Abstract
Tumor necrosis factor-related apoptosis-inducing ligand (TRAIL) is a promising agent for anticancer therapy; however, non-small-cell lung carcinoma (NSCLC) cells are relatively TRAIL resistant. Identification of small molecules that can restore NSCLC susceptibility to TRAIL-induced apoptosis is meaningful. We found here that rotenone, as a mitochondrial respiration inhibitor, preferentially increased NSCLC cells sensitivity to TRAIL-mediated apoptosis at subtoxic concentrations, the mechanisms by which were accounted by the upregulation of death receptors and the downregulation of c-FLIP (cellular FLICE-like inhibitory protein). Further analysis revealed that death receptors expression by rotenone was regulated by p53, whereas c-FLIP downregulation was blocked by Bcl-XL overexpression. Rotenone triggered the mitochondria-derived reactive oxygen species (ROS) generation, which subsequently led to Bcl-XL downregulation and PUMA upregulation. As PUMA expression was regulated by p53, the PUMA, Bcl-XL and p53 in rotenone-treated cells form a positive feedback amplification loop to increase the apoptosis sensitivity. Mitochondria-derived ROS, however, promote the formation of this amplification loop. Collectively, we concluded that ROS generation, Bcl-XL and p53-mediated amplification mechanisms had an important role in the sensitization of NSCLC cells to TRAIL-mediated apoptosis by rotenone. The combined TRAIL and rotenone treatment may be appreciated as a useful approach for the therapy of NSCLC that warrants further investigation.
Similar content being viewed by others
Main
Tumor necrosis factor-related apoptosis-inducing ligand (TRAIL) has emerged as a promising cancer therapeutic because it can selectively induce apoptosis in tumor cells in vitro, and most importantly, in vivo with little adverse effect on normal cells.1 However, a number of cancer cells are resistant to TRAIL, especially highly malignant tumors such as lung cancer.2, 3 Lung cancer, especially the non-small-cell lung carcinoma (NSCLC) constitutes a heavy threat to human life. Presently, the morbidity and mortality of NSCLC has markedly increased in the past decade,4 which highlights the need for more effective treatment strategies.
TRAIL has been shown to interact with five receptors, including the death receptors 4 and 5 (DR4 and DR5), the decoy receptors DcR1 and DcR2, and osteoprotegerin.5 Ligation of TRAIL to DR4 or DR5 allows for the recruitment of Fas-associated protein with death domain (FADD), which leads to the formation of death-inducing signaling complex (DISC) and the subsequent activation of caspase-8/10.6 The effector caspase-3 is activated by caspase-8, which cleaves numerous regulatory and structural proteins resulting in cell apoptosis. Caspase-8 can also cleave the Bcl-2 inhibitory BH3-domain protein (Bid), which engages the intrinsic apoptotic pathway by binding to Bcl-2-associated X protein (Bax) and Bcl-2 homologous antagonist killer (BAK). The oligomerization between Bcl-2 and Bax promotes the release of cytochrome c from mitochondria to cytosol, and facilitates the formation of apoptosome and caspase-9 activation.7 Like caspase-8, caspase-9 can also activate caspase-3 and initiate cell apoptosis. Besides apoptosis-inducing molecules, several apoptosis-inhibitory proteins also exist and have function even when apoptosis program is initiated. For example, cellular FLICE-like inhibitory protein (c-FLIP) is able to suppress DISC formation and apoptosis induction by sequestering FADD.8, 9, 10, 11
Until now, the recognized causes of TRAIL resistance include differential expression of death receptors, constitutively active AKT and NF-κB,12, 13 overexpression of c-FLIP and IAPs, mutations in Bax and BAK gene.2 Hence, resistance can be overcome by the use of sensitizing agents that modify the deregulated death receptor expression and/or apoptosis signaling pathways in cancer cells.5 Many sensitizing agents have been developed in a variety of tumor cell models.2 Although the clinical effectiveness of these agents needs further investigation, treatment of TRAIL-resistant tumor cells with sensitizing agents, especially the compounds with low molecular weight, as well as prolonged plasma half-life represents a promising trend for cancer therapy.
Mitochondria emerge as intriguing targets for cancer therapy. Metabolic changes affecting mitochondria function inside cancer cells endow these cells with distinctive properties and survival advantage worthy of drug targeting, mitochondria-targeting drugs offer substantial promise as clinical treatment with minimal side effects.14, 15, 16 Rotenone is a potent inhibitor of NADH oxidoreductase in complex I, which demonstrates anti-neoplastic activity on a variety of cancer cells.17, 18, 19, 20, 21 However, the neurotoxicity of rotenone limits its potential application in cancer therapy. To avoid it, rotenone was effectively used in combination with other chemotherapeutic drugs to kill cancerous cells.22
In our previous investigation, we found that rotenone was able to suppress membrane Na+,K+-ATPase activity and enhance ouabain-induced cancer cell death.23 Given these facts, we wonder whether rotenone may also be used as a sensitizing agent that can restore the susceptibility of NSCLC cells toward TRAIL-induced apoptosis, and increase the antitumor efficacy of TRAIL on NSCLC. To test this hypothesis, we initiated this study.
Results
Rotenone sensitizes NSCLC cell lines to TRAIL-induced apoptosis
Four NSCLC cell lines including A549, H522, H157 and Calu-1 were used in this study. As shown in Figure 1a, the apoptosis induced by TRAIL alone at 50 or 100 ng/ml on A549, H522, H157 and Calu-1 cells was non-prevalent, indicating that these NSCLC cell lines are relatively TRAIL resistant. Interestingly, when these cells were treated with TRAIL combined with rotenone, significant increase in cell apoptosis was observed. To examine whether rotenone was also able to sensitize normal cells to TRAIL-mediated apoptosis, peripheral blood mononuclear cell (PBMC) isolated from human blood were used. As a result, rotenone failed to sensitize human PBMC to TRAIL-induced apoptosis, indicating that the sensitizing effect of rotenone is tumor cell specific. Of note, the apoptosis-enhancing effect of rotenone occurred independent of its cytotoxicity, because the minimal dosage required for rotenone to cause toxic effect on NSCLC cell lines was 10 μM, however, rotenone augmented TRAIL-mediated apoptosis when it was used as little as 10 nM.
Effects of rotenone and TRAIL on NSCLC and PBMC cell apoptosis. (a) A549, H522, H157, Calu-1 and human PBMC cells were treated with rotenone at concentrations ranging from 10 ng/ml to 10 000 nM, or TRAIL at 50, 100 ng/ml, or both for 8 h, after treatment, cells were collected and the occurrence of apoptosis was measured by annexin V/PI double staining method. (b) NSCLC cells, including A549, H522, H157 and Calu-1 cells, were treated with rotenone at 1 μM, TRAIL at 100 ng/ml or both for 8 h, after treatment, cells were fixed and stained with Hoechst, then observed under Carl Zeiss fluorescent microscope (Carl Zeiss, Jena, Germany). Scale bars, 20 μM. (c) A549 cells were treated with rotenone at 1 μM, TRAIL at 100 ng/ml or both for 8 h, after treatment, cells were collected and stained with FITC-conjugated substrates for caspase-3, -8 and -9. FITC fluorescent intensity within each sample was measured by flow cytometry. The representative overlaid histograms of three independent experiments with similar results are shown. (d) A549 or H522 cells were preincubated with 4 μM z-VAD.fmk for 30 min, and further treated with TRAIL (100 ng/ml) and rotenone (1 μM) for 8 h. After treatment, cell apoptosis was assessed by annexin V/PI double staining method. Each experiment was performed at least in triplicate
To further confirm the effect of rotenone, cells were stained with Hoechst and observed under fluorescent microscope (Figure 1b). Consistently, the combined treatment of rotenone with TRAIL caused significant nuclear fragmentation in A549, H522, H157 and Calu-1 cells. Rotenone or TRAIL treatment alone, however, had no significant effect.
Caspases activation is a hallmark of cell apoptosis. In this study, the enzymatic activities of caspases including caspase-3, -8 and -9 were measured by flow cytometry by using FITC-conjugated caspases substrate (Figure 1c). As a result, rotenone used at 1 μM or TRAIL used at 100 ng/ml alone did not cause caspase-3, -8 and -9 activation. The combined treatment, however, significantly increased the enzymatic activities of them. Moreover, A549 or H522 cell apoptosis by TRAIL combined with rotenone was almost completely suppressed in the presence of z-VAD.fmk, a pan-caspase inhibitor (Figure 1d). All of these data indicate that both intrinsic and extrinsic pathways are involved in the sensitizing effect of rotenone on TRAIL-mediated apoptosis in NSCLC.
Upregulation of death receptors expression is required for rotenone-mediated sensitization to TRAIL-induced apoptosis
Sensitization to TRAIL-induced apoptosis has been explained in some studies by upregulation of death receptors,24 whereas other results show that sensitization can occur without increased TRAIL receptor expression.25 As such, we examined TRAIL receptors expression on NSCLC cells after treatment with rotenone. Rotenone increased DR4 and DR5 mRNA levels in A549 cells in a time or concentration-dependent manner (Figures 2a and b), also increased DR4 and DR5 protein expression levels (Supplementary Figure S1). Notably, rotenone failed to increase DR5 mRNA levels in H157 and Calu-1 cells (Supplementary Figure S2). To observe whether the increased DR4 and DR5 mRNA levels finally correlated with the functional molecules, we examined the surface expression levels of DR4 and DR5 by flow cytometry. The results, as shown in Figure 2c demonstrated that the cell surface expression levels of DR4 and DR5 were greatly upregulated by rotenone in either A549 cells or H522 cells.
Effect of rotenone on DR4 and DR5 expression levels in NSCLC cells. A549 cells were treated with rotenone at 1 μM for 0, 2, 4 and 6 h, respectively, in a, or treated with rotenone at 0, 10, 100 and 1000 ng/ml for 6 h in b, after treatment, the total cellular mRNAs were extracted by TRIZOL agent, RT-PCR analysis was performed to measure the DR4 and DR5 mRNA levels. GAPDH was included as a control. The quantitative results of DR4 and DR5 mRNA expression levels are shown on the right panel; the mRNA expression levels of DR4 and DR5 in controls were arbitrarily set at 100%. (c) A549 or H522 cells were treated with rotenone at 1 μM for 8 h, after treatment, cells were collected, incubated with DR4 or DR5 monoclonal antibody and FITC-conjugated secondary IgGs. The representative histograms of three independent experiments with similar results are shown. (d) A549 or H522 cells were transfected with control siRNA, DR4 or DR5 siRNAs for 24 h, then treated with rotenone (1 μM) and TRAIL (100 ng/ml) for 8 h. After treatment, cells were collected, and cell apoptosis was measured by annexin V/PI double staining. Each experiment was performed in triplicate. **P<0.01, ***P<0.001, as compared with cells transfected with control siRNA
To analyze whether the upregulation of DR4 and DR5 is a ‘side-effect’, or contrarily, necessary for rotenone-mediated sensitization to TRAIL-induced apoptosis, we blocked upregulation of the death receptors by small interfering RNAs (siRNAs) against DR4 and DR5 (Supplementary Figure S3). The results showed that blocking DR4 and DR5 expression alone significantly reduced the rate of cell apoptosis in A549 cells (Figure 2d). However, the highest inhibition of apoptosis was observed when upregulation of both receptors was blocked in parallel, thus showing an additive effect of blocking DR4 and DR5 at the same time. Similar results were also obtained in H522 cells.
Rotenone-induced p53 activation regulates death receptors upregulation
TRAIL receptors DR4 and DR5 are regulated at multiple levels. At transcriptional level, studies suggest that several transcriptional factors including NF-κB, p53 and AP-1 are involved in DR4 or DR5 gene transcription.2 The NF-κB or AP-1 transcriptional activity was further modulated by ERK1/2, JNK and p38 MAP kinase activity. Unexpectedly, we found here that none of these MAP kinases inhibitors were able to suppress the apoptosis mediated by TRAIL plus rotenone (Figure 3a). To find out other possible mechanisms, we observed that rotenone was able to stimulate p53 phosphorylation as well as p53 protein expression in A549 and H522 cells (Figure 3b). As a p53-inducible gene, p21 mRNA expression was also upregulated by rotenone treatment in a time-dependent manner (Figure 3c). To characterize the effect of p53, A549 cells were transfected with p53 siRNA. The results, as shown in Figure 3d-1 demonstrated that rotenone-mediated surface expression levels of DR4 and DR5 in A549 cells were largely attenuated by siRNA-mediated p53 expression silencing. Control siRNA, however, failed to reveal such effect. Similar results were also obtained in H522 cells (Figure 3d-2). Silencing of p53 expression in A549 cells also partially suppressed the apoptosis induced by TRAIL plus rotenone (Figure 3e).
Involvement of p53 in the apoptosis-enhancing effect of rotenone. (a) A549 cells were pretreated with 20 μM PD98059, SB203580 and SP60025 for 30 min, further treated with TRAIL (100 ng/ml) plus rotenone (1 μM ) for an additional 8 h, after treatment, cell apoptosis was analyzed by annexin V/PI double staining method. (b) A549 or H522 cells were treated with rotenone at 1 μM for 0, 2, 4, 6 and 8 h, respectively, after treatment, cell were collected and western blot analysis was performed to measure the protein expression levels of p53 as well as its phosphorylation. (c) A549 cells were treated with rotenone at 1 μM for 0, 2, 4, 6 and 8 h, respectively. After treatment, RT-PCR was performed to measure p21 mRNA level. The quantitative result is shown on the right panel; the p21 mRNA expression in control cells was arbitrarily set at 1.0. (d) A549 cells in (d-1) or H522 cells in (d-2) were transfected with control or p53 siRNAs for 24 h, after transfection, cells were further treated with rotenone at 1 μM for 8 h. After treatment, cells were collected and the surface expression levels of DR4 and DR5 on A549 and H522 cells were measured by flow cytometry. ***P<0.001, ###P<0.001, as compared with cells transfected with control siRNA. (e) A549 cells were transfected with control or p53 siRNAs for 24 h, after transfection, cells were further treated with TRAIL at 100 nM plus rotenone at 1 μM for 8 h. After treatment, cells were collected and apoptosis was measured by annexin V/PI double staining method. **P<0.01, as compared with cells transfected with control siRNA
Rotenone suppresses c-FLIP expression and increases the sensitivity of A549 cells to TRAIL-induced apoptosis
The c-FLIP protein has been commonly appreciated as an anti-apoptotic molecule in death receptor-mediated cell apoptosis. In this study, rotenone treatment led to dose-dependent downregulation of c-FLIP expression, including c-FLIPL and c-FLIPs in A549 cells (Figure 4a-1), H522 cells (Figure 4a-2), H441 and Calu-1 cells (Supplementary Figure S4). To test whether c-FLIP is essential for the apoptosis enhancement, A549 cells were transfected with c-FLIPL-overexpressing plasmids. As shown in Figure 4b-1, the apoptosis of A549 cells after the combined treatment was significantly reduced when c-FLIPL was overexpressed. Similar results were also obtained in H522 cells (Figure 4b-2).
Effect of rotenone on c-FLIP expression in NSCLC cells. A549 cells (a-1) or in H522 cells in (a-2) were treated with rotenone at 0, 10, 100, 1000 and 10 000 nM, respectively, for 8 h. After treatment, cells were lyzed and subjected to western blot analysis. The quantitative result is shown below, and the protein abundance of c-FLIPL and c-FLIPs in control cells were arbitrarily set at 100%. A549 cells (b-1) or H522 cells in (b-2) were transfected with 2 μg mock plasmid or c-FLIPL-overexpressing plasmid for 24 h, further treated with 100 ng TRAIL plus rotenone at 1 μM for 8 h. After treatment, cells were collected and apoptosis was measured by annexin V/PI double staining. Each experiment was performed in triplicate. **P<0.01, ***P<0.001, as compared with mock plasmid transfected cells
Bcl-XL is involved in the apoptosis enhancement by rotenone
Notably, c-FLIP downregulation by rotenone in NSCLC cells was irrelevant to p53 signaling (data not shown). To identify other mechanism involved, we found that anti-apoptotic molecule Bcl-XL was also found to be downregulated by rotenone in a dose-dependent manner (Figure 5a). Notably, both Bcl-XL and c-FLIPL mRNA levels remained unchanged in cells after rotenone treatment (Supplementary Figure S5). Bcl-2 is homolog to Bcl-XL. But surprisingly, Bcl-2 expression was almost undetectable in A549 cells. To examine whether Bcl-XL is involved, A549 cells were transfected with Bcl-XL-overexpressing plasmid. As compared with mock transfectant, cell apoptosis induced by TRAIL plus rotenone was markedly suppressed under the condition of Bcl-XL overexpression (Figure 5b). To characterize the mechanisms, surface expression levels of DR4 and DR5 were examined. As shown in Figure 5c, the increased surface expression of DR4 and DR5 in A549 cells, or in H522 cells were greatly reduced after Bcl-XL overexpression (Figure 5c). In addition, Bcl-XL overexpression also significantly prevented the downregulation of c-FLIPL and c-FLIPs expression in A549 cells by rotenone treatment (Figure 5d).
Involvement of Bcl-XL in the apoptosis enhancement by rotenone. (a) A549 cells were treated with rotenone at 0, 10, 100, 1000 and 10 000 nM for 8 h, after treatment, western blot analysis was performed to measure Bcl-XL and Bcl-2 expression levels, tubulin was included as a loading control. The quantitative analysis of Bcl-XL protein expression in rotenone-treated A549 cells is shown on the right panel. The protein expression of Bcl-XL in control cells was arbitrarily set at 100%. **P<0.01, ***P<0.001, as compared with control cells. (b) A549 cells were transfected with mock, or Bcl-XL-overexpressing plasmid for 24 h, then treated with TRAIL (100 ng/ml) plus rotenone (1 μM ) for 8 h. Cell apoptosis was measured by annexin V/PI double staining. ***P<0.001, as compared with mock transfectant. (c) A549 or H522 cells were transfected with mock or Bcl-XL plasmids for 24 h, after transfection, cells were treated with rotenone at 1 μM for an additional 8 h, surface expression levels of DR4 and DR5 receptors were measured by flow cytometric analysis. ***P<0.001, ###P<0.001, as compared with mock transfectant. (d) A549 cells were transfected with mock or Bcl-XL plasmids for 24 h, after transfection, cells were treated with rotenone at 1 μM for an additional 8 h, western blot analysis was performed to examine the c-FLIPL and c-FLIPs expression levels
Rotenone suppresses the interaction between BCL-XL/p53 and increases PUMA transcription
Lines of evidence suggest that Bcl-XL has a strong binding affinity with p53, and can suppress p53-mediated tumor cell apoptosis.26 In this study, FLAG-tagged Bcl-XL and HA-tagged p53 were co-transfected into cells; immunoprecipitation experiment was performed by using FLAG antibody to immunoprecipitate HA-tagged p53. As a result, we found that at the same amount of p53 protein input, rotenone treatment caused a concentration-dependent suppression of the protein interaction between Bcl-XL and p53 (Figure 6a). Rotenone also significantly suppressed the interaction between endogenous Bcl-XL and p53 when polyclonal antibody against p53 was used to immunoprecipitate cellular Bcl-XL (Figure 6b). Recent study highlighted the importance of PUMA in BCL-XL/p53 interaction and cell apoptosis.27 We found here that rotenone significantly increased PUMA gene transcription (Figure 6c) and protein expression (Figure 6d) in NSCLC cells, but not in transformed 293T cell. Meanwhile, this effect was attenuated by silencing of p53 expression (Figure 6e).
Effects of rotenone on Bcl-XL/p53 interaction and PUMA expression. (a) A549 cells were transfected with FLAG-tagged Bcl-XL and HA-tagged p53 for 24 h, after transfection, cells were further treated with rotenone at 0, 10, 100 and 1000 nM for 8 h. After treatment, immunoprecipitation experiments were performed by using anti-FLAG IgG. The immunopellets were subject to western blot analysis by using anti-HA IgG. Cellular protein input of HA-tagged p53 and FLAG-tagged Bcl-XL was also shown, GAPDH was included as a loading control. (b) A549 cells were treated with rotenone at 1 μM for 8 h, after treatment, cells were lyzed and immunoprecipitation experiments were performed by using anti-p53 IgG to pull down p53 and the associated proteins. Cellular protein input of p53 and Bcl-XL was also shown, GAPDH was included as a loading control. (c) The 293T, H522, A549 and Calu-1 cells were treated with rotenone at 1 μM for 8 h, after treatment, cells were collected and RT-PCR analysis was performed to measure PUMA mRNA expression levels, GAPDH was included as a loading control. (d) A549 cells were treated with rotenone at 1, 10, 100, 1000 and 10 000 nM for 8 h, after treatment, western blot analysis was performed to examine the PUMA expression levels. (e) A549 were transfected with control or p53 siRNAs for 24 h, after transfection, cells were treated with rotenone at 1 μM for an additional 8 h, western blot analysis was performed to examine the PUMA protein expression. The experiments were performed at least in triplicate
Mitochondria-derived ROS are responsible for the apoptosis-enhancing effect of rotenone
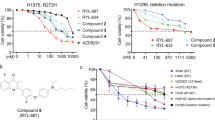
As an inhibitor of mitochondrial respiration, rotenone was found to induce reactive oxygen species (ROS) generation in a variety of transformed or non-transformed cells.20, 22 Consistently, by using 2',7'-dichlorofluorescin diacetate (DCFH) for the measurement of intracellular H2O2 and dihydroethidium (DHE) for O2.−, we found that rotenone significantly triggered the .generation of H2O2 (Figure 7a) and O2.− (Figure 7b) in A549 and H522 cells. To identify the origin of ROS production, we first incubated cells with diphenylene iodonium (DPI), a potent inhibitor of plasma membrane NADP/NADPH oxidase. The results showed that DPI failed to suppress rotenone-induced ROS generation (Figure 7c). Then, we generated A549 cells deficient in mitochondria DNA by culturing cells in medium supplemented with ethidium bromide (EB). These mtDNA-deficient cells were subject to rotenone treatment, and the result showed that rotenone-induced ROS production were largely attenuated in A549 ρ° cells, but not wild-type A549 cells, suggesting ROS are mainly produced from mitochondria (Figure 7d). Notably, the sensitizing effect of rotenone on TRAIL-induced apoptosis in A549 cells was largely dependent on ROS, because the antioxidant N-acetylcysteine (NAC) treatment greatly suppressed the cell apoptosis, as shown in annexin V/PI double staining experiment (Figure 7e), cell cycle analysis (Figure 7f) and caspase-3 cleavage activity assay (Figure 7g). Finally, in A549 cells stably transfected with manganese superoxide (MnSOD) and catalase, apoptosis induced by TRAIL and rotenone was partially reversed (Figure 7h). All of these data suggest that mitochondria-derived ROS, including H2O2 and O2.−, are responsible for the apoptosis-enhancing effect of rotenone.
Involvement of ROS in rotenone-mediated apoptosis enhancement. (a and b) A549 or H522 cells were treated with rotenone at 1 μM for 0, 1, 2, 3 and 4 h, respectively. After treatment, cells were collected and stained with DCFH for measurement of H2O2 generation in a, or with DHE for measurement of O2.− in b by using flow cytometry. (c) A549 cells were pretreated with DPI at 10 μM for 1 h, further treated with rotenone at 1 μM for 4 h, after treatment, cells were stained with DCFH for measurement of H2O2, or with DHE for O2.−, respectively. (d) A549 WT or ρ° cells were treated with rotenone at 1 μM for 4 h, after treatment, cells were stained with DCFH for measurement of H2O2, or with DHE for O2.−, respectively. ***P<0.001, ###P<0.001, as compared with wild-type A549 cells. (e–g) A549 cells were preincubated with NAC at 5 mM for 30 min, further treated with TRAIL (100 ng/ml) plus rotenone (1 μM) for 8 h, cell apoptosis was measured by annexin V/PI double staining in (e), cell cycle analysis was performed by flow cytometry in (f), the caspase-3 cleavage activity was measured by flow cytometry by using FITC-conjugated substrate in (g). (h) A549 were transfected with mock, catalase or MnSOD-overexpressing plasmids for 24 h, after transfection, cells were treated with 100 ng TRAIL plus 1 μM rotenone for an additional 8 h, cell apoptosis was measured by annexin V/PI double staining method. ***P<0.001, as compared with control cells
Rotenone promotes BCl-XL degradation and PUMA transcription in ROS-dependent manner
To understand why ROS are responsible for the apoptosis-enhancing effect of rotenone, we found that rotenone-induced suppression of BCL-XL expression can be largely reversed by NAC treatment (Figure 8a). To examine whether this effect of rotenone occurs at posttranslational level, we used cycloheximide (CHX) to halt protein synthesis, and found that the rapid degradation of Bcl-XL by rotenone was largely attenuated in A549 ρ0 cells (Figure 8b). Similarly, rotenone-induced PUMA upregulation was also significantly abrogated in A549 ρ0 cells (Figure 8c). Finally, A549 cells were inoculated into nude mice to produce xenografts tumor model. In this model, the therapeutic effect of TRAIL combined with rotenone was evaluated. Notably, in order to circumvent the potential neurotoxic adverse effect of rotenone, mice were challenged with rotenone at a low concentration of 0.5 mg/kg. The results, as shown in Figure 8d revealed that while TRAIL or rotenone alone remained unaffected on A549 tumor growth, the combined therapy significantly slowed down the tumor growth. Interestingly, the tumor-suppressive effect of TRAIL plus rotenone was significantly attenuated by NAC (P<0.01). After experiment, tumors were removed and the caspase-3 activity in tumor cells was analyzed by flow cytometry. Consistently, the caspase-3 cleavage activities were significantly activated in A549 cells from animals challenged with TRAIL plus rotenone, meanwhile, this effect was attenuated by NAC (Figure 8e). The similar effect of rotenone also occurred in NCI-H441 xenografts tumor model (Supplementary Figure S6).
Involvement of ROS in rotenone-induced Bcl-XL degradation and PUMA expression. (a) A549 cells were preincubated with NAC at 5 mM for 30 min, further treated with rotenone (1 μM) for 8 h, western blot analysis was performed to examine Bcl-XL expression levels. GAPDH was included as a control. The quantitative result of three independent experiments showing similar results was shown on the bottom panel. The Bcl-XL expression in control cells was arbitrarily set at 1.0. ***P<0.001, as compared with control cells. (b) A549 WT or ρ° cells were treated with CHX at 5 μM to halt protein synthesis, Bcl-XL expression in cells after CHX treatment for 2 or 4 h was measured by western blot analysis. A549 WT or ρ° cells were also incubated in the absence or presence of rotenone at 1 μM to examine the effect of rotenone on the degradation of Bcl-XL. (c) A549 WT or ρ° cells were treated with rotenone at 0, 100 and 1000 nM, respectively, after treatment, western blot analysis was performed to examine PUMA protein expression, GAPDH was included as a control. The quantitative result of western blot analysis is shown on the bottom panel. (d) A549 cells were inoculated into nude mice to produce xenografts model. Animals were challenged with 100 μg TRAIL, 0.5 mg/kg rotenone or both once per 3 days for 21 consecutive days. For NAC treatment, animals were challenged with 600 mg/kg NAC once per 2 days. The tumor growth curve was plotted. **P<0.01, TRAIL+rotenone versus TRAIL+rotenone+NAC groups. (e) After experiment, the tumors were removed, and the caspase-3 activity in tumor cells was measured by flow cytometry by using FITC-conjugated caspase-3 substrate. The percentage of cells with activated caspase-3 activity in three independent experiments is indicated
Discussion
Restoration of cancer cells susceptibility to TRAIL-induced apoptosis is becoming a very useful strategy for cancer therapy.28 In this study, we provided evidence that rotenone increased the apoptosis sensitivity of NSCLC cells toward TRAIL by mechanisms involving ROS generation, p53 upregulation, Bcl-XL and c-FLIP downregulation, and death receptors upregulation. Among them, mitochondria-derived ROS had a predominant role. Although rotenone is toxic to neuron, increasing evidence also demonstrated that it was beneficial for improving inflammation,29 reducing reperfusion injury,30, 31 decreasing virus infection32 or triggering cancer cell death. We identified here another important characteristic of rotenone as a tumor sensitizer in TRAIL-based cancer therapy, which widens the application potential of rotenone in disease therapy.
As Warburg proposed the cancer ‘respiration injury’ theory, increasing evidence suggest that cancer cells may have mitochondrial dysfunction, which causes cancer cells, compared with the normal cells, are under increased generation of ROS.33 The increased ROS in cancer cells have a variety of biological effects.34, 35, 36 We found here that rotenone preferentially increased the apoptosis sensitivity of cancer cells toward TRAIL, further confirming the concept that although tumor cells have a high level of intracellular ROS, they are more sensitive than normal cells to agents that can cause further accumulation of ROS.37
Cancer cells stay in a stressful tumor microenvironment including hypoxia, low nutrient availability and immune infiltrates. These conditions, however, activate a range of stress response pathways to promote tumor survival and aggressiveness.38 In order to circumvent TRAIL-mediated apoptotic clearance, the expression levels of DR4 and DR5 in many types of cancer cells are nullified, but interestingly, they can be reactivated when cancer cells are challenged with small chemical molecules. Furthermore, those small molecules often take advantage of the stress signaling required for cancer cells survival to increase cancer cells sensitivity toward TRAIL. For example, the unfolded protein response (UPR) has an important role in cancer cells survival, SHetA2, as a small molecule, can induce UPR in NSCLC cell lines and augment TRAIL-induced apoptosis by upregulating DR5 expression in CHOP-dependent manner.39 Here, we found rotenone manipulated the oxidative stress signaling of NSCLC cells to increase their susceptibility to TRAIL. These facts suggest that cellular stress signaling not only offers opportunity for cancer cells to survive, but also renders cancer cells eligible for attack by small molecules. A possible explanation is that depending on the intensity of stress, cellular stress signaling can switch its role from prosurvival to death enhancement. As described in this study, although ROS generation in cancer cells is beneficial for survival, rotenone treatment further increased ROS production to a high level that surpasses the cell ability to eliminate them; as a result, ROS convert its role from survival to death.
Regulation of DR4 or DR5 expression is rather complex, mapping the promoter region of DR5 reveals multiple potential transcription factor binding sites including p53.40 DR5 was first cloned as a p53-responsive gene.41 In this study, we found rotenone-induced upregulation of DR5 was attenuated under the reduced expression of p53 by siRNA, suggesting that p53 may be an important trans-acting factor involved in DR5 transcription by rotenone. Notably, rotenone-induced DR4 upregulation was also mediated by p53, but no putative p53 binding sites are found on the sequence of DR4 promoter.42 We thus speculate that p53 may have an indirect stimulatory effect on DR4 transcription. Previous study suggests the sensitivity to TRAIL therapy was not closely associated with the expression levels of TRAIL receptors in some types of cancer.25 We found here that rotenone was able to increase DR4 and DR5 mRNA transcripts, protein expression and surface expression, furthermore, silencing of DR4 and DR5 expression in A549 cells led to significant suppression of cell apoptosis induced by TRAIL plus rotenone. All of these data suggest the upregulation of DR4 and DR5 receptors in NSCLC cells is important for the increased sensitivity by rotenone.
Both c-FLIPL and c-FLIPS are short-lived proteins and their expression levels are subject to regulation by ubiquitin/proteasome-mediated degradation.43 In general, c-FLIP expression is associated with chemoresistance. The downregulation of c-FLIP using antisense oligonucleotides44 or siRNA sensitizes cancer cells to chemotherapeutic agent-induced apoptosis, whereas overexpression of c-FLIP protects cells from apoptosis induced by certain cancer therapeutic agents such as etoposide and cisplatin.45, 46, 47, 48 In this study, rotenone decreased c-FLIPL and c-FLIPs expression levels in NSCLC, overexpression of c-FLIPL partially suppressed the apoptosis by TRAIL plus rotenone, rotenone-induced downregulation of c-FLIPL was abrogated by Bcl-XL overexpression, all of these facts suggest that c-FLIP downregulation involves as an important mechanism for the apoptosis enhancement by rotenone. Interestingly, c-FLIP inhibits the extrinsic pathway of cell death, whereas Bcl-XL inhibits the intrinsic pathway. Suppression of c-FLIP downregulation by Bcl-XL suggests a possible crosstalk between intrinsic pathway and extrinsic pathway in rotenone-treated NSCLC. Whether Bcl-XL regulates c-FLIP expression at transcriptional or posttranslational level needs more experiments to verify.
Reports have shown that cellular Bcl-XL protein level is regulated by phosphorylation and ubiquitin-dependent proteasomal degradation,49, 50 or by deamidation process,51 or by apoptotic cleavage.52 Here, rotenone caused ROS-dependent Bcl-XL degradation, raising a possibility that rotenone may induce the phosphorylation of Bcl-XL and the subsequent Bcl-XL ubiquitination and degradation, because previous study has demonstrated that ROS regulate phosphorylation and ubiquitination of Bcl-2 family proteins.53
The p53 is a well-known tumor suppressor, which has a key role in cellular stress response pathway. Bcl-2 family proteins are binding targets of p53, PUMA can release p53 from Bcl-XL/p53 complex and allow BAK and Bax to induce mitochondrial permeability.26, 27, 54 We found here rotenone stimulated ROS-dependent PUMA transcription, which in turn destroyed the Bcl-XL/p53 interaction. Both Bcl-XL and p53 were involved in death receptors expression. We presume that the overexpressed Bcl-XL may sequester p53 molecules and suppress its stimulatory effect on death receptors. As such, the effect of Bcl-XL on death receptors expression may be indirect. As PUMA expression was regulated by p53, it seems reasonable to conclude here that under the treatment of rotenone, PUMA, Bcl-XL and p53 form a positive feedback amplification loop to increase the apoptosis sensitivity. Mitochondria-derived ROS, however, promote the formation of this amplification loop.
MDM2 is a specific E3 ligase for p53 that can induce p53 ubiquitination and degradation.55 The strategy by disrupting the interaction between p53 and MDM2 has proven to be valuable for drug development.56 Following this fact, screening small molecules that can disrupt the interaction between p53/Bcl-XL is also promising for drug development. We noticed here that the reduced interaction between p53 and Bcl-XL by rotenone may be largely accounted by PUMA upregulation, however, the possibility that rotenone may have a direct inhibitory effect on the interaction between p53 and Bcl-XL cannot be excluded.
In this study, it seemed that rotenone can sensitize both p53 wt (A549) and p53 null (Calu-1) NSCLC cells to TRAIL-mediated apoptosis. Moreover, rotenone increased the sensitivity of other p53 null NSCLC cells, including H526, H1437, H727, H441, H1299 and H510A to TRAIL (Supplementary Figure S7). As the majority of especially metastatic NSCLC cancers harbors p53 mutations, the therapeutic use of rotenone–TRAIL combination in p53 null NSCLC cancers may be of great significance.
In sum, the results of this study suggest that targeting mitochondrial respiration by rotenone is a promising approach to overcome the resistance of NSCLC cells to TRAIL treatment.
Materials and Methods
Reagents
Human recombinant TRAIL was cloned, and expressed in Escherichia coli BL21 (DE3) cells as previously described.57 Dulbecco’s modified Eagle’s medium (DMEM), lipofectamine 2000 and Trypan blue were from Invitrogen (Carlsbad, CA, USA). Rotenone, CHX and NAC were from Sigma (St. Louis, MO, USA). SP600125, PD98058 and SB203580 were from Calbiochem (San Diego, CA, USA). H2DCFDA and DHE fluorescent dyes were obtained from Molecular Probes (Eugene, OR, USA).
Cell culture and preparation of human PBMCs
The human NSCLC cell lines including A549, H522, Calu-1 and H157 were purchased from American Type Culture Collection (ATCC, Manassas, VA, USA), and maintained in DMEM, which was supplemented with 10% heat-inactivated FBS, at 37 °C in a humidified incubator with 5% CO2. Human PBMCs were isolated by Ficoll/Paque (Pharmacia, Uppsala, Sweden) density gradient centrifugation of heparinized blood obtained from healthy adult donors. Cells were harvested from the inter-phase layer, washed in PBS twice and then resuspended in RPMI 1640 medium containing 10% (v/v) FBS, 10 mM glutamine and 50 U/ml penicillin/streptomycin. Cells were cultured in six-well tissue culture plates. Cell viability was assessed by Trypan blue.
Plasmids
The expressing plasmids including FLAG-tagged Bcl-XL, HA-tagged p53 and c-FLIPL were all generated in our lab by RT-PCR and subcloning method. Plasmid DNA was purified by plasmid DNA Maxi Purification Kit (Promega, Madison, WI, USA). The purified DNA was diluted to 1 mg/ml and stored at −20 °C until use. The plasmids were introduced into cells by PolyJet transfection reagent based on manufacturer’s instruction.
RNA interference
A549 cells were transfected with the siRNA for human TRAIL receptor DR4 and DR5,58 or for human p53.29 The transfection of siRNA was conducted in a 24-well plate using lipofectamine 2000. Forty-eight hours after the transfection, cells were treated with rotenone alone, TRAIL alone or their combination. Gene silencing effect was evaluated by western blot analysis and apoptosis was measured by annexin V/PI staining method.
Semiquantitative RT-PCR
Total RNA was isolated using TRIZOL agent and chloroform extraction according to the seller’s RNA extraction protocol. The PCR primers used to amplify DR4, DR5, PUMA, p21 and GAPDH were listed as follows: DR4: sense, 5′-TTGTGTCCACCAGGATCTCA-3′, antisense, 5′-GTCACTCCAGGGCGTACAAT-3′; DR5: sense, 5′-ACTCCTGGAATGACTACCTG-3′, antisense, 5′-ATCCCAAGTGAACTTGAGCC-3′; p21: sense, 5′-GGAACTTCGACTTTGTCACC-3′, antisense, 5′-AAGGCAGAAGATGTAGAGCG-3′; GAPDH: sense, 5′-CACCATCTTCCAGGAGCGAG-3′, antisense, 5′-GCAGGAGGCATTGCTGAT-3′; PUMA: sense, 5′-ACCTCAACGCACAGTACGAG-3′, antisense, 5′-GTATGCTACATGGTGCAGAG-3′.
Western blot analysis
The immunoblot analysis was performed as previously described.59 Briefly, A549 cells after treatment were harvested and lysed, and the cleared lysate was separated by SDS-PAGE. After electrophoresis, proteins were transferred to PVDF membranes. The membranes were first hybridized with primary antibodies, and then with a horseradish peroxidase-conjugated anti-mouse, or anti-rabbit IgG secondary Ab (Sigma). FLAG, HA, GAPDH and tubulin antibodies were from Santa Cruz Biotechnology (Santa Cruz, CA, USA). Abs to p53, phosphor-p53, PUMA, c-FLIPL, c-FLIPs and Bcl-XL were from Cell Signaling Technology (Beverly, MA, USA). The immune blots were developed using enhanced chemiluminescence system (Amersham Pharmacia Biotech, Amersham, UK).
Measurement of caspase-3, -8 and -9 activities by flow cytometry
The caspase-3, -8 and -9 activities in NSCLC cells were determined by using CaspGLOW Green caspase-3, -8 and -9 staining kits (Biovision, Palo Alto, CA, USA) based on manufacturer’s instructions.
Measurement of surface DR4 and DR5 expression levels by flow cytometry
A549 cells (2 × 106) before or after treatment were harvested and washed with ice-cold PBS. For measurement of antibody binding, cells were probed with primary antibody for 1 h. After incubation, cells were washed and resuspended in a 100 μl binding buffer containing 2 μl FITC-conjugated secondary IgG for another 1 h. Cells were then harvested, washed and resuspended in a 400 μl ice-cold PBS and analyzed with flow cytometry (Becton Dickinson, Franklin Lakes, NJ, USA). Immediately before flow cytometric analysis, PI (1 μg/ml) was added in each sample. Cells were divided into two population based on their permeability to PI, PI permeable or PI impermeable cells, which were individually gated to measure the mean FITC fluorescence in FL1 channel.
Generation of stable cell lines transfected with human catalase or MnSOD cDNA
The full-length cDNA of human catalase and MnSOD were amplified by using RT-PCR reaction and cloned in frame into bicistronic pIRES-EGFP eukaryotic expression vector containing the gene encoding GFP. These constructs (pIRES-EGFP-catalase, pIRES-EGFP-MnSOD) were stably transfected into A549 cells by lipofectamine 2000. Transfected clones, derived from single colonies, were selected by limiting dilution in DMEM medium containing G418 (1.5 mg/ml) and screened for EGFP expression by flow cytometry, and further screened for catalase or MnSOD overexpression by western blot analysis. Control cell lines were generated by stably transfecting A549 cells with the pIRES-EGFP vector.
Generation of A549 ρ° cells
A549 cell lacking mitochondrial DNA (ρ°) were generated by growing A549 cells in DMEM medium supplied with 10% fetal calf serum, 2 mM L-glutamine, 1 mM pyruvate, 50 μg/ml uridine, 25 mM glucose and 50 ng/ml EB for 5–6 weeks.60 After selection, the cells were grown in the same medium without EB. Oxygen consumption was measured with a Clark-type oxygen electrode, and no oxygen uptake was observed for A549 ρ° cells.
Measurement of intracellular H2O2 and O2.−
Generation of intracellular H2O2 was measured using H2DCFDA upon oxidation to the fluorescent derivative 2’–7’-dichlorofluorescin (DCF) by reactions with H2O2. Following treatment, cells were collected and resuspended in 500 μl DMEM containing 2% FBS and 10 μM H2DCFDA for 20 min at 37 °C. Subsequently, cells were washed with PBS and analyzed with flow cytometry (Becton Dickinson, Oxford, UK) with excitation set at 488 nm and emission at 530 nm. Immediately before flow cytometry analysis, PI (1 μg/ml) was added to exclude the non-viable cells. For measurement of intracellular O2.−, cells after treatment were loaded with 5 μM DHE, analyzed with flow cytometry in FL3 channel.23
Assessment of cell apoptosis by annexin V and PI double staining
The annexin V/PI double staining method was performed based on previously described method.59 In brief, A549 cells after treatment were stained with EGFP-tagged annexin V (1 : 2000), PI was added immediately before analysis by flow cytometry. EGFP fluorescence emission was measured in the FL1 channel and PI fluorescence was measured in the FL3 channel after cell doublets were excluded by pulse processing. Five thousand cells were counted per sample. The data were analyzed with BD CellQuest software (Franklin Lakes, NJ, USA).
Xenograft tumor models and in vivo delivery of drugs
A549 cells were grown in DMEM medium with 10% heat-inactivated FBS and 2 mM L-glutamine. Cells were suspended in calcium and magnesium-free PBS (pH 7.35–7.45), and implanted subcutaneously in the right flank of each mouse. The cell concentrations for implantation were 5 × 106 cells/0.2 ml per mouse. Once tumor volume reached >50 mm3, animals were randomized so that all groups had similar starting mean tumor volumes. Tumor measurements were taken three times per week. Animals were individually monitored throughout the experiment. Tumor volume (mm3) was calculated using the ellipsoid formula: [D*(d2)]2, where ‘D’ represents the largest diameter of the tumor, and ‘d’ represents the smallest diameter. After establishment of A549 xenografts, animals were injected intraperitoneally with 100 μg TRAIL, or rotenone at 0.5 mg/kg, or both once per 3 days for 21 consecutive days. For NAC treatment, animals were challenged with NAC at 600 mg/kg once per 2 days. At the end of the experiments, animals were killed and the tumors were removed, homogenized and stained with 1–2 μl FITC-DEVD-FMK for detection of caspase-3 activities.
Statistical analysis
The results were expressed as mean±S.E.M. The statistical analysis involving two groups was performed by means of Student's t-test. Analysis of variance followed by Dunnett’s multiple comparison tests was used to compare more than two groups. All data were processed with SPSS 10.0 software (SPSS, Chicago, IL, USA).
Abbreviations
- c-FLIP:
-
cellular FLICE-like inhibitory protein
- DHE:
-
dihydroethidium
- DISC:
-
death-inducing signaling complex
- DPI:
-
diphenylene iodonium
- DR4/DR5:
-
death receptor 4/5
- EB:
-
ethidium bromide
- FADD:
-
Fas-associated protein with death domain
- MnSOD:
-
manganese superoxide
- NAC:
-
N-acetylcysteine
- NSCLC:
-
non-small-cell lung carcinoma
- PBMC:
-
peripheral blood mononuclear cells
- ROS:
-
reactive oxygen species
- TRAIL:
-
tumor necrosis factor-related apoptosis-inducing ligand
- UPR:
-
unfolded protein response
References
Kelley SK, Ashkenazi A . Targeting death receptors in cancer with Apo2L/TRAIL. Curr Opin Pharmacol 2004; 4: 333–339.
Stegehuis JH, de Wilt LH, de Vries EG, Groen HJ, de Jong S, Kruyt FA . TRAIL receptor targeting therapies for non-small cell lung cancer: current status and perspectives. Drug Resist Updates 2010; 13: 2–15.
Thorburn A, Behbakht K, Ford H . TRAIL receptor-targeted therapeutics: resistance mechanisms and strategies to avoid them. Drug Resist Updates 2008; 11: 17–24.
Siegel R, Ma J, Zou Z, Jemal A . Cancer statistics, 2014. CA Cancer J Clin 2014; 64: 9–29.
Oldenhuis CN, Stegehuis JH, Walenkamp AM, de Jong S, de Vries EG . Targeting TRAIL death receptors. Curr Opin Pharmacol 2008; 8: 433–439.
Lavrik I, Golks A, Krammer PH . Death receptor signaling. J Cell Sci 2005; 118: 265–267.
Jin Z, El-Deiry WS . Overview of cell death signaling pathways. Cancer Biol Ther 2005; 4: 139–163.
Wang S, El-Deiry WS . TRAIL and apoptosis induction by TNF-family death receptors. Oncogene 2003; 22: 8628–8633.
Bodmer JL, Schneider P, Tschopp J . The molecular architecture of the TNF superfamily. Trends Biochem Sci 2002; 27: 19–26.
Schneider P, Bodmer JL, Thome M, Hofmann K, Holler N, Tschopp J . Characterization of two receptors for TRAIL. FEBS Lett 1997; 416: 329–334.
Ozoren N, El-Deiry WS . Defining characteristics of types I and II apoptotic cells in response to TRAIL. Neoplasia 2002; 4: 551–557.
Eid MA, Lewis RW, Abdel-Mageed AB, Kumar MV . Reduced response of prostate cancer cells to TRAIL is modulated by NFkappaB-mediated inhibition of caspases and Bid activation. Int J Oncol 2002; 21: 111–117.
Martelli AM, Tazzari PL, Tabellini G, Bortul R, Billi AM, Manzoli L et al. A new selective AKT pharmacological inhibitor reduces resistance to chemotherapeutic drugs, TRAIL, all-trans-retinoic acid, and ionizing radiation of human leukemia cells. Leukemia 2003; 17: 1794–1805.
Armstrong JS . Mitochondria: a target for cancer therapy. Br J Pharmacol 2006; 147: 239–248.
Decaudin D, Marzo I, Brenner C, Kroemer G . Mitochondria in chemotherapy-induced apoptosis: a prospective novel target of cancer therapy (review). Int J Oncol 1998; 12: 141–152.
Loureiro R, Mesquita KA, Oliveira PJ, Vega-Naredo I . Mitochondria in cancer stem cells: a target for therapy. Recent Pat Endocr Metab Immune Drug Discov 2013; 7: 102–114.
Roy D, Kabiraj P, Pal R . EF24 prevents rotenone-induced estrogenic status alteration in breast cancer. Cell Biol Int 2014; 38: 511–519.
Goncalves AP, Videira A, Maximo V, Soares P . Synergistic growth inhibition of cancer cells harboring the RET/PTC1 oncogene by staurosporine and rotenone involves enhanced cell death. J Biosci 2011; 36: 639–648.
Goncalves AP, Maximo V, Lima J, Singh KK, Soares P, Videira A . Involvement of p53 in cell death following cell cycle arrest and mitotic catastrophe induced by rotenone. Biochimica et Biophysica Acta 2011; 1813: 492–499.
Deng YT, Huang HC, Lin JK . Rotenone induces apoptosis in MCF-7 human breast cancer cell-mediated ROS through JNK and p38 signaling. Mol Carcinogenesis 2010; 49: 141–151.
Lee J, Huang MS, Yang IC, Lai TC, Wang JL, Pang VF et al. Essential roles of caspases and their upstream regulators in rotenone-induced apoptosis. Biochem Biophys Res Commun 2008; 371: 33–38.
Pelicano H, Feng L, Zhou Y, Carew JS, Hileman EO, Plunkett W et al. Inhibition of mitochondrial respiration: a novel strategy to enhance drug-induced apoptosis in human leukemia cells by a reactive oxygen species-mediated mechanism. J Biol Chem 2003; 278: 37832–37839.
Yin W, Li X, Feng S, Cheng W, Tang B, Shi YL et al. Plasma membrane depolarization and Na,K-ATPase impairment induced by mitochondrial toxins augment leukemia cell apoptosis via a novel mitochondrial amplification mechanism. Biochem Pharmacol 2009; 78: 191–202.
Liu X, Yue P, Zhou Z, Khuri FR, Sun SY . Death receptor regulation and celecoxib-induced apoptosis in human lung cancer cells. J Natl Cancer Inst 2004; 96: 1769–1780.
Frese S, Pirnia F, Miescher D, Krajewski S, Borner MM, Reed JC et al. PG490-mediated sensitization of lung cancer cells to Apo2L/TRAIL-induced apoptosis requires activation of ERK2. Oncogene 2003; 22: 5427–5435.
Bharatham N, Chi SW, Yoon HS . Molecular basis of Bcl-X(L)-p53 interaction: insights from molecular dynamics simulations. PloS One 2011; 6: e26014.
Follis AV, Chipuk JE, Fisher JC, Yun MK, Grace CR, Nourse A et al. PUMA binding induces partial unfolding within BCL-xL to disrupt p53 binding and promote apoptosis. Nat Chem Biol 2013; 9: 163–168.
Prasad S, Kim JH, Gupta SC, Aggarwal BB . Targeting death receptors for TRAIL by agents designed by Mother Nature. Trends Pharmacol Sci 2014; 35: 520–536.
Zhang LH, Jia YL, Lin XX, Zhang HQ, Dong XW, Zhao JM et al. AD-1, a novel ginsenoside derivative, shows anti-lung cancer activity via activation of p38 MAPK pathway and generation of reactive oxygen species. Biochimica et Biophysica Acta 2013; 1830: 4148–4159.
Lin FL, Hsu JL, Chou CH, Wu WJ, Chang CI, Liu HJ . Activation of p38 MAPK by damnacanthal mediates apoptosis in SKHep 1 cells through the DR5/TRAIL and TNFR1/TNF-alpha and p53 pathways. Eur J Pharmacol 2011; 650: 120–129.
Sai Y, Chen J, Wu Q, Liu H, Zhao J, Dong Z . Phosphorylated-ERK 1/2 and neuronal degeneration induced by rotenone in the hippocampus neurons. Environ Toxicol Pharmacol 2009; 27: 366–372.
Wender PA, Cribbs CM, Koehler KF, Sharkey NA, Herald CL, Kamano Y et al. Modeling of the bryostatins to the phorbol ester pharmacophore on protein kinase C. Proc Natl Acad Sci USA 1988; 85: 7197–7201.
Pelicano H, Carney D, Huang P . ROS stress in cancer cells and therapeutic implications. Drug Resist Updates 2004; 7: 97–110.
Hileman EA, Achanta G, Huang P . Superoxide dismutase: an emerging target for cancer therapeutics. Expert Opin Ther Targets 2001; 5: 697–710.
Trachootham D, Alexandre J, Huang P . Targeting cancer cells by ROS-mediated mechanisms: a radical therapeutic approach? Nat Rev Drug Discov 2009; 8: 579–591.
Weinberg F, Hamanaka R, Wheaton WW, Weinberg S, Joseph J, Lopez M et al. Mitochondrial metabolism and ROS generation are essential for Kras-mediated tumorigenicity. Proc Natl Acad Sci USA 2010; 107: 8788–8793.
Zhou Y, Hileman EO, Plunkett W, Keating MJ, Huang P . Free radical stress in chronic lymphocytic leukemia cells and its role in cellular sensitivity to ROS-generating anticancer agents. Blood 2003; 101: 4098–4104.
Fulda S, Gorman AM, Hori O, Samali A . Cellular stress responses: cell survival and cell death. Int J Cell Biol 2010; 2010: 214074.
Lin YD, Chen S, Yue P, Zou W, Benbrook DM, Liu S et al. CAAT/enhancer binding protein homologous protein-dependent death receptor 5 induction is a major component of SHetA2-induced apoptosis in lung cancer cells. Cancer Res 2008; 68: 5335–5344.
Takimoto R, El-Deiry WS . Wild-type p53 transactivates the KILLER/DR5 gene through an intronic sequence-specific DNA-binding site. Oncogene 2000; 19: 1735–1743.
Wu GS, Burns TF, McDonald ER 3rd, Jiang W, Meng R, Krantz ID et al. KILLER/DR5 is a DNA damage-inducible p53-regulated death receptor gene. Nat Genet 1997; 17: 141–143.
Surget S, Chiron D, Gomez-Bougie P, Descamps G, Menoret E, Bataille R et al. Cell death via DR5, but not DR4, is regulated by p53 in myeloma cells. Cancer Res 2012; 72: 4562–4573.
Chang L, Kamata H, Solinas G, Luo JL, Maeda S, Venuprasad K et al. The E3 ubiquitin ligase itch couples JNK activation to TNFalpha-induced cell death by inducing c-FLIP(L) turnover. Cell 2006; 124: 601–613.
Lin Y, Liu X, Yue P, Benbrook DM, Berlin KD, Khuri FR et al. Involvement of c-FLIP and survivin down-regulation in flexible heteroarotinoid-induced apoptosis and enhancement of TRAIL-initiated apoptosis in lung cancer cells. Mol Cancer Ther 2008; 7: 3556–3565.
Liu X, Yue P, Schonthal AH, Khuri FR, Sun SY . Cellular FLICE-inhibitory protein down-regulation contributes to celecoxib-induced apoptosis in human lung cancer cells. Cancer Res 2006; 66: 11115–11119.
Rogers KM, Thomas M, Galligan L, Wilson TR, Allen WL, Sakai H et al. Cellular FLICE-inhibitory protein regulates chemotherapy-induced apoptosis in breast cancer cells. Mol Cancer Ther 2007; 6: 1544–1551.
Longley DB, Wilson TR, McEwan M, Allen WL, McDermott U, Galligan L et al. c-FLIP inhibits chemotherapy-induced colorectal cancer cell death. Oncogene 2006; 25: 838–848.
Abedini MR, Qiu Q, Yan X, Tsang BK . Possible role of FLICE-like inhibitory protein (FLIP) in chemoresistant ovarian cancer cells in vitro. Oncogene 2004; 23: 6997–7004.
Park K, Lee JH . Bcl-XL protein is markedly decreased in UVB-irradiated basal cell carcinoma cell lines through proteasome-mediated degradation. Oncol Rep 2009; 21: 689–692.
Ji L, Chen Y, Liu T, Wang Z . Involvement of Bcl-xL degradation and mitochondrial-mediated apoptotic pathway in pyrrolizidine alkaloids-induced apoptosis in hepatocytes. Toxicol Appl Pharmacol 2008; 231: 393–400.
Dho SH, Deverman BE, Lapid C, Manson SR, Gan L, Riehm JJ et al. Control of cellular Bcl-xL levels by deamidation-regulated degradation. PLoS Biol 2013; 11: e1001588.
Emanuele S, D'Anneo A, Bellavia G, Vassallo B, Lauricella M, De Blasio A et al. Sodium butyrate induces apoptosis in human hepatoma cells by a mitochondria/caspase pathway, associated with degradation of beta-catenin, pRb and Bcl-XL. Eur J Cancer 2004; 40: 1441–1452.
Li D, Ueta E, Kimura T, Yamamoto T, Osaki T . Reactive oxygen species (ROS) control the expression of Bcl-2 family proteins by regulating their phosphorylation and ubiquitination. Cancer Sci 2004; 95: 644–650.
Yu J, Zhang L . PUMA, a potent killer with or without p53. Oncogene 2008; 27 Suppl 1: S71–S83.
Haupt Y, Maya R, Kazaz A, Oren M . Mdm2 promotes the rapid degradation of p53. Nature 1997; 387: 296–299.
Devine T, Dai MS . Targeting the ubiquitin-mediated proteasome degradation of p53 for cancer therapy. Curr Pharm Design 2013; 19: 3248–3262.
Cao L, Du P, Jiang SH, Jin GH, Huang QL, Hua ZC . Enhancement of antitumor properties of TRAIL by targeted delivery to the tumor neovasculature. Mol Cancer Ther 2008; 7: 851–861.
Frese S, Frese-Schaper M, Andres AC, Miescher D, Zumkehr B, Schmid RA . Cardiac glycosides initiate Apo2L/TRAIL-induced apoptosis in non-small cell lung cancer cells by up-regulation of death receptors 4 and 5. Cancer Res 2006; 66: 5867–5874.
Yin W, Cheng W, Shen W, Shu L, Zhao J, Zhang J et al. Impairment of Na(+),K(+)-ATPase in CD95(APO-1)-induced human T-cell leukemia cell apoptosis mediated by glutathione depletion and generation of hydrogen peroxide. Leukemia 2007; 21: 1669–1678.
King MP, Attardi G . Isolation of human cell lines lacking mitochondrial DNA. Methods Enzymol 1996; 264: 304–313.
Acknowledgements
This project was sponsored by Natural Science Fund of China (31071250, 81473293, 81171843 and J1103521), open project of National Key Lab of Natural Medicines (SKLNMKF201303), open project of National Key Lab of Drug Discovery (SIMM1106KF-01), program for New Century Excellent Talents in University (NECT-10-0187), the Fundamental Research Funds for the Central Universities (14360013, 14330006, 14330029 and 14330040), Jiangsu ‘333’ project and Nanjing ‘321’ talents project to WY. Part of this work was initiated in Dr. Hua Zi-Chun’s lab.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by M Agostini
Supplementary Information accompanies this paper on Cell Death and Disease website
Supplementary information
Rights and permissions
Cell Death and Disease is an open-access journal published by Nature Publishing Group. This work is licensed under a Creative Commons Attribution 4.0 International Licence. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons licence, users will need to obtain permission from the licence holder to reproduce the material. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0
About this article
Cite this article
Shi, YL., Feng, S., Chen, W. et al. Mitochondrial inhibitor sensitizes non-small-cell lung carcinoma cells to TRAIL-induced apoptosis by reactive oxygen species and Bcl-XL/p53-mediated amplification mechanisms. Cell Death Dis 5, e1579 (2014). https://doi.org/10.1038/cddis.2014.547
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2014.547
This article is cited by
-
Mechanistic Investigations of the Mitochondrial Complex I Inhibitor Rotenone in the Context of Pharmacological and Safety Evaluation
Scientific Reports (2017)
-
Reversal of the Apoptotic Resistance of Non-Small-Cell Lung Carcinoma towards TRAIL by Natural Product Toosendanin
Scientific Reports (2017)
-
ROS homeostasis and metabolism: a dangerous liason in cancer cells
Cell Death & Disease (2016)
-
Migration and invasion of drug-resistant lung adenocarcinoma cells are dependent on mitochondrial activity
Experimental & Molecular Medicine (2016)