh. However, Cos2-Cos2 FRET was decreased with Hh treatment, suggesting that in the presence of Hh, the recruitment of the Cos2/Fu complex to Smo may induce a conformational change of the Cos2 dimer (Figure 1E-1F and Supplementary information, Figure S2).
The Cos2/Fu complex switches the binding region on Smo in the presence of Hh signal
To further investigate which region of the Smo C-tail contributes to the interaction between Smo and the Cos2/Fu complex, we examined the co-localization of Smo with Cos2 or Fu under different conditions. First, we co-expressed Smo-CFPC and Cos2-YFPC when endogenous Fu was knocked down (Figure 2A, 2A′ and Supplementary information, Figure S3A), or we co-transfected with Smo-CFPC and Fu-YFPN in the absence of endogenous Cos2 (Figure 2B, 2B′, Supplementary information, Figure S3B). We found that, in the presence of Hh, the overexpression of Fu or the knockdown of endogenous Fu did not affect the co-localization of Smo with Cos2 on the membrane. However, in the absence of Hh, the percentage of Smo that co-localized with Cos2 decreased in the endogenous Fu knockdown sample compared with the overexpressed Fu sample (Figure 2A and 2A′). In contrast, in the absence of Hh, the co-localization of Fu and Smo in the endogenous Cos2 knockdown condition was greater than the co-localization of Cos2 and Smo in the endogenous Fu knockdown condition (Figure 2A, 2A′, 2B and 2B′). In the presence of Hh, the Smo plasma membrane localization was decreased in the absence of Cos2 (Figure 2B, compare Smo plasma membrane localization with or without Cos2 RNAi in the presence of Hh), and the majority of Fu protein failed to co-localize with Smo on the plasma membrane in the absence of Cos2 (Figure 2B and 2B′). These results, together with the results shown in Figure 1A, imply that Fu contributes to the partial co-localization of the Cos2/Fu complex with Smo in the absence of Hh, whereas Cos2 is responsible for the recruitment of the Cos2/Fu complex to the plasma membrane by activated Smo in the presence of Hh. Although previous research had provided some biochemical evidence for the binding region of Smo with Cos2 11, the physiological evidence for the interacting region of Smo with Cos2 is lacking. As the binding of the Cos2/Fu complex with Smo may be due to a positional switch in response to Hh, we attempted to determine the region of the Smo C-tail that contributes to its binding with the Cos2/Fu complex. In Smo C-tail, a region of three phosphorylation clusters (amino acids 661 to 818), is important for Smo activity, while the last 59 amino acids of Smo are critical for its binding to Fu 9, 19. We generated two Smo mutants, SmoΔ974 and SmoΔ818, in which the Smo C-tail fragment from either amino acid 974 or 818 was deleted. We found that, with or without Hh, Fu failed to co-localize in these two Smo mutants when the endogenous Cos2 was knocked down. In addition, no co-localization between Fu and SmoSDΔ974 or SmoSDΔ818 was observed under the same condition (Figure 2C). With co-transfected Cos2, however, Fu co-localized with SmoSDΔ974 or with SmoΔ974 when Hh was present (Supplementary information, Figure S4A). Fu failed to co-localize with SmoSDΔ818 or with SmoΔ818 regardless of the presence or absence Hh (Supplementary information, Figure S4A). These data suggest that Cos2 likely binds to a different region of the Smo C-tail, which is important for the binding of the Cos2/Fu complex to Smo in response to Hh signaling, and its binding region is different from Fu. We further dissected the specific region of Smo that is responsible for Cos2 binding. In the absence of Hh, Cos2 did not co-localize with wild-type Smo or Smo mutants when the endogenous Fu was knocked down (Figure 2A, 2D and Supplementary information, Figure S4B). In the presence of Hh, however, Cos2 co-localized with both SmoΔ974 and SmoSDΔ974 on the cell membrane regardless of the level of Fu. In contrast, Cos2 failed to co-localize with SmoSDΔ818 or with SmoΔ818 in the presence of Hh, suggesting that amino acids 818-974 of Smo are important for the physiological interaction between Smo and Cos2 in the presence of Hh (Figure 2D and Supplementary information, Figure S4B). The FRET result in Figure 1C also suggests that the C-terminal region of Smo is far from the Cos2 C-terminus in the absence of Hh. Taken together, these data suggest that the Cos2/Fu complex interacts with Smo at different regions in the presence or absence of Hh. In the presence of Hh, Cos2 mediates the interaction between the Cos2/Fu complex and Smo by binding to the region of Smo between amino acids 818 and 974; in the absence of Hh, the C-terminal part of Smo (amino acids 974-1 035) interacts with Fu.
The binding regions of Smo with Cos2 and Fu in the absence or presence of Hh. (A) S2 cells were transfected with Smo-CFPC and Cos2-YFPC with control RNAi and Fu, or the endogenous Fu knocked down by dsRNA in the absence or presence of Hh. (B) S2 cells were transfected with Smo-CFPC and Fu-YFPN with control RNAi and Cos2, or the endogenous Cos2 was knocked down by dsRNAi in the absence or presence of Hh. Then, the co-localization of Smo with Cos2 (A) or Fu (B) was examined. (A′) Quantification of Smo co-localized with Cos2 by using dsRNA of Fu to knockdown endogenous Fu. (B′) Quantification of Smo co-localized with Fu by using dsRNA of Cos2 to knockdown endogenous Cos2. (C, D) S2 cells were transfected with indicated Smo mutant constructs and Fu construct with endogenous Cos2 knocked down (C), or Cos2 construct with endogenous Fu knocked down (D), treated with or without Hh, respectively. (E) FRET efficiency from the indicated constructs in S2 cells treated with or without Hh (mean ± sd, n ≥ 15).
The C-tail of Smo keeps a closed and inactivated conformation in the absence of Hh. In this condition, the Smo C-terminal region from amino acids 656 to 755 self-interacts with the region from amino acids 860 to 1 035 17. Therefore, in the absence of Hh, the binding region of Cos2 on the Smo C-tail is likely masked by the self-interaction between these two Smo C-tail regions. As the interaction between Fu and the C-terminal region of Smo in the absence of Hh may place Cos2 in close proximity to the third cytoplasm loop of Smo, we further tested this possibility by inserting a CFP into the third internal loop of SmoWT, SmoSA or SmoSD to generate SmoL3-CFP, SmoL3SA-CFP and SmoL3SD-CFP, respectively, and co-transfecting them with Cos2-YFPC in S2 cells. We found that FRET between SmoL3-CFP and Cos2-YFPC changed from high to low along with Hh stimulation. FRET between SmoL3SA-CFP and Cos2-YFPC was high and FRET between SmoL3SD-CFP and Cos2-YFPC was low (Figure 2E). These results indicate that Cos2 is close to the third internal loop of Smo when Fu interacts with the C-terminal end of Smo in the absence of Hh.
Hh promotes Fu dimerization
As both Smo and Cos2 form dimers, and the Cos2/Fu complex interacts with Smo following Hh treatment, we investigated whether Fu forms a dimer in response to Hh. We examined FRET efficiency by co-transfecting Fu-CFPN and Fu-YFPN in S2 cells together with Smo and Cos2 with or without Hh treatment. FRET between Fu-CFPN and Fu-YFPN was low in the absence of Hh but increased dramatically in the presence of Hh. The different phosphor-mimetic forms of Smo progressively increased the basal FRET between Fu-CFPN and Fu-YFPN in a graded manner (Figure 3A). In wing imaginal discs, Hh also induced a significant increase in FRET between Fu-CFPN and Fu-YFPN at the P-compartment (we used Ms1096 to drive Fu protein expression in the wing pouch) and the A/P boundary (to generate a more convincing result, we used Ptc gal4 to drive Fu protein expression at the A/P boundary in a separate experiment), compared with that at the A-compartment (Figure 3B and Supplementary information, Figure S5). We also measured FRET between Fu-CFPC and Fu-YFPC in S2 cells with similar results (Supplementary information, Figure S6). Therefore, Fu forms a dimer in response to Hh or graded Smo activity. We next examined whether Fu dimerization is dependent on Smo and/or Cos2. We found that FRET efficiency between Fu-CFPN and Fu-YFPN was decreased when Cos2 or Smo expression was inhibited by RNAi, suggesting that the dimerization of Fu induced by Hh or graded Smo activity is Smo and Cos2 dependent (Figure 3C-3D and Supplementary information, Figure S7A-S7B).
The dimerization of Fu is induced by Hh and is Smo and Cos2 dependent. (A, B) FRET efficiency of Fu-CFPN and Fu-YFPN in S2 cells (mean ± sd, n ≥ 15) (A), or in wing discs (mean ± sd, n ≥ 10) (B). (C, D) In the absence/presence of Hh, FRET efficiency of Fu-CFPN and Fu-YFPN co-expressing Smo (C) or Cos2 (D) with control RNAi, Cos2 RNAi (C) or Smo RNAi (D), to knockdown the endogenous protein. (E) Smo-CFPC, Cos2-YFPC or Cos2ΔMD-YFPC and Myc-Fu were co-transfected in S2 cells, with or without Hh treatment, to detect the effects of the Cos2 motor domain on the co-localization of Smo and the Cos/Fu complex. Myc-Fu was immunostained (blue). (E′) Quantification of Smo that co-localized with Cos2 or Cos2ΔMD in the absence or presence of Hh. (F) FRET efficiency of Smo-CFPC and Cos2-YFPC or Cos2ΔMD-YFPC with Myc-Fu co-transfected in S2 cells with or without Hh treatment. (G) FRET efficiency of Smo-CFPC and Cos2-YFPC with control RNAi, or the endogenous Fu knocked down by Fu RNAi in the absence/presence of Hh, respectively.
Cos2 is a kinesin-related protein whose motor domain is important for its transportation along the microtubules 20, 21, 22, 23. We generated a Cos2-YFPC mutant, Cos2ΔMD-YFPC, in which the motor domain was deleted, and examined the co-localization of Smo and Cos2ΔMD. We found that Cos2ΔMD still co-localized with Fu but this Cos2ΔMD/Fu complex failed to be recruited to the membrane following Hh treatment (Figure 3E and 3E′). Moreover, in the presence of Hh, FRET between Smo-CFPC and Cos2ΔMD-YFPC was significantly lower than that between Smo-CFPC and Cos2WT-YFPC (Figure 3F). Therefore, Cos2 transportation mediated by its motor domain is important for the recruitment of the Cos2/Fu complex to the plasma membrane by Smo in response to Hh signaling. Given the fact that Cos2 moves along microtubules and is released in the presence of Hh 20, 21, 23, our results imply that this movement of Cos2 along microtubules may bring Fu in proximity to Smo and then dissociate from microtubules and interact with Smo in the presence of Hh.
We next investigated whether Fu plays a role in the recruitment of the Cos2/Fu complex to the membrane by Hh. We observed a 33% reduction of FRET between Smo and Cos2 when endogenous Fu expression is inhibited by RNAi (Figure 3G and Supplementary information, Figure S7C). Given that the loss of endogenous Fu did not affect the co-localization of Cos2 and Smo on the plasma membrane following Hh treatment (Figure 2A), Fu may only play a minor role in the recruitment of the Cos2/Fu complex by Smo.
Dimerized Fu activates the Hh pathway independently of Smo and Cos2
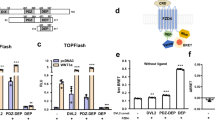
As the formation of Fu dimer is regulated by Hh and requires Smo and Cos2, Smo may transduce the Hh by inducing Fu dimerization, in turn activating the Hh pathway. To test this hypothesis, we investigated whether dimerized Fu could activate downstream gene expression by generating two Fu dimerization systems. The FKBP-based Fu dimerization system is dependent on a synthetic dimerizing agent (AP20187) that crosslinks two molecules carrying FKBP domains based on its sub-nanomolar binding affinity 24, 25. Fv2-Fu contains two FK506-binding domains and can be induced to form a dimer or oligomer upon AP20187 treatment. As the FKBP-AP20187 system is not suitable for in vivo experiments, we employed DivIVA protein to generate another Fu dimerization system. DivIVA (abbreviated to Div) is a coil-coiled domain-containing protein that forms constitutive dimers or oligomers 26. We therefore generated a Div-Fu fusion protein to examine the in vivo function of dimerized Fu. In addition, several Fu mutants were also generated and examined using these two dimerization systems. FuG13V is a kinase-dead and non-functional mutant in which Gly 13 is replaced with Val 15, 27. Fu kinase (FuK) only contains the Fu kinase domain (amino acids 1-306 of Fu), while FuK* carries both the Fu kinase domain and the Su(fu) binding region (amino acids 1-436) 7. We first used the ptc-luciferase assay, a transcriptional reporter for Hh activity, to test the pathway activation by various Fu dimers. We found that either Fv2-Fu treated by AP20187 or Div-Fu was able to activate ptc gene expression (Figure 4A-4B). The dimerized FuK domain had similar activity to full-length Fu, while dimerized FuK* induced the highest reporter activity, which might be due to its phosphorylation on Su(fu) (Figure 4A; see below).
Induced Fu dimerization triggers the Hh pathway, which is independent of Smo and Cos2. (A-E) The ptc-luc reporter assay shows the pathway activity changes in S2 cells transfected with Ci155 and Su(fu). Error bars, 1 sd (triplicate wells). S2 cells were co-transfected with the indicated Div-based Fu expression constructs (A), or FKBP-based Fu constructs and treated with or without AP20187 (B). (C) S2 cells were co-transfected with Fv2-FuG13V or Fv2-Fu, and treated with the graded AP20187. Along with the concentration change of the synthetic agent (AP20187), the ptc-luc reporter activity increased from low to high when the cells were transfected with Fv2-Fu (red line, Fv2-Fu; blue line, Fv2-FuG13V). (D, E) With Smo or Cos2 RNAi to knockdown endogenous protein expression, in both Fu dimerization system based on FKBP or Div, dimerized Fu could trigger ptc-luciferase reporter activity bypassing Smo and Cos2.
FKBP-based dimerization system has a unique advantage. The different concentrations of AP20187 can induce Fu dimerization at different levels and may mimic Hh gradient. Following transfection with Fv2-Fu and treatment with increased concentration of AP20187 in S2 cells, ptc luciferase reporter activity was induced in a graded manner, suggesting that graded Fu dimerization may activate the Hh pathway to varying amounts (Figure 4C). As Smo may transduce a graded Hh signal across the plasma membrane by inducing Fu dimerization and leading to pathway activation, we examined whether FKBP- or Div-induced Fu dimerization could activate downstream gene expression independently of Smo and Cos2. We found that dimerized FuFL or FuK* induced ptc reporter expression in the absence of Smo or Cos2, while the activity of dimerized FuK was partially dependent on Smo (Figure 4D-4E).
Dimerization of Fu triggers its kinase activity and promotes auto-phosphorylation and the phosphorylation of Cos2 and Su(fu)
Fu is the Hh pathway-specific Ser/Thr kinase and is a major member of the Cos2/Fu complex. Previous studies have suggested that Cos2 might be a phosphorylation target of Fu 7, 20, 21, 28, but the mechanism by which Fu kinase activity is regulated remains unknown. Notably, many kinases, such as receptor tyrosine kinases, form a dimer, and then show kinase activity upon ligand binding 29. To examine whether dimerized Fu has kinase activity and to determine its phosphorylation target(s), we performed immunoprecipitation-based phosphorylation experiments in the two dimerization systems. We found that dimerized Fv2-Fu or Fv2-FuK* but not Fv2-FuG13V immunoprecipitated products from S2 cells that were auto-phosphorylated after dimerization in immunoprecipitated product (Figure 5A and Supplementary information, Figure S8A). The dimerized Fu, based on Div-dimerization system, was auto-phosphorylated (Supplementary information, Figure S8B). To determine whether Cos2 can be phosphorylated by dimerized Fu, we used similar kinase assays. In an in vitro kinase assay, dimerized Fu (Fv2-Fu and Div-Fu), but not Fu G13V, phosphorylated Cos2 (Figure 5B1 and Supplementary information, Figure S8C). FuK and FuK* also displayed phosphorylation activity towards Cos2 after dimerization (Figure 5B2-5B3). Therefore, Fu following dimerization is auto-phosphorylated, and can phosphorylate Cos2.
The dimerization of Fu promotes Fu auto-phosphorylation and Cos2 and Su(fu) phosphorylation. (A) Phosphorylation mobility shift of dimerized Fv2-Fu as induced by a synthetic agent (AP20187) in vitro with ATP. Fv2-FuG13V served as a control. Phosphatase was used to dephosphorylate the phosphorylated protein. (B) Phosphorylation of Cos2 by dimerized Fu. In Div-based dimerization system, dimerized Fu can phosphorylate Cos2 in vitro. S2 cells were transfected with Flag-Cos2, Div-FuG13V, Div-Fu, Div-FuKG13V, Div-FuK*G13V or Div-FuK*, respectively. Then, Cos2 or different Div-Fu fusion proteins were concentrated by immunoprecipitation, respectively. The in vitro kinase assays were followed. Phosphatase was used to dephosphorylate the phosphorylated proteins. Div-Fu (B1), Div-FuK (B2) and Div-FuK* (B3) can phosphorylate Cos2 but not the G13V mutants. (C) Phosphorylation of Flag-Su(fu) by activated Fu in an in vitro Div-Fu system. Su(fu) can be phosphorylated by Div-Fu and Div-FuK* but not Div-FuK. In (A-C), the mobility shift band of target proteins were labeled by stars. (D) Smo-CFPC(green), Cos2WT/SA-YFPC (red) and Myc-FuWT/G13V (blue) were co-transfected in S2 cells in the absence/presence of Hh, to detect the effect of Cos2 phosphorylation or Fu kinase activity on co-localization between Smo and Cos2/Fu complex.
Su(fu) binds Fu in the absence of Hh, while, in the presence of Hh, the function of Su(fu) is inhibited by activated Fu 7. Although Su(fu) has been shown to be phosphorylated in response to Hh 13, how this phosphorylation is regulated remains unknown. We next investigated whether Su(fu) phosphorylation is directly affected by dimerized Fu. We found that Su(fu) is phosphorylated by dimerized Div-Fu or Fv2-Fu in an in vitro kinase assay (Figure 5C and Supplementary information, Figure S8D). Moreover, dimerized FuK*, which contains a Su(fu) binding region, but not FuK, which lacks a Su(fu) binding site, phosphorylated Su(fu), indicating that the binding of Fu with Su(fu) is required for Su(fu) phosphorylation (Figure 5C).
To investigate whether phosphorylation of Fu and Cos2 affects the interaction of Cos2/Fu complex with Smo, we examined the co-localization of Smo and Cos2/Fu. A previous study suggested that Fu activity regulated Smo plasma membrane accumulation 15. We also found that FuG13V could attenuate the Smo cell membrane accumulation (Figure 5D). However, we found that Cos2SA (a phosphorylation-defective mutant of Cos2 in which Cos2 amino acid 572 was mutated from Ser to Ala) could not affect the recruitment of the Cos2/Fu complex by Smo after Hh treatment to the plasma membrane (Figure 5D). This result suggests that the phosphorylation status of Cos2 does not affect the interaction between Smo and the Cos2/Fu complex. It also implies that the phosphorylation of Cos2 is likely a downstream event that occurs after the recruitment of Cos2/Fu to the plasma membrane by Smo in the presence of Hh.
Dimerized Fu promotes the dissociation of Ci155 from Cos2 and Su(fu) and Ci155 nuclear translocation
In the presence of Hh, Ci155 dissociates from the Cos2/Fu complex 22, 30. To investigate whether Fu dimerization plays a role in this event, we examined the binding between Ci155 and Cos2 in response to a Hh gradient mimicked by Smo phosphor-mimetic mutants, or by dimerized Fu. We found that the binding between Cos2 and Ci155 became weaker with an increased Hh gradient as mimicked by graded Smo activity (Figure 6A). Importantly, dimerized Fu but not FuG13V induced a similar dissociation of Cos2 and Ci155 to the graded Smo activity, suggesting that Hh-induced Fu dimerization dissociates Ci155 from the Cos2/Fu complex (Figure 6B-6B′).
Phosphorylated Cos2 and Su(fu) by dimerized Fu promote the dissociation of Ci155 from the Cos2/Fu complex or Su(fu) and its nuclear translocation. (A, B-B′, C, D-D″) S2 cells were transfected with Myc-Ci155, Flag-Cos2 (A-B′) or Flag-Su(fu) (C-D′), and the indicated constructs. Cell extracts were immunoprecipitated with anti-Flag antibodies, followed by immunoblotting with indicated antibodies. Western blot for Flag-Cos2 or Flag-Su(fu) is serving as loading control, respectively. The Ci155 expression in the lysate is detected with anti-Myc antibody. (A) S2 cells were transfected with Flag-Cos2, Myc-Ci155 and Smo phosphor-mimetic mutants, which mimicked Hh gradient. (B, B′) FKBP- or Div-based Fu constructs were transfected into S2 cells with Flag-Cos2 and Myc-Ci155. (C) S2 cells were transfected with Flag-Su(fu), Myc-Ci155 and Smo phosphor-mimetic mutants. SmoSD, which mimics the high Hh concentration, but not SmoSD1 and SmoSD12, could induce the dissociation between Su(fu) and Ci155. (D, D′) The dimerized Fu (Fv2-Fu with AP20187 treatment or Div-Fu) decreased the association between Su(fu) and Ci155, compared with Fv2-FuG13V with or without AP20187 treatment, Fv2-Fu without AP20187 treatment, or Div-FuG13V. (D′′) Div-FuK* enhanced dissociation of Su(fu) with Ci155, compared with Div-FuK, Div-FuKG13V and Div-FuK*G13V. (E) S2 cells were transfected with HA-Ci155, or HA-Ci155 and Flag-Su(fu), and treated with Hh or LMB, respectively, or with Hh and LMB treatment. The antibody detecting full-length Ci was used to stain for Ci155 (red), antibody recognizing Flag was used for immunostaining Flag-Su(fu) (green), and the nuclei were stained with DAPI (blue). The nuclei were circled by white line. (F, G) The percentage of S2 cells with Ci155 nuclear localization after co-transfected with Smo phosphor-mimetic mutants to mimic Hh gradient (F), or with dimerized Fu. FKBP- or Div-based dimerized Fu could antagonize Su(fu) and promote Ci155 nuclear translocation (G). In F and G, pUAST and Hh treatment served as negative and positive control, respectively.
Su(fu), which binds Ci/Gli and inhibits the activity of Ci155/Gli, dissociates from Ci155/Gli in response to a high concentration of Hh 7, 31. We next investigated the effect of either the Hh gradient or Fu dimerization on the association of Su(fu) and Ci155. We found that low or medium Hh concentrations, which were mimicked by SmoSD1 or SmoSD12, did not affect the association of Su(fu) and Ci155. However, high concentrations of Hh, mimicked by SmoSD, enhanced the dissociation of Ci155 from Su(fu) (Figure 6C). Similarly, dimerized Fu caused a dissociation of Ci and Su(fu) (Figure 6D-6D′). In addition, Div-FuK* mutant, but not the Div-FuK mutant, promoted the dissociation of Su(fu) and Ci155 (Figure 6D″). Collectively, these data suggest that dimerized Fu phosphorylates Su(fu), leading to the dissociation of Ci155 and Su(fu).
Hh not only prevents the processing of Ci155, but also promotes the nuclear translocation of accumulated Ci155 32, 33, 34, and in the presence of high levels of Hh signaling, the E3 Roadkill/Hib can inhibit Ci nuclear import 35. To determine how Hh and Fu dimerization regulates Ci155 nuclear translocation, we used LMB to block Ci155 nuclear export and Su(fu) to inhibit Ci155 nuclear translocation and then examined Ci155 accumulation in the nucleus. We found that very little Ci155 entered the nucleus without Hh treatment, but this increased dramatically when cells were treated with Hh (Figure 6E). With LMB, almost all Ci155 was located in the nucleus in the absence of Hh, an effect that was blocked by co-transfecting Su(fu). Hh treatment or co-transfecting SmoSD antagonized the negative effect of Su(fu) and promoted Ci155 nuclear translocation (Figure 6E and Supplementary information, Figure S9A). The nuclear translocation of Ci155 was gradually enhanced with increased Smo phosphorylation levels and Hh gradient as mimicked by Smo mutants (Figure 6F and Supplementary information, Figure S9A). Similarly, dimerized Fu, but not dimerized FuG13V, promoted Ci155 nuclear translocation (Figure 6G and Supplementary information, Figure S9B). Taken together, these results suggest that Fu dimerization induced by Hh promotes the dissociation of Ci155 and Cos2 or Su(fu), and promotes nuclear translocation of Ci155.
Dimerization of Fu promotes pathway activation
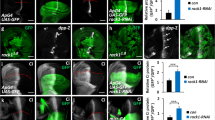
To further determine whether the dimerization of Fu is sufficient to activate the Hh pathway, we examined the activation of the Hh pathway by dimerized Fu in vivo. Because we failed to induce Fu dimerization by feeding the flies AP20187, an approach that has been successfully employed in a mouse system 24, 25, we used the Div-based Fu dimerization system for in vivo experiments. We found that wing imaginal discs expressing UAS-Div-Fu by act>CD2>Gal4, displayed a dpp-lacZ expression in clones at the A-compartment (Figure 7B). In contrast, Div or Div-FuG13V failed to activate dpp-lacZ (Figure 7A and Supplementary information, Figure S10A-S10A″). Moreover, both Div-FuK and Div-FuK* activated dpp-lacZ, with more extensive expression observed in those that expressed Div-FuK* (Figure 7C, 7D). Conversely, Div-FuKG13V, Div-FuK*G13V or wild-type Fu failed to activate dpp expression (Supplementary information, Figure S10B-S10D″). However, the ptc expression levels are not very strong, suggesting that the dimerization of Fu alone could not mimic the highest levels of Hh signaling. Div-Fu, Div-FuK and Div-FuK*, but not Div, also activated ptc expression in the A-compartment and Div-FuK* appeared to have a higher activity than Div-FuK (Figure 7E-7H). Fu, Div-FuG13V, Div-FuKG13V or Div-FuK*G13V failed to induce ptc expression in the A-compartment (data not shown). Altogether, although dimerization of Fu alone does not seem to fully activate the Hh pathway, these data still suggest that Fu dimerization promotes Hh pathway activity in vivo.
In vivo activities of dimerized Fu variants. (A-H) Wing discs expressing the indicated dimerized Fu variants by act>CD2>Gal4. The immunostainings show the expression of Fu variants, dpp-lacZ (dppZ) (A-D) and endogenous Ptc (E-H). In the Div-Fu, Div-FuK and Div-FuK* clones, which were marked by green, dppZ or Ptc (blue) expression was turned on in A-compartment (B, F, Div-Fu; C and G, Div-FuK; D, H, Div-FuK*). However, as the control, DivIVA protein failed to upregulate dppZ or Ptc expression (A, E).
Discussion
The Hh family of morphogens controls distinct cellular outcomes through graded signal activities 1, 17, 36, 37. As one of the main morphogens, Hh forms a gradient when it diffuses from Hh secreting cells to receiving cells. In Drosophila wing imaginal discs, a Hh gradient is required for the patterning of the adult wing. As a long-range morphogen, low concentrations of Hh activate dpp and collier (col) expression, whereas as a short-range morphogen, high Hh concentrations activate ptc and en expression 38, 39, 40. However, how the Hh gradient regulates differential gene expression remains elusive. The seven-transmembrane protein Smo is thought to relay different levels of Hh activity through differential phosphorylation and the graded conformational change of its carboxyl-terminal cytoplasmic tail 9, 17, 41. Previous studies on the Cos2/Fu complex suggested that Smo recruits the Cos2/Fu complex to membranes by physically interacting with the Cos2/Fu complex through its C-tail 11, 22. In the present study, we found that Smo recruits the Cos2/Fu complex to the membrane through the interaction between Smo and Cos2, which is regulated by graded Hh or Smo C-tail phosphorylation levels. Moreover, the motor domain of Cos2 is important for this recruitment. Upon Hh binding with Ptc, the inhibition of Ptc on Smo is terminated. Under this condition, the Smo C-tail interacts with Cos2 at amino acids 818 to 974, and Fu fails to co-localize with the Smo C-tail in the absence of Cos2. Cos2 appears to be essential for the interaction between Cos2/Fu and Smo in the presence of Hh. Fu is important for the interaction between Cos2/Fu and Smo in the absence of Hh. Taken together with data from FRET, these findings suggest that the Cos2/Fu complex interacts with Smo at different regions in the presence or absence of a Hh signal. In the presence of Hh, Cos2 mediates the interaction between the Cos2/Fu complex and Smo by binding to the region of Smo from amino acids 818 to 974; in the absence of Hh, the C-terminal part of Smo (aa 974-1 035) interacts with Fu.
Recent studies have suggested that Fu kinase activity also plays a very important role in germline stem cell fate control by generating a gradient BMP response 42. Given that previous studies demonstrated that the Hh pathway is not required in germline cells, and the loss-of-function of Fu leads to early germ cell proliferation 43, the mechanism of Fu activation in the germline might be different and Hh independent. Our data indicate that Fu forms a dimer in response to Hh through Cos2/Smo interactions; the formation of a Fu dimer is Smo- and Cos2-dependent and is regulated by both graded Hh and the graded phosphorylation of the Smo C-tail. In a previous study, it was suggested that membrane tethering can activate Fu 44. We also tested whether the membrane-tethered form Fu forms a dimer by FRET and whether our Div-Fu associates with the plasma membrane. The results suggested that the membrane-tethered form Fu used by Claret et al. might form a dimer and our Div-Fu did not associate with the plasma membrane in the absence of Hh (data not shown). This implies that in the presence of Hh, the Fu membrane association induced by Smo and Cos2 might increase the possibility of dimerization. As a Hh pathway specific Ser/Thr kinase, how does Fu act in the Hh pathway? We found that dimerized Fu is auto-phosphorylated and can phosphorylate both Cos2 and Su(fu). With changes in the Hh gradient, Cos2 is phosphorylated under low Hh concentrations, however, only upon a high Hh concentration could Su(fu) be phosphorylated. FKBP- or Div-based Fu dimers could induce the phosphorylation of Cos2 and Su(fu). This suggests that Cos2 and Su(fu) are direct phosphorylation substrates of activated Fu. We observed that AP20187, a chemical dimer inducer, induced a graded Fu dimerization and pathway activation, which suggests that the Hh gradient may be conveyed by graded Fu dimerization. In agreement with this conclusion, we found that the dimerized Fu kinase domain could activate the Hh pathway, partially bypassing Smo and Cos2, while the dimerized FuFL or Fu kinase domain with a Su(fu) binding site (FuK*), could completely bypass the functional requirement of Cos2 and Smo to activate downstream gene expression. The different potency in pathway activation by FuK and FuK* is likely due to the Su(fu) binding region carried by FuK*, which can mediate the phosphorylation of Su(fu), leading to Ci155 release and its nuclear translocation. Our in vivo data indicate that a Div-based Fu dimer could turn on downstream gene dpp and ptc expression in the A-compartment. In the presence of low levels of Hh, which are mimicked by SmoSD1 and SmoSD12, the phosphorylation of Cos2 by dimerized Fu contributes to the dissociation of Ci155 from Cos2/Fu complex. As Su(fu) is not phosphorylated by dimerized Fu in the presence of a low concentration of Hh, Ci155 dissociates from Cos2 but not from Su(fu), leading to dpp expression. Under this condition, Fu forms a weak dimer compared with those at high Hh concentration. In the presence of a high Hh concentration, which is mimicked by SmoSD, Fu forms a strong dimer and phosphorylates Su(fu), which terminates the inhibition of Su(fu) on Ci155 and causes the dissociation of Ci155 from Cos2 and Su(fu). Released Ci155 translocates to the nucleus and activates the expression of the ptc gene.
Taken together, our study suggests that, upon Hh binding with the Ptc receptor, the Hh gradient is translated to a graded Smo phosphorylation, which induces Fu dimerization at different levels through the interaction between Cos2/Fu complex and activated Smo. Fu dimerization conveys the Hh gradient by phosphorylating Cos2 and/or Su(fu), which results in the nuclear translocation of Ci155, leading to the differential activation of downstream gene expression (Supplementary information, Figure S11). This research suggests roles for specific phosphorylation events of Fu, Cos2 and Su(fu) in Hh signaling transduction and also sheds light on how the phosphorylation of Hh pathway components regulates their activity, the dissociation of the Cos2/Fu complex and Ci155, and the subcellular location of Ci155 to result in different downstream gene expression patterns in response to Hh signaling. Furthermore, we have just begin to uncover how a morphogen gradient controls one cell type to generate diverse cell types in metazoan development.
Materials and Methods
Constructs, cell culture, transfection, immunoprecipitation, western blot analysis and small molecule treatment
All constructs described in this study were made from Drosophila genes and generated using the pUAST vector. Amino acid substitutions were generated using PCR-based mutagenesis. S2 cells were cultured in the Schneider's Drosophila Medium (Invitrogen) with 10% fetal bovine serum, 100 U/ml penicillin and 100 μg/ml streptomycin as described. Hh conditional medium was obtained from Hh stable cell line of S2 cells after 24 h induction by 0.7 mM CuSO4, and was added to cells at 50% of the medium for 24 h stimulation before cells were harvested. Transfection was carried out using the Calcium Phosphate Transfection Kit (Specialty Media) according to the manufacturer's instructions. Immunoprecipitation and western blot analysis were performed using standard protocols as previously described 20, 30. The antibodies used were mouse anti-Myc (Abmart); mouse anti-Flag, M2 (Sigma); mouse anti-HA, F7 (Santa Cruz); and rat anti-Ci, 2A 45. To induce Fu dimerization based on FKBP, the indicated constructs were transfected into S2 cells. At 10 or 20 h for phosphorylation and luciferase assays before harvested, respectively, cells were treated with or without AP20187 (50 nM; kindly provided by ARIAD Pharmaceuticals). In gradient treatment assay, the concentration of AP20187 changed from 0 nM to 50 nM. Leptomycin B (LMB, Calbiochem) was used at 5 nM to treat transfected cells for 2 h before cells were harvested. In Ci155 dissociation assay, to inhibit Ci155 degradation, MG132 (Sigma) was applied at 50 μM to treat cells for 6 h before cells were harvested.
Immunoprecipitation based kinase assay
S2 cells were transfected with indicated Fv2- or Div-based Fu constructs, Flag-tagged Cos2 or Su(fu), respectively. Cells were then harvested and treated with standard immunoprecipitation worksheet, respectively. The beads were washed with 1 ml kinase assay buffer for three times at 4 ºC, 10 min for each time. The immunoprecipitated products for FKBP-Fu and Cos2 or Su(fu) were mixed with 50 μM ATP and 100 nM AP20187 in kinase buffer, and then incubated at 30 ºC for 60 min. For Div-Fu phosphorylation assay, immunoprecipitated products for Div-Fu and Cos2 or Su(fu) were mixed with 50 μM ATP in kinase buffer, and then incubated at 30 ºC for 60 min. After the reaction, the samples would be treated for western blot following the standard protocol. The kinase assay buffer consists of 20 mM Tris-HCl, pH 7.5; 20 mM β-glycerophosphate; 10 mM MgCl2; 50 mM NaCl; 100 μM orthovanadate; 0.5 mM NaF; 0.1 mM Na3VO4 and proteinase inhibitor (Sigma Cop.). For conforming that the mobility shift was caused by the protein phosphorylation, the phosphatase (Fermentas Cop.) was used in the kinase assay.
RNAi, immunostaining, luciferase reporter assay and FRET analysis
The double-strand RNA was synthesized using the in vitro Transcription T7 Kit from TakaRa. After cells were transfected for 24 h, the culture medium was changed to Serum Free Medium with 20-50 μg dsRNA/106 cells for 1-6 h starvation. Then fresh medium with serum was added and cells were cultured for 24-36 h. The targeting amino acids and primer sequences are as follows: Fu, targeting amino acids 302-525, Fu-RNAi-F, 5′-TAATACGACTCACTATAGGGAGATTTGGCGAGTCGCGACAGG-3′, Fu-RNAi-R, 5′-TAATACGACTCACTATAGGGAGATGTTCAAGAAGGCTAGCCACTC-3′; Cos2, targeting amino acids 621-841, Cos-RNAi-F, 5′-TAATACGACTCACTATAGGGAGACAGCCGCCTCCGCCCATCGAT-3′, Cos-RNAi-R, 5′-TAATACGACTCACTATAGGGAGACGTAATCCTAGCCTGTACTGC-3′; GFP, targeting amino acids 2-201, GFP-RNAi-F, 5′-TAATACGACTCACTATAGGGAGAGTGAGCAAGGGCGAGGAGC-3′, GFP-RNAi-R, 5′-TAATACGACTCACTATAGGGAGAGGTAGTGGTTGTCGGGCAGC-3′; Renilla-RNAi-F, 5′-GATCACTAATACGACTCACTATAGGGATGACTTCGAAAGTTTATGATCCAG-3′, Renilla-RNAi-R, 5′-GATCACTAATACGACTCACTATAGGGTTATCTTGATGCTCATAGCTATAATG-3′; Smo, targeting 201-862bp of Smo cDNA, Smo-RNAi-F, 5′-TTAATACGACTCACTATAGGGAGACAGGGACGACAAACCCTGGTTTG-3′, Smo-RNAi-R, 5′-TTAATACGACTCACTATAGGGAGAGATACTTGTTGGCATTCTTCC-3′ 13. Cell staining was carried out as described 9. Imaginal dick immunostaining was performed as described 12. Primary antibodies used in this study were mouse anti-Myc (Abmart); mouse anti-Flag, M2 (Sigma); mouse anti-GFP 1:100 (Abmart); rat anti-Ci, A245; rabbit anti-βGal (Cappel). The Ci-responsive ptc-Luc reporter was described earlier 17. In all luciferase assays, 1 × 106 S2 cells were transfected with 300 ng ptc-luc reporter, 6 ng Renilla, 300 ng Gal4, 100 ng Ci155, 100 ng Su(fu) and 300 ng indicated constructs in each well of a 24-well plate. GFP RNAi was used as control in the indicated luciferase assay. The luciferase activity was tested after 48 h using the dual-reporter luciferase system on a 96 luminometer (Promega). The normalized data are expressed as mean of at least five replicates obtained from at least three independent experiments. For FRET analysis of cultured cells, CFP- and YFP-tagged constructs were transfected into S2 cells together with an ub-Gal4 expression vector. Transfected cells were treated with or without Hh conditional medium. For maximal Hh signal strength, a UAS-Hh expression construct was also included in the transfection. Cells were washed with PBS, fixed with 4% formaldehyde for 20 min, and mounted on slides in 80% glycerol. Each data set was based on 15-20 individual cells. In each cell, three to five regions of interest in photobleached area were selected for analysis. In the absence of Hh, only limited Smo colocalizes with Cos2/Fu complex. Under this condition, only areas, in which Smo co-localized with Cos2/Fu complex, were selected for collecting FRET signal. For FRET assay of wing discs, transgenes were expressed with MS1096 for analysis of A or P compartment and A/P boundary cells. Late third instar larvae wing discs were dissected in PBS, fixed in 4% formaldehyde and mounted on slides in 80% glycerol. Each data set was based on 5-10 individual discs. For collecting FRET change information, Leica LAS SP5 confocal microscope was employed. CFP was excited at 458 nm wavelength and the emission was collected through a BP 480-520 filter. YFP was excited at 514 nm wavelength and the emission was collected through a BP 535-590 nm filter. CFP signal was obtained once before (BP) and after (AP) photobleaching YFP using the full power of 514 nm laser line for 1-2 min at the top half of each cell or selected dick area, leaving the bottom half as the internal control. The intensity change of CFP was analyzed using the ImageJ software (NIH). The efficiency of FRET was calculated using the formula: FRET% = [(CFPAP−CFPBP)/CFPAP] × 100.
Real-time PCR assay
We used real-time PCR to identify the RNAi efficiency for Cos2, Fu and Smo because of the absence of antibodies for endogenous protein. In this assay, S2 cells treated by the indicated Cos2, Fu or Smo dsRNA or control Renilla dsRNA were harvested and used to prepare the total RNA using standard RNA isolation protocol. Then the RNA products were used to synthesize the first strand cDNA using the Kit (#K1622) from Fermentas Company. Finally, the real-time PCR was done using the SYBR green master mix (QPK-201) from TOYOBO on the instrument of ABI-7500 (Applied Biosystem Cop.). The target regions of genes in the real-time PCR were different from the RNAi target regions. The primers used in this assay were tested to have good specificity and proliferation rate. The 2-ΔΔCt method was used for the relative quantification 46. Rp49 was used as the internal control for the mRNA level normalization 47.
Transgenes
Transgenic lines were generated using pUAST vector by standard P-element mediated transformation and were as follows: UAS-Fu-CFPC, UAS-Fu-YFPC, UAS-Smo-CFPC 17, UAS-Cos2-CFPC, UAS-Cos2-YFPC, UAS-Div, UAS-Div-Fu, UAS-Div-FuK, UAS-Div-FuK*, UAS-Div-FuG13V, UAS-Div-FuKG13V and UAS-Div-FuK*G13V. Fu-G13V corresponds to a substitution of Gly 13 by Val within Fu catalytic domain. Multiple independent transgenic lines were tested for activity. MS1096, dpp-lacZ, Ptc-Gal4 and AG-Gal4, dpp-lacZ have been described 9.
References
Ingham PW, McMahon AP . Hedgehog signaling in animal development: paradigms and principles. Genes Dev 2001; 15:3059–3087.
Taipale J, Beachy PA . The Hedgehog and Wnt signalling pathways in cancer. Nature 2001; 411:349–354.
Jiang J, Hui CC . Hedgehog signaling in development and cancer. Dev Cell 2008; 15:801–812.
Pospisilik JA, Schramek D, Schnidar H, et al. Drosophila genome-wide obesity screen reveals hedgehog as a determinant of brown versus white adipose cell fate. Cell 2010; 140:148–160.
Varjosalo M, Taipale J . Hedgehog: functions and mechanisms. Genes Dev 2008; 22:2454–2472.
Alcedo J, Ayzenzon M, Von Ohlen T, Noll M, Hooper JE . The Drosophila smoothened gene encodes a seven-pass membrane protein, a putative receptor for the hedgehog signal. Cell 1996; 86:221–232.
Aikin RA, Ayers KL, Therond PP . The role of kinases in the Hedgehog signalling pathway. EMBO Rep 2008; 9:330–336.
Zhang C, Williams EH, Guo Y, Lum L, Beachy PA . Extensive phosphorylation of Smoothened in Hedgehog pathway activation. Proc Natl Acad Sci USA 2004; 101:17900–17907.
Jia J, Tong C, Wang B, Luo L, Jiang J . Hedgehog signalling activity of Smoothened requires phosphorylation by protein kinase A and casein kinase I. Nature 2004; 432:1045–1050.
Apionishev S, Katanayeva NM, Marks SA, Kalderon D, Tomlinson A . Drosophila Smoothened phosphorylation sites essential for Hedgehog signal transduction. Nat Cell Biol 2005; 7:86–92.
Jia J, Tong C, Jiang J . Smoothened transduces Hedgehog signal by physically interacting with Costal2/Fused complex through its C-terminal tail. Genes Dev 2003; 17:2709–2720.
Ruel L, Rodriguez R, Gallet A, Lavenant-Staccini L, Therond PP . Stability and association of Smoothened, Costal2 and Fused with Cubitus interruptus are regulated by Hedgehog. Nat Cell Biol 2003; 5:907–913.
Lum L, Zhang C, Oh S, et al. Hedgehog signal transduction via Smoothened association with a cytoplasmic complex scaffolded by the atypical kinesin, Costal-2. Mol Cell 2003; 12:1261–1274.
Ogden SK, Ascano M Jr, Stegman MA, et al. Identification of a functional interaction between the transmembrane protein Smoothened and the kinesin-related protein Costal2. Curr Biol 2003; 13:1998–2003.
Liu Y, Cao X, Jiang J, Jia J . Fused-Costal2 protein complex regulates Hedgehog-induced Smo phosphorylation and cell-surface accumulation. Genes Dev 2007; 21:1949–1963.
Ascano M Jr, Robbins DJ . An intramolecular association between two domains of the protein kinase Fused is necessary for Hedgehog signaling. Mol Cell Biol 2004; 24:10397–10405.
Zhao Y, Tong C, Jiang J . Hedgehog regulates smoothened activity by inducing a conformational switch. Nature 2007; 450:252–258.
Piston DW, Kremers GJ . Fluorescent protein FRET: the good, the bad and the ugly. Trends Biochem Sci 2007; 32:407–414.
Malpel S, Claret S, Sanial M, et al. The last 59 amino acids of Smoothened cytoplasmic tail directly bind the protein kinase Fused and negatively regulate the Hedgehog pathway. Dev Biol 2007; 303:121–133.
Robbins DJ, Nybakken KE, Kobayashi R, et al. Hedgehog elicits signal transduction by means of a large complex containing the kinesin-related protein costal2. Cell 1997; 90:225–234.
Sisson JC, Ho KS, Suyama K, Scott MP . Costal2, a novel kinesin-related protein in the Hedgehog signaling pathway. Cell 1997; 90:235–245.
Zhang W, Zhao Y, Tong C, et al. Hedgehog-regulated Costal2-kinase complexes control phosphorylation and proteolytic processing of Cubitus interruptus. Dev Cell 2005; 8:267–278.
Farzan SF, Ascano M Jr, Ogden SK, et al. Costal2 functions as a kinesin-like protein in the hedgehog signal transduction pathway. Curr Biol 2008; 18:1215–1220.
Gazdoiu S, Yamoah K, Wu K, et al. Proximity-induced activation of human Cdc34 through heterologous dimerization. Proc Natl Acad Sci USA 2005; 102:15053–15058.
Clackson T, Yang W, Rozamus LW, et al. Redesigning an FKBP-ligand interface to generate chemical dimerizers with novel specificity. Pro Natl Acad Sci USA 1998; 95:10437–10442.
Rigden MD, Baier C, Ramirez-Arcos S, et al. Identification of the coiled-coil domains of Enterococcus faecalis DivIVA that mediate oligomerization and their importance for biological function. J Biochem 2008; 144:63–76.
Raisin S, Ruel L, Ranieri N, Staccini-Lavenant L, Therond PP . Dynamic phosphorylation of the kinesin Costal-2 in vivo reveals requirement of fused kinase activity for all levels of hedgehog signalling. Dev Biol 2010; 344:119–128.
Nybakken KE, Turck CW, Robbins DJ, Bishop JM . Hedgehog-stimulated phosphorylation of the kinesin-related protein Costal2 is mediated by the serine/threonine kinase fused. J Biol Chem 2002; 277:24638–24647.
Schlessinger J . Cell signaling by receptor tyrosine kinases. Cell 2000; 103:211–225.
Chen CH, von Kessler DP, Park W, et al. Nuclear trafficking of Cubitus interruptus in the transcriptional regulation of Hedgehog target gene expression. Cell 1999; 98:305–316.
Humke EW, Dorn KV, Milenkovic L, Scott MP, Rohatgi R . The output of Hedgehog signaling is controlled by the dynamic association between Suppressor of Fused and the Gli proteins. Genes Dev 2010; 24:670–682.
Wang QT, Holmgren RA . Nuclear import of cubitus interruptus is regulated by hedgehog via a mechanism distinct from Ci stabilization and Ci activation. Development 2000; 127:3131–3139.
Wang QT, Holmgren RA . The subcellular localization and activity of Drosophila cubitus interruptus are regulated at multiple levels. Development 1999; 126:5097–5106.
Wang G, Amanai K, Wang B, Jiang J . Interactions with Costal2 and suppressor of fused regulate nuclear translocation and activity of cubitus interruptus. Genes Dev 2000; 14:2893–2905.
Seong KH, Akimaru H, Dai P, et al. Inhibition of the nuclear import of cubitus interruptus by roadkill in the presence of strong hedgehog signal. PloS One 2010; 5:e15365.
Briscoe J, Sussel L, Serup P, et al. Homeobox gene Nkx2.2 and specification of neuronal identity by graded Sonic hedgehog signalling. Nature 1999; 398:622–627.
Hooper JE, Scott MP . Communicating with Hedgehogs. Nat Rev 2005; 6:306–317.
Lecuit T, Brook WJ, Ng M, et al. Two distinct mechanisms for long-range patterning by Decapentaplegic in the Drosophila wing. Nature 1996; 381:387–393.
Nellen D, Burke R, Struhl G, Basler K . Direct and long-range action of a DPP morphogen gradient. Cell 1996; 85:357–368.
Strigini M, Cohen SM . Formation of morphogen gradients in the Drosophila wing. Sem Cell Dev Biol 1999; 10:335–344.
Zhao Y, Tong C, Jiang J . Transducing the Hedgehog signal across the plasma membrane. Fly 2007; 1:333–336.
Xia L, Jia S, Huang S, et al. The Fused/Smurf complex controls the fate of Drosophila germline stem cells by generating a gradient BMP response. Cell 2010; 143:978–990.
Narbonne-Reveau K, Besse F, Lamour-Isnard C, Busson D, Pret AM . fused regulates germline cyst mitosis and differentiation during Drosophila oogenesis. Mech Dev 2006; 123:197–209.
Claret S, Sanial M, Plessis A . Evidence for a novel feedback loop in the Hedgehog pathway involving Smoothened and Fused. Curr Biol 2007; 17:1326–1333.
Motzny CK, Holmgren R . The Drosophila cubitus interruptus protein and its role in the wingless and hedgehog signal transduction pathways. Mech Dev 1995; 52:137–150.
Livak KJ, Schmittgen TD . Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 2001; 25:402–408.
Gesellchen V, Kuttenkeuler D, Steckel M, Pelte N, Boutros M . An RNA interference screen identifies Inhibitor of Apoptosis Protein 2 as a regulator of innate immune signalling in Drosophila. EMBO Rep 2005; 6:979–984.
Acknowledgements
We thank Dr Jin Jiang (University of Texas Southwestern Medical Center at Dallas, USA), Dr Robert Holmgren (Northwestern University, USA), the ARIAD Pharmaceuticals and Developmental Studies Hybridoma Bank at the university of Iowa for reagents. We thank Dr Xiaochen Wang (National Institute of Biological Sciences, China), Dr Min Wu (Wuhan University, China) and Dr Zhaocai Zhou (Institute of Biochemistry and Cell Biology, CAS, China) for critical comments on the manuscript. This work was supported by grants from the National Natural Science Foundation of China (30623003, 30971647 and 30971646), the National Key Scientific Program of China (2011CB943902 and 2010CB912100), Shanghai Pujiang Program (09PJ1410900) and also from the “Strategic Priority Research Program” of the Chinese Academy of Sciences (XDA01010405 and XDA01010406). Y Z and L Z are the scholars of the Hundred Talents Program of the Chinese Academy of Sciences.
Author information
Authors and Affiliations
Corresponding authors
Additional information
( Supplementary information is linked to the online version of the paper on the Cell Research website.)
Supplementary information
Supplementary information, Figure S1
FRET assay of wing discs expressing Smo and Cos2 transgenes. (PDF 473 kb)
Supplementary information, Figure S2
FRET assay of wing discs expressing Cos2 transgenes. (PDF 457 kb)
Supplementary information, Figure S3
RNAi efficiency for Fu and Cos2. (PDF 50 kb)
Supplementary information, Figure S4
Localization of Cos2 or Fu with indicated Smo mutations. (PDF 311 kb)
Supplementary information, Figure S5
FRET assay of wing discs expressing Fu transgenes. (PDF 633 kb)
Supplementary information, Figure S6
FRET assay of S2 cells expressing the indicated Fu constructs. (PDF 154 kb)
Supplementary information, Figure S7
RNAi efficiency for Cos2, Smo and Fu. (PDF 71 kb)
Supplementary information, Figure S8
Dimerization of Fu promotes Fu auto-phosphorylation, Cos2 and Su(fu) phosphorylation. (PDF 213 kb)
Supplementary information, Figure S9
Smo phosphor-mimetic mutation and dimerized Fu promotes Ci155 nucleus translocation. (PDF 581 kb)
Supplementary information, Figure S10
Wild type Fu and dimerized Fu mutations failed to turn on Hh pathway. (PDF 465 kb)
Supplementary information, Figure S11
Working model for Fu transducing Hh morphogen gradient. (PDF 45 kb)
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0
About this article
Cite this article
Zhang, Y., Mao, F., Lu, Y. et al. Transduction of the Hedgehog signal through the dimerization of Fused and the nuclear translocation of Cubitus interruptus. Cell Res 21, 1436–1451 (2011). https://doi.org/10.1038/cr.2011.136
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2011.136