Abstract
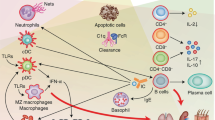
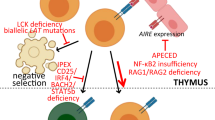
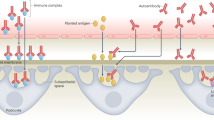
Immunological tolerance loss is fundamental to the development of autoimmunity; however, the underlying mechanisms remain elusive. Immune tolerance consists of central and peripheral tolerance. Central tolerance, which occurs in the thymus for T cells and bone marrow for B cells, is the primary way that the immune system discriminates self from non-self. Peripheral tolerance, which occurs in tissues and lymph nodes after lymphocyte maturation, controls self-reactive immune cells and prevents over-reactive immune responses to various environment factors. Loss of tolerance results in autoimmune disorders, such as systemic lupus erythematosus (SLE), rheumatoid arthritis (RA), type 1 diabetes (T1D) and primary biliary cirrhosis (PBC). The etiology and pathogenesis of autoimmune diseases are highly complicated. Both genetic predisposition and epigenetic modifications are implicated in the loss of tolerance and autoimmunity. In this review, we will discuss the genetic and epigenetic influences on tolerance breakdown in autoimmunity. Genetic and epigenetic influences on autoimmune diseases, such as SLE, RA, T1D and PBC, will also be briefly discussed.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Carvalheiras G, Faria R, Braga J, Vasconcelos C. Fetal outcome in autoimmune diseases. Autoimmun Rev 2012; 11: A520–530.
Alarcon-Riquelme ME. Recent advances in the genetics of autoimmune diseases. Ann N Y Acad Sci 2007; 1110: 1–9.
Strickland FM, Richardson BC. Epigenetics in human autoimmunity. Epigenetics in autoimmunity - DNA methylation in systemic lupus erythematosus and beyond. Autoimmunity 2008; 41: 278–286.
Jerne NK. The somatic generation of immune recognition. 1971. Eur J Immunol 2004; 34: 1234–1242.
Walker LS, Abbas AK. The enemy within: keeping self-reactive T cells at bay in the periphery. Nat Rev Immunol 2002; 2: 11–19.
Cheng MH, Anderson MS. Monogenic autoimmunity. Annu Rev Immunol 2012; 30: 393–427.
Finnish-German AC. An autoimmune disease, APECED, caused by mutations in a novel gene featuring two PHD-type zinc-finger domains. Nat Genet 1997; 17: 399–403.
Nagamine K, Peterson P, Scott HS, Kudoh J, Minoshima S, Heino M et al. Positional cloning of the APECED gene. Nat Genet 1997; 17: 393–398.
Liston A, Lesage S, Wilson J, Peltonen L, Goodnow CC. Aire regulates negative selection of organ-specific T cells. Nat Immunol 2003; 4: 350–354.
Hori S, Nomura T, Sakaguchi S. Pillars Article: Control of Regulatory T Cell Development by the Transcription Factor Foxp3. Science 2003; 299: 1057–1061J Immunol 2017; 198: 981–985.
Fontenot JD, Gavin MA, Rudensky AY. Foxp3 programs the development and function of CD4+CD25+ regulatory T cells. Nat Immunol 2003; 4: 330–336.
Khattri R, Cox T, Yasayko SA, Ramsdell F. An essential role for Scurfin in CD4+CD25+ T regulatory cells. Nat Immunol 2003; 4: 337–342.
Ramsdell F, Ziegler SF. FOXP3 and scurfy: how it all began. Nat Rev Immunol 2014; 14: 343–349.
Salomon B, Bluestone JA. Complexities of CD28/B7: CTLA-4 costimulatory pathways in autoimmunity and transplantation. Annu Rev Immunol 2001; 19: 225–252.
Klocke K, Sakaguchi S, Holmdahl R, Wing K. Induction of autoimmune disease by deletion of CTLA-4 in mice in adulthood. Proc Natl Acad Sci U S A 2016; 113: E2383–2392.
Ueda H, Howson JM, Esposito L, Heward J, Snook H, Chamberlain G et al. Association of the T-cell regulatory gene CTLA4 with susceptibility to autoimmune disease. Nature 2003; 423: 506–511.
Lenardo M, Chan KM, Hornung F, McFarland H, Siegel R, Wang J et al. Mature T lymphocyte apoptosis—immune regulation in a dynamic and unpredictable antigenic environment. Annu Rev Immunol 1999; 17: 221–253.
Fischer A, Rieux-Laucat F, Le Deist F. Autoimmune lymphoproliferative syndromes (ALPS): models for the study of peripheral tolerance. Rev Immunogenet 2000; 2: 52–60.
Northrop JK, Thomas RM, Wells AD, Shen H. Epigenetic remodeling of the IL-2 and IFN-gamma loci in memory CD8 T cells is influenced by CD4 T cells. J Immunol 2006; 177: 1062–1069.
Bruniquel D, Schwartz RH. Selective, stable demethylation of the interleukin-2 gene enhances transcription by an active process. Nat Immunol 2003; 4: 235–240.
Makar KW, Wilson CB. DNA methylation is a nonredundant repressor of the Th2 effector program. J Immunol 2004; 173: 4402–4406.
Medvedovic J, Ebert A, Tagoh H, Busslinger M. Pax5: a master regulator of B cell development and leukemogenesis. Adv Immunol 2011; 111: 179–206.
Bandyopadhyay S, Dure M, Paroder M, Soto-Nieves N, Puga I, Macian F. Interleukin 2 gene transcription is regulated by Ikaros-induced changes in histone acetylation in anergic T cells. Blood 2007; 109: 2878–2886.
Fields PE, Kim ST, Flavell RA. Cutting edge: changes in histone acetylation at the IL-4 and IFN-gamma loci accompany Th1/Th2 differentiation. J Immunol 2002; 169: 647–650.
Avni O, Lee D, Macian F, Szabo SJ, Glimcher LH, Rao A. T(H) cell differentiation is accompanied by dynamic changes in histone acetylation of cytokine genes. Nat Immunol 2002; 3: 643–651.
Wei G, Wei L, Zhu J, Zang C, Hu-Li J, Yao Z et al. Global mapping of H3K4me3 and H3K27me3 reveals specificity and plasticity in lineage fate determination of differentiating CD4+ T cells. Immunity 2009; 30: 155–167.
Lin YC, Jhunjhunwala S, Benner C, Heinz S, Welinder E, Mansson R et al. A global network of transcription factors, involving E2A, EBF1 and Foxo1, that orchestrates B cell fate. Nat Immunol 2010; 11: 635–643.
Kuchen S, Resch W, Yamane A, Kuo N, Li Z, Chakraborty T et al. Regulation of microRNA expression and abundance during lymphopoiesis. Immunity 2010; 32: 828–839.
Singleton AB, Hardy J, Traynor BJ, Houlden H. Towards a complete resolution of the genetic architecture of disease. Trends Genet 2010; 26: 438–442.
Long H, Yin H, Wang L, Gershwin ME, Lu Q. The critical role of epigenetics in systemic lupus erythematosus and autoimmunity. J Autoimmun 2016; 74: 118–138.
Shlomchik MJ, Craft JE, Mamula MJ. From T to B and back again: positive feedback in systemic autoimmune disease. Nat Rev Immunol 2001; 1: 147–153.
Jarvinen P, Aho K. Twin studies in rheumatic diseases. Semin Arthritis Rheum 1994; 24: 19–28.
Karassa FB, Trikalinos TA, Ioannidis JP. The role of FcgammaRIIA and IIIA polymorphisms in autoimmune diseases. Biomed Pharmacother 2004; 58: 286–291.
Kyogoku C, Langefeld CD, Ortmann WA, Lee A, Selby S, Carlton VE et al. Genetic association of the R620W polymorphism of protein tyrosine phosphatase PTPN22 with human SLE. Am J Hum Genet 2004; 75: 504–507.
Suarez A, Lopez P, Mozo L, Gutierrez C. Differential effect of IL10 and TNF{alpha} genotypes on determining susceptibility to discoid and systemic lupus erythematosus. Ann Rheum Dis 2005; 64: 1605–1610.
Rosado S, Rua-Figueroa I, Vargas JA, Garcia-Laorden MI, Losada-Fernandez I, Martin-Donaire T et al. Interleukin-10 promoter polymorphisms in patients with systemic lupus erythematosus from the Canary Islands. Int J Immunogenet 2008; 35: 235–242.
Sestak AL, Nath SK, Sawalha AH, Harley JB. Current status of lupus genetics. Arthritis Res Ther 2007; 9: 210.
Tarassi K, Carthy D, Papasteriades C, Boki K, Nikolopoulou N, Carcassi C et al. HLA-TNF haplotype heterogeneity in Greek SLE patients. Clin Exp Rheumatol 1998; 16: 66–68.
Sullivan KE, Suriano A, Dietzmann K, Lin J, Goldman D, Petri MA. The TNFalpha locus is altered in monocytes from patients with systemic lupus erythematosus. Clin Immunol 2007; 123: 74–81.
Kim HY, Lee SH, Yang HI, Park SH, Cho CS, Kim TG et al. TNFB gene polymorphism in patients with systemic lupus erythematosus in Korean. Korean J Intern Med 1995; 10: 130–136.
Prokunina L, Castillejo-Lopez C, Oberg F, Gunnarsson I, Berg L, Magnusson V et al. A regulatory polymorphism in PDCD1 is associated with susceptibility to systemic lupus erythematosus in humans. Nat Genet 2002; 32: 666–669.
Parks CG, Hudson LL, Cooper GS, Dooley MA, Treadwell EL St, Clair EW et al. CTLA-4 gene polymorphisms and systemic lupus erythematosus in a population-based study of whites and African-Americans in the southeastern United States. Lupus 2004; 13: 784–791.
Zhang Y, Zhao M, Sawalha AH, Richardson B, Lu Q. Impaired DNA methylation and its mechanisms in CD4(+)T cells of systemic lupus erythematosus. J Autoimmun 2013; 41: 92–99.
Yung RL, Richardson BC. Role of T cell DNA methylation in lupus syndromes. Lupus 1994; 3: 487–491.
Zhou Y, Lu Q. DNA methylation in T cells from idiopathic lupus and drug-induced lupus patients. Autoimmun Rev 2008; 7: 376–383.
Lu Q, Kaplan M, Ray D, Ray D, Zacharek S, Gutsch D et al. Demethylation of ITGAL (CD11a) regulatory sequences in systemic lupus erythematosus. Arthritis Rheum 2002; 46: 1282–1291.
Kaplan MJ, Deng C, Yang J, Richardson BC. DNA methylation in the regulation of T cell LFA-1 expression. Immunol Invest 2000; 29: 411–425.
Lu Q, Wu A, Richardson BC. Demethylation of the same promoter sequence increases CD70 expression in lupus T cells and T cells treated with lupus-inducing drugs. J Immunol 2005; 174: 6212–6219.
Mi XB, Zeng FQ. Hypomethylation of interleukin-4 and -6 promoters in T cells from systemic lupus erythematosus patients. Acta Pharmacol Sin 2008; 29: 105–112.
Lal G, Zhang N, van der Touw W, Ding Y, Ju W, Bottinger EP et al. Epigenetic regulation of Foxp3 expression in regulatory T cells by DNA methylation. J Immunol 2009; 182: 259–273.
Zhao M, Sun Y, Gao F, Wu X, Tang J, Yin H et al. Epigenetics and SLE: RFX1 downregulation causes CD11a and CD70 overexpression by altering epigenetic modifications in lupus CD4+ T cells. J Autoimmun 2010; 35: 58–69.
Li Y, Zhao M, Yin H, Gao F, Wu X, Luo Y et al. Overexpression of the growth arrest and DNA damage-induced 45alpha gene contributes to autoimmunity by promoting DNA demethylation in lupus T cells. Arthritis Rheum 2010; 62: 1438–1447.
Li Y, Huang C, Zhao M, Liang G, Xiao R, Yung S et al. A possible role of HMGB1 in DNA demethylation in CD4+ T cells from patients with systemic lupus erythematosus. Clin Dev Immunol 2013; 2013: 206298.
Hu N, Qiu X, Luo Y, Yuan J, Li Y, Lei W et al. Abnormal histone modification patterns in lupus CD4+ T cells. J Rheumatol 2008; 35: 804–810.
Zhou Y, Qiu X, Luo Y, Yuan J, Li Y, Zhong Q et al. Histone modifications and methyl-CpG-binding domain protein levels at the TNFSF7 (CD70) promoter in SLE CD4+ T cells. Lupus 2011; 20: 1365–1371.
Hedrich CM, Rauen T, Tsokos GC. cAMP-responsive element modulator (CREM)alpha protein signaling mediates epigenetic remodeling of the human interleukin-2 gene: implications in systemic lupus erythematosus. J Biol Chem 2011; 286: 43429–43436.
Zhang Z, Song L, Maurer K, Petri MA, Sullivan KE. Global H4 acetylation analysis by ChIP-chip in systemic lupus erythematosus monocytes. Genes Immun 2010; 11: 124–133.
Zhao S, Wang Y, Liang Y, Zhao M, Long H, Ding S et al. MicroRNA-126 regulates DNA methylation in CD4+ T cells and contributes to systemic lupus erythematosus by targeting DNA methyltransferase 1. Arthritis Rheum 2011; 63: 1376–1386.
Pan W, Zhu S, Yuan M, Cui H, Wang L, Luo X et al. MicroRNA-21 and microRNA-148a contribute to DNA hypomethylation in lupus CD4+ T cells by directly and indirectly targeting DNA methyltransferase 1. J Immunol 2010; 184: 6773–6781.
Ding S, Liang Y, Zhao M, Liang G, Long H, Zhao S et al. Decreased microRNA-142-3p/5p expression causes CD4+ T cell activation and B cell hyperstimulation in systemic lupus erythematosus. Arthritis Rheum 2012; 64: 2953–2963.
Tang Q, Yang Y, Zhao M, Liang G, Wu H, Liu Q et al. Mycophenolic acid upregulates miR-142-3P/5P and miR-146a in lupus CD4+T cells. Lupus 2015; 24: 935–942.
Liu Y, Dong J, Mu R, Gao Y, Tan X, Li Y et al. MicroRNA-30a promotes B cell hyperactivity in patients with systemic lupus erythematosus by direct interaction with Lyn. Arthritis Rheum 2013; 65: 1603–1611.
Tobon GJ, Youinou P, Saraux A. The environment, geo-epidemiology, and autoimmune disease: Rheumatoid arthritis. J Autoimmun 2010; 35: 10–14.
Fan W, Liang D, Tang Y, Qu B, Cui H, Luo X et al. Identification of microRNA-31 as a novel regulator contributing to impaired interleukin-2 production in T cells from patients with systemic lupus erythematosus. Arthritis Rheum 2012; 64: 3715–3725.
Alarcon-Segovia D, Alarcon-Riquelme ME, Cardiel MH, Caeiro F, Massardo L, Villa AR et al. Familial aggregation of systemic lupus erythematosus, rheumatoid arthritis, and other autoimmune diseases in 1,177 lupus patients from the GLADEL cohort. Arthritis Rheum 2005; 52: 1138–1147.
Orozco G, Rueda B, Martin J. Genetic basis of rheumatoid arthritis. Biomed Pharmacother 2006; 60: 656–662.
Chung IM, Ketharnathan S, Thiruvengadam M, Rajakumar G. Rheumatoid Arthritis: The Stride from Research to Clinical Practice. Int J Mol Sci 2016, 17: pii:E900.
Bayley R, Kite KA, McGettrick HM, Smith JP, Kitas GD, Buckley CD et al. The autoimmune-associated genetic variant PTPN22 R620W enhances neutrophil activation and function in patients with rheumatoid arthritis and healthy individuals. Ann Rheum Dis 2015; 74: 1588–1595.
Rong C, Hu W, Wu FR, Cao XJ, Chen FH. Interleukin-23 as a potential therapeutic target for rheumatoid arthritis. Mol Cell Biochem 2012; 361: 243–248.
Abdul-Sater AA, Edilova MI, Clouthier DL, Mbanwi A, Kremmer E, Watts TH. The signaling adaptor TRAF1 negatively regulates Toll-like receptor signaling and this underlies its role in rheumatic disease. Nat Immunol 2017; 18: 26–35.
Kurko J, Besenyei T, Laki J, Glant TT, Mikecz K, Szekanecz Z. Genetics of rheumatoid arthritis - a comprehensive review. Clin Rev Allergy Immunol 2013; 45: 170–179.
Duffau P, Menn-Josephy H, Cuda CM, Dominguez S, Aprahamian TR, Watkins AA et al. Promotion of Inflammatory Arthritis by Interferon Regulatory Factor 5 in a Mouse Model. Arthritis Rheumatol 2015; 67: 3146–3157.
Swierkot J, Nowak B, Czarny A, Zaczynska E, Sokolik R, Madej M et al. The Activity of JAK/STAT and NF-kappaB in Patients with Rheumatoid Arthritis. Adv Clin Exp Med 2016; 25: 709–717.
Julian B, Gao K, Harwood BN, Beinborn M, Kopin AS. Mutation-Induced Functional Alterations of CCR6. J Pharmacol Exp Ther 2017; 360: 106–116.
Fan L, Zong M, Gong R, He D, Li N, Sun LS et al. PADI4 Epigenetically Suppresses p21 Transcription and Inhibits Cell Apoptosis in Fibroblast-like Synoviocytes from Rheumatoid Arthritis Patients. Int J Biol Sci 2017; 13: 358–366.
Richardson B, Scheinbart L, Strahler J, Gross L, Hanash S, Johnson M. Evidence for impaired T cell DNA methylation in systemic lupus erythematosus and rheumatoid arthritis. Arthritis Rheum 1990; 33: 1665–1673.
Nakano K, Whitaker JW, Boyle DL, Wang W, Firestein GS. DNA methylome signature in rheumatoid arthritis. Ann Rheum Dis 2013; 72: 110–117.
Karouzakis E, Gay RE, Michel BA, Gay S, Neidhart M. DNA hypomethylation in rheumatoid arthritis synovial fibroblasts. Arthritis Rheum 2009; 60: 3613–3622.
Nile CJ, Read RC, Akil M, Duff GW, Wilson AG. Methylation status of a single CpG site in the IL6 promoter is related to IL6 messenger RNA levels and rheumatoid arthritis. Arthritis Rheum 2008; 58: 2686–2693.
Wood NC, Symons JA, Dickens E, Duff GW. In situ hybridization of IL-6 in rheumatoid arthritis. Clin Exp Immunol 1992; 87: 183–189.
Karouzakis E, Rengel Y, Jungel A, Kolling C, Gay RE, Michel BA et al. DNA methylation regulates the expression of CXCL12 in rheumatoid arthritis synovial fibroblasts. Genes Immun 2011; 12: 643–652.
Gillespie J, Savic S, Wong C, Hempshall A, Inman M, Emery P et al. Histone deacetylases are dysregulated in rheumatoid arthritis and a novel histone deacetylase 3-selective inhibitor reduces interleukin-6 production by peripheral blood mononuclear cells from rheumatoid arthritis patients. Arthritis Rheum 2012; 64: 418–422.
Toussirot E, Abbas W, Khan KA, Tissot M, Jeudy A, Baud L et al. Imbalance between HAT and HDAC activities in the PBMCs of patients with ankylosing spondylitis or rheumatoid arthritis and influence of HDAC inhibitors on TNF alpha production. PLoS One 2013; 8: e70939.
Huber LC, Brock M, Hemmatazad H, Giger OT, Moritz F, Trenkmann M et al. Histone deacetylase/acetylase activity in total synovial tissue derived from rheumatoid arthritis and osteoarthritis patients. Arthritis Rheum 2007; 56: 1087–1093.
Kawabata T, Nishida K, Takasugi K, Ogawa H, Sada K, Kadota Y et al. Increased activity and expression of histone deacetylase 1 in relation to tumor necrosis factor-alpha in synovial tissue of rheumatoid arthritis. Arthritis Res Ther 2010; 12: R133.
Araki Y, Tsuzuki Wada T, Aizaki Y, Sato K, Yokota K, Fujimoto K et al. Histone Methylation and STAT-3 Differentially Regulate Interleukin-6-Induced Matrix Metalloproteinase Gene Activation in Rheumatoid Arthritis Synovial Fibroblasts. Arthritis Rheumatol 2016; 68: 1111–1123.
Maciejewska-Rodrigues H, Karouzakis E, Strietholt S, Hemmatazad H, Neidhart M, Ospelt C et al. Epigenetics and rheumatoid arthritis: the role of SENP1 in the regulation of MMP-1 expression. J Autoimmun 2010; 35: 15–22.
Wada TT, Araki Y, Sato K, Aizaki Y, Yokota K, Kim YT et al. Aberrant histone acetylation contributes to elevated interleukin-6 production in rheumatoid arthritis synovial fibroblasts. Biochem Biophys Res Commun 2014; 444: 682–686.
Stanczyk J, Pedrioli DM, Brentano F, Sanchez-Pernaute O, Kolling C, Gay RE et al. Altered expression of MicroRNA in synovial fibroblasts and synovial tissue in rheumatoid arthritis. Arthritis Rheum 2008; 58: 1001–1009.
Alivernini S, Kurowska-Stolarska M, Tolusso B, Benvenuto R, Elmesmari A, Canestri S et al. MicroRNA-155 influences B-cell function through PU.1 in rheumatoid arthritis. Nat Commun 2016; 7: 12970.
Pauley KM, Satoh M, Chan AL, Bubb MR, Reeves WH, Chan EK. Upregulated miR-146a expression in peripheral blood mononuclear cells from rheumatoid arthritis patients. Arthritis Res Ther 2008; 10: R101.
Nakamachi Y, Kawano S, Takenokuchi M, Nishimura K, Sakai Y, Chin T et al. MicroRNA-124a is a key regulator of proliferation and monocyte chemoattractant protein 1 secretion in fibroblast-like synoviocytes from patients with rheumatoid arthritis. Arthritis Rheum 2009; 60: 1294–1304.
Fulci V, Scappucci G, Sebastiani GD, Giannitti C, Franceschini D, Meloni F et al. miR-223 is overexpressed in T-lymphocytes of patients affected by rheumatoid arthritis. Hum Immunol 2010; 71: 206–211.
Lu MC, Yu CL, Chen HC, Yu HC, Huang HB, Lai NS. Increased miR-223 expression in T cells from patients with rheumatoid arthritis leads to decreased insulin-like growth factor-1-mediated interleukin-10 production. Clin Exp Immunol 2014; 177: 641–651.
Stankov K, Benc D, Draskovic D. Genetic and epigenetic factors in etiology of diabetes mellitus type 1. Pediatrics 2013; 132: 1112–1122.
Noble JA, Erlich HA. Genetics of type 1 diabetes. Cold Spring Harb Perspect Med 2012; 2: a007732.
Thomson G, Valdes AM, Noble JA, Kockum I, Grote MN, Najman J et al. Relative predispositional effects of HLA class II DRB1-DQB1 haplotypes and genotypes on type 1 diabetes: a meta-analysis. Tissue Antigens 2007; 70: 110–127.
Balic I, Angel B, Codner E, Carrasco E, Perez-Bravo F. Association of CTLA-4 polymorphisms and clinical-immunologic characteristics at onset of type 1 diabetes mellitus in children. Hum Immunol 2009; 70: 116–120.
Sharp RC, Abdulrahim M, Naser ES, Naser SA. Genetic Variations of PTPN2 and PTPN22: Role in the Pathogenesis of Type 1 Diabetes and Crohn's Disease. Front Cell Infect Microbiol 2015; 5: 95.
Marwaha AK, Panagiotopoulos C, Biggs CM, Staiger S, Del Bel KL, Hirschfeld AF et al. Pre-diagnostic genotyping identifies T1D subjects with impaired Treg IL-2 signaling and an elevated proportion of FOXP3+IL-17+ cells. Genes Immun 2017; 18: 15–21.
Nepom GT, Buckner JH. A functional framework for interpretation of genetic associations in T1D. Curr Opin Immunol 2012; 24: 516–521.
Morahan G. Insights into type 1 diabetes provided by genetic analyses. Curr Opin Endocrinol Diabetes Obes 2012; 19: 263–270.
Park Y, Lee HS, Park Y, Min D, Yang S, Kim D et al. Evidence for the role of STAT4 as a general autoimmunity locus in the Korean population. Diabetes Metab Res Rev 2011; 27: 867–871.
Li Y, Zhao M, Hou C, Liang G, Yang L, Tan Y et al. Abnormal DNA methylation in CD4+ T cells from people with latent autoimmune diabetes in adults. Diabetes Res Clin Pract 2011; 94: 242–248.
Bell CG, Teschendorff AE, Rakyan VK, Maxwell AP, Beck S, Savage DA, Genome-wide DNA. methylation analysis for diabetic nephropathy in type 1 diabetes mellitus. BMC Med Genomics 2010; 3: 33.
Miao F, Chen Z, Zhang L, Liu Z, Wu X, Yuan YC et al. Profiles of epigenetic histone post-translational modifications at type 1 diabetes susceptible genes. J Biol Chem 2012; 287: 16335–16345.
Chen SS, Jenkins AJ, Majewski H. Elevated plasma prostaglandins and acetylated histone in monocytes in Type 1 diabetes patients. Diabet Med 2009; 26: 182–186.
Miao F, Smith DD, Zhang L, Min A, Feng W, Natarajan R. Lymphocytes from patients with type 1 diabetes display a distinct profile of chromatin histone H3 lysine 9 dimethylation: an epigenetic study in diabetes. Diabetes 2008; 57: 3189–3198.
Sebastiani G, Grieco FA, Spagnuolo I, Galleri L, Cataldo D, Dotta F. Increased expression of microRNA miR-326 in type 1 diabetic patients with ongoing islet autoimmunity. Diabetes Metab Res Rev 2011; 27: 862–866.
Salas-Perez F, Codner E, Valencia E, Pizarro C, Carrasco E, Perez-Bravo F. MicroRNAs miR-21a and miR-93 are down regulated in peripheral blood mononuclear cells (PBMCs) from patients with type 1 diabetes. Immunobiology 2013; 218: 733–737.
Yang M, Ye L, Wang B, Gao J, Liu R, Hong J et al. Decreased miR-146 expression in peripheral blood mononuclear cells is correlated with ongoing islet autoimmunity in type 1 diabetes patients 1miR-146. J Diabetes 2015; 7: 158–165.
Hirschfield GM, Gershwin ME. The immunobiology and pathophysiology of primary biliary cirrhosis. Annu Rev Pathol 2013; 8: 303–330.
Poupon R. Primary biliary cirrhosis: a 2010 update. J Hepatol 2010 52: 745–758.
Selmi C, Mayo MJ, Bach N, Ishibashi H, Invernizzi P, Gish RG et al. Primary biliary cirrhosis in monozygotic and dizygotic twins: genetics, epigenetics, and environment. Gastroenterology 2004; 127: 485–492.
Begovich AB, Klitz W, Moonsamy PV. Van de Water J, Peltz G, Gershwin ME. Genes within the HLA class II region confer both predisposition and resistance to primary biliary cirrhosis. Tissue Antigens 1994; 43: 71–77.
Hirschfield GM, Liu X, Xu C, Lu Y, Xie G, Lu Y et al. Primary biliary cirrhosis associated with HLA, IL12A, and IL12RB2 variants. N Engl J Med 2009; 360: 2544–2555.
Donaldson PT, Baragiotta A, Heneghan MA, Floreani A, Venturi C, Underhill JA et al. HLA class II alleles, genotypes, haplotypes, and amino acids in primary biliary cirrhosis: a large-scale study. Hepatology 2006; 44: 667–674.
Onishi S, Sakamaki T, Maeda T, Iwamura S, Tomita A, Saibara T et al. DNA typing of HLA class II genes; DRB1*0803 increases the susceptibility of Japanese to primary biliary cirrhosis. J Hepatol 1994; 21: 1053–1060.
Invernizzi P, Battezzati PM, Crosignani A, Perego F, Poli F, Morabito A et al. Peculiar HLA polymorphisms in Italian patients with primary biliary cirrhosis. J Hepatol 2003; 38: 401–406.
Joshita S, Umemura T, Nakamura M, Katsuyama Y, Shibata S, Kimura T et al. STAT4 gene polymorphisms are associated with susceptibility and ANA status in primary biliary cirrhosis. Dis Markers 2014; 2014: 727393.
Nakamura M, Nishida N, Kawashima M, Aiba Y, Tanaka A, Yasunami M et al. Genome-wide association study identifies TNFSF15 and POU2AF1 as susceptibility loci for primary biliary cirrhosis in the Japanese population. Am J Hum Genet 2012; 91: 721–728.
Dong M, Li J, Tang R, Zhu P, Qiu F, Wang C et al. Multiple genetic variants associated with primary biliary cirrhosis in a Han Chinese population. Clin Rev Allergy Immunol 2015; 48: 316–321.
Mells GF, Floyd JA, Morley KI, Cordell HJ, Franklin CS, Shin SY et al. Genome-wide association study identifies 12 new susceptibility loci for primary biliary cirrhosis. Nat Genet 2011; 43: 329–332.
Schuster C, Gerold KD, Schober K, Probst L, Boerner K, Kim MJ et al. The Autoimmunity-Associated Gene CLEC16A Modulates Thymic Epithelial Cell Autophagy and Alters T Cell Selection. Immunity 2015; 42: 942–952.
Selmi C, Cavaciocchi F, Lleo A, Cheroni C, De Francesco R, Lombardi SA et al. Genome-wide analysis of DNA methylation, copy number variation, and gene expression in monozygotic twins discordant for primary biliary cirrhosis. Front Immunol 2014; 5: 128.
Lleo A, Liao J, Invernizzi P, Zhao M, Bernuzzi F, Ma L et al. Immunoglobulin M levels inversely correlate with CD40 ligand promoter methylation in patients with primary biliary cirrhosis. Hepatology 2012; 55: 153–160.
Hu Z, Huang Y, Liu Y, Sun Y, Zhou Y, Gu M et al. beta-Arrestin 1 modulates functions of autoimmune T cells from primary biliary cirrhosis patients. J Clin Immunol 2011; 31: 346–355.
Padgett KA, Lan RY, Leung PC, Lleo A, Dawson K, Pfeiff J et al. Primary biliary cirrhosis is associated with altered hepatic microRNA expression. J Autoimmun 2009; 32: 246–253.
Sasaki M, Ikeda H, Sato Y, Nakanuma Y. Decreased expression of Bmi1 is closely associated with cellular senescence in small bile ducts in primary biliary cirrhosis. Am J Pathol 2006; 169: 831–845.
Sander S, Bullinger L, Klapproth K, Fiedler K, Kestler HA, Barth TF et al. MYC stimulates EZH2 expression by repression of its negative regulator miR-26a. Blood 2008; 112: 4202–4212.
Banales JM, Saez E, Uriz M, Sarvide S, Urribarri AD, Splinter P et al. Up-regulation of microRNA 506 leads to decreased Cl-/HCO3- anion exchanger 2 expression in biliary epithelium of patients with primary biliary cirrhosis. Hepatology 2012; 56: 687–697.
Ananthanarayanan M, Banales JM, Guerra MT, Spirli C, Munoz-Garrido P, Mitchell-Richards K et al. Post-translational regulation of the type III inositol 1,4,5-trisphosphate receptor by miRNA-506. J Biol Chem 2015; 290: 184–196.
Acknowledgements
This work was supported by the Hunan Provincial Natural Science Foundation of China (No.14JJ1009) and the National Natural Science Foundation of China (No. 81522038 and No. 81220108017).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Zhang, P., Lu, Q. Genetic and epigenetic influences on the loss of tolerance in autoimmunity. Cell Mol Immunol 15, 575–585 (2018). https://doi.org/10.1038/cmi.2017.137
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cmi.2017.137
This article is cited by
-
T cell expressions of aberrant gene signatures and Co-inhibitory receptors (Co-IRs) as predictors of renal damage and lupus disease activity
Journal of Biomedical Science (2024)
-
Sexual dimorphism during integrative endocrine and immune responses to ionizing radiation in mice
Scientific Reports (2024)
-
Proteomic analysis of lysine 2-hydroxyisobutyryl in SLE reveals protein modification alteration in complement and coagulation cascades and platelet activation Pathways
BMC Medical Genomics (2023)
-
The critical importance of epigenetics in autoimmune-related skin diseases
Frontiers of Medicine (2023)
-
Bridging autoinflammatory and autoimmune diseases
The Egyptian Journal of Internal Medicine (2021)