Abstract
The Set1 family of histone H3 lysine 4 (H3K4) methyltransferases is highly conserved from yeast to human. Here we show that the Set1 complex (Set1C) directly binds RNA in vitro through the regions that comprise the double RNA recognition motifs (dRRM) and N-SET domain within Set1 and its subunit Spp1. To investigate the functional relevance of RNA binding, we performed UV RNA crosslinking (CRAC) for Set1 and RNA polymerase II in parallel with ChIP-seq experiments. Set1 binds nascent transcripts through its dRRM. RNA binding is important to define the appropriate topology of Set1C distribution along transcription units and correlates with the efficient deposition of the H3K4me3 mark. In addition, we uncovered that Set1 binds to different classes of RNAs to levels that largely exceed the levels of binding to the general population of transcripts, suggesting the Set1 persists on these RNAs after transcription. This class includes RNAs derived from SET1, Ty1 retrotransposons, specific transcription factors genes and snRNAs (small nuclear RNAs). We propose that Set1 modulates adaptive responses, as exemplified by the post-transcriptional inhibition of Ty1 retrotransposition.
Similar content being viewed by others
Introduction
Highly conserved histone proteins undergo several types of covalent modifications including acetylation, methylation, phosphorylation, ubiquitylation, SUMOylation, citrullination and ADP-ribosylation [1]. These modifications that are deposited and removed by specific chromatin-modifying enzymes can either directly alter the chromatin architecture or create docking sites that facilitate the binding of specific domains present in chromatin readers [2]. These readers in turn recruit chromatin remodeling enzymes or additional chromatin modifiers to shape chromatin landscapes that regulate DNA accessibility [3]. Among these marks, methylation of lysine 4 on histone H3 (H3K4) has aroused considerable interest [4]. In mammals, this modification is catalyzed by at least six different complexes that differ by their catalytic SET domain subunit (Set1a, Set1b, Mll1, Mll2, Mll3 and Mll4) [5] but share a protein module comprises WDR5, RbBP5, ASH2L and DPY-30, which binds to the catalytic SET domain and stimulate H3K4 methyltransferase activity [6]. Each complex contains additional factors specifying their recruitment to chromatin and their biological effect [7].
In Saccharomyces cerevisiae, all H3K4 methylation is carried out by a complex called COMPASS (for complex of proteins associated with Set1) or Set1C (for Set1 Complex) [8]. The catalytic subunit Set1 acts as a scaffold for seven other components (Swd1 (mammalian homolog RbBP5), Swd2 (WDR82), Swd3 (WDR5), Bre2 (ASHL2), Sdc1 (DPY-30), Spp1 (CFP1) and Shg1 (BOD1)) [9]. Swd1, Swd3, Bre2 and Sdc1 associate with the SET domain of Set1 to form the SETc that is minimally sufficient to methylate free H3 in vitro [9], whereas Spp1 and Shg1 directly associate to Set1 by binding to the N-SET domain and the Set1 central region, respectively [10]. The loss of individual Set1C subunits differentially affects Set1 stability, complex integrity, the pattern of global H3K4 methylation and the distribution of H3K4 methylation marks along active genes [11]. The WD40 repeat protein Swd2 is the only essential subunit of Set1C and its depletion strongly affects Set1 stability and H3K4 methylation [9]. Swd2 also belongs to the APT complex (for ‘associated with Pta1’), which is part of the cleavage polyadenylation factor [12]. Several studies suggested a functional link between Set1C and 3′-end formation/termination [13] but it remains unknown how the binding of Swd2 to either of the two complexes (Set1C and APT complex) is regulated. Other regions outside of the SET domain have been reported to regulate Set1 catalytic activity, including the N-SET domain, the double RNA recognition motif (dRRM) and a centrally located auto-inhibitory domain, but the mechanism underlying such regulation is still elusive [14].
Genome-wide studies in yeast indicate that active transcription is characteristically accompanied by histone H3K4 trimethylation (H3K4me3) at the 5′-end of genes and by H3K4 di- and monomethylation (H3K4me2 and H3K4me1) at downstream nucleosomes [15]. H3K4me3 can also be found at the 3′-end of a number of genes most likely reflecting the presence of antisense ncRNAs (non-coding RNAs) [16]. These H3K4 methylation patterns correlate with Set1 occupancy that is higher at the 5′-end of coding regions of highly transcribed RNA polymerase II (RNAPII) genes and decreases at more distal nucleosomes [17]. Set1 has been reported to associate with the elongation complex in the early stages of the transcription cycle, which is thought to contribute to the prevalence of H3K4me3 at the 5′-end of active genes. Recruitment occurs when the carboxyl terminal domain of RNAPII is preferentially phosphorylated at the serine in the fifth position (Ser5) of its heptad repeats, which has been reported to depend on the Paf1 complex [18]. However, direct interactions that underpin the recruitment of Set1C to actively transcribed genes remain to be characterized. Although interaction of Set1C with chromatin was proposed to be mediated by the interaction of Swd2 with ubiquitylated H2B (H2Bub) [19], this model has been challenged by in vitro reconstitution experiments showing that the Swd2-deficient Set1C can methylate chromatinized H3K4 in an H2B ubiquitylation-dependent manner [20]. Thus, current models to explain Set1 recruitment and the establishment of H3K4 methylation along genes in vivo still need to be improved.
Set1 contains two tandem RRMs, RRM1 and RMM2 (dRRM) [14]. We previously reported that Set1 RRM1 contains the canonical RRM-fold but lacks some typical RNA-binding features. Consistently, RRM1 is necessary but not sufficient for Set1 to bind RNA in vitro and RRM2 was also shown to be required [21]. Deletion or mutation of RRM1 has been shown to lead to decreased H3K4me3 in the 5′ regions of active genes along with an increase in H3K4me2 [14], opening the possibility that a potential RNA-binding activity of Set1 could regulate Set1 occupancy and/or the distribution of H3K4 methylation [22].
Here, we show that Set1 binds directly RNA and that its dRRM and N-SET, as well as Spp1, contribute to Set1 RNA binding in vitro in the context of a reconstituted Set1C. By combining ChIP-seq and CRAC experiments of Set1 and Set1 mutants that have lost the ability to bind RNA, we show that Set1 RNA-binding activity mediated by its dRRM does not affect Set1 recruitment to chromatin per se but maintains Set1 in the 5′ region of genes. We propose that RNA binding to Set1 increases the time of residency of Set1C in the proximity of chromatin allowing additional time for H3K4 trimethylation in the 5′-end of genes. Our results also indicate that Set1 strongly associates, presumably post-transcriptionally to transcripts produced by specific classes of genes, including snRNAs small nuclear RNAs, Ty1 and adaptive response genes. In particular, we show that Ty1 retrotransposition is negatively regulated by Set1 at a post-transcriptional level.
Results
Binding of RNA in vitro by reconstituted Set1C involves the dRRM and N-SET domains of Set1
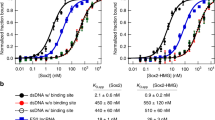
Our previous results suggested that purified Set1 RRM1-RRM2 (dRRM) binds RNA in vitro [21]. We reconstituted the whole Set1C [20] to analyze Set1 RNA binding in the context of the complex form of Set1 with its associated subunits. Set1C was purified from insect cells expressing FLAG-fused full-length or truncated Set1 together with the seven other subunits (Figure 1a and b). Purified complexes lacking specific domains of Set1 were incubated with in vitro transcribed and purified GAL1 and GAL10 mRNAs. Interaction was probed by electrophoretic mobility shift assay (Figure 1c and d). We found that reconstituted Set1C was able to directly bind GAL1 and GAL10 mRNAs in vitro. In agreement with our previous results, a truncated Set1 lacking the two RRM motifs (C569) was unable to bind RNA. Surprisingly, we found that the C762 fragment encompassing the N-SET and the SET domains (Figure 1a) was able to bind RNA in contrast to C569 and C938 that could not. These results suggest that N-SET contributes to Set1 RNA binding, which might be inhibited by the region between residues 569 and 762 (Figure 1c and d).
Purified Set1C binds directly to RNA in vitro. (a) A schematic diagram of Set1 and derived fragments with predicted RRM1, RRM2, N-SET and post-SET (hatched box) domains and associated subunits. FL indicates full-length. (b) Sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS–PAGE) and Coomassie blue staining of purified Set1Cs reconstituted with baculoviruses expressing FLAG-Set1 or FLAG-Set1 fragments and untagged subunits. FLAG-Set1 polypeptides are marked by asterisks. (c, d) Radiolabeled GAL1 (c) and GAL10 (d) transcripts were subjected to in vitro RNA electrophoretic mobility shift assays with 0.5 (lanes 2, 4, 6, 8, 10 and 12) or 2.5 (lanes 3, 5, 7, 9, 11 and 13) pmoles of indicated Set1Cs.
To further confirm these results, we introduced in the full-length Set1 the Y271F272/AA mutation previously shown to decrease the RNA-binding activity of dRRM in vitro [21], as well as a deletion of the dRRM, or the N-SET domain, and a combination of these mutations [21] (Figure 2a and b). Consistent with Figure 1, either the Y271F272/AA mutation or dRRM deletion strongly affected Set1C RNA-binding in vitro (Figure 2c and d). Deleting only the N-SET domain also significantly decreased RNA binding suggesting that the N-SET domain is important for RNA binding (Figure 2c and d). However, we did not detect interaction of the N-SET domain alone with RNA (data not shown) indicating that the N-SET domain is not sufficient to bind RNA. As expected, combining alterations of the dRRM with the N-SET deletion abolished Set1 RNA-binding activity in vitro. We next assessed whether Set1C subunits contribute by themselves to the RNA-binding activity of Set1C by monitoring the RNA-binding activity of each subunit. None of the Set1C subunits were found to bind GAL10 mRNA (Supplementary Figure S1A and B). Finally, as the N-SET domain binds Spp1, we asked whether only omitting Spp1 in Set1C reconstitution also affected RNA-binding activity. The results shown in Figure 2e indicated that Spp1 was required for Set1C to bind RNA despite the fact that Spp1 by itself does not bind RNA.
Set1C RNA binding requires dRRM, the N-SET domain, and Spp1. (a) A schematic representation of Set1, Set1YF/AA and Set1 deletion mutants with predicted RRM1, RRM2, N-SET and post-SET (hatched box) domains and associated subunits. (b) Sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS–PAGE) and Coomassie blue staining of purified Set1Cs reconstituted with baculoviruses expressing FLAG-Set1 or FLAG-Set1 fragments and untagged subunits. (c, d) Radiolabeled GAL1 (c) and GAL10 (d) transcripts were subjected to in vitro RNA electrophoretic mobility shift assays with indicated Set1Cs. (e) Binding of Set1C and Set1C lacking Spp1 to radiolabeled GAL10. Set1C containing Set1∆N-SET is also shown.
Collectively, these results show that the fully reconstituted Set1C has the ability to bind mRNAs in vitro. Unexpectedly, not only the dRRM motif but also the N-SET domain and Spp1 contributed to the ability of Set1 to bind RNA. Therefore, Set1C RNA binding requires the presence of multiple protein surfaces comprising the dRRM, as well as N-SET domain and Spp1.
Altering Set1 dRRM affects Set1 distribution along genes
Before addressing Set1 binding to RNA in vivo, we performed ChIP-seq analysis to determine the genome-wide occupancy of Set1 and Set1 mutants. ChIP-seq experiments were carried out from set1∆ cells expressing N-terminal tagged (Z-tag-Tev-6HIS) version of Set1 (PTH-Set1) [23]. Because in cells grown in SC-LEU-TRP medium, the amount of PTH-Set1 was similar to that of endogenous Set1 (Supplementary Figure S2A) we used these conditions for ChIP-seq and all subsequent experiments.
To address the importance of the Set1 domains involved in RNA-binding in vitro for chromatin binding, we performed ChIP-seq experiments of PTH-Set1 and of its mutant forms (set1YF/AA, set1∆dRRM, set1∆N-SET and set1∆dRRM,∆N-SET) with an anti-Set1 mouse monoclonal antibody (anti-Set1 mAb) [24] that recognizes a Set1 epitope between residues 700 and 761 (Supplementary Figure S2B and C). In parallel, we also performed ChIP-seq of a Myc-Set1 strain with an anti-Myc antibody (9E10). The occupancy profiles for PTH-Set1 and Myc-Set1 at selected genes were overall similar and both datasets were highly correlated (Supplementary Figure S3A, B and C). Set1 occupancy was maximum beyond the H3K4me3 peak and upstream of the H3K4me2 peak and was slightly 3′-shifted relative to RNAPII occupancy (Supplementary Figure S3D), in agreement with previous results indicating that Set1 and its subunits are recruited at the 5′ region of active genes transcribed by RNAPII [17].
Set1 protein amount was controlled in the different mutant strains by western blot (Figure 3a). We reproducibly observed a reduction of the PTH-Set1YF/AA amount by about 1.3–1.5-fold in all experiments [21], whereas deleting dRRM increased the stability of Set1. PTH-Set1 was functional, as it supported wild-type levels of H3K4me3 when expressed in a set1∆ background. In contrast, H3K4me3 was globally abolished when the dRRM and N-SET functions were compromised (Figure 3b). To evaluate the importance of the Set1 domains characterized in vitro, we next assessed the occupancy of PTH-Set1 and mutant forms by ChIP-seq experiments.
RRM but not N-SET regulates the genome-wide occupancy of Set1. (a, b) W303 set1∆::TRP1 pRS415-nHTP-SET1 (and SET1 mutant forms) cells were grown in SC -TRP-LEU. Protein levels of Set1 (a) and methylated histone H3 (b) were verified by western blots using anti-Set1 mAb and anti-H3K4me1, me2, and me3 antibodies, respectively. A Rap1 loading control is shown. (c) Average enrichment profiles of PTH-Set1 and PTH-Set1 mutants in genes>1500 bp. Read counts were normalized to read counts per million of mapped reads. ChIP-seq experiments were performed with the anti-Set1 mAb from set1∆::TRP1 cells expressing PTH-Set1 and PTH-Set1 mutants from the pRS415-nHTP (grown in SD -TRP -LEU). (d) Normalized occupancy profiles of Set1 at the indicated genes. Graphs were normalized to 10 million mapped reads for each mutant.
Surprisingly, deleting dRRM or mutating it in PTH-Set1YF/AA did not affect the distribution of the individual ChIP signals calculated per each mRNA-coding gene (Supplementary Figure S4 for PTH-Set1YF/AA, and data not shown). To analyze the chromatin distribution of Set1 and its mutant derivatives in more details, we generated normalized occupancy profiles over large genes, which allows a better spatial resolution of recruitment regions. As shown in Figure 3c, the aggregate signal for Set1YF/AA and Set1∆dRRM shows reduced occupancy in the 5′ region of genes relative to wild-type (WT) Set1, which is compensated by an average increase in the 3′-end of genes to generate the observed unchanged overall signal on a per gene basis. This trend is illustrated in Figure 3d for individual genes. Deletion of the N-SET domain had no detectable impact in the distribution of signals (data not shown) or the average profile of the signal (Figure 3c).
These results, together with the strong impact of the Set1YF/AA mutation on in vivo RNA binding described below, indicate that the dRRM domain of Set1 is not required for the overall recruitment of Set1 to chromatin, but is essential for its normal distribution along genes. Importantly, the N-SET domain that is essential for Set1C catalytic activity (Figure 3b) and is required to bind RNA in vitro, is not involved in the recruitment and positioning of Set1.
Set1 binding to RNA in vivo is determined by both co-transcriptional and post-transcriptional components
We next used the CRAC procedure to analyze in vivo the genome-wide RNA binding of Set1 and Set1YF/AA whose RNA-binding activity is compromised in vitro to detect in vivo RNA-protein interactions [25]. Briefly, tagged RNA-binding proteins are UV crosslinked to their targets in vivo and purified by three sequential steps of affinity selection, two of which are under denaturing conditions. The associated RNA is isolated and sequenced. We used the same PTH-Set1 and PTH-set1YF/AA constructs and growth conditions used for the ChIP-seq. A non-crosslinked sample was processed in parallel as a control for specificity. A spike in control was generated by adding to the S. cerevisiae cultures 0.5% of S. pombe cells expressing a non-relevant HTP-tagged protein that binds RNA and that was purified with S. cerevisiae Set1. The number of reads mapping to the S. pombe genome was used for normalization. We also monitored RNAPII distribution by the same CRAC technique, which provides high-resolution information about the level of transcription.
We identified 2543 mRNAs displaying high-confidence Set1 RNA crosslinking sites for which the number of reads obtained for the crosslinked sample (Set1 CL) was >5-fold over the no-crosslinked sample (Set1 No-CL) (Supplementary Table S1). Set1 was found to bind mRNAs but also several other transcript classes (Supplementary Figure S5). Set1 binds at least partially during transcription, as witnessed by the significant representation of intronic RNAs in the crosslinked material (see below). For assessing to what extent Set1 binds the RNA during transcription, we sought correlations between the Set1 CRAC signal and the levels of RNAPII occupancy as determined by RNAPII CRAC (Figure 4a) for all mRNA-coding genes. We also compared the Set1 CRAC signal with Set1 recruitment to chromatin as determined by ChIP-Seq (Figure 4b). We reasoned that if binding to the RNA were co-transcriptional, theses datasets should be highly correlated. Consistent with this notion, binding of Set1 to the RNA correlated remarkably well both with RNAPII CRAC (Figure 4a, r2=0.68; P=3E-22) and recruitment of Set1 to chromatin as measured by ChIP (Figure 4b, r2=0.37; P=9E-10).
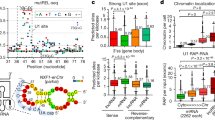
Set1 binding to RNA occurs via two different modes and requires its dRRM. (a) Dispersion plot showing high correlation between the Set1 and RNAPII CRAC signals. The determination coefficient (r2) of the linear, least squares regression is indicated. (b) Dispersion plot of Set1 CRAC signals versus Set1 ChIP-Seq (IP/input). (c) Distribution of the Set1 CRAC/Set1 ChIP ratios for all mRNAs coding genes. The average, mode and median are indicated to highlight the asymmetry of the distribution. The shaded area corresponds to the percentage of features whose Set1 CRAC/ChIP ratios are included in a range symmetrically positioned around the mode. (d) Distribution of the Set1/RNAPII CRAC ratios in cells expressing PTH-Set1 and PTH-Set1YF/AA.
The distribution of Set1 CRAC/ChIP ratios was clearly not symmetric as could have been expected for a homogeneous population with random variability (Figure 4c). Rather, it was markedly skewed toward high values (compare the difference between the mode, the median and the average in Figure 4c) with only 63% of the population symmetrically distributed around the mode (shaded area) and the remaining values tailing over a wide range of higher ratios. This suggests the existence of at least two classes of genes: one major, for which the levels of RNA binding relative to chromatin-associated Set1 are relatively homogeneous; the second displaying levels of Set1 binding to RNA that are generally higher than expected based on the sole co-transcriptional interaction. Overall, these analyses strongly suggest that the levels of Set1 binding to the RNA detected by CRAC are generally dominated by a co-transcriptional component but also contain a post-transcriptional component that might predominate for some genes (Supplementary Table S1). Snapshots of the second class of mRNA are shown in Supplementary Figure S6A and B. Interestingly, the feature with the highest Set1 crosslinking signal was the SET1 mRNA, which is fully consistent with the notion that it associates with the Set1C containing Set1, Swd1, Spp1 and Shg1 during its co-translational assembly [26]. We further sought to determine whether genes encoding this specific class of transcripts (Supplementary Table S1) have physical and/or functional associations. Evidence that many of the proteins encoded by these genes are linked in reliable networks stemmed from computational analysis [27]. Gene ontology analysis revealed that genes whose transcripts were strongly bound by Set1 included some involved in chromosome segregation and many transcription factors (DNA-binding proteins) involved in adaptive responses (Supplementary Figure S6C).
Mutation of the RNA-binding domain in Set1YF/AA led to a marked decrease in the Set1 CRAC signal, which affected uniformly the whole population of Set1 targets as indicated by a general shift in the distribution of Set1/RNAPII CRAC ratios in the set1YF/AA mutant relative to WT (Figure 4d). These data demonstrate that the YF/AA mutation in Set1 dramatically affect the interaction of Set1 with mRNAs in vivo, although this interaction was not totally abolished but partially maintained with a different topology (see below). Set1 was found to bind with similar set1YF/AA dependency stable unannotated transcripts, cryptic unstable transcripts and sno/snRNAs (Figure 5a–c). The first two classes interact with Set1 to a somewhat lower extent even when normalized to the RNAPII CRAC signal, maybe because these RNAs are unstable and the post-transcriptional component might contribute to less to the CRAC signal. Many sno- and snRNAs appear to be bound by Set1 post-transcriptionally, as suggested by the large distribution of Set1/RNAPII CRAC values (Figure 5c) and indicated by the general lack of signal in the regions of the precursor (Figure 5d for the U1, U2 and U4 snRNAs and data not shown). Interestingly, spliceosomal snRNAs were among the strongest binders (Figure 5d), possibly suggesting a role of Set1 in splicing.
Set1 binds to different classes of RNA. (a, b, c) Normalized distribution of Set1/RNAPII CRAC ratios in cells expressing PTH-Set1 and PTH-Set1YF/AA for the indicated classes of RNA. The position in the distribution of each RNAPII-transcribed snRNAs is indicated in C. The distribution of ratios for mRNAs (Figure 4d) is shown in each graph. (d) Snapshots for Set1, Set1YF/AA and PolI CRAC normalized signals for the indicated spliceosomal RNAs.
Aggregated distribution of Set1 binding on RNAs
We profiled the distribution of RNA-associated Set1 for different features aligned on the transcription start site. As co-transcriptional RNA binding at any given position is likely to be strongly dependent on the level of transcription, we plotted in parallel RNAPII occupancy as defined by the CRAC signal. As shown in Figure 6a, binding of Set1 to the RNA was slightly delayed relative to the appearance of the RNAPII signal (see inset in Figure 6a), which resulted in a relative Set1/RNAPII signal building up in the first 100–250 nt of transcription. This is consistent with the notion that Set1 is recruited co-transcriptionally to the RNA and suggests that it binds the nascent transcript after interacting directly with the polymerase. In the set1YF/AA mutant, the CRAC signal was markedly reduced over most of the length of the transcription unit, particularly in the 5′ region.
Distribution of Set1-binding sites on RNA. (a, b) Metagene analysis of Set1 and Set1YF/AA RNA-binding signals on mRNAs as observed by CRAC compared with polymerase occupancy (a) features aligned on transcription start site; (b) features aligned on wPAS. Insets contain zooms of relevant regions. (c) Snapshots for Set1, Set1YF/AA, and PolI CRAC normalized signals at the 5′ region of BAP2 and RPL17-A mRNAs. (d, e) Metagene analysis of Set1 and Set1YF/AA RNA-binding signals, (d) on intronless and intron-containing genes, the distribution of intron position is indicated as well as the average position of introns; (e) to cryptic unstable transcripts (CUTs) compared with size matched open reading frame (ORF).
To assess the distribution of the Set1 signal in the 3′ region of genes we first calculated a positionally weighted average p(A) site (wPAS) for every gene (see Materials and Methods section). This was necessary to improve the quality of the alignment in the 3′-end of genes as most genes have multiple polyadenylation sites. To this end, we used T-fill data [28] and assigned to every p(A) addition site a weight depending on the intensity of the signal at that position. This was used to generate a positionally weighted average p(A) site (wPAS). As shown in Figure 6b, the polymerase signal declines in this region, either because of multiple sites of termination or to increased speed. Interestingly, the Set1 signal is maintained and actually slightly increases immediately before the wPAS. Intriguingly, this increase is maintained in the YF/AA mutant, to the point that the signals for the mutant and WT Set1 are identical in this region. This surprising observation indicates that the YF/AA mutation does not affect the binding to the RNA in this region of the transcripts. Whether this 3′ peak is mainly because of the co-transcriptional or post-transcriptional binding of Set1 to the RNA cannot be determined from these experiments. Note that if the 3′ peak were formed co-transcriptionally, its intensity relative to the polymerase signal, which decreases in this region (Figure 6b), would be higher than in other regions of the RNA (see Discussion). After the wPAS, both the WT and mutant Set1 signals decrease steadily (Figure 6b, inset), indicating that they are significantly above background within the range of the transcription unit. The presence of a Set1 3′ peak can be readily observed at individual genes, most prominently in the mutant for which the signal before the peak is generally lower (Figure 6c; see also snapshots for the SET1, SLK19, and SWI1 loci in Supplementary Figure S6), indicating that this behavior is not limited to a small set of genes. These data are compatible both with increased co-transcriptional recruitment of Set1 in the region of termination and with a post-transcriptional binding to the mRNA in the immediate vicinity of the poly(A) site. Importantly and surprisingly, in both cases the interaction with the RNA is not dependent on dRRM.
Prompted by the strong binding of Set1 to snRNAs, we assessed the profile of Set1 binding to intron-containing RNAs by comparing it with size matched mRNA-coding genes. As shown in Figure 6d, Set1 bound intronic transcripts with similar or even better efficiency than non-intronic RNAs, causing a slight downstream shift of the 5′-peak of Set1 binding. Binding of the Set1YF/AA was similarly affected at intron-containing genes, as well as to the general population. We also assessed binding to cryptic unstable transcripts, a class of transcripts that are unstable in WT yeast because they are rapidly degraded in the nucleus [29], which we compared with matched size small open reading frames. Set1 binding to these features was lower than at small open reading frames, even when normalization to RNAPII was applied (Figure 6e) to account for the generally different levels of transcription. This could be due either to specificities residing in the sequence of the cryptic unstable transcript, or to nuclear degradation of these RNAs.
Set1 represses Ty1 retrotransposition post-transcriptionally
Among the mRNAs that were strongly bound by Set1, presumably post-transcriptionally, we also found Ty1 retrotransposon (Figure 7a). Binding of Set1 to Ty1 mRNA was not affected by the YF/AA mutation suggesting that Set1 binding to Ty mRNA does not involve its dRRM (Figure 7a). The Set1YF/AA mutation had no major effect on steady-state Ty1 mRNA levels (Figure 7b) as previously reported for the set1∆ mutant [30,31]. This indicates that Set1 binding does not affect Ty1 mRNA expression or stability. To assess whether Set1 affects Ty1 retrotransposition, we performed a typical retrotransposition assay based on a Ty1 element marked with a his3AI reporter gene on a plasmid, which confers His+ prototrophy to cells upon retrotransposition (Figure 7c). In the absence of Set1, the frequency of Ty1 retrotransposition significantly increased (Figure 7d), whereas no change in Ty1HIS3 mRNA levels was observed (Figure 7e). This indicates that Set1 can repress Ty1 mobility at a post-transcriptional stage. In contrast, Set1-YF/AA, which retains the ability to bind Ty1 mRNAs, repressed Ty1 retrotransposition as efficiently as WT Set1. These results suggest that Set1 binding to Ty1 mRNA could impair Ty1 mRNA export, translation or encapsidation, all essential steps to Ty1 retrotransposition efficiency. Of note, the less than twofold decrease in Ty1-his3AI mRNA levels observed in the set1-YF/AA mutant may not affect Ty1 retrotransposition (Figure 7e), as much more Ty1 mRNAs are produced than effective transposition events occurring in cells [32]. However, we cannot exclude that the slight defect in Ty1-his3AI mRNA levels may mask a slight increase in Ty1 cDNA integration that could be facilitated by the modification of the histone methylation status of the yeast genome in the set1-YF/AA mutant.
Set1 represses Ty1 retrotransposition post-transcriptionally. (a) Example of the Set1 and Set1YF-AA binding on Ty1-1 mRNA. The Set1 CRAC and ChIP profiles are shown. (b) Global expression of endogenous Ty1 retrotransposons in SET1, set1Δ or set1-YF/AA yeast cells, monitored by quantifying Ty1 mRNA levels by RT-qPCR (normalized to 25 S rRNA values; mean±s.d.; n=3). (c) Ty1 retrotransposition assay from a plasmid expressing a Ty1 element tagged with the his3AI reporter gene [33]. An intron is inserted in the HIS3 gene in an antisense orientation in a spliceable orientation in the Ty1 transcript resulting in a Ty1 complementary DNA (cDNA) bearing a functional HIS3 gene. The cDNA can then be integrated into the host genome. Cells that sustain a Ty1-HIS3 retrotransposition event give rise to His+ colonies [32]. The position of the qPCR amplicons used to amplify all the Ty1 mRNAs in (b) and the Ty1 reporter mRNA specifically expressed from the plasmid in (e) are indicated (Ty1 and Ty1-HIS3 amplicons, respectively). (d) Frequency of Ty1-his3AI retrotransposition in SET1, set1Δ or set1-YF/AA yeast cells (number of His+ prototrophs divided by the total number of cells; mean±s.d.; n=3). *p≤0.05 (Welch’s t-test). (e) Plasmid Ty1-his3AI expression in SET1, set1Δ or set1-YF/AA yeast cells monitored by quantifying Ty1-HIS3 mRNA levels by RT-qPCR (normalized to 25S rRNA values; mean±s.d.; n=3).
Reduced H3K4me3 levels are due to defective recruitment or positioning of Set1 during transcription
Although we showed that the Set1YF/AA mutation only marginally affects the recruitment of Set1 to chromatin on a genome-wide scale, at the gene level a variegated range of cases exists. In some instances, a strong RNA-binding defect translates into a marginal effect on recruitment (for example, MOT3, Figure 8), in other cases (for example, PMA1 and ENO1) recruitment to chromatin is affected in spite of a moderate effect on in vivo crosslinking to the RNA as revealed by the CRAC signal. Although it is unclear why in these latter particular cases, the Set1YF/AA mutation affects Set1 occupancy, we exploited these individual differences to address the role of the nascent RNA and Set1 recruitment in H3K4 methylation. As shown in Figure 8b, in all these three cases H3K4me3 was found to be strongly reduced, indicating that neither the recruitment to chromatin (MOT3) nor the crosslinking to the RNA alone (PMA1 and ENO1, Figure 8) are sufficient to promote methylation.
Reduction of Set1 occupancy and RNA-binding activity in Set1YF/AA correlates with reduced H3K4me3 levels. (a) ChIP-seq and CRAC signals of PTH-Set1 (Set1) and PTH-Set1YF/AA (Set1 YF/AA) at representative genes. ChIP-seq bedGraphs were generated by normalization to 10 million mapped reads for each sample. CRAC signals were normalized as described above. (b) H3K4me3, me2, me1 occupancies at the indicated genes in W303 set1∆::TRP1 pRS415-nHTP-SET1 and SET1YF/AA strains. Levels of H3K4me3, me2 and me1 are normalized to total H3. Positions of the primers used for the ChIP-qPCR for each representative gene are indicated. Errors bars represent the s.d. from three independent experiments. (c) Recombinant chromatin template containing fully ubiquitylated H2B (H2Bub) was subjected to in vitro HMT assays with purified Set1C and Set1C YF/AA in the absence and presence of purified GAL10 RNA [0.1 (lanes 2 and 7), 0.2 (lanes 3 and 8), 0.5 (lanes 4 and 9) and 2 (lanes 5 and 10) molar ratio to Set1C. H3 methylated status was monitored by western blots with indicated antibodies.
The general strong decrease in methylation when dRRM is mutated might be due to the defective positioning of the protein along transcription units, to an allosteric requirement for RNA interaction or to a general inactivation of the methylation function of Set1 by the Set1-YF/AA mutation. To distinguish between these possibilities, we analyzed in vitro the histone methyltransferase (HMT) activity of the Set1C containing the Set1YF/AA mutation in the presence or absence of RNA. As shown in Figure 8c, Set1C YF/AA reproducibly displayed a higher HMT activity compared with WT on a recombinant chromatin template containing ubiquitylated H2B. This indicates that the Set1YF/AA not only retained full HMT activity but its in vitro activity was even enhanced. Addition of purified GAL1 RNA did not improve the activity of Set1C, and actually inhibited its function in a concentration-dependent manner. As expected, it had no effect when added to Set1C YF/AA (Figure 8c). This strongly suggests that the interaction with RNA is not required for activating the HMT function of Set1, and is consistent with the possibility that RNA might negatively regulate its activity.
Together, these results strongly suggest that binding to the RNA is important to define the appropriate topology of Set1C distribution along transcription units, which is important for the deposition of the H3K4me3 mark.
Discussion
In this study, we first showed with in vitro studies that the interaction between Set1C and RNA is direct. Both the dRRM and N-SET domains of Set1 contribute to Set1 RNA-binding activity in vitro in the context of the reconstituted complex. Although structural data indicated that dRRM has the canonical structure to bind RNA [14], binding to RNA of the C762 fragment alone was unexpected because none of the C762 constituents (N-SET, SET and post-SET domains and Spp1, Bre2, Sdc1, Swd3 and Swd1) has a canonical RNA-binding motif. In addition, no direct interaction of Set1C subunits and N-SET domain alone (data not shown) with RNA was observed, strongly suggesting that interaction with RNA is mediated by a composite surface potentially involving all or some components associated with the C762 fragment. Interestingly, addition of residues 569–762 to the C762 fragment, which are known to have an inhibitory effect on Set1 methyltransferase activity [10] inhibited the RNA-binding activity of the C762 fragment, suggesting that Set1 binding to RNA could regulate the methyltranferase activity of Set1.
The involvement of the N-SET domain in RNA binding is of particular interest as this domain acts as central regulatory region of Set1C by its ability to bind Spp1 [34] and Swd1 [20]. The N-SET domain also mediates an Spp1-dependent interaction with the SET domain and its associated subunits, an interaction that likely regulates Set1 methyltransferase activity [35]. Other studies have indicated a cross-talk between Swd2 and Spp1 suggesting a complex regulation mediated by Set1C subunits for interaction between the C- and the N-terminal regions of Set1C [21]. In this work, we found that omitting Spp1 in the reconstitution assay strongly decreases Set1C RNA-binding activity. Therefore, in the context of the full-length Set1, it is possible that dRRM, Spp1 and the N-SET cooperate to stabilize interaction with RNA. In vivo, inhibition of the C762 RNA-binding activity by the region of Set1 encompassing amino acids 569–762 remains an open question.
We performed ChIP-seq experiments with the same Set1 mutants studied in vitro using an anti-Set1 mAb [24]. We provide a high-resolution map of Set1 occupancy indicating that the peak of Set1 occupancy is shifted 3′ to the peak of H3K4me3 and slightly shifted with respect to RNAPII average occupancy. Analysis of ChIP-seq signals revealed that set1YF/AA mutation did not affect Set1 recruitment to chromatin per se but rather regulated Set1C distribution by maintaining Set1 in the 5′ region of genes. In addition, we show that the N-SET domain that is required to bind RNA in vitro is not involved in recruitment of Set1 to chromatin, whereas it is essential for Set1C catalytic activity. Assessing the functional importance of the RNA-binding activity of the N-SET remains a challenge for future studies.
To assess whether Set1 binds RNA in vivo, we performed CRAC experiments using Set1 and Set1YF/AA. Our high-resolution and strand-specific CRAC analysis shows that Set1 binds to RNA in vivo thereby extending and providing a physiological facet to our in vitro analysis with the reconstituted complex. CRAC analysis is expected to detect binding to the RNA both during and after transcription. In our experiments, the occurrence of co-transcriptional binding is demonstrated by the observation that Set1 binds to intronic regions and is also strongly suggested by the highly significant correlation with Set1 and RNAPII occupancy (as determined by ChIP and CRAC, respectively) for most features. Although it is formally possible that the levels of transcription and/or Set1 chromatin occupancy also impact to some extent the post-transcriptional binding to the RNA, we favor the hypothesis that a co-transcriptional component dominates in directing Set1 binding to the RNA for the largest fraction of the population. Binding of Set1 to the nascent transcript is not sequence specific and occurs preferentially at the 5′-end of RNAs, with a peak that is slightly shifted downstream relative to the maximum of RNAPII occupancy as detected by CRAC.
Our results indicate that the substitution of residues Y271 and F272, which are part of the hydrophobic core of RRM1 and predicted to be important for maintaining the structure of whole dRRM [36], strongly decreased Set1 RNA binding, particularly in the 5′ region of RNAs in vivo. This result combined with our ChIP-seq data supports the notion that Set1 is recruited via protein–protein interactions and subsequently contacts the nascent RNA 5′-region via its dRRM. Transfer of Set1C to the nascent RNA once the latter emerges from the elongation complex would contribute to position Set1 predominantly to the 5′ regions of genes.
The set1YF/AA mutation markedly affects the deposition of the H3K4me3 mark at the 5′-end of genes for all genes tested consistent with the fact that global H3K4me3 is strongly reduced in the set1YF/AA mutant. Importantly, the set1YF/AA mutation in the context of reconstituted Set1C enhanced the methyltransferase activity of Set1C assayed on recombinant chromatin containing ubiquitylated H2B.
There might be several mechanisms by which the set1YF/AA mutation affects H3K4me3. Interaction with the nascent RNA might be important to increase the persistence of Set1C in the proximity of chromatin to allow additional time for H3K4 trimethylation in the 5′-end of genes, whereas transcription proceeds at its normal speed. It is possible that binding to the RNA activates allosterically the HMT activity of the protein. However, we showed that in vitro addition of the RNA does not activate Set1 but actually inhibits H3K4 methylation, in a manner that depends on the RNA-binding activity of Set1. It is not clear whether this inhibition detected in vitro is physiologically relevant, but the possibility exists that binding to the RNA could regulate Set1 activity in some phases of the H3 methylation process. The simultaneous or sequential interaction of Set1C with the polymerase and the nascent RNA might constitute a quality control strategy to ensure deposition of the H3K4m3 mark only to regions of active transcription. Whether RNA binding to N-SET could contribute to such a process remains to be determined.
Metagene analysis experiments revealed that Set1 was also crosslinked to mRNA at the 3′-end of the molecules, showing a 3′ peak immediately before the poly(A) addition site. Importantly, formation of this 3′ RNA-binding peak was fully insensitive to the set1YF/AA mutation suggesting that the binding of Set1 at poly(A) sites is independent of its dRRM. Interestingly, although the level of this peak was low relative to the levels of Set1 at the 5′-end of genes, its intensity relative to the polymerase signal (that is low in this region) is the highest over the whole transcription unit, suggesting that it might have functional significance.
The dRRM-independent formation of the 3′ Set1 peak may occur via a 3′ recruitment of Set1 by Swd2, an essential subunit of Set1C that also belongs to the APT complex, a subcomplex of the cleavage and polyadenylation factor that is involved in mRNA and snoRNA 3'-end formation [36,37]. These observations might suggest a functional link between Set1C and 3′-end formation/termination but it remains unknown how the binding of Swd2 to either of the two complexes (Set1C and APT) is regulated. Swd2 was shown to directly interact with the N-terminus of Set1 [20]. Consistent with this, Swd2 recruitment to the 5′ region of genes is reduced when SET1 is deleted [38] but whether Swd2 contributes to recruit Set1C at the vicinity of the poly(A) site to signal cleavage and polyadenylation remains to be determined.
We also observed a striking enrichment of Set1 to snRNA, suggesting that Set1 could fulfill a function in signaling splicing events. Consistent with this notion we observed that Set1 is enriched within introns, even when the signal is evaluated relative to the RNA polymerase (that also increases in introns).
Interestingly, cryptic unstable transcripts were under-represented when compared with matched size small mRNAs, opening the possibility that at early stages of ncRNA (non-coding RNA) transcription, dRRM may compete with Nab3 RRM (of the Nrd1 complex) for the recognition of sequences in the nascent transcript. This might explain the more efficient termination of ncRNA observed in cells lacking Set1 [39,40].
Finally, we show that Set1 binds to a class of transcripts to an extent that cannot be justified by a co-transcriptional component alone, or, at least, not by the same co-transcriptional component that holds for the majority of the population. We therefore suggest that Set1 can bind RNAs after transcription or that binding occurs during transcription but additionally persists in this class of genes. The very high level of Set1 bound to SET1 mRNA was previously uncovered [26] and probably reflects the co-translational assembly mode of Set1C. In this model, SET1 mRNA is bound by the nascent Set1 protein that emerges from the ribosome through indirect interactions with the translation machinery.
Set1 also binds post-transcriptionally to other transcripts. Interestingly, many of these factors are transcription factors some of which are functionally related. For instance, Mot3 and Rox1, which are transcriptional repressors of genes encoding cell wall proteins [36] and of hypoxic genes [37] are functionally linked with Msn2-Msn4 in the osmostress response [36] and with Sok2 in the ergosterol biosynthetic pathway [41]. Interestingly, Set1 was previously described to be required for the expression of genes in the ergosterol biosynthetic pathway [42]. Strikingly, Set1 binds also to mRNA of several genes functionally related to chromosome segregation, in line with recent results linking Set1 to mitotic spindle assembly [43,44]. However, the steady-state levels of these transcripts were not affected by deletion of SET1, at least in rich medium (YPD) [39], suggesting that Set1 might affect the expression of these genes at levels that do not involve mRNA synthesis or degradation rates, or that the impact of Set1 is revealed only under defined growth or stress conditions.
Our results also show that Set1 binds to Ty1 mRNA and repress Ty1 mobility at a post-transcriptional stage. Our data uncover a new function of Set1C as a repressor of Ty1 mobility and add another layer of regulation by Set1 to previous studies showing that Set1 had a synergistic role with the histone H4 metyhyltransferase Set5 in repressing transcription of Ty transposable elements [23]. Although Ty1 retrotransposition can alter yeast genome integrity and is consequently a highly controlled process, release of Ty1 repression is supposed to contribute to genome evolution and cell adaptation to stress [45]. Therefore, it would be interesting to determine whether Set1C repression could be alleviated under stress conditions that are known to stimulate Ty1 retrotransposition [46,47].
Deciphering the role of the post-transcriptional binding of Set1 to RNAs will reveal unexpected function of Set1C that might explain the incredibly complex genetic interaction map of Set1 [48].
Materials and methods
Purification of recombinant Set1C and subunits, in vitro RNA electrophoretic mobility shift assay, and in vitro HMT assay
Preparation of FLAG-tagged recombinant Set1C and subunits were as described [20]. For radiolabeled RNA probe preparation, DNA duplexes containing T7 promoter sequence ( 5′-TAATACGACTCACTATAGGG-3′) followed by 100-nucleotide sequence encoding GAL1 or GAL10 mRNA starting from the transcription start site were generated by PCR. GAL1 and GAL10 RNA were transcribed by T7 RNA polymerase according to the manufacturer’s instructions (Promega, Fitchburg, WI, USA) and then purified by gel elution method. After removing 5′ phosphate by Antarctic phosphatase, the 5′-end of RNA was radiolabeled by T4 polynucleotide kinase using [γ-32P]ATP and then purified by Sephadex column (iNtRon, Seongnam, South Korea) and gel elution method. For in vitro RNA electrophoretic mobility shift assay, reactions containing purified Set1C or individual subunits and 0.25 pmole of radiolabeled RNA in 20 μl reaction buffer (10 mM Tris-Cl (pH 7.5), 1 mM EDTA, 5 mM MgCl2, 50 mM K-glutamate, 5% glycerol, 1 mM DTT and two units of RNasin) were incubated at room temperature for 30 min. The samples were resolved by electrophoresis at 4 °C on 5% polyacrylamide gels in 1×TBE buffer and subjected to autoradiography. For in vitro HMT assay, 40 μl reactions containing 350 ng (histone amount) recombinant chromatin assembled as described in Kim et al. (2013) with H2Bub-containing histone octamer [49], purified Set1C and 100 μM S-adenosylmethionine were incubated at 30 °C for 2 h. Proteins were resolved by sodium dodecyl sulfate–polyacrylamide gel electrophoresis and subjected to western blots.
Strains, constructs and growth conditions
For reconstitution of Set1Cs, SET1, set1 mutants and Set1C subunits genes were subcloned in pFASTBAC1 with or without a FLAG tag [20]. Baculoviruses were generated according to the manufacturer’s instruction (Gibco-Invitrogen, Waltham, MA, USA). Sf9 cells were infected with combinations of baculoviruses and proteins/complexes were affinity purified on M2 agarose (Sigma, St Louis, MO, USA) as described [20].
Yeast strains and primers used in this study are described in Supplementary Tables S2 and S3, respectively (see Supplementary Information). Full-length SET1 and SET1 mutants were cloned into in pRS415-nHTP [23]. Expression of the resulting constructs (Z-tag—TEV cleavage site—His6—SET1) is under the control of MET25 promoter. The pRS415-nHTP-SET1 (or SET1 mutants) were transformed into W303 set1∆::TRP1 strain. Plasmid pRS415-nHTP-SET1 complements all the tested set1∆-associated phenotypes of the set1∆::TRP1 strain. For ChIP-seq and CRAC experiments, W303 set1∆::TRP1 pRS415-nHTP-SET1 (or SET1 mutants) cells were grown in SC-TRP-LEU. RNAPII CRAC experiments were performed from W303 cells expressing Rpb1-HTP and grown in SC-TRP. Construction of fully functional chromosomally encoded Myc9-tagged Set1 is described in Dehe et al. [10].
CRAC analyses
Cells in exponential phase were crosslinked with a Megatron for 100 s (Set1) and 50 s (Rpb1), harvested by centrifugation, resuspended in 2.4 volume/g of cells of TN150 buffer (50 mM Tris pH 7.8, 150 mM NaCl, 0.1% NP-40 and 5 mM beta mercaptoethanol) supplemented with protease inhibitors (complete, Mini, EDTA-free Protease Inhibitor Cocktail). This suspension was flash frozen in droplets and cells were mechanically broken using the Mixer Mill MM 400 by doing five cycles of 3 min at 20 Hz. A non-crosslinked sample was treated in parallel as a background control.
Powders were thawed and the resulting extracts were treated for one hour at 25 °C with DNase I (165 U/g of cells) to solubilize chromatin and then clarified by centrifugation for 20′ at 20 000 g at 4 °C. Subsequent purifications steps were performed essentially as described with minor modifications from Granneman et al. [50]. For both nPTH-Set1 and Rpb1-HTP strains, adaptors were modified in order to sequence RNA molecules from the 3′-end.
The RNA was recovered after proteinase K treatment and reverse transcribed using specific primers. The resulting complementary DNA was used to perform multiple PCR reactions in a final volume of 25 μl using the following conditions: 0.4 μM of each primers 0.2 mM dNTP, 2.5 U LA Taq DNA polymerase from Takara, 1X LA PCR Buffer II and 2 μl of complementary DNA per reaction with the programme: 2′ at 95 °C, (30′′ at 95 °C, 45′′ at 58 °C, 1′ at 72 °C) × 13 cycles, 5′ at 72 °C. PCR were pooled and treated with 200 U of Exonuclease I (NEB) per milliliter of PCR reaction for 1 h at 37 °C. After Exonuclease I inactivation for 20′ at 80 °C, DNA was purified on PCR clean up columns (NucleoSpin Gel and PCR Clean-up, Macherey-Nagel, Düren, Germany) and sequenced using Illumina technology (San Diego, CA, USA). Primers are indicated in Supplementary Table S2.
Samples were demultiplexed using the pyBarcodeFilter script from the pyCRAC utility suite. Subsequently, the 3′ adaptor is clipped with Cutadapt and the resulting insert is quality trimmed from the 3′-end using Trimmomatic rolling mean clipping (window size=5, minimum quality=25). At this stage, the pyCRAC script pyFastqDuplicateRemover is used to collapse PCR duplicates and ensure each insert is represented only once. Each unique insert in our library is associated with a six-nucleotides random tag within the 5′ adaptor. The resulting sequences are reverse complemented with Fastx_reverse_complement (part of the fastx toolkit [51]), and mapped to the R64 genome (sgd) with bowtie2 (-N 1 –f).
Read counts were normalized relative to reads derived from an S. pombe spike that was added to S. cerevisiae cells before the crosslinking step. The S. pombe spike cells contain a non-relevant protein tagged with the same HTP tag that was co-purified with the S. cerevisiae material. The positionally weighted average poly(A) addition site (wPAS) for every gene was calculated by weighting the position of each poly(A) site using its intensity and calculating an average position.
ChIP-seq, data processing and ChIP-qPCR
ChIP of Myc-Set1 and PTH-Set1 were performed as previously described [52] with 9E10 (anti-MYC, Santa Cruz Biotechnology, Dallas, TX, USA) and anti-Set1 monoclonal antibodies (P Nagy, University of Toronto, Toronto, Canada). Libraries were prepared from fragmented DNA using the Chip-seq MicroPlex Library Preparation Kit v2 samples preparation (Diagenode, Seraing, Belgium) according to the manufacturer’s instructions. In all, 2 ng from IP samples were used as the starting material. Each library was barcoded using MicroPlex Single Index (Diagenode): iPCRtagT5, T6, T7 and T8 and amplified for 10 and 6 cycles for IP and input samples, respectively. Each library was quantified on Qubit with Qubit dsDNA HS Assay Kit (Life Technologies, Carlsbad, CA, USA) and then, size distribution was examined on the Bioanalyser with High Sensitivity DNA chip (Agilent, Santa Clara, CA, USA), to ensure that the samples have the proper size, no adaptor contamination and to estimate sample molarity. Each library was diluted to 4 nM and then pulled together at equimolar ratio. Libraries were denatured according to the manufacturer’s instruction and sequenced on a mid-output flow cell (130 M clusters) using the NextSeq 500/550 High Output v2 150 cycles kit (Illumina), in paired-end 75/7 nt mode, according to the manufacturer’s instructions. In all, 148 million (M) paired-end reads were generated (34–39 M per sample) with 93% >= Q30.
ChIP-Seq data quality was assessed using FastQC. FasQC: a quality control tool for high-throughput sequence data. Available online at: http://www.bioinformatics.babraham.ac.uk/projects/fastqc. Sequencing reads (FastQ format) were mapped to the Saccharomyces cerevisiae genome (sacCer3) using BFAST alignment tool with default parameters [53] (PMID 19907642) to obtain a Binary Alignment Mapped (BAM) file. The sorted BAM files were used to determine average profiles of ChIP-Seq read density using ngs.plot software [54], (PMID 24735413) around the transcription start site. Read counts were normalized to the total number of million uniquely mapped reads or to read count per million of mapped reads (RPM). The RPM values allow samples to be compared regardless of differences in sequencing depth. To generate BedGraphs for visualization on genome browsers, ChIP-Seq BAM files were processed using HOMER package. The tag directory for each sample was then created using the makeTagDirectory tool and the corresponding BedGraph was generated using makeUCSCfile tool with default options. Only uniquely mappable reads (non-secondary alignment) were considered to create BedGraphs with a normalization to 10 million mapped reads for each sample. To compare Set1 binding positions on RNA with Set1 occupancy on genes of interest, the Multicov command from Bedtools [55] was used to obtain read counts within each gene.
For ChIP-qPCR, samples were prepared as previously described [52]. DNA was analyzed by real-time qPCR using SYBR Green Premix Ex Taq (Takara, Mountain View, CA, USA) in a Rotor Gene 6000 (Corbett Research, Labgene, Archamps, France). Primers are listed in Supplementary Table S2. The following antibodies were used: anti-H3 (Abcam1791, Cambridge, UK), anti-H3K4me2 (Abcam-ab7766, Cambridge, UK), anti-H3K4me3 (Abcam-ab8580), anti-Myc 9E10 (Santa Cruz Biotechnology-sc-40) and anti-Rap1 (V. Géli’s laboratory, Marseille, France).
Data access
ChIP-seq data sets (PTH-Set1, PTH-Set1YF/AA, PTH-Set1∆dRRM, PTH-Set1∆NSET, PTH-Set1∆RRM ∆NSET, PTH-vector, Input PTH-Set1 and input PTH-Set1YF/AA), as well as CRAC sequences generated during this work are deposited to the NCBI Gene Expression Omnibus (GEO; http://www.ncbi.nlm.nih.gov/geo/) under the accession number GSE104486 (GSE104484 for chiPseq datasets and GSE104485 for CRAC datasets).
Ty1 transposition assays
The pOY1 URA3, centromeric vector carrying a Ty1-his3AI reporter element expressed from its own promoter was previously described [33]. Total RNAs were extracted from yeast cultures after 4 h or 8 h at 20 °C, for Ty1 mRNAs or Ty1-his3AI mRNAs respectively, using the Nucleospin RNA II kit (Macherey-Nagel) and reverse transcribed with Superscript-II reverse transcriptase (Invitrogen, Waltham, MA, USA). Complementary DNA quantification was achieved by real-time PCR with a LightCycler 480 system (Roche, Basel, Switzerland) using SYBR Green incorporation according to the manufacturer’s instructions. The amounts of the mRNAs of interest were normalized relative to 25S ribosomal RNA values. Primers used are described in Supplementary Table S2.
To estimate the frequency of Ty1his3AI mobility [32], overnight liquid cultures were grown at 30 °C from an individual clone in HC medium (Hartwell’s synthetic complete) [56], lacking uracil and supplemented with 2% glucose. Each culture was diluted to OD 0.01 in HC-URA medium and grown to saturation at 20 °C, which is permissive for Ty1 transposition. In all, 3 ml of each culture were plated on two HC agar plates lacking histidine. Cell titer was determined by plating 10 000-fold diluted cultures on YEPD rich medium. Plates were incubated for 3 days at 30 °C and counted to determine the fraction of [HIS+] prototrophs. Ty1 retrotransposition frequencies were defined as the mean of 3 experiments, each one performed with four independent clones.
References
Rothbart SB, Strahl BD . Interpreting the language of histone and DNA modifications. BBA Gene Regul Mech 2014; 1839: 627–643.
Suganuma T, Workman JL . Histone modification as a reflection of metabolism. Cell Cycle 2016; 15: 481–482.
Tessarz P, Kouzarides T . Histone core modifications regulating nucleosome structure and dynamics. Nat Rev Mol Cell Biol 2014; 15: 703–708.
Shilatifard A . Chromatin modifications by methylation and ubiquitination: implications in the regulation of gene expression. Annu Rev Biochem 2006; 75: 243–269.
Ruthenburg AJ, Allis CD, Wysocka J . Methylation of lysine 4 on histone H3: intricacy of writing and reading a single epigenetic mark. Mol Cell 2007; 25: 15–30.
Ernst P, Vakoc CR . WRAD: enabler of the SET1-family of H3K4 methyltransferases. Brief Funct Genomics 2012; 11: 217–226.
Smith E, Shilatifard A . The chromatin signaling pathway: diverse mechanisms of recruitment of histone-modifying enzymes and varied biological outcomes. Mol Cell 2010; 40: 689–701.
Briggs SD, Bryk M, Strahl BD et al. Histone H3 lysine 4 methylation is mediated by Set1 and required for cell growth and rDNA silencing in Saccharomyces cerevisiae. Genes Dev 2001; 15: 3286–3295.
Roguev A, Schaft D, Shevchenko A et al. The Saccharomyces cerevisiae Set1 complex includes an Ash2 homologue and methylates histone 3 lysine 4. EMBO J 2001; 20: 7137–7148.
Dehe PM, Dichtl B, Schaft D et al. Protein interactions within the Set1 complex and their roles in the regulation of histone 3 lysine 4 methylation. J Biol Chem 2006; 281: 35404–35412.
Dehé P-M, Géli V . The multiple faces of Set1This paper is one of a selection of papers published in this Special Issue, entitled 27th International West Coast Chromatin and Chromosome Conference, and has undergone the Journal's usual peer review process. Biochem Cell Biol 2006; 84: 536–548.
Dichtl B, Blank D, Sadowski M, Hübner W, Weiser S, Keller W . Yhh1p/Cft1p directly links poly(A) site recognition and RNA polymerase II transcription termination. EMBO J 2002; 21: 4125–4135.
Terzi N, Churchman LS, Vasiljeva L, Weissman J, Buratowski S . H3K4 trimethylation by Set1 promotes efficient termination by the Nrd1-Nab3-Sen1 pathway. Mol Cell Biol 2011; 31: 3569–3583.
Schlichter A, Cairns BR . Histone trimethylation by Set1 is coordinated by the RRM, autoinhibitory, and catalytic domains. EMBO J 2005; 24: 1222–1231.
Pokholok DK, Harbison CT, Levine S et al. Genome-wide map of nucleosome acetylation and methylation in yeast. Cell 2005; 122: 517–527.
Kirmizis A, Santos-Rosa H, Penkett CJ et al. Arginine methylation at histone H3R2 controls deposition of H3K4 trimethylation. Nature 2007; 449: 928–932.
Ng HH, Robert F, Young RA, Struhl K . Targeted recruitment of Set1 histone methylase by elongating Pol II provides a localized mark and memory of recent transcriptional activity. Mol Cell 2003; 11: 709–719.
Mueller CL, Jaehning JA . Ctr9, Rtf1, and Leo1 are components of the Paf1/RNA polymerase II complex. Mol Cell Biol 2002; 22: 1971–1980.
Lee J-S, Shukla A, Schneider J et al. Histone crosstalk between H2B monoubiquitination and H3 methylation mediated by COMPASS. Cell 2007; 131: 1084–1096.
Kim J, Kim J-A, McGinty RK et al. The n-SET domain of Set1 regulates H2B ubiquitylation-dependent H3K4 methylation. Mol Cell 2013; 49: 1121–1133.
Trésaugues L, Dehé P-M, Guérois R et al. Structural characterization of Set1 RNA recognition motifs and their role in histone H3 lysine 4 methylation. J Mol Biol 2006; 359: 1170–1181.
Hampsey M, Reinberg D . Tails of intrigue: phosphorylation of RNA polymerase II mediates histone methylation. Cell 2003; 113: 429–432.
Martín GM, King DA, Green EM et al. Set5 and Set1 cooperate to repress gene expression at telomeres and retrotransposons. Epigenetics 2014; 9: 513–522.
Nagy PL, Griesenbeck J, Kornberg RD, Cleary ML . A trithorax-group complex purified from Saccharomyces cerevisiae is required for methylation of histone H3. Proc Natl Acad Sci USA 2002; 99: 90–94.
Bohnsack MT, Tollervey D, Granneman S . Identification of RNA helicase target sites by UV cross-linking and analysis of cDNA. Meth Enzymol 2012; 511: 275–288.
Halbach A, Zhang H, Wengi A et al. Cotranslational assembly of the yeast SET1C histone methyltransferase complex. EMBO J 2009; 28: 2959–2970.
Szklarczyk D, Franceschini A, Kuhn M et al. The STRING database in 2011: functional interaction networks of proteins, globally integrated and scored. Nucleic Acids Res 2011; 39: D561–D568.
Wyers F, Rougemaille M, Badis G et al. Cryptic pol II transcripts are degraded by a nuclear quality control pathway involving a new poly(A) polymerase. Cell 2005; 121: 725–737.
Wilkening S, Pelechano V, Järvelin AI et al. An efficient method for genome-wide polyadenylation site mapping and RNA quantification. Nucleic Acids Rese 2013; 41: e65–e65.
Berretta J, Pinskaya M, Morillon A . A cryptic unstable transcript mediates transcriptional trans-silencing of the Ty1 retrotransposon in S. cerevisiae. Genes Dev 2008; 22: 615–626.
Martín GM, King DA, Garcia-Nieto PE, Morrison AJ . Transcriptome profiling of Set5 and Set1 methyltransferases: tools for visualization of gene expression. GDATA 2014; 2: 216–218.
Curcio MJ, Garfinkel DJ . Regulation of retrotransposition in Saccharomyces cerevisiae. Mol Microbiol 1991; 5: 1823–1829.
Lee BS, Lichtenstein CP, Faiola B et al. Posttranslational inhibition of Ty1 retrotransposition by nucleotide excision repair/transcription factor TFIIH subunits Ssl2p and Rad3p. Genetics 1998; 148: 1743–1761.
Mersman DP, Du H-N, Fingerman IM, South PF, Briggs SD . Charge-based interaction conserved within histone H3 lysine 4 (H3K4) methyltransferase complexes is needed for protein stability, histone methylation, and gene expression. J Biol Chem 2012; 287: 2652–2665.
Vitaliano-Prunier A, Menant A, Hobeika M, Géli V, Gwizdek C, Dargemont C . Ubiquitylation of the COMPASS component Swd2 links H2B ubiquitylation to H3K4 trimethylation. Nat Cell Biol 2008; 10: 1365–1371.
Martínez-Montañés F, Rienzo A, Poveda-Huertes D, Pascual-Ahuir A, Proft M . Activator and repressor functions of the Mot3 transcription factor in the osmostress response of Saccharomyces cerevisiae. Eukaryotic Cell 2013; 12: 636–647.
Klinkenberg LG, Mennella TA, Luetkenhaus K, Zitomer RS . Combinatorial repression of the hypoxic genes of Saccharomyces cerevisiae by DNA binding proteins Rox1 and Mot3. Eukaryotic Cell 2005; 4: 649–660.
Soares LM, Buratowski S . Yeast Swd2 is essential because of antagonism between Set1 histone methyltransferase complex and APT (associated with Pta1) termination factor. J Biol Chem 2012; 287: 15219–15231.
Margaritis T, Oreal V, Brabers N et al. Two distinct repressive mechanisms for histone 3 lysine 4 methylation through promoting 3'-end antisense transcription. Madhani HD, ed. PLoS Genet 2012; 8: e1002952.
Castelnuovo M, Zaugg JB, Guffanti E et al. Role of histone modifications and early termination in pervasive transcription and antisense-mediated gene silencing in yeast. Nucleic Acids Res 2014; 42: 4348–4362.
Hongay C, Jia N, Bard M, Winston F . Mot3 is a transcriptional repressor of ergosterol biosynthetic genes and is required for normal vacuolar function in Saccharomyces cerevisiae. EMBO J 2002; 21: 4114–4124.
South PF, Harmeyer KM, Serratore ND, Briggs SD . H3K4 methyltransferase Set1 is involved in maintenance of ergosterol homeostasis and resistance to Brefeldin A. Proc Natl Acad Sci USA 2013; 110: E1016–E1025.
Schibler A, Koutelou E, Tomida J et al. Histone H3K4 methylation regulates deactivation of the spindle assembly checkpoint through direct binding of Mad2. Genes Dev 2016; 30: 1187–1197.
Beilharz TH, Harrison PF, Miles DM et al. Coordination of cell cycle progression and mitotic spindle assembly involves histone H3 lysine 4 methylation by Set1/COMPASS. Genetics 2017; 205: 185–199.
Curcio MJ, Lutz S, Lesage P . The Ty1 LTR-retrotransposon of budding yeast, Saccharomyces cerevisiae. Microbiol Spectr 2015; 3: 1–35.
Morillon A, Springer M, Lesage P . Activation of the Kss1 invasive-filamentous growth pathway induces Ty1 transcription and retrotransposition in Saccharomyces cerevisiae. Mol Cell Biol 2000; 20: 5766–5776.
Todeschini A-L, Morillon A, Springer M, Lesage P . Severe adenine starvation activates Ty1 transcription and retrotransposition in Saccharomyces cerevisiae. Mol Cell Biol 2005; 25: 7459–7472.
Costanzo M, VanderSluis B, Koch EN et al. A global genetic interaction network maps a wiring diagram of cellular function. Science 2016; 353: aaf1420–aaf1420.
McGinty RK, Köhn M, Chatterjee C, Chiang KP, Pratt MR, Muir TW . Structure-activity analysis of semisynthetic nucleosomes: mechanistic insights into the stimulation of Dot1L by ubiquitylated histone H2B. ACS Chem Biol 2009; 4: 958–968.
Granneman S, Kudla G, Petfalski E, Tollervey D . Identification of protein binding sites on U3 snoRNA and pre-rRNA by UV cross-linking and high-throughput analysis of cDNAs. Proc Natl Acad Sci USA 2009; 106: 9613–9618.
Weiner A, Hsieh T-HS, Appleboim A et al. High-resolution chromatin dynamics during a yeast stress response. Mol Cell 2015; 58: 371–386.
Ostrow AZ, Viggiani CJ, Aparicio JG, Aparicio OM ChIP-Seq to analyze the binding of replication proteins to chromatinIn: Vengrova S, Dalgaard J (eds). DNA Replication Vol 1300 Methods in Molecular Biology New York, NY, USA: Springer New York. 2015, 155–168.
Homer N, Merriman B, Nelson SF . BFAST: an alignment tool for large scale genome resequencing. Creighton C, ed. PLoS ONE 2009; 4: e7767.
Shen L, Shao N, Liu X, Nestler E . ngs.plot: quick mining and visualization of next-generation sequencing data by integrating genomic databases. BMC Genomics 2014; 15: 284.
Quinlan AR, Hall IM . BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 2010; 26: 841–842.
Servant G, Pennetier C, Lesage P . Remodeling yeast gene transcription by activating the Ty1 long terminal repeat retrotransposon under severe adenine deficiency. Mol Cell Biol 2008; 28: 5543–5554.
Acknowledgements
We deeply thank Peter Nagy and Bichtien Rouse for providing the Set1 mAb and Markus Bohnsack and Khaterine Sloan for pRS415-nHTP. We thank Luis Soares and Steve Buratowski for Set1 constructs, Tom Muir for ubiquiytlated H2B, and Nir Friedman and Alon Appleboim for communicating H3K4 Exo-ChIP profiles. We thank Julie Drogat, Ghislain Bidaud and Nicolas Fernandez for their involvement in the ChIP-seq procedures and Lorant Székvögyi for discussions. We also thank Judith Recht for critical reading of the manuscript. Work in VG laboratory is supported by the ‘Ligue contre le Cancer’ (Equipe Labéllisée). Bioinformatic support and computing sources at CRCM were provided by the CIBI and DISC platforms (CRCM Integrative Bioinformatics and Datacentre IT and Scientific Computing). High-throughput sequencing was performed at the TGML Platform, supported by grants from Inserm, GIS IBiSA, Aix Marseille Université, and ANR-10-INBS-0009-10. AE is supported by a fellowship from the Canceropole PACA. Work in the DL laboratory was supported by the CNRS, the ANR (‘TermCut’ grant) and the FRM. TC and DC are supported by French MRT fellowships. Work in the JK laboratory was supported by grants from the National Research Foundation of Korea (2012M3A9C6049938, 2015K1A3A1A21000296 and 2015R1A1A1A05001593). Collaboration between VG and JK laboratories is supported by the PHC STAR exchange programme (34325UF).
Author information
Authors and Affiliations
Contributions
JJ performed all the in vitro reconstitution experiments. PL and AE-k performed all the ChIP-seq and ChIP-qPCR experiments and analysis. DC, MB and TC performed the CRAC experiments and analyzed the data. FJ constructed and characterized strains. AB and PL performed Ty1 retrotransposition experiments. VG, JK and DL analyzed the data, wrote the manuscript and directed the work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
( Supplementary information is linked to the online version of the paper on the Cell Discovery website.)
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Luciano, P., Jeon, J., El-kaoutari, A. et al. Binding to RNA regulates Set1 function. Cell Discov 3, 17040 (2017). https://doi.org/10.1038/celldisc.2017.40
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/celldisc.2017.40
This article is cited by
-
The COMPASS subunit Spp1 protects nascent DNA at the Tus/Ter replication fork barrier by limiting DNA availability to nucleases
Nature Communications (2023)
-
Diverse and dynamic forms of gene regulation by the S. cerevisiae histone methyltransferase Set1
Current Genetics (2023)
-
Roles of RNA-binding proteins in neurological disorders, COVID-19, and cancer
Human Cell (2022)
-
The Set1 N-terminal domain and Swd2 interact with RNA polymerase II CTD to recruit COMPASS
Nature Communications (2020)