Abstract
MicroRNAs (miRNAs), which are small noncoding RNA molecules, play important roles in the post-transcriptional regulation process. The microRNA-21 gene (miR-21) has been reported to be highly expressed in various solid tumors, including breast cancer. Bone morphogenetic protein-6 (BMP-6) has been identified as an inhibitor of breast cancer epithelial-mesenchymal transition (EMT) through rescuing E-cadherin expression. We initiated experiments to identify the relationships between miR-21 and BMP-6 in breast cancer progression. Real-time PCR analysis showed that miR-21 expression was very high in MDA-MB-231 cells that expressed little BMP-6. A reverse correlation between BMP-6 and miR-21 was also determined in breast cancer tissue samples. Moreover, BMP-6 inhibited miR-21 transcription in MDA-MB-231 cells. In order to investigate how BMP-6 inhibited the miR-21 promoter (miPPR-21), we constructed a series of miPPR-21 reporters. Luciferase assay results indicated that BMP-6 inhibited miPPR-21 activity through the E2-box and AP-1-binding sites. We also demonstrated that both δEF1 and TPA induced miR-21 expression. Using site-directed mutation and CHIP assay, we found that δEF1 induced miPPR-21 activity by binding to the E2-box on miPPR-21. Moreover, TPA triggered miPPR-21 activity through the AP-1 binding sites. BMP-6 treatment significantly reduced the binding of these factors to miPPR-21 by decreasing the expression of δEF1 and c-Fos/c-Jun. We also demonstrated that BMP-6-induced downregulation of miR-21 modified the activity of PDCD4 3′UTR and inhibited MDA-MB-231 cell invasion. δEF1 overexpression and TPA induction blocked this inhibitory effect of BMP-6. In conclusion, BMP-6-induced inhibition of miR-21 suggests that BMP-6 may function as an anti-metastasis factor by a mechanism involving transcriptional repression of miR-21 in breast cancer.
Similar content being viewed by others
Introduction
Post-transcriptional regulation has been realized to be an important mechanism in regulation of gene expression. Several hundred genes in our genome encode small functional RNA molecules, collectively called microRNAs (miRNAs) 1. More than 700 human miRNAs have been experimentally identified. Computational predictions of miRNA targets suggest that up to 30% of human protein-coding genes may be regulated by miRNAs 2. This makes miRNAs one of the most abundant classes of regulatory genes in humans. miRNAs are now predicted to be a key layer of post-transcriptional control within the networks of gene expression regulation.
Among human diseases, it has been shown that miRNAs are aberrantly expressed or mutated in cancer, suggesting that they may function as a novel class of oncogenes or tumor suppressor genes 3. Although several reports have shown examples of mammalian miRNA genes regulated by known transcription factors, it still remains largely unknown how human miRNA expression is regulated at the transcriptional level 4. Recent reports have identified that microRNA-21 gene (miR-21) is a unique miRNA overexpressed in nine types of solid tumors, including lung, breast, stomach, prostate, colon, brain, head and neck, esophagus, and pancreas carcinomas 5. Of these carcinomas, breast cancer is a leading cause of cancer death in women worldwide. More than one million new cases of breast cancer are diagnosed annually 6. Moreover, distant metastases remain the most common type of breast cancer recurrence and are often the cause of death in breast cancer patients. For example, metastases result in more than 40 000 deaths per year in the US alone 7. Recently, studies suggested that miR-21 could play important roles in invasion and metastasis in different types of cancers by inhibiting its target genes (e.g. tropomyosin 1 (TPM1) and programmed cell death 4 (PDCD4)) 5, 8, 9. Elucidating the regulatory mechanisms of miR-21 expression in breast cancer would provide novel clues for therapeutic strategies.
Bone morphogenetic protein-6 (BMP-6) belongs to the TGF-β superfamily. The traditional BMP-6 signaling pathway is Smad-dependent 10. In addition to the Smad pathway, BMP-6 is also known to activate and cross-talk with other pathways, such as the MAPK pathway 11. In addition to its effect on inducing new bone formation, BMP-6 has been shown to be involved in numerous biological processes. In our previous studies, we found that BMP-6 was abnormally expressed and regulated by estrogen receptor alpha in breast cancer cells 12. Moreover, we confirmed that BMP-6 induced E-cadherin expression by repressing δEF1 in MDA-MB-231 cells 13. E-cadherin has been well documented as a key player in breast cancer EMT. Loss of E-cadherin expression correlates with tumor invasiveness and metastasis 14, whereas restoration of E-cadherin expression reverses EMT progression. Taken together, these results suggest that BMP-6 might play an important role in breast cancer invasion and metastasis.
Here, we discovered a negative correlation between the expressions of BMP-6 and miR-21 in breast cancer tissue samples. The expression of miR-21 was high in MDA-MB-231 cells that expressed a very low level of BMP-6 transcripts. Moreover, BMP-6 inhibited miR-21 expression in MDA-MB-231 cells at the transcriptional level, an effect that was mediated via reduction of the expression of δEF1 and c-Fos/c-Jun. BMP-6 thus inhibited δEF1 and c-Fos/c-Jun binding to the miPPR-21. BMP-6 could also modify PDCD4 3′UTR activity and block MDA-MB-231 cell invasion by inhibiting miR-21.
Results
BMP-6 inhibits miR-21 expression at the transcriptional level
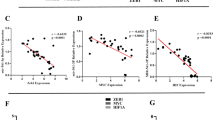
Recent reports indicated that MDA-MB-231 breast cancer cells, which have low endogenous BMP-6 expression, expressed a high level of mature miR-21 8. Consistent with these observations, we used real-time PCR in this study to show that miR-21 exhibited high level of expression in the metastatic MDA-MB-231 cells (Figure 1A). It is well known that bone is the most frequently targeted organ of breast cancer metastasis 7. Growth factors that are involved in bone development, such as TGF-β and BMPs, also play important roles in the breast cancer metastatic process 13, 15. Here, we detected miR-21 and BMP-6 expression levels in matched normal and breast tumor tissues by real-time PCR. We collected 12 pairs of matched advanced breast tumor specimens with lymph node metastasis, and found that miR-21 expression was much higher in tumor tissue than in the corresponding normal tissues, whereas BMP-6 expression level was much higher in normal tissues (Figure 1B). After normalizing the data, there was an average of nearly a five-fold increase in the level of miR-21 in tumor tissues compared with the matched normal tissues. These results from clinical samples suggest that miR-21 may play a significant role in breast cancer metastasis, a finding that is consistent with previous reports 8.
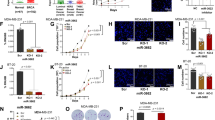
BMP-6 inhibited miR-21 expression at the transcriptional level. (A) Different breast cancer cell lines were cultured and total RNA was extracted using TRIZOL (Invitrogen). Real-time PCR indicated that the miR-21 expression level was much higher in the metastatic breast cancer cell line MDA-MB-231 than in other cell lines. (B) Fresh-frozen human breast cancer samples were obtained and total RNA was extracted using TRIZOL (Invitrogen). Real-time PCR results showed a much higher miR-21 expression level in the tumor tissues than in the corresponding normal tissues, whereas the BMP-6 expression level was higher in normal tissues. The expression level of miR-21 and BMP-6 in normal breast tissue was normalized to 1. (C) MDA-MB-231 cells were plated into a 24-well plate and treated with different concentrations of BMP-6. Total RNA was extracted 24 h after treatment. Real-time PCR results indicated that 200 ng/ml BMP-6 obviously inhibited miR-21 expression. (D) BMP-6 inhibited miR-21 expression in a time-dependent manner. MDA-MB-231 cells were plated into a 24-well plate for 24 h, and then BMP-6 (200 ng/ml) was added. Total mRNA was extracted at different time points after treatment, and miR-21 expression was examined by real-time PCR. (E) 1.4-kb miPPR-21-containing plasmids were transfected into MDA-MB-231 cells, and BMP-6 (200 ng/ml) was added 5 h after transfection. After another 24 h, cells were harvested, and luciferase assay results confirmed that BMP-6 treatment obviously inhibited miPPR-21 activity.
We further investigated whether there is an association between the expression of BMP-6 and miR-21 in breast cancer. We treated MDA-MB-231 cells with BMP-6. Interestingly, we found that BMP-6 inhibited miR-21 expression in a dose-dependent manner, with a BMP-6 dose of 200 ng/ml obviously inhibiting miR-21 expression (Figure 1C). The inhibitory effect of BMP-6 was also time-dependent, exhibiting maximum repression at 24 h (Figure 1D). Since the transcriptional start site of miR-21 has been established 4, we constructed a 1.4-kb promoter of miR-21 to investigate whether BMP-6 inhibited miPPR-21 activity. Results from luciferase assays indicated that BMP-6 markedly inhibited miPPR-21 activity (Figure 1E). These results indicate that BMP-6 may transcriptionally inhibit miR-21 expression.
BMP-6 inhibits miPPR-21 activity through repression of δEF1 expression
In order to clarify the regulatory mechanism by which BMP-6 affects miR-21 expression, we analyzed miPPR-21 using the online tools TRANSFAC (http://www.gene-regulation.com) and TESS (http://www.cbil.upenn.edu/cgi-bin/tess/tess). Since we did not find a typical BMP response element (BRE) or a Smad binding element (SBE) on this promoter, we proposed that BMP-6 might influence miR-21 expression by affecting other transcription factors. Based on our online analysis, we found that there was an E2-box, whose sequence is CACCTG, located in the promoter region (Figure 2A). Our previous results indicated that BMP-6 inhibited the expression of δEF1, which could bind to the E2-box directly 13. To investigate whether δEF1 regulated miR-21 expression, we overexpressed δEF1 in MDA-MB-231 cells. Real-time PCR results demonstrated that δEF1 significantly increased miR-21 expression. However, upon transfection of a siδEF1 expression plasmid, the δEF1-induced activation of miR-21 expression was totally abolished (Figure 3A). These data suggest that δEF1 is a transcriptional activator of miR-21. We found similar results in the luciferase experiments. δEF1 overexpression induced miPPR-21 activity in a dose-dependent manner. In contrast, siδEF1 decreased the promoter activity (Figure 3B and 3C). We further examined whether BMP-6 affected this δEF1-induced promoter activity. As shown in Figure 3B and 3C, δEF1-induced miPPR-21 activity was evidently weakened in the presence of BMP-6. With increasing of BMP-6 dosage, the weakening effect was more obvious. Further, δEF1 overexpression in MDA-MB-231 cells in the presence of BMP-6 rescued the miR-21 expression (Figure 3D). Collectively, the above results confirmed our hypothesis that BMP-6 inhibited miR-21 expression, an effect that could be mediated through repression of δEF1.
miPPR-21 was analyzed using the online tools TRANSFAC (http://www.gene-regulation.com) and TESS (http://www.cbil.upenn.edu/cgi-bin/tess/tess). (A) An E2-box and three AP-1-binding sites were found on the promoter. No typical BRE elements were located on this promoter. In order to study the effects of the E2-box and AP-1 elements in BMP-6-induced miR-21 repression, a series of deleted miPPR-21s were constructed and named F1, F2, F3, F4, and F5. (B) A series of mutated miPPR-21s were constructed using the QuikChange Lightning Site-Directed Mutagenesis Kit (Stratagene) following the instruction manual. These plasmids were named F1m, F2m, F2m1, F2m2, and F2m3, as described in the figure. Primers used for mutation were designed using the QuikChange Primer Design Program (http://www.stratagene.com/sdmdesigner/default.aspx). The primers used in construction and mutation are described in Supplementary information, Table S1.
BMP-6 inhibited miPPR-21 activity by repressing δEF1. (A) δEF1 and siδEF1 expression plasmids were transfected into MDA-MB-231 cells. Total RNA was extracted at different time points. Real-time PCR results indicated that δEF1 induced miR-21 expression, whereas additional siδEF1 weakened δEF1-induced miR-21 upregulation. (B, C) miPPR-21 F1, δEF1, and siδEF1 expression plasmids were transfected into MDA-MB-231 cells following the experimental design. After 24 h, cells were harvested. Luciferase assay results indicated that δEF1 increased miPPR-21 F1 activity, whereas siδEF1 exhibited the negative effect. BMP-6 treatment had a similar effect with siδEF1. The inhibitory effect of BMP-6 was dose-dependent. (D) BMP-6 expression plasmids were transfected into MDA-MB-231 cells. At 24 h after the first transfection, different amounts of δEF1 expression plasmids were transfected again, and total RNA was extracted 24 h later. Real-time PCR results indicated that δEF1 overexpression rescued the miR-21 expression, which was inhibited by BMP-6 treatment. (E) miPPR-21s F1 and F1m, in which the E2-box was mutated, together with the δEF1 expression plasmids were transfected into MDA-MB-231 cells. BMP-6 (200 ng/ml) was added 5 h after transfection. Cells were harvested 24 h later. Luciferase assay results indicated that BMP-6-mediated inhibition of miPPR-21 was partially abolished when the E2-box was mutated.
How does BMP-6 affect δEF1-induced miPPR-21 activity? We further investigated whether BMP-6 affects δEF1 binding to the E2-box on the miPPR-21. We mutated the E2-box sequence from CACCTG to CGAATG so that δEF1 could not bind to the mutated site. Luciferase results indicated that δEF1-induced miPPR-21 activity was completely abolished. This result confirmed that δEF1 induces miR-21 expression and its promoter activity by binding to the E2-box directly (Figure 3E). Moreover, with E2-box mutation, the inhibitory effect of BMP-6 on miPPR-21 was also weakened (Figure 3E). Using a Q-CHIP assay, we sought to determine whether BMP-6 decreases δEF1 binding to the miPPR-21. The results showed that the binding of δEF1 to miPPR-21 was decreased by BMP-6 induction (Figure 4E and 4F).
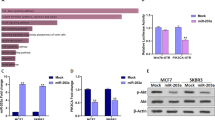
BMP-6 inhibited miPPR-21 activity by reducing c-Fos and c-Jun expression. (A) The miPPR-21 F2 was transfected into MDA-MB-231 cells. TPA (100 ng/ml) and BMP-6 (200 ng/ml) were added 5 h after transfection. Cells were harvested 24 h later; luciferase assay results indicated that BMP-6 inhibited the truncated miPPR-21 F2 activity in the absence or presence of TPA treatment. (B) A series of truncated miPPR-21s (F2, F3, F4, and F5) were transfected into MDA-MB-231 cells. Cells were treated with TPA (100 ng/ml), BMP-6 (200 ng/ml), or both, after 5 h. Cells were harvested 24 h later; luciferase assay results indicated that the inhibitory effect of BMP-6 depended on the AP-1 elements. (C) miPPR-21 F2 and another four promoters (F2m1, F2m2, F2m3, and F2m) that had different amounts of mutated-AP-1 sites were transfected into MDA-MB-231 cells. After 5 h, cells were treated with TPA (100 ng/ml), BMP-6 (200 ng/ml), or both. Cells were harvested 24 h later; luciferase assay results indicated that the inhibitory effect of BMP-6 was abolished when all AP-1 elements were mutated. (D) MDA-MB-231 cells were plated into six-well plates and treated with BMP-6 (200 ng/ml). Cells were harvested 24 h later to extract protein. Western blot results indicated that BMP-6 obviously inhibited c-Fos and c-Jun expression. (E, F) MDA-MB-231 cells were plated into 10-cm dishes and BMP-6 (200 ng/ml) was added 24 h later. Cells were harvested and CHIP assays were processed using the Chromatin Immunoprecipitation (CHIP) Assay Kit (Upstate). CHIP assay results indicated that BMP-6 obviously decreased binding of δEF1 and c-Fos/c-Jun to the endogenous miPPR-21. The binding of the control without BMP-6 treatment was normalized to 1.
BMP-6 inhibits miPPR-21 activity by repressing c-Fos and c-Jun
Our experiments indicate that BMP-6 decreases δEF1 binding to the E2-box, and thus inhibits miPPR-21 activity. However, as shown in Figure 3E, we found that E2-box mutation did not completely abolish the inhibitory effect of BMP-6 on miPPR-21 activity. We proposed that there might be other signaling pathways mediating BMP-6-induced transrepression of miPPR-21 activity. According to a previous report 4, there are three AP-1-binding elements located upstream of the TATA box on the miPPR-21 (Figure 2A). Moreover, the results from our previous microarray assay indicate that BMP-6 overexpression in MDA-MB-231 cells strongly inhibits c-Fos and c-Jun transcription (data not shown). It is known that the c-Fos/c-Jun complex plays a critical role among different components of AP-1. TPA, which is a specific AP-1 activator, induces miPPR-21 activity by activating AP-1-binding sites, and the c-Fos/c-Jun dimer shows the strongest induction effect on miPPR-21 activity 4. Based on these results, we propose that, along with the E2-box, BMP-6 might also inhibit miPPR-21 activity through AP-1-binding elements. In order to exclude the effect of δEF1, we deleted the E2-box from the 1.4-kb miPPR-21 and constructed a series of truncated miPPR-21s that contained different AP-1-binding elements (Figure 2A). Luciferase assay results indicated that TPA obviously induced the miPPR-21 activity, whereas BMP-6 treatment could inhibit the promoter activity in the absence or presence of TPA (Figure 4A). We sequentially deleted each of the AP-1-binding sites and repeated the luciferase assays. The results show that when all the three AP-1-binding sites were deleted, BMP-6 lost its inhibitory effect on miPPR-21; so did the induction of miPPR-21 activity by TPA (Figure 4B).
In order to confirm whether the inhibitory effect of BMP-6 is dependent on the AP-1-binding sites, we constructed another set of miPPR-21s (Figure 2B). We mutated the AP-1-binding elements, individually or in combination. The results indicated that when any single or any two AP-1-binding elements were mutated, the negative effect of BMP-6 was weakened but did not vanish. When all three AP-1 sites were mutated, however, neither BMP-6 nor TPA exhibited an effect on miPPR-21 activity (Figure 4C). Based on the previous microarray results, we hypothesized that the AP-1-dependent inhibition by BMP-6 is due to BMP-6 repressing the expression of the AP-1 complex components, as well as decreasing the binding of these components to the AP-1 sites on the miPPR-21. We investigated whether BMP-6 affects c-Fos or c-Jun expression at the protein level. Western blotting experiments demonstrated that BMP-6 treatment obviously inhibited both c-Fos and c-Jun expression (Figure 4D). In addition, we investigated whether this inhibitory effect by BMP-6 results in reduction of the binding of both c-Fos and c-Jun to the miPPR-21. Q-CHIP assay confirmed that (1) both c-Fos and c-Jun bound to the miPPR-21, and (2) BMP-6 treatment significantly reduced this binding (Figure 4E and 4F). With BMP-6 treatment, the binding rates were only 40% and 60% compared with the control group. Taking these findings together, we elementarily clarified the molecular mechanism of BMP-6-mediated inhibition of miR-21 expression in breast cancer.
BMP-6 inhibits miR-21-induced invasion in vitro
Having clarified how BMP-6 inhibits miR-21 expression, we investigated whether BMP-6 influences the biological effect of miR-21 in breast cancer. miR-21 has been reported to induce invasion in MDA-MB-231 cells by inhibiting several tumor suppressor genes 8. Our previous results indicated that BMP-6 rescues E-cadherin expression, which is a critical gene in breast cancer invasion and metastasis, through inhibition of δEF1 in MDA-MB-231 cells 16. We investigated whether BMP-6 could rescue miR-21-targeted gene expression, and thus inhibit MDA-MB-231 invasion in vitro. We constructed a miR-21 expression plasmid and validated its expression in HEK293T cells (Supplementary information, Figure S1). The luciferase results indicated that BMP-6 reversed the inhibitory effect of miR-21 on PDCD4 3′UTR activity (Figure 5A). BMP-6 also increased PDCD4 3′UTR activity in MDA-MB-231 cells, whereas miR-21 overexpression abolished this effect of BMP-6 (Figure 5B). Consistent with previously reported results 8, Boyden chamber assay showed that miR-21 induced MDA-MB-231 invasion in vitro. In contrast, BMP-6 decreased miR-21-induced invasion of MDA-MB-231 cells (Figure 5C and Supplementary information, Figure S2A). Moreover, the inhibitory effect of BMP-6 on miR-21-induced invasion was also mediated through decreasing δEF1 and c-Fos/c-Jun expression. Either δEF1 overexpression or TPA treatment could abolish this inhibitory effect of BMP-6 (Figure 5D and Supplementary information, Figure S2B).
BMP-6 reversed the miR-21 biological effect in vitro. (A) MDA-MB-231 cells were transfected with PDCD4 3′UTR plasmids or miR-21 expression plasmids or treated with BMP-6 (200 ng/ml). Cells were harvested 24 h later. Luciferase assay results indicated that miR-21 obviously inhibited PDCD4 3′UTR activity and BMP-6 rescued miR-21-inhibited PDCD4 3′UTR activity. (B) PDCD4 3′UTR plasmids were transfected into MDA-MB-231 cells. Cells were treated with BMP-6 (200 ng/ml) 5 h later. Cells were harvested 24 h after treatment. Luciferase results indicated that BMP-6 treatment increased the PDCD4 3′UTR activity. With the miR-21 expression plasmid co-transfected with the PDCD4 3′UTR plasmid, BMP-6-induced activity was inhibited. (C) MDA-MB-231 cells were transfected with miR-21 expression plasmids or the vector for 24 h and cells were digested and counted in a Boyden chamber assay. The Boyden chamber assay results indicated that BMP-6 inhibited miR-21-induced MDA-MB-231 invasion. (D) MDA-MB-231 cells treated with BMP-6 (200 ng/ml) were transfected with the δEF1 expression plasmid or treated with TPA (100 ng/ml). After 24 h, cells were digested and counted. The Boyden chamber assay results indicated that both δEF1 overexpression and TPA treatment increased MDA-MB-231 invasion.
In summary, this study demonstrated that BMP-6 could inhibit miR-21 transcription in MDA-MB-231 cells. Moreover, this effect was mediated through inhibition of δEF1 and c-Fos/c-Jun expression by BMP-6. Meanwhile, BMP-6 also decreased the binding of δEF1 and c-Fos/c-Jun to the miPPR-21. Since miR-21 has been confirmed as an oncogene that facilitates breast cancer metastasis, our research provides a novel function of BMP-6, namely, acting as a miR-21 inhibitor in breast cancer progression. This regulation mechanism by BMP-6 is likely to be clinically significant in breast cancer progression and metastasis research.
Discussion
Metastases are the major source of morbidity and mortality in breast cancer patients. Breast cancer – the second most common metastatic cancer – frequently metastasizes to the bone, lung, liver, and brain 17. Because of this clinical importance, detection and treatment of breast cancer metastases have been urgently researched. However, little is known about the mechanisms of cancer metastasis in distinct tissues. Several studies have reported that breast cancer produces a variety of growth factors that can play important roles in cancer progression and metastasis. Moreover, as an important post-transcriptional regulator, some miRNAs have been found not only to regulate diverse cellular processes, including differentiation, proliferation, and apoptosis 18, but also to cause cancer cells to invade and metastasize 19.
miR-21, the only miRNA that was overexpressed in nine types of solid tumors including breast cancer 5, was focused on recently. The genes targeted by miR-21 have been under intense study because miRNAs generally function by downregulating target gene mRNAs. The phosphatase and tensin homolog deleted on chromosome 10 (PTEN) gene was first determined to be a potential miR-21 target based on its well-characterized roles in tumor biology 16. Researchers subsequently reported that other tumor suppressor genes, such as TPM1 20 and PDCD4 9, 21, were targets of miR-21. All these genes encode tumor suppressors that are involved in breast cancer migration and invasion 22. Correspondently, miR-21 also has a role in invasion and metastasis 8.
Just like those traditional protein-encoding genes, miRNAs were reported to be transcribed by RNA polymerase II to produce a pri-miRNA. This process has also been reported to be regulated by known transcription factors. But it remains largely unknown how human miRNA expression itself is regulated at the transcriptional level.
BMP-6, an important growth factor involved in the regulation of bone development, has been found to be a multifunctional growth factor. We have reported that BMP-6 acts as an anti-apoptotic factor during H2O2-induced neuron apoptosis 23 and stress-induced breast cancer cell apoptosis 24. Moreover, we recently reported that BMP-6 upregulated E-cadherin expression by repressing δEF1 in MDA-MB-231 breast cancer cells, providing evidence that BMP-6 might restore E-cadherin-mediated cell-to-cell adhesion during breast tumorigenesis 13. In this report, we studied the interaction between BMP-6 and miR-21. Our results demonstrated the novel finding that BMP-6 treatment inhibited miR-21 expression. We thus investigated whether BMP-6 affects miPPR-21 activity directly. After analysis of the miPPR-21 using online tools, we did not find the typical BMP response element (BRE) or Smad binding element (SBE). We moved on to study whether BMP-6 inhibited the miPPR-21 activity via other signal pathways. The results of transcriptional factor searching indicated that there are three AP-1-binding sites and an E2-box located on the miPPR-21.
Our previous report indicated that BMP-6 repressed δEF1 expression and thus rescued E-cadherin expression. Traditionally, δEF1 was identified as a transcriptional inhibitor, acting through interactions with co-repressors or competition with activators for DNA binding sites. However, it has been reported that δEF1 could also be a transcriptional activator 25. Here, we reported that δEF1 elicited transcriptional activation of miPPR-21. BMP-6 inhibited both δEF1 expression and δEF1 binding to the miPPR-21, resulting in downregulation of miR-21.
Besides δEF1, we also found that BMP-6 inhibited miR-21 expression and miPPR-21 activity through the AP-1 signaling pathway. Our previous results showed that BMP-6 inhibited c-Fos and c-Jun expression. It has also been reported that during TPA-induced miR-21 expression through AP-1, c-Fos and c-Jun are the principal contributors among the AP-1 components 4. In this study, we further confirmed that BMP-6 inhibited c-Fos and c-Jun protein expression. Meanwhile, a CHIP assay demonstrated that BMP-6 decreased the binding of c-Fos and c-Jun to the miPPR-21.
Taken together, our results suggested that BMP-6 might be a potential miR-21 inhibitor in breast cancer progression. BMP-6 not only inhibited miR-21 expression but also repressed miR-21-induced breast cancer invasion in vitro. Moreover, our research provided evidence that BMP-6 rescued miR-21-inhibited PDCD4 3′UTR activity. Understanding the regulatory mechanisms of BMP-6 that function in breast cancer progression would help to further elucidate the mechanism of miR-21-induced metastasis, and will lead to a factor that may facilitate the clinical treatment and metastasis prevention of breast cancer.
Materials and Methods
Human tissue samples
Fresh-frozen human breast cancer samples were obtained from the Tissue Banking Facility, which is jointly supported by TMUCIH (Tianjin Medical University Cancer Institute and Hospital) and NFCR (National Foundation for Cancer Research). This study was approved by the institutional ethics committee.
Cell lines and culture conditions
Human breast cancer cell lines MDA-MB-231 and HEK293 were obtained from the American Type Culture Collection. All cell lines were maintained in DMEM supplemented with 10% fetal bovine serum (FBS).
Real-time PCR assay
Total RNA (2 μg) was reverse-transcribed using random hexamers. The cDNA was PCR-amplified by rTaq (TAKARA) using the primers listed in Supplementary information, Table S1. GAPDH cDNA amplification was used as an internal control. Reaction conditions were as follows: 95 °C/3 min; 40 cycles (95 °C/30 s; 60 °C/60 s).
Luciferase reporter assay
TSS of miR-21 was described previously 4. miPPR-21 fragments were PCR-amplified from human blood genomic DNA. The fragments were cloned into pGL4.10 (Promega). The mutant miPPR-21 inserts were generated by the QuikChange Lightning Site-Directed Mutagenesis Kit (Stratagene). The pri-miR-21 fragment was cloned into pSilencer4.1 (Ambion). The PDCD4 3′UTR fragment was a kind gift from Professor H Allgayer (University of Heidelberg, Germany). MDA-MB-231 cells or 293 cells (8 × 105 cells) were transfected with 400 or 200 ng of plasmid, respectively, and 100 ng of the control vector harboring Renilla luciferase using Lipofectamine 2000 (Invitrogen). TPA (100 ng/ml) was added to the culture 24 h after transfection. Firefly and Renilla activities were measured consecutively using the Dual Luciferase Assay kit (Promega) 5 h after addition of either TPA or vehicle. All primers used in the experiments are listed in Supplementary information, Table S1.
Chromatin immunoprecipitation
CHIP assays were performed using reagents commercially obtained from Upstate and were conducted essentially according to the manufacturer's instructions. The antibodies used in these experiments were anti-δEF1 (sc-81428, Santa Cruz), anti-c-Jun (sc-1694, Santa Cruz), anti-c-Fos (sc-52, Santa Cruz), and anti-mouse normal IgG (sc-2025, Santa Cruz). The immunoprecipitates were washed and eluted in a solution containing 1% SDS and 0.1 M NaHCO3 and incubated overnight at 65 °C with 20 μl of 5 M NaCl for reverse cross-linking. DNA was purified, and each fragment was PCR-amplified using the primer pairs listed in Supplementary information, Table S1.
Immunoblotting
Total protein extracts were prepared by boiling the cells in 1.5× SDS sample buffer for 10 min, separated by SDS-Tricine-PAGE, and transferred onto Hybond ECL nitrocellulose membranes (Amersham). Immunoblotting was performed either by incubating the membrane overnight at 4 °C with the antibodies anti-c-Jun (sc-1694, Santa Cruz) or anti-c-Fos (sc-52, Santa Cruz) or by incubating for 1 h at room temperature (RT) with anti-actin (A5316, Sigma). Secondary antibodies conjugated to horseradish peroxidase were incubated with the membranes for 1 h at RT after three washes with phosphate-buffered saline (PBS) containing Tween-20. Signals were detected using ECL reagent (Amersham).
Invasion assay
Transwell 24-well chambers with 8 μm pore size (Costar) were used as directed by the manufacturer. The membranes were pre-coated with collagen matrix (Sigma), which was reconstituted by adding 0.5 ml serum-free medium to the well for 2 h. To assess the invasion ability of the cells, 2.5 × 104 cells in 0.5 ml medium containing 1% FBS were placed into the upper compartment of the wells and 0.75 ml of medium containing 10% FBS was placed in the lower compartment. The transwell chambers were incubated for 16 h at 37 °C in a 5% CO2 incubator. Cell penetration through the membrane was detected by staining the cells on the porous membrane with 0.25% crystal violet. To quantify the data, we washed the chamber twice with PBS, and then used 33% acetic acid to wash off the excess crystal violet. Crystal violet remaining on the membranes was measured on a spectrophotometer at A570.
Statistical analysis
The statistical significance of the in vitro studies was analyzed using Student's t-test. Differences with P values of less than 0.01 are considered significant. All experiments were done in triplicate and repeated at least three times.
Supplementary information is linked to the online version of the paper on the Cell Research website.)
References
Sassen S, Miska EA, Caldas C . MicroRNA: implications for cancer. Virchows Arch 2008; 452:1–10.
Lewis BP, Burge CB, Bartel DP . Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 2005; 120:15–20.
Iorio MV, Ferracin M, Liu CG, et al. MicroRNA gene expression deregulation in human breast cancer. Cancer Res 2005; 65:7065–7070.
Fujita S, Ito T, Mizutani T, et al. miR-21 Gene expression triggered by AP-1 is sustained through a double-negative feedback mechanism. J Mol Biol 2008; 378:492–504.
Lu Z, Liu M, Stribinskis V, et al. MicroRNA-21 promotes cell transformation by targeting the programmed cell death 4 gene. Oncogene 2008; 27:4373–4379.
Parkin DM, Bray F, Ferlay J, Pisani P . Global cancer statistics, 2002. CA Cancer J Clin 2005; 55:74–108.
Rugo HS . The importance of distant metastases in hormone-sensitive breast cancer. Breast 2008; 17 Suppl 1:S3–S8.
Zhu S, Wu H, Wu F, et al. MicroRNA-21 targets tumor suppressor genes in invasion and metastasis. Cell Res 2008; 18:350–359.
Asangani IA, Rasheed SA, Nikolova DA, et al. MicroRNA-21 (miR-21) post-transcriptionally downregulates tumor suppressor Pdcd4 and stimulates invasion, intravasation and metastasis in colorectal cancer. Oncogene 2008; 27:2128–2136.
Tang SJ, Hoodless PA, Lu Z, et al. The Tlx-2 homeobox gene is a downstream target of BMP signalling and is required for mouse mesoderm development. Development 1998; 125:1877–1887.
Derynck R, Zhang YE . Smad-dependent and Smad-independent pathways in TGF-beta family signalling. Nature 2003; 425:577–584.
Zhang M, Wang Q, Yuan W, et al. Epigenetic regulation of bone morphogenetic protein-6 gene expression in breast cancer cells. J Steroid Biochem Mol Biol 2007; 105:91–97.
Yang S, Du J, Wang Z, et al. BMP-6 promotes E-cadherin expression through repressing δEF1 in breast cancer cells. BMC Cancer 2007; 7:211.
Birchmeier W, Behrens J . Cadherin expression in carcinomas: role in the formation of cell junctions and the prevention of invasiveness. Biochim Biophys Acta 1994; 1198:11–26.
Buijs JT, Henriquez NV, van Overveld PG, et al. TGF-beta and BMP7 interactions in tumour progression and bone metastasis. Clin Exp Metastasis 2007; 24:609–617.
Meng F, Henson R, Wehbe-Janek H, et al. MicroRNA-21 regulates expression of the PTEN tumor suppressor gene in human hepatocellular cancer. Gastroenterology 2007; 133:647–658.
Oxmann D, Held-Feindt J, Stark AM, et al. Endoglin expression in metastatic breast cancer cells enhances their invasive phenotype. Oncogene 2008; 27:3567–3575.
Croce CM, Calin GA . miRNAs, cancer, and stem cell division. Cell 2005; 122:6–7.
Ma L, Weinberg RA . MicroRNAs in malignant progression. Cell Cycle 2008; 7:570–572.
Zhu S, Si ML, Wu H, Mo YY . MicroRNA-21 targets the tumor suppressor gene tropomyosin 1 (TPM1). J Biol Chem 2007; 282:14328–14336.
Frankel LB, Christoffersen NR, Jacobsen A, et al. Programmed cell death 4 (PDCD4) is an important functional target of the microRNA miR-21 in breast cancer cells. J Biol Chem 2008; 283:1026–1033.
Varga AE, Stourman NV, Zheng Q, et al. Silencing of the tropomyosin-1 gene by DNA methylation alters tumor suppressor function of TGF-beta. Oncogene 2005; 24:5043–5052.
Du J, Zhu Y, Chen X, et al. Protective effect of bone morphogenetic protein-6 on neurons from H2O2 injury. Brain Res 2007; 1163:10–20.
Du J, Yang S, Wang Z, et al. Bone morphogenetic protein 6 inhibit stress-induced breast cancer cells apoptosis via both Smad and p38 pathways. J Cell Biochem 2008; 103:1584–1597.
Dillner NB, Sanders MM . Transcriptional activation by the zinc-finger homeodomain protein δ EF1 in estrogen signaling cascades. DNA Cell Biol 2004; 23:25–34.
Acknowledgements
The PDCD4 3′UTR plasmid was a kind gift of Professor H Allgayer (University of Heidelberg, Germany). This work was supported by a grant from the National Natural Science Foundation of China to S Yang (No. 30700471).
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary information, Figure S1
miR-21 overexpression effect validation. (PDF 61 kb)
Supplementary information, Figure S2
BMP-6 reversed miR21-induced invasion in vitro. (PDF 65 kb)
Rights and permissions
About this article
Cite this article
Du, J., Yang, S., An, D. et al. BMP-6 inhibits microRNA-21 expression in breast cancer through repressing δEF1 and AP-1. Cell Res 19, 487–496 (2009). https://doi.org/10.1038/cr.2009.34
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2009.34
Keywords
This article is cited by
-
Inhibition of triple negative breast cancer metastasis and invasiveness by novel drugs that target epithelial to mesenchymal transition
Scientific Reports (2021)
-
Integrated nicking enzyme-powered numerous-legged DNA walker prepared by rolling circle amplification for fluorescence detection of microRNA
Microchimica Acta (2021)
-
MicroRNAs are Necessary for BMP-7-induced Dendritic Growth in Cultured Rat Sympathetic Neurons
Cellular and Molecular Neurobiology (2019)
-
Transregulation of microRNA miR-21 promoter by AP-1 transcription factor in cervical cancer cells
Cancer Cell International (2019)
-
microRNA-21 and hypertension
Hypertension Research (2018)