Abstract
Bats are natural reservoir hosts and sources of infection of several microorganisms, many of which cause severe human diseases. Because of contact between bats and other animals, including humans, the possibility exists for additional interspecies transmissions and resulting disease outbreaks. The purpose of this article is to supply an overview on the main pathogens isolated from bats that have the potential to cause disease in humans.
Similar content being viewed by others
Facts
-
Bats are an important reservoirs of different pathogenic agents, and many of them have already caused disease outbreaks worldwide.
-
More than 200 viruses have been associated with bats, and almost all are RNA viruses probably owing to their great ability to adapt to changing environmental conditions through a higher genetic variability.
-
Bacteria in bats and their putative threat to humans remain poorly studied.
Open questions
-
The understanding of the mechanisms of viral persistence in bats remains unclear.
-
Extensive studies are needed to improve our understanding of bat–human interactions in order to design new control measures in the future.
-
Strategies on surveillance and monitoring of disease outbreaks in bat populations need to be further developed, in particular where bats and humans are in close contact.
Bats, mammals of the Chiroptera order, are present all over the world with the exception of the Arctic, the Antarctic and a few oceanic islands. Bats are the only mammals with the ability to fly and are present in >1100 different species.1 Bats are essential members of the global ecosystem and humans benefit from their presence in many ways. They are involved in seed dispersal and pollination activity: tropical bats are vital – as an example – in rebuilding cut down forests and in the pollination of wild plants as bananas, avocados and dates. Furthermore, the flying mammals are the major predators of night insects, including crop and human pests.1 Finally, their guano, which is rich in nitrogen, is used as biological fertilizer.
Despite the multiple benefits attributed to these animals, since the ancient times – through myths and misapprehensions – bats have gained a bad reputation in the general public. The classical literature is full of examples in which bats are associated with evil and darkness. The roman poet Ovid narrates in the Metamorphoses that the god Bacchus transforms the daughters of Mineus, king of Boeotia, into bats as a punishment for profanating his celebration (Figure 1 and Box A). In the Divine Comedy (Inferno – Canto XXXIV), Dante Alighieri, the father of the italian language, describes the devil Lucifer as bearing large bat wings (Figure 2 and Box B). In the eighteenth century, scientists called ‘vampire’ a bat that fed on blood, giving rise to the myth of human vampires that sucked blood from other men and could transform into bats. The Irish novelist Bram Stoker with his novel, Dracula (1897) did nothing but make this belief famous worldwide. More recently, Bob Kane, an American comic book artist, ideated a character called Batman a positive character that, however, disguises in a bat costume to scare his enemies: ‘Criminals are a superstitious cowardly lot. So my disguise must be able to strike terror into their hearts. I must be a creature of the night, black, terrible..... ‘Just then a huge bat flies in the open window’. A bat! That’s it! It’s an omen. I shall become a bat! In: Detective Comics no 33, 1939.
Bats, however, can be involuntarily dangerous to humans. Indeed, they are natural reservoir hosts and sources of infection for several microorganisms, including pathogens that can cause severe human diseases, and are more frequently implicated in zoonotic virus emergencies.1,2 Bats are widespread in urban areas and come in close contact with both domestic animals and humans, contaminating houses with guano and urine, additionally, humans occasionally encroach into bat habitats.3 Their characteristic ecology undoubtedly influences the maintenance and transmission of microorganisms within the colony and directly or indirectly to humans.1,3
Microbial transmission within bat colonies is promoted by the behavior of several species of these mammals aggregating in crowded roosts. Bats can transmit infectious agents to humans through intermediate hosts, which are in close contact with humans.4 These intermediate hosts can be infected in many ways, including ingestion of food partially digested by bats. Frugivorous bats, in fact, cannot ingest wide amounts of food because of the aerodynamics of flight,5 therefore, they extract nutrients by chewing fruits and spitting the residues. This partially digested food dropped on the ground can then be ingested by other animals and is a potential infectious source. A similar modality of viral transmission has also been described for insectivorous bats.5 Bats can also directly infect humans.4 This can occur through ingestion of infected bat meat, as in some areas bats are a food source, or through bat’s bite as in the case of rabies virus.
Other features of bat’s life make them a good host for infectious agents. The fact that several species of bats hibernate during the winter is one of these. Although the role of the hibernation in infection dynamics has not been widely studied, it is likely that this condition can contribute to the maintenance of pathogens in the cold weather (see below: white-nose syndrome (WNS)). Moreover, unlike many other small mammals, bats live >30 years. Their long life makes them a great reservoir for pathogens and gives them many occasions to transmit them to other species. In addition, some species of bats migrate – also for distances >1000 km – allowing them to spread diseases in big areas and acquire new microorganisms.6,7
Finally, most pathogens are not dangerous for bats and can therefore survive for long time in the host without killing it. Indeed, despite the fact that bats are infected with more different zoonotic viruses per host – with the exception of rabies virus and other Lyssaviruses – these are apparently not pathogenic for them, suggesting that they may control viral replication more efficiently than other mammals. It has been hypothesized that there may be a relationship between flight and low virulence.8 During flight, bats show an increase in the metabolic rate and in body temperature, comparable to the fever response rendering replication of infectious agents, which are temperature sensitive, less favorable.
In this review, we give a comprehensive overview of the microorganisms and viruses isolated from bats and their possible role in human disease.
Viruses
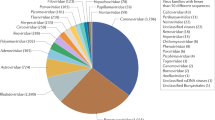
Bats are recognized as important reservoirs of different families of viruses, most of which are emerging as human pathogens, such as Ebola and Marburg viruses, Severe Acute Respiratory Syndrome (SARS) and Middle East Respiratory Syndrome (MERS) coronaviruses. More than 200 viruses have been associated with bats, and almost all are RNA viruses probably owing to their great ability to adapt to changing environmental conditions through a higher genetic variability.3,9 In fact, RNA viruses have higher mutation rates compared with DNA viruses as the viral RNA polymerases lack proofreading activity. Furthermore, RNA viruses with segmented genomes have the ability to modify their genome through genetic re-assortment (i.e., Orthomyxoviruses). Below we report some examples of human infectious diseases associated with bat viruses.
Rhabdoviridae
Rhabdoviridae contain six genera, including Lyssavirus, the most important bat-associated virus. At least 14 species of the Lyssavirus genus can be detected in bats, which are considered the ancestral hosts for these viruses. Lyssaviruses can be found worldwide and can be classified using different criteria, such as genetic distance, antigenic patterns, geographical distribution and host range.10,11 The characteristic bullet-shaped virus, transmitted to humans through the bite of infected animals, causes an acute, and frequently fatal, encephalitic disease.
The first report of a transmission of a viral disease from bats to humans was in 1911 and related to the rabies virus (RABV) belonging to the Lyssavirus genus.12 Carini12 suggested a link between rabies infection and hematophagous bats, known as vampires, in Central and South America. Several years later, rabies was also detected in non-hematophagous bat species.13 Although RABV is found worldwide in several terrestrial hosts, its presence in bats is observed only in the Americas. In Europe, four different Lyssaviruses have been isolated from bats: European Bat Lyssavirus type 1 (EBLV-1) and European Bat Lyssavirus type 2 (EBLV-2), Bokeloh Bat Lyssavirus (BBLV) and West Caucasian Bat Virus (WCBV).14 Recently, a new putative Lyssavirus in bat, named Lleida Bat Lyssavirus (LLBV), was found in Spain.15 To date, no human exposure to LLBV has been reported. EBLV-1, with the sub-types EBLV-1a and EBLV-1b, is the most isolated type throughout Europe. In addition, spillover infections by EBLV-1 in other mammals were also observed.13,14 The type 2 of EBLV is believed to be less virulent than type 113 and is found less frequently being present only in few countries and human contamination has been reported only in two cases.14 Two other members of this family are found in bats but significantly less frequently than the previous ones: BBLV isolated in Germany and France13,16,17; WCBV isolated once in the Caucasus Mountains but also detected in Kenya in seropositive bats, suggesting a greater geographical distribution.14,18 Australian Bat Lyssavirus (ABLV) is the first endemic lyssavirus identified in Australia and is phylogenetically related to RABV and EBLV1.10,19 ABLV has been identified in all flying fox species on Australia’s land mass. Three fatal human infections by ABLV have been reported. Additionally, other viruses of this family detected in bats are summarized in Table 1.
Paramyxoviridae
Paramyxoviridae constitute a wide viral family that includes human and animal pathogens. Several bat-borne paramyxoviruses have been recognized such as parainfluenza type 2 virus, Mapuera, Menangle and Tioman viruses and two infectious agents of emerging diseases, such as Nipah and Hendra viruses.20 Nipah and Hendra viruses, classified as the genus Henipavirus, are capable of causing severe, potentially fatal diseases in humans.20 Fruit bats of the Pteropus genus are the common reservoir hosts of the Nipah and Hendra viruses.20
Nipah virus (NiV) first emerged in 1998 in Malaysia, causing an outbreak of respiratory illness and encephalitis in pigs.21 Pig-to-human transmission of Nipah virus – associated with severe febrile encephalitis – was described and it was thought to occur through close contact with infected animals. Although uncommon, human-to-human transmission of virus was also described.21 In two other outbreaks in Bangladesh and India, an intermediate animal host was not identified, suggesting bat-to-human and human-to-human transmissions.
Hendra virus (HeV) causes a fatal respiratory disease in both humans and horses.20,22 Several outbreaks of HeV have occurred in Australia. Horse is the intermediate host and the virus is likely transmitted via ingestion of feed, pasture or water contaminated with urine, saliva and feces of infected bats. Horse-to-human transmission occurs when there is close contact with ill animals.20 To date, human-to-human transmission has not been observed.
Coronaviridae
Coronaviruses (CoVs) prior to the SARS outbreak were only known to be the second cause of the common cold after rhinoviruses. At least four different species can cause mild, self-limiting upper respiratory tract infections in humans: alphacoronaviruses HCoV-229E and HCoV-NL63, and betacoronaviruses HCoV-HKU1 and HCoV-OC43. More recently, two more additional pathogenic human-CoV were identified: Severe Acute Respiratory Syndrome Coronavirus (SARS-CoV) and Middle East Respiratory Syndrome Coronavirus (MERS-CoV).23 SARS-CoV was first identified in China in February 2003, and 4 months later, >8000 cases had been reported with about 800 deaths in 27 different countries worldwide.24 SARS-CoV has a wide host range and it is associated with wildlife meat industry. The natural history of the virus involves bats as primary hosts that then transmitted it to the intermediate amplifying hosts – as mask palm civets and raccoon dogs – that then could spread it to humans.23,25 Human-to-human transmission follows and can lead to large numbers of infected patients and is considered the main route of transmission in large-scale epidemics.9
MERS-CoV is phylogenetically related to SARS-CoV and share with SARS-CoV the origin in bats.23,26,27 Several CoVs have been identified in insectivorous and frugivorous bat species in various countries, indicating that bats may represent an important reservoir of these viruses.23 MERS-CoV was first identified in Saudi Arabia in 2012 and then spread to other countries causing hundreds of deaths.26,28 Clinical features of MERS-CoV are similar to SARS-CoV, although this virus has also been associated with several extrapulmonary manifestations, such as severe renal complications. Recent studies have indicated that dromedary camels may be the intermediate hosts and potential source of the virus for humans.26,29 In addition, the first experimental infection of bats with MERS-CoV has been described. The virus maintains the ability to replicate in the host without clinical signs of disease, supporting the general hypothesis that bats are the ancestral reservoir for MERS-CoV.30 Human-to-human transmission has also been reported. Based on epidemiological data, both animal-to-human and human-to-human transmission are considered to be important elements in MERS outbreak.26
Filoviridae
Ebolavirus and Marburgvirus are two genera of the family Filoviridae, responsible for severe, often fatal, hemorrhagic fever diseases in humans and other primates.31 The first report on the Marburgvirus first was in 196732 in German laboratory workers in Marburg who contacted it from African monkeys imported from Uganda. In 1976, a virus with similar characteristics but immunologically distinct was isolated in the Northern Democratic Republic of Congo and it was named Ebolavirus.32 Both viruses have generated several epidemics during the past years.31 Recently, in 2014, the largest ever registered Ebola epidemic started in West Africa and has affected several countries with >10 000 confirmed cases and thousands of deaths (Source CDC Atlanta, USA: 2014 Ebola outbreak in West Africa, updated 22 September 2015). The natural reservoirs for Marburgvirus and Ebolavirus are both fruit and insectivorous bat species, indicating that these filoviruses are multihost parasites.31,33 The virus is transmitted to humans through contact with body fluids – mainly blood and feces – and dead bodies of infected bats. Other animals such as monkeys and apes can also develop the disease and in turn transmit it to humans. Epidemics usually are a consequence of human-to-human transmission of the virus (Figure 3).
A third filovirus species of the new Cuevavirus genus, named Lloviu virus, was recently detected in insectivorous bats in Spain.34 Lloviu virus, genetically distinct from the others, is the first filovirus detected in Europe that was not imported from Africa. Unlike the other two species, this virus may be virulent in bats.34 As this virus has not been yet isolated, its capacity to infect other mammalian cells or to cause disease in humans remains to be determined.
Orthomyxoviridae
Orthomyxoviridae are enveloped segmented RNA viruses that include five genera of which influenza A virus is the most preponderant pathogen in humans. It causes respiratory tract infections, resulting in moderate-to-severe disease and occasionally death. Influenza A viruses are divided into subtypes on the basis of two surface glycoproteins, namely, hemagglutinin (H) and neuraminidase (N). Influenza A virus is an uncommon promiscuous virus with a wide host range, including humans, pigs and birds. Recently, two new subtypes evolutionarily distinct from all others – H17N10 and H18N11 – were detected in different fruit bat species in Central and South America.35,36 It has been observed that, although H17N10 subtype is phylogenetically separate from all other subtypes, the virus genome is compatible with genetic exchange with human influenza A viruses, suggesting a potential re-assortment capability between subtypes and the consequent ability to generate highly pathogenic hybrid forms.35 More recently, serological evidence of influenza A virus subtypes other than H17N10 and H18N11 were reported in African frugivorous bats.37 In particular, about 30% antibody detection rate was found against avian H9 subtype known to cause infections in humans worldwide.38 These data, albeit preliminary, suggest that bats might represent asymptomatic mammalian carriers of influenza A viruses.37 Thus, similar to other pathogens, bats may represent a considerable reservoir for these viruses.
Bunyaviridae
Hantavirus genus (from Hantan river in South Korea) is constituted of several emerging segmented RNA viruses that can cause human infections, including severe and lethal diseases such as hemorrhagic fever with renal syndrome and hantavirus cardiopulmonary syndrome.39,40 Rodents have long been believed to be the primary reservoirs of hantaviruses; however, a wider range of mammalian hosts, including insectivorous bats, has been reported.39,40 The evolutionary history of this genus is characterized by relatively frequent cross-species transmission that is also considered a major force in its evolution. The first hantavirus isolated from bats was the Hantaan virus, the etiological agent of hemorrhagic fever with renal syndrome.41 Successively, hantaviruses were identified in other bat species, but to date, however, no bat-to-human transmission of hantaviruses has been observed.39
Reoviridae
Mammalian orthoreovirus of the genus Orthoreovirus can cause mild respiratory or gastrointestinal illness to severe diseases, including encephalitis and diarrhea. The virus is present in different serotypes throughout the world and has been isolated from several mammals, including humans.42 Mammalian orthoreoviruses were also isolated in several bat species, suggesting an extensive distribution of the virus in these animals.42,43,44 Several evidence suggest that bats may act as the natural reservoir of these viruses.42 Although bat-origin orthoreoviruses have been isolated from human patients, the zoonotic potential of these viruses is still unclear.43,44,45
Other viruses
Several other mammalian viruses have been detected in bats for which the zoonotic potential or host range is unclear.9,46–48 An example are Poxviruses – important infectious agents of both humans and animals and capable of infecting multiple host species and to induce cross-species infections that were also recently identified in bats.49 Another example is the Dengue virus, an arthropod-borne virus belonging to the Flavivirus genus (Flaviviridae) that includes several relevant human pathogens associated with encephalitis and hemorrhagic fevers. Despite the fact that Flaviviridae are the second most frequent viruses found in bats and Dengue virus has been described in several bat species worldwide, the role of these animals in the dynamics of viral spreading remain insufficiently understood.50
Bacteria
Unlike viruses, bacteria in bats and their putative threat to humans remain poorly studied.51 Here we report some examples of bacteria responsible for common human and animal infections that have occasionally been detected in bats.
Bartonella spp.
Bartonellosis is a globally emerging zoonotic bacterial disease.52 Bartonella sp. is a Gram-negative bacterium transmitted through the bite of hematophagous arthropod vectors. Several species have been identified in domestic and wild animals, including bats.53–55 Recently, two species of Bartonella – B. mayotimonensis and B. naantaliensis – were detected from both the peripheral blood of bats and in their ectoparasites, suggesting that bats might be a source of the human bacterial pathogens.54 More recently, it has been reported the presence of closely related Bartonella genotypes in fruit bats and their associated bat flies in Madagascar, suggesting the transmission of a potentially zoonotic pathogen by bat fly vectors.56
Pasteurella spp.
Pasteurella is commonly spread among animals as part of the normal microbiota of the oral, nasopharyngeal and upper respiratory tract.57 This genus comprises opportunistic pathogen species that can cause endemic disease and are associated with epizootic outbreaks.57 Animal bites and nasal secretions are the most likely sources of transmission to humans. In bats, various Pasteurella species – mainly P. multocida – have been identified as the main pathogens of several localized and systemic infections.51,58 The predominant source of infections appears to be wounds caused by the bite of domestic predators. However, a recent study from Wisconsin in USA reported for the first time an outbreak of acute pasteurellosis from P. multocida in wild bats without associated traumatic injures.59
Leptospira sp.
Leptospira has worldwide distribution and its transmission to humans is primarily through exposure to water contaminated with the urine of infected animals.60 Bacterium harbors in several wild and domestic hosts, colonizes their kidneys and it is eliminated in their urine. The presence of Leptospira in bats has been demonstrated in several studies.51,61,62 However, the potential role of bats in human leptospirosis is questionable.63,64 In a case report, the patient’s history of bat exposure supports the idea that bats are a reservoir of the bacterium and can serve as a vector in disease transmission to humans.64
Enterobacteriaceae
Several members of the Enterobacteriaceae family – responsible for a variety of human illnesses – were isolated from bats.51,65–67 A number of studies reported that Salmonella serotypes isolated from bats have similar characteristics to those found from livestock and humans, indicating that bats can be locally important in the epidemiology of salmonellosis in human and domestic livestock.51,66 Two of these serotypes, S. typhimurium and S. enteritidis, are a frequent cause of human and animal diseases.
Escherichia coli strain has also been frequently isolated from bats.51,66–68 It is to emphasize the high percentage of multiresistance of these class of pathogens to several classes of antimicrobials51,66,68 that is a major and increasing global health-care problem. Antimicrobial resistance was also observed in domestic and wild animals, with an increased incidence of resistance in both pathogenic and endogenous bacteria.69 Resistant pathogens can then be transmitted to humans and bats can therefore contribute to the spreading of resistant bacteria.
Several other genera – such as Yersinia, Campylobacter, Vibrio – have been identified in bats, but their impact on these animals remains mostly unknown.51
Fungi
Histoplasma capsulatum
H. capsulatum is a dimorphic pathogenic fungus of mammals, which causes pulmonary and systemic infections in humans and it is acquired via inhalation of the fungal spores. This microorganism is commonly found in soil associated with great amounts of birds’ droppings or bats guano. Although bats are considered the main reservoir and dispersers of this fungus in the environment, their role in spreading H. capsulatum remains unclear.70 It has, however, been observed that subjects occupationally exposed to bat sites, such as miners, geologists or farmers who use bat guano as fertilizer, have high risk of infection and can develop severe clinical forms of histoplasmosis.70,71
Pseudogymnoascus destructans
Although implications in human health for this microorganism are not known, it is important to write a few words on an emerging fungal disease, named WNS, responsible for the deaths of millions of bats in North America. It is caused by the psychrophilic (cold-loving) fungus P. destructans that infects the skin of bats – especially the wings – during the winter months while they are in hibernation.72 Unlike other dermatophytes, which colonize the outer layer of the skin, P. destructans is able to invade the living tissue of the host causing the characteristic severe skin lesions.73 In addition P. destructans increases the frequency of periodic arousals in bats, resulting in premature consumption of stored fat essential to survive the winter leading to death within 4 months of infection.72 Recently, it has been observed that bacteria of the Pseudomonas genus – isolated from the skin of bats – inhibit the growth of the fungus in vitro.74 Additional in vivo studies will tell us whether in the future they could be used as biological control agents to protect bats exposed to P. destructans.
Conclusions and Future Perspective
Emergence of new infectious diseases correlates with socio-economic, environmental and ecological factors and are a major public health problem as well as an important burden on economies worldwide.75 Most of these are caused by zoonotic pathogens originating in wildlife and then spreading to humans. Bats are an important reservoir of several pathogenic agents, mainly viruses, and many of them have already caused disease outbreaks worldwide. The increasing rate of bat-associated infections is also supported by an expanding overlap between bat and human habitats. Recently, to increase the knowledge of bat-associated viruses, a database has been constructed (http://www.mgc.ac.cn/DBatVir).76 DbatVir analyzes the virome diversity of bats as well as the ecological and epidemiological data to examine and track current and future bat-related transmissible diseases. To date, DbatVir has collected information on 5717 bat-associated animal viruses detected from 207 bat species in 77 different countries (update on 2 march 2016). Strategies on surveillance and monitoring of disease outbreaks in bat populations need to be further developed, in particular where bats and humans are in close contact. Extensive studies are also needed to improve our understanding of bat–human interactions to design new control measures in future. Furthermore, the identification of new human pathogens requires a continuous study to monitor the potential impact of these animals in their diffusion.
References
Calisher CH, Childs JE, Field HE, Holmes KV, Schountz T . Bats: important reservoir hosts of emerging viruses. Clin Microbiol Rev 2006; 19: 531–545.
Brook CE, Dobson AP . Bats as 'special' reservoirs for emerging zoonotic pathogens. Trends Microbiol 2015; 23: 172–180.
Hayman DT, Bowen RA, Cryan PM, McCracken GF, O'Shea TJ, Peel AJ et al. Ecology of zoonotic infectious diseases in bats: current knowledge and future directions. Zoonoses Public Health 2013; 60: 2–21.
Han HJ, Wen HL, Zhou CM, Chen FF, Luo LM, Liu JW et al. Bats as reservoirs of severe emerging infectious diseases. Virus Res 2015; 205: 1–6.
Dobson AP . Virology. What links bats to emerging infectious diseases? Science 2005; 310: 628–629.
Krauel JJ, McCracken GF . Recent advances in bat migration research. In: Adams RA, Pedersen SC (eds), Bat Evolution, Ecology, and Conservation. Springer Science: New York, NY, USA, 2013, 293–313.
McGuire LP, Fenton MB, Guglielmo CG . Phenotypic flexibility in migrating bats: seasonal variation in body composition, organ sizes and fatty acid profiles. J Exp Biol 2013; 216: 800–808.
O' Shea TJ, Cryan PM, Cunningham AA, Fooks AR, Hayman DT, Luis AD et al. Bat flight and zoonotic viruses. Emerg Infect Dis 2014; 20: 741–745.
Moratelli R, Calisher CH . Bats and zoonotic viruses: can we confidently link bats with emerging deadly viruses? Mem Inst Oswaldo Cruz 2015; 110: 1–22.
Banyard AC, Evans JS, Luo TR, Fooks AR . Lyssaviruses and bats: emergence and zoonotic threat. Viruses 2014; 6: 2974–2990.
Costa LJ, Fernandes ME . Rabies: knowledge and practices regarding rabies in rural communities of the Brazilian Amazon Basin. PLoS Negl Trop Dis 2016; 10: e0004474.
Carini A . Sur une grande épizootie de rage. Ann Inst Pasteur 1911, 843–846.
Johnson N, Vos A, Freuling C, Tordo N, Fooks AR, Muller T . Human rabies due to lyssavirus infection of bat origin. Vet Microbiol 2010; 142: 151–159.
McElhinney LM, Marston DA, Leech S, Freuling CM, van der Poel WH, Echevarria J et al. Molecular epidemiology of bat lyssaviruses in Europe. Zoonoses Public Health 2013; 60: 35–45.
Arechiga Ceballos N, Vazquez Moron S, Berciano JM, Nicolas O, Aznar Lopez C, Juste J et al. Novel lyssavirus in bat, Spain. Emerg Infect Dis 2013; 19: 793–795.
Freuling CM, Abendroth B, Beer M, Fischer M, Hanke D, Hoffmann B et al. Molecular diagnostics for the detection of Bokeloh bat lyssavirus in a bat from Bavaria, Germany. Virus Res 2013; 177: 201–204.
Picard-Meyer E, Servat A, Robardet E, Moinet M, Borel C, Cliquet F . Isolation of Bokeloh bat lyssavirus in Myotis nattereri in France. Arch Virol 2013; 158: 2333–2340.
Kuzmin IV, Niezgoda M, Franka R, Agwanda B, Markotter W, Beagley JC et al. Possible emergence of West Caucasian bat virus in Africa. Emerg Infect Dis 2008; 14: 1887–1889.
Weir DL, Annand EJ, Reid PA, Broder CC . Recent observations on Australian bat lyssavirus tropism and viral entry. Viruses 2014; 6: 909–926.
Clayton BA, Wang LF, Marsh GA . Henipaviruses: an updated review focusing on the pteropid reservoir and features of transmission. Zoonoses Public Health 2013; 60: 69–83.
Tee KK, Takebe Y, Kamarulzaman A . Emerging and re-emerging viruses in Malaysia, 1997-2007. Int J Infect Dis 2009; 13: 307–318.
Plowright RK, Foley P, Field HE, Dobson AP, Foley JE, Eby P et al. Urban habituation, ecological connectivity and epidemic dampening: the emergence of Hendra virus from flying foxes (Pteropus spp.). Proc Biol Sci 2011; 278: 3703–3712.
Drexler JF, Corman VM, Drosten C . Ecology, evolution and classification of bat coronaviruses in the aftermath of SARS. Antiviral Res 2014; 101: 45–56.
Weinstein RA . Planning for epidemics--the lessons of SARS. N Engl J Med 2004; 350: 2332–2334.
Guan Y, Zheng BJ, He YQ, Liu XL, Zhuang ZX, Cheung CL et al. Isolation and characterization of viruses related to the SARS coronavirus from animals in southern China. Science 2003; 302: 276–278.
Chan JF, Lau SK, To KK, Cheng VC, Woo PC, Yuen KY . Middle East respiratory syndrome coronavirus: another zoonotic betacoronavirus causing SARS-like disease. Clin Microbiol Rev 2015; 28: 465–522.
Razanajatovo NH, Nomenjanahary LA, Wilkinson DA, Razafimanahaka JH, Goodman SM, Jenkins RK et al. Detection of new genetic variants of Betacoronaviruses in endemic frugivorous bats of Madagascar. Virol J 2015; 12: 42.
Zaki AM, van Boheemen S, Bestebroer TM, Osterhaus AD, Fouchier RA . Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N Engl J Med 2012; 367: 1814–1820.
de Wit E, Munster VJ . MERS-CoV: the intermediate host identified? Lancet Infect Dis 2013; 13: 827–828.
Munster VJ, Adney DR, van Doremalen N, Brown VR, Miazgowicz KL, Milne-Price S et al. Replication and shedding of MERS-CoV in Jamaican fruit bats (Artibeus jamaicensis). Sci Rep 2016; 6: 21878.
Olival KJ, Hayman DT . Filoviruses in bats: current knowledge and future directions. Viruses 2014; 6: 1759–1788.
Beer B, Kurth R, Bukreyev A . Characteristics of Filoviridae: Marburg and Ebola viruses. Naturwissenschaften 1999; 86: 8–17.
Leendertz SA, Gogarten JF, Dux A, Calvignac-Spencer S, Leendertz FH . Assessing the evidence supporting fruit bats as the primary reservoirs for Ebola viruses. Ecohealth 2015; 13: 18–25.
Negredo A, Palacios G, Vazquez-Moron S, Gonzalez F, Dopazo H, Molero F et al. Discovery of an ebolavirus-like filovirus in europe. PLoS Pathog 2011; 7: e1002304.
Tong S, Li Y, Rivailler P, Conrardy C, Castillo DA, Chen LM et al. A distinct lineage of influenza A virus from bats. Proc Natl Acad Sci USA 2012; 109: 4269–4274.
Tong S, Zhu X, Li Y, Shi M, Zhang J, Bourgeois M et al. New world bats harbor diverse influenza A viruses. PLoS Pathog 2013; 9: e1003657.
Freidl GS, Binger T, Muller MA, de Bruin E, van Beek J, Corman VM et al. Serological evidence of influenza A viruses in frugivorous bats from Africa. PLoS One 2015; 10: e0127035.
Wong SS, Yuen KY . Avian influenza virus infections in humans. Chest 2006; 129: 156–168.
Holmes EC, Zhang YZ . The evolution and emergence of hantaviruses. Curr Opin Virol 2015; 10: 27–33.
Bennett SN, Gu SH, Kang HJ, Arai S, Yanagihara R . Reconstructing the evolutionary origins and phylogeography of hantaviruses. Trends Microbiol 2014; 22: 473–482.
Kim GR, Lee YT, Park CH . A new natural reservoir of hantavirus: isolation of hantaviruses from lung tissues of bats. Arch Virol 1994; 134: 85–95.
Wang L, Fu S, Cao L, Lei W, Cao Y, Song J et al. Isolation and identification of a natural reassortant mammalian orthoreovirus from least horseshoe bat in China. PLoS One 2015; 10: e0118598.
Kohl C, Lesnik R, Brinkmann A, Ebinger A, Radonic A, Nitsche A et al. Isolation and characterization of three mammalian orthoreoviruses from European bats. PLoS One 2012; 7: e43106.
Lelli D, Moreno A, Lavazza A, Bresaola M, Canelli E, Boniotti MB et al. Identification of Mammalian orthoreovirus type 3 in Italian bats. Zoonoses Public Health 2013; 60: 84–92.
Steyer A, Gutierrez-Aguire I, Kolenc M, Koren S, Kutnjak D, Pokorn M et al. High similarity of novel orthoreovirus detected in a child hospitalized with acute gastroenteritis to mammalian orthoreoviruses found in bats in Europe. J Clin Microbiol 2013; 51: 3818–3825.
Dacheux L, Cervantes-Gonzalez M, Guigon G, Thiberge JM, Vandenbogaert M, Maufrais C et al. A preliminary study of viral metagenomics of French bat species in contact with humans: identification of new mammalian viruses. PLoS One 2014; 9: e87194.
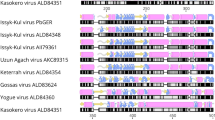
Sano K, Okazaki S, Taniguchi S, Masangkay JS, Puentespina Jr R, Eres E et al. Detection of a novel herpesvirus from bats in the Philippines. Virus Genes 2015; 51: 136–139.
Bonvicino CR, Moreira MA, Soares MA . Hepatitis B virus lineages in mammalian hosts: potential for bidirectional cross-species transmission. World J Gastroenterol 2014; 20: 7665–7674.
Baker KS, Murcia PR . Poxviruses in bats... so what? Viruses 2014; 6: 1564–1577.
Sotomayor-Bonilla J, Chaves A, Rico-Chavez O, Rostal MK, Ojeda-Flores R, Salas-Rojas M et al. Dengue virus in bats from southeastern Mexico. Am J Trop Med Hyg 2014; 91: 129–131.
Mühldorfer K . Bats and bacterial pathogens: a review. Zoonoses Public Health 2013; 60: 93–103.
Eremeeva ME, Gerns HL, Lydy SL, Goo JS, Ryan ET, Mathew SS et al. Bacteremia, fever, and splenomegaly caused by a newly recognized bartonella species. N Engl J Med 2007; 356: 2381–2387.
Bai Y, Kosoy M, Recuenco S, Alvarez D, Moran D, Turmelle A et al. Bartonella spp. in bats, Guatemala. Emerg Infect Dis 2011; 17: 1269–1272.
Veikkolainen V, Vesterinen EJ, Lilley TM, Pulliainen AT . Bats as reservoir hosts of human bacterial pathogen, Bartonella mayotimonensis. Emerg Infect Dis 2014; 20: 960–967.
Lei BR, Olival KJ . Contrasting patterns in mammal-bacteria coevolution: bartonella and leptospira in bats and rodents. PLoS Negl Trop Dis 2014; 8: e2738.
Brook CE, Bai Y, Dobson AP, Osikowicz LM, Ranaivoson HC, Zhu Q et al. Bartonella spp. in fruit bats and blood-feeding ectoparasites in Madagascar. PLoS Negl Trop Dis 2015; 9: e0003532.
Wilson BA, Ho M . Pasteurella multocida: from zoonosis to cellular microbiology. Clin Microbiol Rev 2013; 26: 631–655.
Mühldorfer K, Schwarz S, Fickel J, Wibbelt G, Speck S . Genetic diversity of Pasteurella species isolated from European vespertilionid bats. Vet Microbiol 2011; 149: 163–171.
Blehert DS, Maluping RP, Green DE, Berlowski-Zier BM, Ballmann AE, Langenberg JA . Acute pasteurellosis in wild big brown bats (Eptesicus fuscus). J Wildl Dis 2014; 50: 136–139.
Adler B, de la Pena Moctezuma A . Leptospira and leptospirosis. Vet Microbiol 2010; 140: 287–296.
Lagadec E, Gomard Y, Guernier V, Dietrich M, Pascalis H, Temmam S et al. Pathogenic Leptospira spp. in bats, Madagascar and Union of the Comoros. Emerg Infect Dis 2012; 18: 1696–1698.
Dietrich M, Muhldorfer K, Tortosa P, Markotter W . Leptospira and bats: story of an emerging friendship. PLoS Pathog 2015; 11: e1005176.
Bessa TA, Spichler A, Chapola EG, Husch AC, de Almeida MF, Sodre MM et al. The contribution of bats to leptospirosis transmission in Sao Paulo City, Brazil. Am J Trop Med Hyg 2010; 82: 315–317.
Vashi NA, Reddy P, Wayne DB, Sabin B . Bat-associated leptospirosis. J Gen Intern Med 2010; 25: 162–164.
Prem Anand AA, Sripathi K . Digestion of cellulose and xylan by symbiotic bacteria in the intestine of the Indian flying fox (Pteropus giganteus). Comp Biochem Physiol A Mol Integr Physiol 2004; 139: 65–69.
Adesiyun AA, Stewart-Johnson A, Thompson NN . Isolation of enteric pathogens from bats in Trinidad. J Wildl Dis 2009; 45: 952–961.
Chaverri G . Aerobic bacterial flora from the digestive tract of the common vampire bat, Desmodus rotundus (Chiroptera: Phyllostomidae). Rev Biol Trop 2006; 54: 717–724.
Costa D, Poeta P, Saenz Y, Vinue L, Coelho AC, Matos M et al. Mechanisms of antibiotic resistance in Escherichia coli isolates recovered from wild animals. Microb Drug Resist 2008; 14: 71–77.
Allocati N, Masulli M, Alexeyev MF, Di Ilio C . Escherichia coli in Europe: an overview. Int J Environ Res Public Health 2013; 10: 6235–6254.
Taylor ML, Chavez-Tapia CB, Rojas-Martinez A, del Rocio Reyes-Montes M, del Valle MB, Zuniga G . Geographical distribution of genetic polymorphism of the pathogen Histoplasma capsulatum isolated from infected bats, captured in a central zone of Mexico. FEMS Immunol Med Microbiol 2005; 45: 451–458.
Santos L, Santos-Martinez G, Magana-Ortiz JE, Puente-Pinon SL . Acute histoplasmosis in three Mexican sewer workers. Occup Med (Lond) 2013; 63: 77–79.
Warnecke L, Turner JM, Bollinger TK, Lorch JM, Misra V, Cryan PM et al. Inoculation of bats with European Geomyces destructans supports the novel pathogen hypothesis for the origin of white-nose syndrome. Proc Natl Acad Sci USA 2012; 109: 6999–7003.
Cryan PM, Meteyer CU, Boyles JG, Blehert DS . Wing pathology of white-nose syndrome in bats suggests life-threatening disruption of physiology. BMC Biol 2010; 8: 135.
Hoyt JR, Cheng TL, Langwig KE, Hee MM, Frick WF, Kilpatrick AM . Bacteria isolated from bats inhibit the growth of Pseudogymnoascus destructans, the causative agent of white-nose syndrome. PLoS One 2015; 10: e0121329.
Jones KE, Patel NG, Levy MA, Storeygard A, Balk D, Gittleman JL et al. Global trends in emerging infectious diseases. Nature 2008; 451: 990–993.
Chen L, Liu B, Yang J, Jin Q . DBatVir: the database of bat-associated viruses. Database (Oxford) 2014; 2014: bau021.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by I Harris
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Allocati, N., Petrucci, A., Di Giovanni, P. et al. Bat–man disease transmission: zoonotic pathogens from wildlife reservoirs to human populations. Cell Death Discovery 2, 16048 (2016). https://doi.org/10.1038/cddiscovery.2016.48
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/cddiscovery.2016.48
This article is cited by
-
Comparative virome analysis of individual shedding routes of Miniopterus phillipsi bats inhabiting the Wavul Galge cave, Sri Lanka
Scientific Reports (2023)
-
COVID-19 pandemic: a health challenge for commoners during the first unlock phase in India
Journal of Public Health (2023)
-
Essential role of Rnd1 in innate immunity during viral and bacterial infections
Cell Death & Disease (2022)
-
Statistical data analysis of risk factor associated with mortality rate by COVID-19 pandemic in India
Modeling Earth Systems and Environment (2022)
-
Sleep quality and anxiety symptoms in Egyptian children and adolescents during COVID-19 pandemic lockdown
Bulletin of the National Research Centre (2021)