Abstract
To better elucidate epigenetic mechanisms that correlate with the dynamic gene expression program observed upon T-cell differentiation, we investigated the genomic landscape of histone modifications in naive and memory CD8+ T cells. Using a ChIP-Seq approach coupled with global gene expression profiling, we generated genome-wide histone H3 lysine 4 (H3K4me3) and H3 lysine 27 (H3K27me3) trimethylation maps in naive, T memory stem cells, central memory cells, and effector memory cells in order to gain insight into how histone architecture is remodeled during T cell differentiation. We show that H3K4me3 histone modifications are associated with activation of genes, while H3K27me3 is negatively correlated with gene expression at canonical loci and enhancers associated with T-cell metabolism, effector function, and memory. Our results also reveal histone modifications and gene expression signatures that distinguish the recently identified T memory stem cells from other CD8+ T-cell subsets. Taken together, our results suggest that CD8+ lymphocytes undergo chromatin remodeling in a progressive fashion. These findings have major implications for our understanding of peripheral T-cell ontogeny and the formation of immunological memory.
Similar content being viewed by others
INTRODUCTION
A naïve T lymphocyte is the smallest unit from which an adaptive cellular immune response can give rise to diverse progeny that collectively provide protection against acute infection and establish long-lasting immunologic memory.1 The diverse progeny of a single naïve T cell display unique phenotypic and functional attributes that distinguish numerous T-cell subsets including short-lived effectors and long-lived memory T cells.2,3,4
Reminiscent of hematopoietic stem cells that can self-renew and replenish all blood cell types,5,6,7 there is increasing evidence for a stem cell-like memory precursor in T cells. The T memory stem cell (TSCM) was recently identified in mice, nonhuman primates, and humans.8,9,10,11 TSCM are characterized by a naïve-like phenotype in conjunction with the expression of the memory markers interleukin-2 receptor β (IL2Rβ) and chemokine CXC receptor (CXCR)3.8 Functionally, TSCM cells maintain robust replicative capacity and are multipotent in their ability to give rise to cytolytic effector and memory progeny.
Elucidating the mechanisms by which a naïve T cell can give rise to a multiplicity of cellular states is critical for understanding the ontogeny of antigen-experienced T cells and the formation of immunological memory. While much of the diversity of cell subsets in a clonal response to infection is attributed to well-described trans- and cis-regulatory elements such as transcription factors and gene enhancers,12,13 epigenetic mechanisms—such as DNA methylation and histone modifications—are increasingly being recognized as critical components in the specification of distinct T-cell subsets that arise from the clonal response of a naïve T cell activated by its cognate antigen.14,15,16,17
The basic structure of eukaryotic chromatin is a nucleosome, which is comprised of 146 base pairs (bp) of DNA wrapped around an octomer of four core histones (H2A, H2B, H3, and H4).18 Post-translational modifications of the amino-terminal tail of histones have previously been shown to alter chromatin structure in lymphocytes and alter gene expression in activated T cells.14 In particular, it has been observed that trimethylation of lysine 27 on histone H3 (H3K27me3) is associated with gene repression, while H3 K4 trimethylation (H3K4me3) correlates with gene expression in T cells.14,15 There remains insufficient information, however, on histone modifications that occur at signature genes associated with memory and effector function during distinct stages of T-cell differentiation, particularly in the recently identified TSCM subset.
In the present analysis, we utilized chromatin immunoprecipitation coupled with massively parallel sequencing to map genome-wide H3K4me3 and H3K27me3 modifications in distinct subsets of CD8+ T cells—naïve (TN), memory stem cell (TSCM), central memory (TCM), and effector memory (TEM)—to gain insight into the epigenetic regulation of antigen-experienced T cells. Our results show dynamic remodeling of histone architecture after activation of naïve T cells, particularly in promoter regions of signature genes associated with memory and effector function, suggesting that chromatin remodeling seems to play a key role in the coordination of multiple gene expression programs that result in a multiplicity of CD8+ T-cell subsets. In addition, the findings support a model of CD8+ T-cell differentiation characterized by progressively increasing permissivity of chromatin at effector-associated genes, and concomitant loss of permissivity at memory-associated genes.
MATERIALS AND METHODS
Mice, cell culture, and isolation of cells
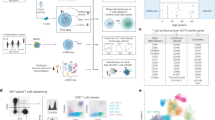
C57BL/6J mice were obtained from Jackson Laboratory (Bar Harbor, ME, USA). Animals were housed and cared for in accordance with the guidelines set forth by the National Institutes of Health (NIH) Animal Care and User Committee. Naïve splenic CD8+ T cells from C57BL/6J mice were isolated by fluorescence-activated cell sorting (FACS) based on CD62L+, CD44−, and SCA-1− phenotype. Central memory (TCM) and effector memory (TEM) T cells were generated in vitro with functional anti-CD3 (1 µg/ml) and anti-CD28 (1 µg/ml) antibody in IL-2-containing (100 IU/ml) complete media. T cells were cultured for 5 days and subsequently fluorescence-activated cell sorting (FACS)-sorted based on CD62L and CD44 expression. T memory stem cell (TSCM) T cells were generated in the same conditions as TCM and TEM with the addition of an inhibitor of GSK-3β (7 µM TWS119 EMD Millipore). TSCM were FACS-sorted based on expression of CD62L, CD44, and SCA-1 as previously described by Gattinoni et al.9
ChIP-Seq
ChIP-Seq assay was performed as described previously. Briefly, 2 × 107 T cells (per subset) were treated with MNase to generate approximately 20% dinucleosomes and 80% mononucleosomes. This yielded approximately 100 ng of DNA. Antibodies against H3K4me3 (ab8580, Abcam) and H3K27me3 (07-449, Upstate) were used. The ChIP DNA fragments were blunt-ended, ligated to Solexa adaptors, and sequenced with the Illumina 1G Genome Analyzer. Chip-Seq data are available in the National Center for Biotechnology Information (NCBI) GEO database under the accession number GSE67881.
Microarray data collection and analysis
Total cellular RNA was extracted from TN, TSCM, TCM, and TEM subsets with TRIzol reagent (Invitrogen) according to manufacturer’s protocol. Approximately 10 µg of RNA was labeled and hybridized to Mouse Gene 1.0 ST arrays (Affymetrix) according to manufacturer’s instructions, and probe-level intensities were called from images using the GeneChip Operating Software (GCOS) v1.1.1 software. The probe intensities stored in CEL files were then background-adjusted, quantile-normalized, and summarized into probeset-level and gene transcript-level expression levels using the RMA algorithm as implemented in the Affymetrix Power Tools’ apt-probeset-summarize tool (used with “-a rma” option and Affymetrix’s MoGene-1_0-st-v1.r4 pgf/clf/qcc/mps files). Two outlier samples revealed by principal component analysis were removed from further analysis. We also excluded gene transcripts flagged with potential cross-hybridization in the transcript annotations file provided by Affymetrix, and those not detected in all four cell types (where a transcript is defined as detected in a cell type if at least half of its exons are detected above background at P < 0.05 in at least half the samples of this cell type (or all samples if only two samples in this cell type), with the detection P computed using apt-probeset-summarize tool with the “-a dabg” option). Since the samples were done in three cell preparation batches, we accounted for batch effects19 by applying a standard adjustment procedure (viz., fitting a linear model of each gene’s expression levels across samples to the batch covariate and taking the intercept-added residuals from this fit as the adjusted expression data for further analyses). Microarray data are available in NCBI GEO database under accession number GSE67825.
Clustering and heatmap visualization of expression data
The gene expression heatmaps in this study were based on hierarchical clustering of the genes (and samples) of interest, using Euclidean distance metric and average-linkage agglomeration (as implemented in the R 2.15 statistical environment). Each gene’s expression levels across samples were Z-score normalized before hierarchical clustering, and the hierarchical clustering dendrogram reordered using the “optimal leaf ordering” algorithm from R package “seriation” to improve visualization of the expression patterns. Results of an independent k-means clustering of genes with k = 4 (as obtained from R’s k-means function using default parameters but 25 random restarts) is shown alongside certain heatmaps to identify gene groups with broadly similar expression patterns.
Identification of H3K4me3 and H3K27me3 islands
We followed a previously published approach to analyze histone modification data.20 Briefly, ChIP-Seq reads of largely 25 bp corresponding to H3K4me3 or H3K27me3 modification levels were obtained with the Solexa Analysis Pipeline. All reads were mapped to the mouse genome (mm8) and reads mapping to multiple places in the genome were removed. We then filtered these genome-mapped reads by retaining only reads that fell in “islands” of enriched ChIP signals found by the SICER tool. Note that SICER relies on a clustering approach and Poisson statistics in fixed-size windows along the genome to find gapped windows or islands of enriched signals. We used settings recommended in SICER v1.1 documentation for using SICER without a control library—specifically, we used counts of unique reads falling in 200 bp windows, gapsize of 1 window for H3K4me3 and 3 windows for H3K27me3, fragment size of 150 bp (which corresponds to shifting mapped reads by 75 bp to center them on the fragment), effective fraction of mm8 genome uniquely mappable by 25 bp reads to 0.778, and finally expected false positives or E-value of 500 islands. The SICER-filtered, mapped reads are called modification tags or simply tags in the text.
Quantitative calculation of H3K4me3 and H3K27me3 levels for all genes
In order to quantify the H3K4me3 or H3K27me3 levels of a particular gene region, we normalized the number of tags falling in that region by the library size (total number of modification tags) and the length of the region in kbp (kilo bp). This normalized tag density was calculated for the promoter region (TSS−2kbp to TSS+2kbp) and the gene body (TSS−2kbp to transcription end) of each gene. All our results and figures, unless otherwise specified, use the promoter region density as a gene’s normalized tag density. In order to plot the tag density profiles of genes, the same procedure was applied to calculate the normalized tag density in smaller windows defined along a gene body (windows inside a gene body of size 5% of the total transcript length, and surrounding windows of size 1 kbp in the 5 kbp region enclosing the gene transcript). For a gene with multiple transcripts with different TSS or transcription end sites, we calculated the normalized tag density of each transcript and took the largest of them as the modification level of the gene.
RESULTS
H3K4me3 and H3K27me3 histone modifications are enriched in promoter regions and correlate with gene expression in CD8+ T-cell subsets
Although it has previously been shown that histone modification and nucleosome positioning correlate with gene transcription in T cells,14,21,22 we sought to elucidate whether distinct subsets of antigen-experienced T cells display a unique histone architecture that correlates with gene expression. To reveal epigenetic features acquired during CD8+ T-cell differentiation, we generated global maps of H3K4me3 and H3K27me3 modifications and compared these with global gene expression profiles in four subsets of peripheral CD8+ T cells: naïve (TN; CD62L+CD44−SCA-1−), memory stem cell (TSCM; CD62L+CD44− SCA-1+), central memory (TCM; CD62L+CD44+), and effector memory (TEM; CD62L−CD44+) T cells. Histone modification levels were quantified as previously published20 by uniquely aligning ChIP-Seq reads to the mouse genome and retaining only those that fall inside islands of enriched signals. The total number of mapped reads before filtering for H3K27me3 for each subset is as follows: (TN = 10,760,085), (TSCM = 12,686,401), (TCM = 12,393,934), and (TEM = 13015897). For H3K4me3, the total number of mapped before filtering is: (TN = 11,062,100), (TSCM = 11,391,608), (TCM = 12,571,378), and (TEM = 12,476,165). After filtering, the total number of mapped reads for H3K27me3 for each subset is as follows: (TN = 518,268) (TSCM = 579,475), (TCM = 649,347), and (TEM = 786,435). For H3K4me3, the total number of island-filtered reads is: (TN = 420,850), (TSCM = 550,534), (TCM = 2,436,594), and (TEM = 3,278,059).
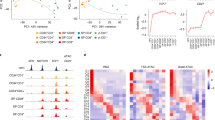
The distribution of H3K4me3 and H3K27me3 modifications along the body of a typical gene revealed an enrichment of histone modification tags at the transcription start site, sparsely distributed tag density along the gene body, and increased density after the transcription end site (Supplemental Figure 1A and B). Given the enrichment at promoter regions, we quantified histone modification levels of individual genes by computing the normalized density of tags in promoter regions, and correlated it with gene expression levels (mean RMA-normalized intensity of 100 genes that were grouped based on their rank of mRNA expression from highest to lowest). Consistent with earlier studies, we observed that H3K4me3 positively correlates with gene expression (Figure 1A) and H3K27me3 negatively correlates with gene expression of antigen-experienced T cells (Figure 1B).
Global correlation between gene expression and histone methylation in CD8+ T-cell subsets. Normalized tag count densities of H3K4me3 and H3K27me3 were plotted against the gene expression of all annotated genes. Each dot (in plots A and B) represents the mean value of 100 genes that were grouped based on their rank of mRNA expression from highest to lowest. (A) Gene expression was positively correlated with H3K4me3 in naïve (TN), stem cell memory (TSCM), central memory (TCM), and effector memory (TEM) CD8+ T-cell subsets. (B) Gene expression was negatively correlated with H3K27me3 in naïve, stem cell memory (TSCM), central memory (TCM), and effector memory (TEM) CD8+ T-cell subsets.
Histone modifications at CD8+ T-cell signature genes
As we were particularly interested in activating and repressive histone modifications at gene loci exhibiting distinct expression patterns across the four CD8+ T-cell subsets under study, we sought to identify signature genes that are differentially expressed (defined using an analysis of variance (ANOVA) test P <= 0.01 after Benjamini–Hochberg correction for multiple testing; see Supplemental Dataset 1) in at least one of the four cell subsets. A hierarchical cluster analysis of the resulting 5916 signature genes demonstrated that TN and TSCM subsets segregate from their TCM and TEM counterparts (Figure 2A). In addition, we observed that H3K4me3 broadly correlates with gene expression and H3K27me3 showed a negative correlation with the expression of these signature genes defining the subsets of antigen-experienced CD8+ T cells (Figure 2B).
Correlation between gene expression and histone methylation in signature genes. (A) Heat map of signature genes (defined as those specific to at least one of the T-cell subsets using an ANOVA test at BH-adjusted P <= 0.01). Genes with similar patterns across the cell types are grouped using k-means algorithm and marked by colors alongside the rows. (B) Scatterplot showing H3K4me3 correlation with median signature gene expression and H3K27me3 correlation with median signature gene repression in a population of naïve (red), stem cell memory (green), central memory (blue), and effector memory (purple) CD8+ T cells.
It was striking that gene expression of two-thirds of signature genes were either progressively upregulated (Figure 2A, purple cluster, 31%) or progressively downregulated (Figure 2A, red cluster, 35%) across subsets of T cells (from naïve T cells → TSCM → TCM → TEM). A progressive pattern in enhanced gene expression (of at least a 1.5-fold change) across distinct T-cell subsets correlated with a trend in either acquisition of permissive H3K4me3 or loss of repressive H3K27me3 histone modifications (Figure 3A), while a progressive downregulation of gene expression (of at least a 1.5-fold change) correlated with a trend in loss of permissive H3K4me3 and gain of repressive H3K27me3 marks (Figure 3B). Increasingly greater H3K4me3 tag density correlated with gene expression for 31% of 271 genes that were progressively upregulated (Figure 3A), while loss of H3K4me3 tag density was consistent with 44% of 77 genes that were progressively downregulated (Figure 3B). In similar fashion, loss of H3K27me3 tag density correlated with 22% of 76 genes that were progressively upregulated (Figure 3A), while increasing gain of H3K27me3 tag density was consistent with 14% of 36 genes that were progressively downregulated (Figure 3B).
Histone methylation in signature genes that show progressive expression. (A) Box plots showing normalized tag density (after subtraction from naïve) of permissive H3K4me3 or repressive H3K27me3 in progressively upregulated signature genes with at least 1.5-fold change in purple cluster of Figure 2, panel A. (B) Box plots showing normalized tag density (after subtraction from naïve) of permissive H3K4me3 or repressive H3K27me3 in progressively downregulated signature genes with at least 1.5-fold change in red cluster of Figure 2, panel A. Color of lines in panels (A) and (B) represent a change of tag density from naïve to stem cell memory T cells and stem cell memory to average of central and effector memory T cells (naïve T cell → TSCM → average of TCM and TEM) in the following directions: red (down–down), yellow (down–up), blue (up–down), purple (up–up), and green (other) where “up” means tag density difference between the two compared subset is >0, and “down” means tag density difference between the two compared subset is <0, and “other” means no difference in tag density in at least one of the two comparisons. In each panel, we excluded genes that have zero normalized tag density in all cell subsets.
Whereas our signature genes distinguish CD8+ T-cell subsets derived using in vitro cultures, a set of genes that distinguish memory vs. effector CD8+ T-cell subsets isolated ex vivo after infection with listeria monocytogenes or vesicular stomatitis virus (Lm-OVA or VSV-OVA) has previously been reported by the Immgen Consortium.23 We found that inspecting the expression data of our T-cell subsets among these Immgen in vivo signature genes (Supplemental Figure 2A) produced a similar clustering of our T-cell subsets as the larger in vitro signature set. The observation that H3K4me3 broadly correlates positively with gene expression and H3K27me3 negatively correlates with gene expression was also repeated in the in vivo signature gene set (Supplemental Figure 2B) and in a number of other previously reported Immgen gene clusters associated with effector cytokines (Supplemental Figure 3) and cell cycle and division (Supplemental Figure 4) of CD8+ T cells. To further validate our data set, we evaluated genes in our T-cell subsets that were recently reported to either be upregulated or downregulated in memory and effector T cells isolated from C57BL/6J mice infected with influenza A virus.24 We observed similar gene expression between data sets, particularly among genes that were dynamically regulated in effector T cells (Supplemental Figures 5A and B, 6). Taken together, the observed corroboration between our data set and the two published data sets addresses, in part, a major limitation in the present study—the in vitro derivation of T cells.
Histone modification of H3K4me3 and H3K27me3 at memory-associated genes
Having observed a global correlation between gene expression and histone methylation, we wanted to next focus our study on canonical genes and enhancer regions associated with T-cell effector function, metabolism, and immunological memory. There is increasing evidence that the Wnt/β-catenin signaling pathway plays a central role in the generation and maintenance of CD8+ T-cell memory.25,27 Expression of transcription factor 7 (Tcf7) and lymphoid enhancer-binding factor 1 (Lef1) have previously been reported to be highest in naïve and central memory subsets, and substantially downregulated in short-lived effector cells.25,27 In the present analysis, naïve T cells showed high levels of H3K4me3 in both the Tcf7 gene body and promoter region, suggesting that chromatin modified by H3K4me3 is competent for transcription and constitutive gene expression in quiescent naïve T cells (Figure 4A and B). After activation TSCM maintained high H3K4me3 marks similar to their naïve counterparts, but we observed progressively fewer activating H3K4me3 marks in the TCM and TEM subsets, respectively. In addition, the repressive H3K27me3 histone modification was absent in the Tcf7 gene body of naïve and TSCM subsets, but measurably higher in the promoter and 3′-untranslated region (UTR) of TCM and TEM subsets. In similar fashion, activating histone marks were observed in the promoter region of Lef1 in naïve and TSCM cells, but the appearance of inhibitory H3K27me3 modifications just proximal to the activating marks were observed in TCM and TEM subsets, suggesting that the chromatin associated with Lef1 becomes progressively less permissive of gene expression with terminal differentiation (Figure 4A and B).
Chromatin remodeling in genes associated with memory. (A) Distribution of H3K27me3 and H3K4me3 modifications at canonical memory-associated genes—transcription factor 7 (Tcf7), lymphoid enhancer binding factor 1 (Lef1), Kruppel-like factor 2 (Klf2), and forkhead box O1 (Foxo1)—in the following antigen-experienced CD8+ T-cell subsets: naïve (TN), stem cell memory (TSCM), central memory (TCM), and effector memory (TEM). All figures depicting the distribution of H3K4me3 and H3K27me3 islands are labeled such that the black arrow represents the direction of gene transcription. Gene structure is downloaded from UCSC Genome Browser and only tags on islands are shown. The islands labeled in red represent H3K4me3 and blue islands represent H3K27me3. Scales are constant among cell subsets. (B) Graph shows normalized tag density of H3K4me3 and H3K27me3 (chromatin remodeling) and RMA-normalized intensity (gene expression) for indicated genes and CD8+ T-cell subsets.
The forkhead O transcription factors (Foxo) have recently been shown to have an essential role in the formation of long-lived memory.28,29,30 Because Foxo1-deficient CD8+ T cells lose the capacity to expand after secondary infection,29 we expected that histone modifications at the Foxo1 locus are less permissive of gene expression with progressive differentiation. Indeed, we observed that activating H3K4me3 marks predominate at both the Foxo1 locus, and its target Kruppel-like factor 2 (Klf2), in minimally differentiated naïve and TSCM, but increasingly acquire inhibitory H3K27me3 marks in TCM and TEM subsets (Figure 4A and B).
Finally, we sought to extend our analysis of histone modifications in select enhancer regions of genes—namely Tcf7 and Klf2—thought to play a particularly important role in the formation of TSCM.8,9 In this analysis, we used a previously published data set obtained with P14 CD8+ effector T cells to evaluate enhancer regions which were identified with H3 monomethylation at lysine 4 (H3K4me1).31 In these putative enhancers of both Tcf7 and Klf2, we observed a progressive accumulation of repressive H3K27me3 marks (Figure 5A) across antigen-experienced subsets that correlated with a reduction of gene expression at these loci (Figure 5B). Collectively, these results suggest that H3K4me3 and H3K27me3 modifications are dynamically regulated after antigen activation at canonical loci associated with immunological memory.
Enhancers of Tcf7 and Klf2 progressively acquire repressive H3K27me3 modifications. (A) Distribution of H3K27me3 and H3K4me3 modifications at putative enhancer regions (identified by enrichment of H3K4me1 marks in data set from ref. 31) of Tcf7 and Klf2 in the following antigen-experienced CD8+ T-cell subsets: naïve (TN), stem cell memory (TSCM), central memory (TCM), and effector memory (TEM).
Histone modification of H3K4me3 and H3K27me3 at genes associated with effector function
Having shown that gene loci central to the formation of T-cell memory correlate with permissive chromatin modifications in naïve and TSCM, but less extensively in TCM and TEM, we were interested in better elucidating the histone modifications at genes known to be critical for T-cell effector function. Graded expression of T-box transcription factor (Tbx21) has previously been shown to direct the fate of memory and short-lived effector T cells.13,32,33,34,35,36,37 In our analysis, naïve CD8+ T cells showed extensive repressive H3K27me3 marks throughout the Tbx21 promoter and gene body, whereas repressive marks were only apparent in the 3′-UTR of TSCM (Figure 6A and B). There were no detectable H3K27me3 modifications in TCM and TEM, but increased H3K4me3 mark at the promoter region. Because previous studies have shown that Tbx21 regulates expression of interferon-gamma (Ifnγ),38,39 we anticipated that the chromatin would be more permissive of gene expression at the Ifnγ loci in TCM and TEM compared to their naïve counterparts. Consistently, naïve and TSCM did not have detectable levels of H3K4me3 histone modifications at this locus. Other canonical effector molecules, namely granzyme B and perforin, were also found to have highly permissive chromatin in the more terminally differentiated subsets as evidenced by high prevalence of activating H3K4me3 marks. Finally, B lymphocyte-induced maturation protein-1 (Blimp-1; encoded by Prdm1) is known to promote terminal differentiation of CD8+ T cells and is a negative regulator of memory40,41,42; accordingly, we observed an increase in activating H3K4me3 modifications in both TCM and TEM compared to naïve and TSCM, along with progressive loss of repressive H3K27me3 histone marks (Figure 4A and B). These results suggest that H3K4me3 and H3K27me3 marks are dynamically modified in a manner that correlates with expression of genes central to the effector program of T cells.
Chromatin remodeling in genes associated with effector function. (A) Distribution of H3K27me3 and H3K4me3 modifications at canonical effector-associated genes—T-box 21 (Tbx21), PR domain containing 1 with ZNF domain (Prdm1), perforin 1 (Prf1), granzyme B (Gzmb), and interferon gamma (Ifng)—in the following antigen-experienced CD8+ T-cell subsets: naïve (TN), stem cell memory (TSCM), central memory (TCM), and effector memory (TEM). (B) Graph shows normalized tag density of H3K4me3 and H3K27me3 (chromatin remodeling) and RMA-normalized intensity (gene expression) for indicated genes and CD8+ T-cell subsets.
Dynamic histone modifications at key metabolic genes
Naïve T cells are primarily fueled by energy from oxidative phosphorylation generated by break down of fatty acids, glucose, and amino acids.43 The metabolic enzyme that controls the rate-limiting step of mitochondrial fatty acid oxidation, carnitine palmitoyltransferase (Cpt1a), accordingly demonstrated marginal alterations in gene expression and histone methylation across all subsets (Figure 7A and B). After activation, relatively quiescent naïve T cells undergo metabolic reprogramming to prepare for rapid proliferation and cytolytic effector function by rapid induction of aerobic glycolysis.43 Consistently, we observed enhanced gene expression in key glycolytic enzymes including hexokinase 2 (Hk2) (Figure 7B), which phosphorylates glucose to produce glucose-6-phosphate, thereby committing glucose to the glycolytic pathway. Because reduced activity of triose phosphate isomerase (Tpi1) can lead to partial inhibition of glycolysis,44 we evaluated its expression and found that it progressively increased across subsets (Figure 7B). Similarly pyruvate kinase (Pkm2)—which catalyzes the last step in glycolysis, and lactate dehydrogenase (Ldha)—which catalyzes interconversion of pyruvate and lactate, also showed progressive upregulation in antigen-experienced subsets (Figure 7B). With the exception of Hk2, gene expression in glycolytic-associated enzymes did not correlate with H3K27me3, which was virtually absent at these loci but did seem to associate with increasingly permissive H3K4me3 (Figure 7A and B). Taken together, these findings suggest that metabolic reprogramming of antigen-experienced T cells is associated with progressive chromatin remodeling that correlates with dynamic gene expression.
Metabolic reprogramming of T cells associated with chromatin remodeling. (A) Distribution of H3K27me3 and H3K4me3 modifications at genes implicated in T-cell metabolism—carnitine palmitoyl transferase (Cpt1a), hexokinase 2 (Hk2), lactate dehydrogenase (Ldha), pyruvate kinase (Pkm2), triose phosphate isomerase (Tpi1)—in the following antigen-experienced CD8+ T-cell subsets: naïve (TN), stem cell memory (TSCM), central memory (TCM), and effector memory (TEM). (B) Graph shows normalized tag density of H3K4me3 and H3K27me3 (chromatin remodeling) and RMA-normalized intensity (gene expression) for indicated genes and CD8+ T-cell subsets.
DISCUSSION
To better understand epigenetic mechanisms that underlie the ontogeny of peripheral CD8+ T cells, we have generated genome-wide histone H3 lysine 4 (H3K4me3) and H3 lysine 27 (H3K27me3) trimethylation maps in distinct subsets of antigen-experienced CD8+ T cells. We show that covalent histone modifications are associated with activation (H3K4me3) and repression (H3K27me3) of gene expression at canonical loci associated with T-cell metabolism, effector function, and memory. Our results also suggest that antigen-experienced CD8+ T cells undergo chromatin remodeling in a progressive fashion—from naïve and TSCM to TCM and TEM—that may have implications for our understanding of peripheral T-cell ontogeny and the formation of immunological memory.
In response to an infectious challenge, a naïve CD8+ T cell clonally expands into distinct lineages, which are preserved in subsequent rounds of cell division.45 The lineages are defined by surface phenotype, metabolism, acquisition of cytolytic function, and the capacity to persist long after an infectious threat has been eradicated (i.e. memory).46,47,48 The predominant model for CD8+ T-cell differentiation is that naïve cells, upon activation by their cognate antigen, clonally expand and differentiate into cytolytic effector T cells.45,49 After clearance of the infection, a small proportion of the effector cells become long-lived memory cells capable of homeostatic proliferation and reactivation with episodic infectious challenges.50
In the context of the present analysis, this would suggest a pattern of chromatin remodeling of ‘closed-open-closed’ and ‘open-closed-open’ with regard to the permissivity of chromatin at gene loci involved in effector function and memory, respectively. Indeed, this has recently been proposed to explain other epigenetic mechanisms such as DNA methylomes observed in antigen-specific naïve and effector CD8+ T cells.17 Other developmental systems, however, do not seem to exhibit reversible chromatin changes as cells mature through a lineage, but more closely resemble a progressive change in chromatin as cells differentiate from stem cells to progenitors and ultimately to mature somatic cells.51,52 In other words, specification of cell fate seems to be accompanied by a discrete transition from a uniquely accessible epigenetic landscape to increasingly restrictive configurations.52 It may be that the adaptive immune system has uniquely evolved to dynamically open and close its chromatin, but this would seem to be exceptional as developmental specification has typically been observed to progress from dynamic remodeling to generalized compaction of histone architecture.
Another possibility is the “stem cell model” of CD8+ T-cell differentiation in which there is progressive and largely irreversible differentiation, proceeding from naïve T cell → TSCM →TCM → TEM→ TEFF.53,54 The TSCM stage, being the most minimally differentiated of the antigen-stimulated T cells,8,11 essentially retains the developmental potential of a naïve cell, including its capacity for marked clonal expansion and homeostatic proliferation, and can therefore provide a continual source of effector progeny during acute and chronic infections. The chromatin modifications observed in the present analysis also support this view as both the naïve and TSCM subsets were found to have activating histone modifications at memory-associated gene loci (e.g. Tcf7, Lef1, Foxo1, and Klf2) and poorly permissive chromatin at loci associated with effector function (e.g. Prf1, Gzmb, and Ifng), glycolytic metabolism (e.g. Hk2, Ldha, Pkm2, and Tpi1), and terminal differentiation (e.g. Tbx21, Prdm1). The finding that genes associated with glycolysis and effector-function are largely associated with a progressive increase in permissive H3K4me3 modifications—and not a profound loss of repressive H3K27me3 marks—is consistent with the recent findings by Russ et al. in which acquisition of H3K4me3 marks at immune-related effector gene promoters was observed with progressive differentiation into effector cytotoxic T cells.24
In summary, the present study provides an initial roadmap of the genome-wide histone H3 lysine 4 (H3K4me3) and H3 lysine 27 (H3K27me3) trimethylation modifications that are respectively permissive and antagonistic toward gene expression in distinct stages of CD8+ T-cell differentiation, including the recently identified T memory stem cell population. Sustained immunity to intracellular pathogens and cancer requires that CD8+ T cells have the capacity for long-lived persistence and effector function long after their initial exposure to antigen. The molecular mechanisms underpinning the durability of antigen-experienced CD8+ T cells has largely focused on differential regulation of gene expression by canonical transcription factors that promote immunological memory. By analyzing the genomic landscape of histone modifications in well-defined CD8+ T cells—naïve, memory stem cell, central memory, and effector memory—this study broadens our view of the mechanisms that dictate the fate of activated T cells to better understand how epigenetic regulation accompanies a transcriptional program that drives and sustains CD8+ T-cell differentiation.
References
Burnet FM . Immunological recognition of self . Science 1961 ; 133 : 307 – 311.
Sallusto F, Lenig D, Forster R, Lipp M, Lanzavecchia A . Two subsets of memory T lymphocytes with distinct homing potentials and effector functions . Nature 1999 ; 401 : 708 – 712.
Romero P et al . Four functionally distinct populations of human effector-memory CD8+ T lymphocytes . J Immunol 2007 ; 178 : 4112 – 4119.
Plumlee CR, Sheridan BS, Cicek BB, Lefrancois L . Environmental cues dictate the fate of individual CD8+ T cells responding to infection . Immunity 2013 ; 39 : 347 – 356.
Spangrude GJ, Heimfeld S, Weissman IL . Purification and characterization of mouse hematopoietic stem cells . Science 1988 ; 241 : 58 – 62.
Weissman IL . Translating stem and progenitor cell biology to the clinic: barriers and opportunities . Science 2000 ; 287 : 1442 – 1446.
Weissman IL . Stem cells: units of development, units of regeneration, and units in evolution . Cell 2000 ; 100 : 157 – 168.
Gattinoni L et al . A human memory T cell subset with stem cell-like properties . Nat Med 2011 ; 17 : 1290 – 1297.
Gattinoni L et al . Wnt signaling arrests effector T cell differentiation and generates CD8+ memory stem cells . Nat Med 2009 ; 15 : 808 – 813.
Zhang Y, Joe G, Hexner E, Zhu J, Emerson SG . Host-reactive CD8+ memory stem cells in graft-versus-host disease . Nat Med 2005 ; 11 : 1299 – 1305.
Lugli E et al . Superior T memory stem cell persistence supports long-lived T cell memory . J Clin Invest 2013 ; 123 : 594 – 599.
Kaech SM, Cui W . Transcriptional control of effector and memory CD8+ T cell differentiation . Nat Rev Immunol 2012 ; 12 : 749 – 761.
Intlekofer AM et al . Effector and memory CD8+ T cell fate coupled by T-bet and eomesodermin . Nat Immunol 2005 ; 6 : 1236 – 1244.
Roh TY, Cuddapah S, Cui K, Zhao K . The genomic landscape of histone modifications in human T cells . Proc Natl Acad Sci U S A 2006 ; 103 : 15782 – 15787.
Araki Y et al . Genome-wide analysis of histone methylation reveals chromatin state-based regulation of gene transcription and function of memory CD8+ T cells . Immunity 2009 ; 30 : 912 – 925.
Shin HM et al . Epigenetic modifications induced by Blimp-1 Regulate CD8(+) T cell memory progression during acute virus infection . Immunity 2013 ; 39 : 661 – 675.
Scharer CD, Barwick BG, Youngblood BA, Ahmed R, Boss JM . Global DNA methylation remodeling accompanies CD8 T cell effector function . J Immunol 2013 ; 191 : 3419 – 3429.
Margueron R, Reinberg D . Chromatin structure and the inheritance of epigenetic information . Nat Rev Genet 2010 ; 11 : 285 – 296.
Leek JT et al . Tackling the widespread and critical impact of batch effects in high-throughput data . Nat Rev Genet 2010 ; 11 : 733 – 739.
Wei G et al . Global mapping of H3K4me3 and H3K27me3 reveals specificity and plasticity in lineage fate determination of differentiating CD4+ T cells . Immunity 2009 ; 30 : 155 – 167.
Schones DE et al . Dynamic regulation of nucleosome positioning in the human genome . Cell 2008 ; 132 : 887 – 898.
Barski A et al . High-resolution profiling of histone methylations in the human genome . Cell 2007 ; 129 : 823 – 837.
Best JA et al . Transcriptional insights into the CD8(+) T cell response to infection and memory T cell formation . Nat Immunol 2013 ; 14 : 404 – 412.
Russ BE et al . Distinct epigenetic signatures delineate transcriptional programs during virus-specific CD8(+) T cell differentiation . Immunity 2014 ; 41 : 853 – 865.
Jeannet G et al . Essential role of the Wnt pathway effector Tcf-1 for the establishment of functional CD8 T cell memory . PNAS 2010 ; 107(21) : 97777 – 82.
Zhao DM et al . Constitutive activation of Wnt signaling favors generation of memory CD8 T cells . J Immunol 2010 ; 184 : 1191 – 1199.
Zhou X, Xue HH . Cutting edge: generation of memory precursors and functional memory CD8+ T cells depends on T cell factor-1 and lymphoid enhancer-binding factor-1 . J Immunol 2012 ; 189 : 2722 – 2726.
Tejera MM, Kim EH, Sullivan JA, Plisch EH, Suresh M . FoxO1 controls effector-to-memory transition and maintenance of functional CD8 T cell memory . J Immunol 2013 ; 191 : 187 – 199.
Kim MV, Ouyang W, Liao W, Zhang MQ, Li MO . The transcription factor Foxo1 controls central-memory CD8+ T cell responses to infection . Immunity 2013 ; 39 : 286 – 297.
Hess Michelini R, Doedens AL, Goldrath AW, Hedrick SM . Differentiation of CD8 memory T cells depends on Foxo1 . J Exp Med 2013 ; 210 : 1189 – 1200.
Kurachi M et al . The transcription factor BATF operates as an essential differentiation checkpoint in early effector CD8+ T cells . Nat Immunol 2014 ; 15 : 373 – 383.
Joshi NS et al . Inflammation directs memory precursor and short-lived effector CD8(+) T cell fates via the graded expression of T-bet transcription factor . Immunity 2007 ; 27 : 281 – 295.
Pipkin ME et al . Interleukin-2 and inflammation induce distinct transcriptional programs that promote the differentiation of effector cytolytic T cells . Immunity 2010 ; 32 : 79 – 90.
Intlekofer AM et al . Requirement for T-bet in the aberrant differentiation of unhelped memory CD8+ T cells . J Exp Med 2007 ; 204 : 2015 – 2021.
Intlekofer AM et al . Anomalous type 17 response to viral infection by CD8+ T cells lacking T-bet and eomesodermin . Science 2008 ; 321 : 408 – 411.
Rao RR, Li Q, Odunsi K, Shrikant PA . The mTOR kinase determines effector versus memory CD8+ T cell fate by regulating the expression of transcription factors T-bet and eomesodermin . Immunity 2010 ; 32 : 67 – 78.
Yeo CJ, Fearon DT . T-bet-mediated differentiation of the activated CD8+ T cell . Eur J Immunol 2011 ; 41 : 60 – 66.
Szabo SJ et al . A novel transcription factor, T-bet, directs Th1 lineage commitment . Cell 2000 ; 100 : 655 – 669.
Szabo SJ et al . Distinct effects of T-bet in TH1 lineage commitment and IFN-gamma production in CD4 and CD8 T cells . Science 2002 ; 295 : 338 – 342.
Rutishauser RL et al . Transcriptional repressor Blimp-1 promotes CD8(+) T cell terminal differentiation and represses the acquisition of central memory T cell properties . Immunity 2009 ; 31 : 296 – 308.
Shin H et al . A role for the transcriptional repressor Blimp-1 in CD8(+) T cell exhaustion during chronic viral infection . Immunity 2009 ; 31 : 309 – 320.
Kallies A, Xin A, Belz GT, Nutt SL . Blimp-1 transcription factor is required for the differentiation of effector CD8(+) T cells and memory responses . Immunity 2009 ; 31 : 283 – 295.
Pearce EL, Poffenberger MC, Chang CH, Jones RG . Fueling immunity: insights into metabolism and lymphocyte function . Science 2013 ; 342 : 1242454 .
Ciriacy M, Breitenbach I . Physiological effects of seven different blocks in glycolysis in Saccharomyces cerevisiae . J Bacteriol 1979 ; 139 : 152 – 160.
Fearon DT . The expansion and maintenance of antigen-selected CD8(+) T cell clones . Adv Immunol 2007 ; 96 : 103 – 139.
Crompton JG et al . Akt inhibition enhances expansion of potent tumor-specific lymphocytes with memory cell characteristics . Cancer Res 2015 ; 75 : 296 – 305.
Obar JJ, Lefrancois L . Memory CD8+ T cell differentiation . Ann N Y Acad Sci 2010 ; 1183 : 251 – 266.
Sukumar M et al . Inhibiting glycolytic metabolism enhances CD8+ T cell memory and antitumor function . J Clin Invest 2013 ; 123 : 4479 – 4488.
Youngblood B, Hale JS, Ahmed R . T-cell memory differentiation: insights from transcriptional signatures and epigenetics . Immunology 2013 ; 139 : 277 – 284.
Bannard O, Kraman M, Fearon D . Pathways of memory CD8+ T-cell development . Eur J Immunol 2009 ; 39 : 2083 – 2087.
Hawkins RD et al . Distinct epigenomic landscapes of pluripotent and lineage-committed human cells . Cell Stem Cell 2010 ; 6 : 479 – 491.
Zhu J et al . Genome-wide chromatin state transitions associated with developmental and environmental cues . Cell 2013 ; 152 : 642 – 654.
Fearon DT, Manders P, Wagner SD . Arrested differentiation, the self-renewing memory lymphocyte, and vaccination . Science 2001 ; 293 : 248 – 250.
Restifo NP, Gattinoni L . Lineage relationship of effector and memory T cells . Curr Opin Immunol 2013 ; 25 : 556 – 563.
Acknowledgements
The authors have no financial conflicts of interest. This research was supported by the Intramural Research Programs of the US National Institutes of Health, National Institute of Allergy and Infectious Diseases, and National Cancer Institute. This study was also funded by a generous gift from Mr. Li Jinyuan, Chairman of the Tiens Group. We thank Brian J. Abraham for useful inputs on histone modification data analysis. Finally, Joseph G. Crompton acknowledges funding support from the Wellcome Trust Translational Medicine and Therapeutics Programme.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors have no financial conflicts of interest.
Additional information
Supplementary Information accompanies the paper on Cellular & Molecular Immunology’s website (http://www.nature.com/cmi).
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Crompton, J., Narayanan, M., Cuddapah, S. et al. Lineage relationship of CD8+ T cell subsets is revealed by progressive changes in the epigenetic landscape. Cell Mol Immunol 13, 502–513 (2016). https://doi.org/10.1038/cmi.2015.32
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cmi.2015.32
This article is cited by
-
Distinct transcriptomic and epigenomic modalities underpin human memory T cell subsets and their activation potential
Communications Biology (2023)
-
Extracellular acidosis restricts one-carbon metabolism and preserves T cell stemness
Nature Metabolism (2023)
-
New genetic and epigenetic insights into the chemokine system: the latest discoveries aiding progression toward precision medicine
Cellular & Molecular Immunology (2023)
-
Glutarate regulates T cell metabolism and anti-tumour immunity
Nature Metabolism (2023)
-
Beta cell-specific CD8+ T cells maintain stem cell memory-associated epigenetic programs during type 1 diabetes
Nature Immunology (2020)