Abstract
Osteosarcoma is a common primary bone tumor in children and adolescents. The drug resistance of osteosarcoma leads to high lethality. Macrophage migration inhibitory factor (MIF) is an inflammation-related cytokine implicated in the chemoresistance of breast cancer. In this study, we isolated a novel androstenedione derivative identified as 3,4-dihydroxy-9,10-secoandrosta-1,3,5,7-tetraene-9,17-dione (DSTD). DSTD could inhibit MIF expression in MG-63 and U2OS cells. The inhibition of MIF by DSTD promoted autophagy by inducing Bcl-2 downregulation and the translocation of HMGB1. N-acetyl-L-cysteine (NAC) and 3-methyladenine (3-MA) attenuated DSTD-induced autophagy but promoted cell death, suggesting that DSTD induced ROS-mediated autophagy to rescue cell death. However, in the presence of chemotherapy drugs, DSTD enhanced the chemosensitivity by decreasing the HMGB1 level. Our data suggest MIF inhibition as a therapeutic strategy for overcoming drug resistance in osteosarcoma.
Similar content being viewed by others
Main
Osteosarcoma, a common primary bone tumor in children and adolescents, is prone to early metastasis through blood.1 Treatment with a combination of surgery and aggressive adjuvant chemotherapy has improved the survival rate of osteosarcoma patients. The 5-year-survival rates of non-metastatic patients have reached a plateau of approximately 70%.2, 3 However, patients with poor responses to chemotherapeutics will undergo local recurrence and metastasis, which reduce the 5-year-survival rates to only 20% despite additional doses or drugs.4, 5 Drug resistance is responsible for the poor prognosis. Attenuating chemoresistance facilitates better treatment of osteosarcoma.6, 7 Novel treatment strategies that combine anticancer drugs with adjuvant agents could improve the antitumor effects.8, 9
In the 1960s, macrophage migration inhibitory factor (MIF) was identified as a pluripotent protein that modulates inflammation.10 Increasing evidence suggests that inflammation is closely related to tumorigenesis.11 MIF plays a bridging role between inflammation and tumorigenesis.12, 13, 14 MIF triggers the activation of the MAPK and PI3K pathways by binding its membrane receptor CD74, resulting in the inhibition of cell apoptosis.15 Recently, MIF was demonstrated to be involved in cell proliferation, differentiation, angiogenesis and tumorigenesis.16, 17, 18 Some evidence has indicated that MIF is abundantly expressed in various cancers and is significantly associated with tumor invasion and metastasis.19, 20, 21 MIF has been well established to be involved in the development of glioblastoma,22 breast cancer,23 bladder cancer24 and colon cancer.20, 25 MIF was also upregulated in osteosarcoma.26, 27 The knockdown of MIF blocked osteosarcoma cell proliferation and invasion.26 However, the effect of MIF on drug resistance in osteosarcoma has not yet been investigated. Wu et al. 23 have revealed that MIF knockdown promoted chemosensitivity by inducing autophagy in breast cancer. In contrast, autophagy reportedly contributed to chemoresistance in osteosarcoma.6 These controversial results prompted us to confirm the role of MIF in drug resistance in osteosarcoma.
In this study, we isolated a novel androstenedione derivative identified as 3,4-dihydroxy-9,10-secoandrosta-1,3,5,7-tetraene-9,17-dione (DSTD). DSTD could inhibit MIF expression in MG-63 and U2OS cells. Both N-acetyl-L-cysteine (NAC) and 3-methyladenine (3-MA) attenuated DSTD-induced autophagy but promoted cell death, suggesting that DSTD induced reactive oxygen species (ROS)-mediated autophagy to rescue cell death. Furthermore, MIF inhibition by DSTD enhances chemosensitivity by downregulating HMGB1 in osteosarcoma cells. Our data suggest MIF inhibition as a therapeutic strategy for overcoming drug resistance in osteosarcoma.
Results
Structural elucidation
A novel compound was isolated from the microbial metabolites of androstadienone produced by Bordetella sp. B4 (Bordetella sp. B4 was kindly provided by Center Laboratory, Provincial Hospital Affiliated to Shandong University). As shown in Figures 1a and b, androstadienones, including androst-4-ene-3,17-dione and androst-1,4-diene-3,17-dione, were transformed to three products. Among these products, 9α-hydroxyandrosta-1,4-diene-3,17-dione (9-OH-ADD) and 4-hydroxy-9,10-secoandrosta-1,3,5-triene-9,17-dione (HSTD) have been reported in previous studies.28, 29 The novel product was characterized as follows: Rf value=0.95 (dichloromethane: petroleum ether: ethyl acetate=6:3:1), 1H NMR (600 MHz, CDCl3, as shown in Supplementary Figure 4S) 1.19 (3H, s, -methyl), 1.75–2.09 (6H, m, skeleton protons), 2.28 (3H, s, -methyl), 2.46–2.75 (4H, m, skeleton protons), 5.41(2H, s, -hydroxy), 6.61 (1H, dd, -ethylene), 6.82 (1H, d, -benzene), 7.01 (1H, d,-ethylene), 7.28 (1H, s, -benzene). 13C NMR (600 MHz, CDCl3, as shown in Supplementary Figure 3S) 218.4 (-carbonyl), 210.8 (-carbonyl), 153.9 (-benzene C3-OH), 141.9 (-benzene C4-OH), 139.1 (-benzene C6-C=C-), 131.2 (-benzene C1-CH3), 127.8 (-ethylene), 115.7 (-ethylene),115.3 (-benzene C2-H), 112.6(-benzene C5-H), 49.6 (-cyclohexane C-CH=CH-), 47.3 (-cyclohexane C-C(=O)-C-), 37.5 (-cyclohexane, C-H), 36.2 (-cyclohexane, C-H), 31.3 (-cyclopentane, C-H), 26.6 (-cyclohexane, C-H), 22.4(-cyclopentane, C-H), 18.3 (-methyl), 13.1(-methyl). Ms (EI) m/z (%): 315 (M+H+, 53), 332 (M+NH4+, 91), 337 (M+Na+, 67). According to the above data, the compound was characterized as DSTD (Figure 1b).
Effect of DSTD on cell viability, cell cytotoxicity and MIF expression in osteosarcoma cells. (a) TLC analysis of androstenedione derivatives transformed by Bordetella sp. B4, lane1-5, androstenediones were transformed for 12, 24, 4872 h, respectively; lane 6, purified HSTD; lane 7, purified DSTD. (b) Transformation pathway of androstenediones by Bordetella sp. B4. (c) Effect of DSTD on viability and cytotoxicity of MG-63 and U2OS osteosarcoma cells. Cells were treated with indicated concentration of DSTD (dissolved in DMSO) for 24 h. Cell viability and cytotoxicity were determined respectively using MTT assay and LDH release assay. (d) Effect of DSTD on MIF expression. Cells were treated with 100 μM DSTD for 24 h. MIF protein level and mRNA level were analyzed respectively by western blot and real-time PCR. These experiments were repeated at least three times. *P<0.01 versus control group
The purified DSTD was dissolved in DMSO to prepare 100 mM stock solution before the experiments. For all experiments using DSTD, the effects of DSTD were corrected using the control with the equal volume of DMSO.
DSTD decreases cell viability and MIF level in osteosarcoma cells
We examined the effect of DSTD on the viability of the osteosarcoma cells lines MG-63 and U2OS. As shown in Figure 1c, the results of MTT assay and lactate dehydrogenase (LDH) release assay indicated that exposure to 100 μM DSTD for 24 h did not significantly affect the cell viabilities and cytotoxicity. Given the immunomodulatory properties of steroidal medicines, we further assayed the potential role of DSTD in the regulation of MIF expression. Figure 1d illustrated that exposing cells to 100 μM DSTD for 24 h resulted in a significant reduction of MIF protein level. The results of real-time PCR revealed that MIF mRNA was decreased after treatment with 100 μM DSTD for 24 h (Figure 1d).
DSTD induced autophagy by inhibiting MIF expression
We assessed the effect of MIF inhibition on signal transduction in osteosarcoma cells. As shown in Figure 2a, the target-specific shRNA of MIF significantly suppressed MIF expression. MIF knockdown led to a large decrease of Bcl-2 and an increase of Bax. MIF knockdown also inhibited ERK phosphorylation. Intriguingly, MIF knockdown decreased the nucleic HMGB1 level but increased the cytosolic HMGB1 level, which suggested that MIF downregulation led to the translocation of HMGB1 from the nucleus to the cytoplasm. DSTD was as effective in inhibiting MIF-dependent signal transduction as MIF knockdown by shRNA interference. Because Bcl-2, ERK and HMGB1 are closely related to autophagy,30, 31 we further tested the ability of DSTD to induce autophagy. Because microtubule-associated protein light chain 3 (LC3) is an important constituent of the autophagosome, LC3 is widely used to determine autophagy.32 A significant conversion of LC3-I to LC3-II was detected after exposing MG-63 and U2OS cells to 100 μM DSTD for 24 h (Figure 2b). The accumulation of LC3-II positively correlated with the treatment time, which suggested that DSTD induced autophagy in a time-dependent manner. DSTD also induced autophagy in a dose-dependent manner (Figure 3a). Bcl-2 and Bax are well known to be closely associated with the generation of ROS.33 Moreover, ROS has been demonstrated to induce the cytosolic translocation of HMGB1.34 Combined with our results, the inhibition of MIF was speculated to be able to induce ROS-mediated autophagy.
Effect of blocking MIF on signaling components and autophagy. (a) Cells were treated with 100 μM DSTD for 24 h or transfected with shRNA targeted for MIF for 48 h. The cytoplasmic and nucleic fractions were separated. Bcl-2, Bax, p-ERK, HMGB1 and Histone H3 were analyzed by western blot. (b) DSTD induced time-dependent autophagy. Cells were treated with 100 μM DSTD for indicated time. LC3 conversion was detected by western blot. These experiments were repeated at least three times
DSTD induces autophagy in U2OS and MG-63 osteosarcoma cells. (a) DSTD induced dose-dependent autophagy. Cells were treated with the indicated concentration of DSTD for 24 h. MIF, Bcl-2, Bax, p-ERK, HMGB1 and LC3 were analyzed by western blot. (b) DSTD induced ROS-mediated autophagy in time-dependent manners. Cells were treated with 100 μM DSTD for indicated time. To determine the effect of NAC on autophagy, after treated with 100 μM DSTD for 24 h, cells continued to be treated with 100 μM DSTD combined with 15 mM NAC (pH 7.4) for 24 h. The cytoplasmic and nucleic fractions were separated. MIF, Bcl-2, Bax, HMGB1, p-ERK, PARP and LC3 were analyzed by western blot. (c) Viability of U2OS and MG-63 cells treated with 100 μM DSTD in the presence or absence of NAC for the indicated time. (d) Cytotoxicity induced by 100 μM DSTD in the presence or absence of NAC for the indicated time. Values are means of at least three independent experiments. *P<0.05 versus control group; #P<0.05 versus group treated with DSTD for 48 h; **P<0.01 versus control group
DSTD induced ROS-mediated autophagy
Because DSTD induced autophagy in a time- and dose-dependent manner, we examined the ability of DSTD to regulate multiple signal molecules in a time- and dose-dependent manner. As shown in Figures 3a and b, the inhibition of MIF by DSTD was dose- and time-dependent in U2OS and MG-63 cells. Increasing the dose of DSTD gradually decreased the Bcl-2 level and increased the Bax level. The accumulation of LC3-II was also dose-dependent. In contrast, DSTD slightly activated ERK phosphorylations and did not significantly influence the total HMGB1 level at concentration of 50 μM for 24 h (Figure 3a). When the cells were exposed to 100 μM DSTD for 24 h, both the phosphorylation of ERK and total HMGB1 level decreased. When the cells were treated with 100 μM DSTD, DSTD induced time-dependent signal transduction. As shown in Figure 3b, as the treatment time increased, DSTD gradually inhibited the expression of Bcl-2 and promoted Bax expression. Meanwhile, DSTD induced the accumulation of PARP and formed cleaved PARP. Interestingly, ERK phosphorylation and nucleic HMGB1 were enhanced transiently at 12 h after DSTD treatment, and then were decreased from 24 h to 48 h after DSTD treatment. Exposure to DSTD decreased total HMGB1 and increased cytoplasmic HMGB1 in a time-dependent manner. All of these results are the characteristics of ROS generation. To confirm that DSTD promoted ROS generation, we also tested the effect of NAC on the DSTD-induced signaling response. As shown in Figure 3b, NAC co-treatment reversed DSTD-induced signaling responses. Notably, DSTD induced the accumulation of PARP and formed cleaved PARP, which could not be reversed by NAC treatment. Consistent with this result, NAC treatment significantly decreased the cell viability (Figure 3c), increased cytotoxicity (Figure 3d) and promoted cell death (Figure 4b). These results suggested that the inhibition of MIF by DSTD induced ROS-mediated autophagy to delay cell death.
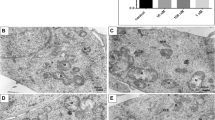
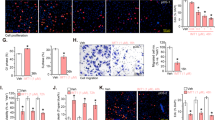
DSTD induces ROS generation, LC3 puncta formation and apoptosis in U2OS and MG-63 osteosarcoma cells. (a) Effect of DSTD on ROS generation and LC3 puncta formation. Cells were treated with 100 μM DSTD for 24 h in the presence or absence of 15 mM NAC. After treatment, cells were stained by DCF-DA or LC3 antibody to determine ROS generation and LC3 puncta formation, respectively. (b) Effect of DSTD on cell apoptosis. After treatment, apoptosis was determined by flow cytometry
To further confirm that DSTD triggered ROS generation, we used an oxidation-sensitive fluorescent reagent (DCF-DA) to track ROS generation in U2OS and MG-63 cells. DSTD induced a dose-related increase in the ROS level, which was reversed by NAC treatment (Figure 4a). We also examined DSTD-induced autophagy by monitoring the formation of LC3 puncta using immunofluorescent staining. The results indicated that the formation of LC3 puncta was distinct and time-dependent (Figure 4a). Consistent with the results in Figure 3b, NAC co-treatment attenuated DSTD-induced autophagy (Figure 4a).
To examine whether DSTD-induced autophagy delayed cell death, we investigated effect of 3-MA (an inhibitor of type III phosphatidylinositol 3-kinase that needed for initiation of autophagy) on cell viability and cytotoxicity. The results demonstrated that 3-MA reduced the level of LC3B-II induced by DSTD in U2OS (Figure 5a) and MG-63 cells (Figure 5b). Pharmacological inhibition of autophagy by 3-MA also suppressed cell viability and increased cell cytotoxicity (Figures 5c and d), suggesting that DSTD-induced autophagy did contribute to delay cell death.
DSTD-mediated autophagy delays cell death. The U2OS (a) and MG-63(b) cells were pretreated with 1 mM 3-MA for 1 h, and was subsequently treated with 100 μM DSTD for another 24 h. LC3 conversion and PARP cleavage were analyzed by Western blot. Cell viability and cytotoxicity in U2OS (c) and MG-63 (d) cells were measured by MTT assay and LDH release assay. **P<0.01 versus control group; *P<0.05 versus group treated with DSTD; #P<0.01 versus group treated with 3-MA
DSTD attenuated drug resistance in osteosarcoma cells
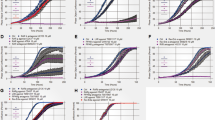
Given the previous finding that implicated autophagy and HMGB1 in drug resistance,6 we tested the effect of DSTD on drug resistance in osteosarcoma cells. As shown in Supplementary Figure 2S, DSTD resulted in slight cell shrinkage, and doxorubicin or cisplatin partly caused cell shrinkage compared with control cells. Notably, large-scale cell death occurred in the presence of DSTD combined with chemotherapy. Consistent with previous reports, doxorubicin and cisplatin reduced the cell viability in a dose-dependent manner (Figure 6). DSTD treatment sensitized osteosarcoma cells to doxorubicin and cisplatin, which was evidenced by the further decrease in the cell viability compared to chemotherapy alone. These results indicated that DSTD attenuated drug resistance in osteosarcoma cells.
HMGB1 overexpression blocked DSTD-induced chemosensitivity during chemotherapy in osteosarcoma cells
To explore the mechanism that DSTD-enhanced chemosensitivity in osteosarcoma cells, we further examined the effect of DSTD on the expression of MIF and HMGB1 during chemotherapy. Because MIF has been involved in chemoresistance in breast cancer,23 we evaluated the effect of doxorubicin and cisplatin on the expression levels of MIF and HMGB1 in the presence or absence of DSTD. Doxorubicin and cisplatin alone did not significantly affect MIF expression but increased the HMGB1 level (Figures 7a and c), which was a result identical to those of previous reports.6 Interestingly, DSTD combined with chemotherapy drugs suppressed HMGB1 expression. As expected, chemotherapy elevated the HMGB1 mRNA level, whereas chemotherapy combined with DSTD significantly decreased the HMGB1 mRNA level (Figures 7b and d), suggesting that DSTD reversed the upregulation of HMGB1 induced by doxorubicin and cisplatin. HMGB1 has been implicated in cancer development and therapy. A series of studies indicate that HMGB1 promotes chemoresistance by regulating autophagy.6 We further investigated the effect of doxorubicin and cisplatin on ERK phosphorylation in the presence or absence of DSTD. Doxorubicin and cisplatin alone did not significantly affect ERK phosphorylation, whereas chemotherapy combined with DSTD significantly decreased ERK phosphorylation (Figures 7a, c and e). Notably, ERK phosphorylation was blocked by DSTD combined with doxorubicin, but was promoted by HMGB1 overexpression in U2OS and Mg-63 cells (Figure 7e). Consistent with a previous report,6 the overexpression of HMGB1 promoted autophagy and significantly decreased the cleaved PARP level (Figure 7e), which was responsible for the lower cell death rate in HMGB1-overexpressing cells (Figure 7f). These finding revealed that autophagy induced by the overexpression of HMGB1 rescued cell death, and DSTD facilitated cell death by decreasing the HMGB1 level, which likely explains how DSTD contributed to chemosensitivity in osteosarcoma cells.
DSTD reversed the upregulation of HMGB1 during chemotherapy in osteosarcoma cells. (a) DSTD reversed Dox-induced HMGB1 expression. Cells were treated with 0.5 μM Dox in the presence or absence of 100 μM DSTD for 24 h. Western blot was used to analyze MIF, HMGB1 and ERK expression using their corresponding antibodies. (b) DSTD decreased mRNA level of HMGB1 during chemotherapy. mRNA level was analyzed by real-time PCR. (c) DSTD reversed Cis-induced HMGB1 expression. Cells were treated with 50 μM Cis in the presence or absence of 100 μM DSTD for 24 h. Western blot was used to analyze MIF, HMGB1 and ERK expression using their corresponding antibodies. (d) DSTD decreased mRNA level of HMGB1 during chemotherapy. mRNA level was analyzed by real-time PCR. (e) Overexpression of HMGB1 blocked DSTD-induced signal transduction during chemotherapy. Cells were transfected with control plasmid and GV230-HMGB1 cDNA for 72 h. HMGB1, pERK, LC3 conversion and PARP cleavage were measured by western blot. (f) Overexpression of HMGB1 decreased DSTD-induced cell death during chemotherapy. Values are means of at least 3 independent experiments. *P<0.05; #P<0.01
Downregulation of MIF inhibited tube formation
Because of the important role of MIF in tube formation, we assessed the effect of MIF inhibition on angiogenesis. Identical to previously reported results,35, 36 MIF knockdown by shRNA interference or inhibition by DSTD suppressed tube formation, in contrast to the control group (Supplementary Figure 1S).
Discussion
Macrophage MIF is a multiple functional cytokine and plays a critical role in modulating inflammation.10 Recent studies have suggested that MIF is abundantly expressed in various cancer cells, such as breast cancer, bladder cancer, colon cancer and osteosarcoma cells.23, 24, 25, 26, 27 ISO-1, an MIF inhibitor, blocks the migration and invasion of several cancer cells.37, 38 MIF knockdown by shRNA interference in breast cancer cells induces autophagy, which promotes chemosensitivity.23 All of these findings indicate that MIF is a potential target in the treatment of tumors. However, the role of MIF in osteosarcoma cells has never been investigated. In the present study, we isolated a novel androstadienone derivative characterized as DSTD. Several steroidal drugs, such as dehydroepiandrosterone (DHEA), dexamethasone (DEX) and hydrocortisone, significantly reduce MIF production.39, 40 Although androstadienone slightly inhibits the MIF level,39 we are still interested in investigating the effect of DSTD on MIF expression. Interestingly, DSTD significantly inhibited the expression of MIF and decreased the viability of osteosarcoma cells, which is consistent with previous reports indicating that the inhibition of MIF decreases cell viability.41, 42
We further delineated the influence of MIF inhibition by DSTD on signal transduction in osteosarcoma cells. Our results showed that DSTD-induced MIF inhibition phenocopied MIF knockdown by shRNA in regulating the downstream proteins. MIF inhibition led to a significant decrease in the Bcl-2 and phosphor-ERK levels, which has been confirmed in dermal fibroblasts and neural stem/progenitor cells.16, 43 Differently, DSTD-induced MIF inhibition enhances transiently ERK phosphorylation and nucleic HMGB1 at 12 h, which might be a protective response to detrimental effect of ROS.44, 45 Interestingly, as the treatment time increased, MIF inhibition promoted the translocation of HMGB1 from the nucleus to the cytoplasm and decreased the total HMGB1 levels. HMGB1 is a highly conserved transcription factor that serves in protein assembly and is normally located in the nucleus.34 HMGB1 is released from the nucleus to the cytoplasm in response to the stimulation of growth factors, cytokines and cellular stresses to activate downstream pathways that are responsible for cell proliferation, invasion, angiogenesis and metastasis.46 Recent studies implicate HMGB1 in autophagy, demonstrating that HMGB1 translocation promotes autophagy.34, 47 Moreover, Bcl-2 is an important regulator in cell growth, apoptosis and autophagy. Bcl-2 has been well documented to block Beclin-1-dependent autophagy by interacting with Beclin-1.48, 49 Because of the critical roles of HMGB1 and Bcl-2 in mediating autophagy, we assessed the ability of MIF inhibition to induce autophagy. We found that DSTD induced time- and dose-dependent autophagy in MG-63 and U2OS cells. Notably, DSTD-induced autophagy could be attenuated by NAC treatment, suggesting that DSTD induced ROS-mediated autophagy. However, NAC treatment could not attenuate PARP cleavage induced by DSTD, which was evidenced by the ability of NAC treatment to further decrease the cell viability and promote cell death. This finding differs from the inability of MIF knockdown to affect cell death in MCF-7 breast cancer cells.23 Our results indicated that DSTD-induced MIF inhibition promoted reversible ROS-mediated autophagy to delay cell death. This finding is consistent with recent reports indicating that HMGB1-mediated autophagy contributes to cell survival and chemoresistance in osteosarcoma cells.6 In contrast, autophagy induced by MIF knockdown promotes cell death and chemosensitivity in MCF-7 breast cancer cells.23 Pretreatment with 3-MA, an inhibitor of type III phosphatidylinositol 3-kinase that is needed for initiation of autophagy, attenuated autophagy induced by DSTD and promoted cell death, suggesting that DSTD-induced autophagy contributed to delay cell death.
The mechanism by which autophagy handles the balance between cytoprotection and cell death is poorly understood. Autophagy plays a protective role in response to most cellular stress. In the case of abnormally expressed autophagy-related proteins, autophagy led to cell death.46 In the present study, DSTD promoted chemosensitivity to doxorubicin and cisplatin even though DSTD-induced autophagy showed a protective effect, suggesting that another pathway was implicated in DSTD-induced chemosensitivity. Because HMGB1 plays a critical role in chemoresistance in leukemia, colon cancer, prostate cancer and osteosarcoma,6, 50, 51, 52 we assessed the effect of DSTD on HMGB1 expression during chemotherapy in osteosarcoma. Consistent with previous reports, doxorubicin and cisplatin enhanced the expression of HMGB1, which suggested that cells expressed HMGB1 protein to counteract the chemotherapeutic damage. Notably, DSTD reversed the upregulation of HMGB1 induced by chemotherapy drugs, which was likely responsible for the DSTD-induced chemosensitivity. To confirm the hypothesis, we tested whether HMGB1 overexpression blocked DSTD-induced chemosensitivity during chemotherapy. Our results showed that HMGB1 overexpression blocked cell death induced by DSTD combined with doxorubicin. DSTD combined with doxorubicin decreased ERK phosphorylation and activated PARP cleavage, which was reversed by HMGB1 overexpression. HMGB1 overexpression also accumulated more LC3 than that induced by DSTD combined with doxorubicin. These results indicated that DSTD-induced chemosensitivity is due to HMGB1 downregulation, HMGB1 induced phosphor-ERK expression and autophagy to delay cell death.
In summary, the present study indicated that an androstenedione derivative, DSTD, could inhibit MIF expression in osteosarcoma cells. MIF inhibition decreased Bcl-2 and induced the translocation of HMGB1 from the nucleus to the cytoplasm, which was responsible for autophagy. DSTD also decreased the phospho-ERK level as the treatment time increased. Both NAC and 3-MA attenuated DSTD-induced autophagy but promoted cell death, suggesting that MIF inhibition induced ROS-mediated autophagy to rescue cell death. In contrast, DSTD promoted chemosensitivity and reduced the HMGB1 level during chemotherapy. HMGB1 overexpression blocked the DSTD-induced chemosensitivity of osteosarcoma cells. These results indicated that DSTD promoted chemosensitivity via the downregulation of HMGB1 during chemotherapy.
Materials and Methods
Antibodies and reagents
The monoclonal antibodies (mAbs) and polyclonal antibodies (pAbs) used in this study were as follows: Anti-MIF (mAb, ab55445), Anti-ERK1/2 (phospho T185+Y187+T202+T204, pAb, ab50011), anti-Bax (pAb, ab7977), anti-HMGB1 (pAb, ab79823), anti-Bcl-2 (pAb, ab7973) and anti-Histone H3 (pAb, ab1791) were purchased from Abcam (Cambridge, MA, USA). Anti-beta-actin (pAb) and anti-GADPH (pAb) was purchased from Anbo Biotechnology Company (San Francisco, CA, USA). Anti-PARP and anti-LC3B (pAb) was purchased from Cell Signaling Technology (Beverly, MA, USA). AnnexinV-FITC and propidium iodide were purchased from BD Pharmingen (San Jose, CA, USA).
Doxorubicin, cisplatin, 3-(4,5-dimethyl thiazol-2-yl)-2,5-diphenyl tetrazolium bromide (MTT), 2′,7′-dichlorofluorescin diacetate (DCF-DA) and 3-Methyladenine (3-MA) were purchased from Sigma Chemical Co. (St. Louis, MO, USA).
Analysis and purification of androstenedione derivatives
Strain B4 of Bordetella sp. was stored and cultured as described in a previous study.53 The androstenedione derivatives were analyzed using thin layer chromatography (TLC). Briefly, the plate was developed in dichloromethane: petroleum ether: ethyl acetate (6 : 3 : 1, v/v/v) and visualized by spraying 10% sulfuric acid (dissolved in ethanol). For purification, the derivatives were separated in 0.5-mm-thick silica gel preparative plates and visualized under ultraviolet light. The derivatives were scraped from the plate and resolved in methanol. After removing methanol using vacuum drying, the crystallized product was identified by mass spectrum (MS) and nuclear magnetic resonance (NMR) analyses.
Cell lines and culture conditions
The human osteosarcoma cell lines (MG-63 and U2OS) were obtained from American Type Culture Collection (ATCC). All cell lines were characterized according to the ATCC instructions. Cells were cultured in Dulbecco’s Modified Eagle’s Medium (Hyclone, Logan, UT, USA) supplemented with 10% fetal bovine serum (Hyclone), 100 U/ml penicillin, 100 U/ml streptomycin and 0.03% L-glutamine at 37 °C in 5% CO2.
Cell viability and LDH release assay
The cells were seeded at a density of 1 × 104 in a 96-well microplate. After the cells reached approximately 80% confluence, they were treated with the indicated concentration of DSTD for 48 h. Subsequently, MTT solution was added and incubated for 4 h at 37 °C. The cell viability was determined by measuring the produced formazan at 490 nm using a SpectraMax M2. LDH release was measured using LDH Release Assay Kit (Beyotime, Shanghai, China). In brief, cells were seeded in 96-well microplates. After reaching about 80% confluence, the cells were subjected to different treatment for indicated time. Subsequently, the microplate was placed directly into centrifuge and supernatant was collected by centrifugation at 400 g for 5 min. 120 μl of supernatant from each well was transferred to a new a 96-well microplate and was mixed with 60 μl LDH working solution at room temperature for 30 min. LDH activity was monitored at 490 nm using a SpectraMax M2. The total cell lysate was set as 100%.
Western blot analysis
The treated cells were harvested and resuspended in RIPA lysis buffer. The whole-cell lysates were obtained by centrifugation, and the concentrations were measured using a bicinchoninic acid assay kit. The cytosol and nuclear protein were isolated using a nuclear and cytoplasmic protein extraction kit (Sangon Biotech, Shanghai, China) according to the manufacturer’s protocol. Briefly, cells were pre-cooled in PBS and scraped in a microfuge tube. After centrifugation at 1000 g for 3 min at 4 °C, the cell pellets were resuspended in 1.0 ml ice-cold buffer A (containing 1 μl DTT, 10 μl PMSF and 1 μl protease inhibitor, which were added before use), followed by vigorous shock for 15 s and incubation on ice for 15 min. After adding 10 μl ice-cold buffer B, the mixture was strongly vortexed for 5 s and incubated on ice for 1 min. After strongly vortexing for 5 s again, the supernatant was centrifuged again at 14000 g and 4 °C for 5 min. The supernatant, which was the cytosolic fraction, and the pellet, which was the nuclear fraction, were collected. The pellet was resuspended in 0.1 ml ice-cold buffer C (containing 1 μl DTT, 10 μl PMSF and 1 μl protease inhibitor per 1 ml of buffer C, which were added before use), vigorously shaken for 10 s and incubated on ice for 40 min. After strong vortexing for 30 s again, the supernatant, which was the nuclear fraction, was collected by centrifugation at 14000 g for 5 min. Equal amounts of proteins were resolved by 12% SDS-PAGE electrophoresis. The proteins were further transferred onto polyvinylidene fluoride membranes. The membranes were blocked with 3% BSA in Tris-buffered saline containing 0.05% Tween-20 (TBST) for 2 h at room temperature and then incubated with the indicated primary antibodies (anti-MIF, anti-Bcl-2, anti-Bax, anti-ERK1/2, anti-LC3B, and anti-PARP, 1 : 1000; anti-HMGB1, anti-actin, and anti-GADPH, 1 : 2000) overnight at 4 °C. After being washed three times with TBST, the membranes were incubated with horseradish peroxidase-conjugated secondary antibodies for 1 h at room temperature. The signals were visualized using the enhanced chemiluminescence detection kit and quantified using an Alpha Imager2200.
Determination of ROS levels
The generated ROS was probed using an oxidation-sensitive fluorescent reagent (DCF-DA). The treated cells were rinsed with cold PBS and then incubated in 10 μM DCF-DA for 30 min at 37 °C. After rinsing three times with PBS, the fluorescence excitation was performed at 488 nm and the emission was detected at 525 nm using a fluorescence microscope (LEICA DMIRE2).
Immunofluorescence
The cells were fixed with 4% paraformaldehyde in PBS for 30 min and permeabilized with 0.2% Triton X-100 in PBS for 15 min. Subsequently, the cells were blocked with 10% normal goat serum for 1 h at room temperature. After being rinsed three times with PBST, the cells were incubated with MAP1-LC3B overnight at 4 °C and then incubated with FITC-conjugated secondary antibody IgG for 2 h. After being rinsed five times with PBST, the cells were visualized using a fluorescence microscope (LEICA DMIRE2).
Analysis of apoptosis
The cells were seeded at a density of 2 × 106 in six-well plates. After reaching 70–80% confluence, the cells were subjected to different treatment for indicated time. Subsequently, the apoptosis was analyzed using a flow cytometer (Beckman Coulter, Chicago, IL, USA). Treated cells were harvested with trypsin without EDTA and phenol red. After washing with PBS three times, the cells were incubated with annexinV-FITC (5 μl, 20 μg/ml) and propidiumiodide (PI, 5 μl, 20 μg/ml) in binding buffer at room temperature for 10 min in the dark. Subsequently, the labeled cells were analyzed by a flow cytometer and the data were processed using WinMDI 2.9 software.
Gene transfection and shRNA interference
shRNAs were synthesized by Genechem (Shanghai, China). The MIF was silenced by infecting human osteosarcoma cells (MG-63 and U2OS) with a lentivirus that expressed shRNA against MIF. To obtain stable interference, the duplex RNAs targeting MIF (sense strand: GACAGGGTCTACATCAACTAT) and control RNAs (sense strand: TTCTCCGAACGTGTCACGT) were inserted into the GV115 vector according to manufacturer’s instructions. To overexpresse HMGB1, GV230-HMGB1 cDNA (Genechem) was transfected into MG-63 and U2OS cells using Lipofectamine 2000 (Invitrogen, San Diego, CA, USA).
Quantitative RT-PCR
The total RNA was extracted from treated cells using Trizol reagent (Takara, Shanghai, China). cDNA was generated by reverse transcribing RNA using the RevertAid First Strand cDNA Synthesis Kit (Fermentas, Hanover, Germany). The cDNA was used as template to amplify HMGB1 with the specific primers (forward 5′- TCAAAGGAGAACATCCTGGCCTGT -3′ and reverse 5′- CTGCTTGTCATCTGCAGCAGTGTT -3′),6 MIF with (forward 5′- GGGGCCATCTTCCGGGTTCA-3′ and reverse 5′- GCCTCGCCTCGGGTCTTTCTTA-3′) and GADPH with the primers (forward 5′-GGTGAAGGTCGGAGTCAACGG-3′ and reverse 5′-GGTCATGAGTCCTTCCACGATACC-3′).6
Tube formation assay
The tube formation assay was performed as previously described.54 Briefly, human umbilical vein endothelium cells lines (HUVECs) expressing MIF shRNA were seeded at 2 × 104 per well in a 96-well microplate coated with Matrigel (60 μl per well, BD Bioscience, San Jose, CA, USA). To assess the effect of DSTD on angiogenesis, the HUVECs were exposed to 100 μM DSTD. After being cultured for 12 h, tube formation was visualized under an inverted fluorescence microscope (Nikon, Tokyo, Japan).
Statistical analysis
For all experiments in this study, the data are represented as the means±S.E. of at least three independent experiments. A student’s t-test was used for statistical comparisons between two groups. One-way ANOVA was used for statistical comparisons among multiple groups. P<0.05 was considered significant.
Abbreviations
- MIF:
-
Macrophage migration inhibitory factor
- DSTD:
-
3, 4-dihydroxy-9, 10-secoandrosta-1, 3, 5, 7-tetraene-9, 17-dione
- HMGB1:
-
High mobility group box 1
- MS:
-
mass spectra
- NMR:
-
nuclear magnetic resonance
- ROS:
-
reactive oxygen species
- NAC:
-
N-acetyl-L-cysteine
References
Endo-Munoz L, Cumming A, Sommerville S, Dickinson I, Saunders NA . Osteosarcoma is characterised by reduced expression of markers of osteoclastogenesis and antigen presentation compared with normal bone. Br J Cancer 2010; 103: 73–81.
Park JY, Kim YW, Park YK . Nrf2 expression is associated with poor outcome in osteosarcoma. Pathology 2012; 44: 617–621.
Endo-Munoz L, Cumming A, Sommerville S, Dickinson I, Saunders NA . Osteosarcoma is characterised by reduced expression of markers of osteoclastogenesisand antigenpresentation compared with normal bone. Br J Cancer 2010; 103: 73–81.
Moriceau G, Roelofs AJ, Brion R, Redini F, Ebetion FH, Rogers MJ et al. Synergistic inhibitory effect of apomine and lovastatin on osteosarcoma cell growth. Cancer 2012; 118: 750–760.
Rosen G, Murphy ML, Huvos AG, Gutierrez M, Marcove RC . Chemotherapy, en bloc resection, and prosthetic bone replacement in the treatment of osteogenic sarcoma. Cancer 1976; 37: 1–11.
Huang J, Ni J, Liu K, Yu Y, Xie M, Kang R et al. HMGB1 promotes drug resistance in osteosarcoma. Cancer Res 2012; 72: 230–238.
Posthumadeboer J, van Egmond PW, Helder MN, de Menezes RX, Cleton-Jansen AM, Beliën JA et al. Targeting JNK-interacting-protein-1 (JIP1) sensitises osteosarcoma to doxorubicin. Oncotarget 2012; 3: 1169–1181.
Zhao YZ, Dai DD, Lu CT, Chen LJ, Lin M, Shen XT et al. Epirubicin loaded with propylene glycol liposomes significantly overcomes multidrug resistance in breast cancer. Cancer Lett 2013; 330: 74–83.
Winiarska M, Bil J, Wilczek E, Wilczynski GM, Lekka M, Engelberts PJ et al. Statins impair antitumor effects of rituximab by inducing conformational changes of CD20. PLoS Med 2008; 5: e64.
Calandra T, Roger T . Macrophage migration inhibitory factor: a regulator of innate immunity. Nat Rev Immunol 2003; 3: 791–800.
Bach JP, Deuster O, Balzer-Geldsetzer M, Meyer B, Dodel R, Bacher M . The role of macrophage inhibitory factor in tumorigenesis and central nervous system tumors. Cancer 2009; 115: 2031–2040.
Bucala R, Donnelly SC . Macrophage migration inhibitory factor: a probable link between inflammation and cancer. Immunity 2007; 26: 281–285.
Nishihira J . Macrophage migration inhibitory factor (MIF): its essential role in the immune system and cell growth. J Interferon Cytokine Res 2000; 20: 751–762.
Bach JP, Rinn B, Meyer B, Dodel R, Bacher M . Role of MIF in inflammation and tumorigenesis. Oncology 2008; 75: 127–133.
Mitchell RA . Mechanisms and effectors of MIF-dependent promotion of tumourigenesis. Cell Signal 2004; 16: 13–19.
Ohta S, Misawa A, Fukaya R, Inoue S, Kanemura Y, Okano H et al. Macrophage migration inhibitory factor (MIF) promotes cell survival and proliferation of neural stem/progenitor cells. J Cell Sci 2012; 125 (Pt 13): 3210–3220.
Madeira MF, Queiroz-Junior CM, Costa GM, Santos PC, Silveira EM, Garlet GP et al. MIF induces osteoclast differentiation and contributes to progression of periodontal disease in mice. Microbes Infect 2012; 14: 198–206.
Asare Y, Schmitt M, Bernhagen J . The vascular biology of macrophage migration inhibitory factor (MIF). Expression and effects in inflammation, atherogenesis and angiogenesis. Thromb Haemost 2013; 109: 391–398.
Lee CY, Su MJ, Huang CY, Chen MY, Hsu HC, Lin CY et al. Macrophage migration inhibitory factor increases cell motility and up-regulates αvβ3 integrin in human chondrosarcoma cells. J Cell Biochem 2012; 113: 1590–1598.
Dessein AF, Stechly L, Jonckheere N, Dumont P, Monté D, Leteurtre E et al. Autocrine induction of invasive and metastatic phenotypes by the MIF-CXCR4 axis in drug-resistant human colon cancer cells. Cancer Res 2010; 70: 4644–4654.
Chen YC, Zhang XW, Niu XH, Xin DQ, Zhao WP, Na YQ et al. Macrophage migration inhibitory factor is a direct target of HBP1-mediated transcriptional repression that is overexpressed in prostate cancer. Oncogene 2010; 29: 3067–3078.
Bacher M, Schrader J, Thompson N, Kuschela K, Gemsa D, Waeber G et al. Up-regulation of macrophage migration inhibitory factor gene and protein expression in glial tumor cells during hypoxic and hypoglycemic stress indicates a critical role for angiogenesis inglioblastoma multiforme. Am J Pathol 2003; 162: 11–17.
Wu MY, Fu J, Xu J, O'Malley BW, Wu RC . Steroid receptor coactivator 3 regulates autophagy in breast cancer cells through macrophage migration inhibitory factor. Cell Res 2012; 22: 1003–1021.
Choudhary S, Hegde P, Pruitt JR, Sielecki TM, Choudhary D, Scarpato K et al. Macrophage migratory inhibitory factor promotes bladder cancer progression via increasing proliferation and angiogenesis. Carcinogenesis 2013; 34: 2891–2899.
He XX, Chen K, Yang J, Li XY, Gan HY, Liu CY et al. Macrophage migration inhibitory factor promotes colorectal cancer. Mol Med 2009; 15: 1–10.
Han I, Lee MR, Nam KW, Oh JH, Moon KC, Kim HS . Expression of macrophage migration inhibitory factor relates to survival in high-gradeosteosarcoma. Clin Orthop Relat Res 2008; 466: 2107–2113.
Cates JM, Friedman DB, Seeley EH, Dupont WD, Schwartz HS, Holt GE et al. Proteomic analysis of osteogenic sarcoma: association of tumour necrosis factor with poor prognosis. Int J Exp Pathol 2010; 91: 335–349.
Sih CJ, Lee SS, Tsong YY, Wang KC . Mechanisms of steroid oxidation by microorganisms. 8. 3, 4-Dihydroxy-9,10-secoandrosta-1,3,5(10)-triene-9,17-dione, an intermediate in the microbiological degradation of ring A of androst-4-ene-3,17-dione. J Biol Chem 1966; 241: 540–550.
Lin Y, Song X, Fu J, Lin J, Qu Y . Microbial transformation of androst-4-ene-3, 17-dione by Bordetella sp. B4 CGMCC 2229. J Chem Technol Biotechnol 2009; 84: 789–793.
Cheng P, Ni Z, Dai X, Wang B, Ding W, Rae Smith A et al. The novel BH-3 mimetic apogossypolone induces Beclin-1-and ROS-mediated autophagy in humanhepatocellular carcinoma cells. Cell Death Dis 2013; 4: e489.
Huang J, Liu K, Yu Y, Xie M, Kang R, Vernon P et al. Targeting HMGB1-mediated autophagy as a novel therapeutic strategy for osteosarcoma. Autophagy 2012; 8: 275–277.
Kabeya Y, Mizushima N, Ueno T, Yamamoto A, Kirisako T, Noda T et al. LC3, a mammalian homologue of yeast Apg8p, is localized in autophagosome membranes after processing. EMBO J 2000; 19: 5720–5728.
Low IC, Kang J, Pervaiz S . Bcl-2: a prime regulator of mitochondrial redox metabolism in cancer cells. Antioxid Redox Signal 2011; 15: 2975–2987.
Tang D, Kang R, Livesey KM, Cheh CW, Farkas A, Loughran P et al. Endogenous HMGB1 regulates autophagy. J Cell Biol 2010; 190: 881–892.
Xu X, Wang B, Ye C, Yao C, Lin Y, Huang X et al. Overexpression of macrophage migration inhibitory factor induces angiogenesis in human breast cancer. Cancer Lett 2008; 261: 147–157.
Hira E, Ono T, Dhar DK, El-Assal ON, Hishikawa Y, Yamanoi A et al. Overexpression of macrophage migration inhibitory factor induces angiogenesis and deteriorates prognosis after radical resection for hepatocellular carcinoma. Cancer 2005; 103: 588–598.
Meyer-Siegler KL, Iczkowski KA, Leng L, Bucala R, Vera PL . Inhibition of macrophage migration inhibitory factor or its receptor (CD74) attenuates growth and invasion of DU-145 prostate cancer cells. J Immunol 2006; 177: 8730–8739.
Winner M, Meier J, Zierow S, Rendon BE, Crichlow GV, Riggs R et al. A novel, macrophage migration inhibitory factor suicide substrate inhibits motility and growth of lung cancer cells. Cancer Res 2008; 68: 7253–7257.
Harding G, Mak YT, Evans B, Cheung J, MacDonald D, Hampson G . The effects of dexamethasone and dehydroepiandrosterone (DHEA) on cytokines and receptor expression in a human osteoblastic cell line: potential steroid-sparing role for DHEA. Cytokine 2006; 36: 57–68.
Balow JE, Rosenthal AS . Glucocorticoid suppression of macrophage migration inhibitory factor. J Exp Med 1973; 137: 1031–1041.
Schrader J, Deuster O, Rinn B, Schulz M, Kautz A, Dodel R et al. Restoration of contact inhibition in human glioblastoma cell lines after MIF knockdown. BMC Cancer 2009; 9: 464.
Rendon BE, Roger T, Teneng I, Zhao M, Al-Abed Y, Calandra T et al. Regulation of human lung adenocarcinoma cell migration and invasion by macrophage migration inhibitory factor. J Biol Chem 2007; 282: 29910–29918.
Kim JY, Kwok SK, Hur KH, Kim HJ, Kim NS, Yoo SA et al. Up-regulated macrophage migration inhibitory factor protects apoptosis of dermal fibroblasts in patients with systemic sclerosis. Clin Exp Immunol 2008; 152: 328–335.
Dunyaporn T, Weiqin L, Marcia AO, Nilsa R, Peng H . Redox Regulation of Cell Survival. Antioxid Redox Signal 2008; 10: 1343–1374.
Macus Tien K . Redox Regulation of Multidrug Resistance in Cancer Chemotherapy: Molecular Mechanisms and Therapeutic Opportunities. Antioxid Redox Signal 2009; 11: 99–133.
Mizushima N, Levine B, Cuervo AM, Klionsky DJ . Autophagy fights disease through cellular self-digestion. Nature 2008; 451: 1069–1075.
Tang D, Kang R, Cheh CW, Livesey KM, Liang X, Schapiro NE et al. HMGB1 release and redox regulates autophagy and apoptosis in cancer cells. Oncogene 2010; 29: 5299–5310.
Pattingre S, Tassa A, Qu X, Garuti R, Liang XH, Mizushima N et al. Bcl-2 antiapoptotic proteins inhibit Beclin 1-dependent autophagy. Cell 2005; 122: 927–939.
Luo S, Rubinsztein DC . Apoptosis blocks Beclin 1-dependent autophagosome synthesis: an effect rescued by Bcl-xL. Cell Death Differ 2010; 17: 268–277.
Liu L, Yang M, Kang R, Wang Z, Zhao Y, Yu Y et al. HMGB1-induced autophagy promotes chemotherapy resistance in leukemia cells. Leukemia 2011; 25: 23–31.
Tang D, Loze MT, Zeh HJ, Kang R . The redox protein HMGB1 regulates cell death and survival in cancer treatment. Autophagy 2010; 6: 1181–1183.
Gnanasekar M, Kalyanasundaram R, Zheng G, Chen A, Bosland MC, Kajdacsy-Balla A . HMGB1: A Promising Therapeutic Target for Prostate Cancer. Prostate Cancer 2013 2013: 157103.
Lin Y, Liu J, Hu Y, Song X, Zhao Y . An antioxidant exopolysaccharide devoid of pro-oxidant activity produced by the soil bacterium Bordetella sp. B4. Bioresour Technol 2012; 124: 245–251.
Chen L, Zhang JJ, Rafii S, Huang XY . Suppression of tumor angiogenesis by Gα13 haploinsufficiency. J Biol Chem 2009; 284: 27409–27415.
Acknowledgements
This study was supported by a grant from the National Natural Science Foundation of China (No. 31370897 and 81000731), Shandong Provincial Natural Science Foundation of China (No. ZR2011HM019), Shandong Provincial science and technology foundation of China (2010GG0020225), and the Promotive Research Fund for Excellent Young and Middle-Aged Scientisits of Shandong Province (BS2010YY045).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by GM Fimia
Supplementary Information accompanies this paper on Cell Death and Disease website
Rights and permissions
Cell Death and Disease is an open-access journal published by Nature Publishing Group. This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Liu, Y., Zhao, L., Ju, Y. et al. A novel androstenedione derivative induces ROS-mediated autophagy and attenuates drug resistance in osteosarcoma by inhibiting macrophage migration inhibitory factor (MIF). Cell Death Dis 5, e1361 (2014). https://doi.org/10.1038/cddis.2014.300
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2014.300
This article is cited by
-
Suppression of heat shock protein 70 by siRNA enhances the antitumor effects of cisplatin in cultured human osteosarcoma cells
Cell Stress and Chaperones (2017)
-
Defective autophagy leads to the suppression of stem-like features of CD271+ osteosarcoma cells
Journal of Biomedical Science (2016)
-
4-cholesten-3-one suppresses lung adenocarcinoma metastasis by regulating translocation of HMGB1, HIF1α and Caveolin-1
Cell Death & Disease (2016)
-
Quercetin blocks t-AUCB-induced autophagy by Hsp27 and Atg7 inhibition in glioblastoma cells in vitro
Journal of Neuro-Oncology (2016)
-
MicroRNA-26b inhibits metastasis of osteosarcoma via targeting CTGF and Smad1
Tumor Biology (2015)