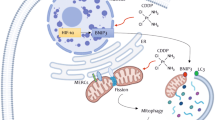
Abstract
Peritoneal carcinomatosis (PC) is the most common secondary cancerous disease, and more effective novel regimens are needed. In this study, we identified a novel combination treatment for PC, chemotherapeutic agent mitomycin C in combination with mTOR (mammalian target of rapamycin) inhibitor rapamycin. We observed that the combination of mitomycin C and rapamycin induced synergistic cytotoxicity and apoptosis, which was mediated through an increase in caspase activation. The combination of mitomycin C and rapamycin inactivated p70 S6 ribosomal kinase (S6K1) and dephosphorylated Bad, leading to dissociation of Bcl-xL from Bak, which resulted in Bak oligomerization, mitochondria dysfunction and cytochrome c release. PF-4708671, a S6K1-specific inhibitor, enhanced the combination treatment-induced apoptosis, whereas S6K1 E389 DeltaCT-HA (S6K1 active form) dramatically decreased the induction of apoptosis. In addition, the combination treatment significantly inhibited LS174T intraperitoneal tumor growth in vivo. This study provides a preclinical rationale for apoptosis induction linked with the mTOR pathway through a combination of chemotherapeutic agents and mTOR inhibitor, and will support this combinatorial strategy to PC patients.
Similar content being viewed by others
Main
Peritoneal carcinomatosis (PC) is the most common secondary cancerous disease to affect the peritoneal cavity following colorectal cancer, appendiceal cancer, ovarian cancer, gastric cancer, or diffuse malignant peritoneal mesothelioma, and is regarded as a lethal condition having poor prognosis with a median survival of ∼6 months.1, 2 Systemic chemotherapy prolongs survival in PC patients, but is less effective than for patients with hepatic metastasis; thus, an increasing number of patients are treated with cytoreductive surgery followed by hyperthermic intraperitoneal chemoperfusion (HIPEC).3 Nevertheless, more effective novel regimens are still needed to improve the survival of PC patients.
Mitomycin C, a potent DNA crosslinker, was used as an anticancer drug and led to pronounced induction of apoptosis, especially through the mitochondrial-dependent pathway.4, 5 It has received a great number of preclinical and clinical studies and is mostly widely used for PC.6 Combination of cytotoxic drugs, such as mitomycin C with novel biological agents, has achieved improvement in animal models and provides additional survival benefits for patients.7, 8, 9
mTOR (mammalian target of rapamycin), a serine/threonine kinase, is ubiquitously involved in cell growth and proliferation. The fact that hyperactivation of mTOR is a common feature of human cancers makes it an attractive target in cancer therapy.10 It has been reported that some agents inactivate mTOR signaling as an early response prior to the execution of apoptosis.11, 12 Recently, it has been found that mTOR can directly control mitochondrial function,13 which raises the possibility that targeting mTOR may regulate the apoptotic pathway. Sirolimus, known as rapamycin, an mTOR inhibitor, potently inhibits S6K1 (S6 ribosomal kinase) activity14 and has been generally well tolerated in clinical trials, often inducing prolonged stable disease or tumor regression, and can be considered as an option for combination with other anticancer agents.15, 16
In this study we observed that the combination of chemotherapy agent mitomycin C and mTOR inhibitor rapamycin induced synergistic cytotoxicity and apoptosis in vitro and in vivo. The combination of mitomycin C and rapamycin inactivated S6K1, not Akt, and resulted in dephosphorylation of Bad, which inactivated its pro-apoptotic function, inducing Bak oligomerization and mitochondrial-dependent apoptotic cell death.
Results
The combination of mitomycin C and rapamycin induced synergistic cytotoxicity and apoptosis
To investigate the effect of the combination of mitomycin C and rapamycin on cell viability, human colorectal carcinoma LS174T cells and LS180 cells were treated with mitomycin C (5 μg/ml) and/or rapamycin (2.5–10 μg/ml) and examined by MTS assay. As shown in Figures 1a and b, mitomycin C induced ∼20–40% cell death in LS174T and LS180. Rapamycin only slightly induced cell death. Rapamycin enhanced the effect of mitomycin C at all the indicated doses in LS174T and in only high dose in LS180 cells. For LS174T, flow cytometry assay shows that the mitomycin C induced ∼29% apoptotic cell death (annexin V+ PI−+annexin V+ PI+) after normalizing with (subtracting) the control group (Figure 1c). Rapamycin induced ∼0–8% cell death at the indicated dose (Figure 1c). Notably, the combination of mitomycin C and rapamcyin induced 56.5–74.5% apoptotic death, indicating the synergistic not additive effect (Figure 1c). Also, an increased activation of caspases and PARP cleavage, the hallmark feature of apoptosis, were detected in both cell lines (Figures 1d and e).
The combination of mitomycin C and rapamycin induced synergistic cytotoxicity and apoptosis. LS174T (a) and LS180 (b) cells were treated with mitomycin C (5 μg/ml) and/or rapamycin (2.5–10 μg/ml) for 24 h. Cell viability was analyzed by MTS assay. Error bars represent S.D. from triplicate experiments. The asterisk (**) represents a statistically significant difference compared with the control group (P<0.01). (c) LS174T cells were treated with mitomycin C (5 μg/ml) and/or rapamycin (2.5–10 μg/ml) for 24 h and cells were stained with fluorescein isothiocyanate (FITC)-Annexin V and propidium iodide (PI). Apoptosis was detected by the flow cytometric assay. After treatment, the cleavage of caspase 8, caspase 9, caspase 3, or PARP was detected by immunoblotting in LS174T (d) and LS180 (e) cells. Actin was used to confirm the equal amount of proteins loaded in each lane
The combination of mitomycin C and rapamycin inactivated S6K1
Jun N-terminal kinases play a critical role in death receptor-initiated extrinsic as well as mitochondrial intrinsic apoptotic pathways.17 Akt as well as S6K1 is a critical mediator of growth factor-induced survival.18, 19 Akt can be activated by phosphorylation within the carboxy terminus at Ser473.20 S6K1 activity most closely correlates with Thr389 phosphorylation.21 As shown in Figures 2a and b, in LS174T and LS180, mitomycin C induced JNK activation dramatically and had no effect on Akt and S6K1, while rapamycin significantly inhibited the activity of S6K1 and had no effect on JNK and Akt activity. Interestingly, the combination potently decreased the activity of S6K1, which might have contributed to the synergistic apoptosis induced by mitomycin C and rapamycin.
The combination of mitomycin C and rapamycin inactivated S6K1. LS174T (a) and LS180 (b) cells were treated with mitomycin C (5 μg/ml) and/or rapamycin (2.5–10 μg/ml) for 24 h and p-Akt/Akt, p-S6K1/S6K1 and p-JNK/JNK were detected by immunoblotting. Actin was used to confirm the equal amount of proteins loaded in each lane
Role of Bad in the combination of mitomycin C and rapamycin-induced apoptosis
It is reported that S6K1 is a dual pathway kinase, signaling cell survival as well as cell growth through different substrates, which include mitochondrial Bad and the ribosomal subunit S6, respectively.14, 22 We observed that the combination treatment significantly dephosphorylated Bad dependent on the dose of rapamycin (Figures 3a and b). Bad was the first BH3-only protein linked to proximal survival signals through phosphorylation by survival kinases; Bad has also been found to help coordinate mitochondrial fuel metabolism and the apoptotic machinery.22 Only the active conformation, nonphosphorylated Bad, heterodimerizes with Bcl-xL or Bcl-2 at mitochondrial membrane sites to promote cell death.23 Figures 3c and d show that Bad phosphorylation decreased its binding with Bcl-xL, and thus pro-apoptotic Bak was sequestered by Bcl-xL in untreated LS174T cells. Interestingly, rapamycin alone was not enough to increase the interaction between Bad and Bcl-xL. However, the ability of Bad binding with Bcl-xL increased dramatically only in the combination treatment, dissociating Bak from Bcl-xL and increasing the apoptosis.
Role of Bad in the combination of mitomycin C and rapamycin-induced apoptosis. LS174T (a) and LS180 (b) cells were treated with mitomycin C (5 μg/ml) and/or rapamycin (2.5–10 μg/ml) for 24 h, p-Bad/Bad were detected by immunoblotting. Actin was used to confirm the equal amount of proteins loaded in each lane. (c) LS174T cells were treated with mitomycin C (5 μg/ml) and/or rapamycin (10 μg/ml) for 24 h. After treatment, cell lysates were immunoprecipitated with anti-Bad antibody or IgG and immunoblotted with anti-Bcl-xL antibody. The presence of Bad and Bcl-xL in the lysates was examined. (d) After treatment, cell lysates were immunoprecipitated with anti-Bak antibody or IgG and immunoblotted with anti-Bcl-xL antibody. The presence of Bak and Bcl-xL in the lysates was examined
The combination of mitomycin C and rapamycin induced Bak oligomerization and mitochondrial dysfunction
It is well known that activation of pro-apoptotic member Bax or Bak forms pores in the mitochondrial outer membrane, which allow cytochrome c to escape into the cytoplasm and induce the caspase cascade.24 LS174T is a Bax null cell line,25 which makes it a good model to study the role of Bak. LS174T cells were treated with mitomycin C (5 μg/ml) and/or rapamycin (10 μg/ml) for 24 h. We observed that Bak oligomerization significantly increased in the combination of mitomycin C and rapamycin (Figure 4a).
The combination of mitomycin C and rapamycin induced Bak oligomerization and mitochondria dysfunction. (a) LS174T cells were treated with mitomycin C (5 μg/ml) and/or rapamycin (10 μg/ml) for 24 h. After treatment, mitochondria were isolated, cross-linked with 1 mM dithiobis (succinimidyl propionate) and subjected to immunoblotting with anti-Bak antibody. Bak monomer (1X) and multimers (2X, 3X, and 4X) are indicated. COX IV was used as a mitochondrial marker. (b) LS174T cells were treated with mitomycin C (2 μg/ml) and/or rapamycin (5 μg/ml) for 24 h. After treatment, mitochondria were stained with MitoTracker. Bak was stained with anti-Bak antibody. Localization of Bak was examined by confocal microscopy. (c) LS174T cells were treated with mitomycin C (5 μg/ml) and/or rapamycin (10 μg/ml) for 24 h. After treatment, cells were stained with JC-1 and analyzed by flow cytometry. (d) Cytochrome c release into cytosol was determined by immunoblotting for cytochrome c in the cytosolic fraction. Actin was used to confirm the equal amount of proteins loaded in each lane
To confirm the role of Bak in the combination-induced apoptosis, LS174T cells were treated with mitomycin C (2 μg/ml) and/or rapamycin (5 μg/ml) for 24 h and then stained with Bak antibody to enable imaging of endogenous Bak with confocal microscopy. Figure 4b shows that Bak was located in the mitochondria. We observed that mitomycin C slightly induced clustering of Bak, while rapamycin had no effect on Bak conformation change, and it was quite obvious that the combination treatment induced Bak clustering, indicating that Bak was activated in the combination treatment.
Mitomycin C has been reported to change the mitochondrial membrane potential.5 In this study we observed that mitomycin C alone impaired mitochondrial membrane potential. The shift from the upper left gate (P2; intact mitochondrial membrane potential) to the lower right gate (P3; impaired mitochondrial membrane potential) was seen. Rapamycin induced modest mitochondrial membrane potential change (Figure 4c). Of note, mitomycin C in combination with rapamycin dramatically impaired mitochondrial membrane potential (a loss of membrane potential) in a dose-dependent manner. Cellular cytochrome c was released from the mitochondria to the cytoplasm and confirmed that the combination treatment significantly induced mitochondria dysfunction (Figure 4d).
Role of S6K1 in the combination of mitomycin C and rapamycin-induced apoptosis
To further determine the role of S6K1 in the combination of mitomycin C and rapamycin-induced apoptosis, PF-4708671, a novel S6K1-specific inhibitor and a useful tool for delineating S6K1-specific roles, was employed. PF-4708671 prevents the S6K1-mediated phosphorylation of S6 protein in response to insulin-like growth factor, while having no effect upon the p90 ribosomal S6 kinase and mitogen- and stress-activated kinases.26
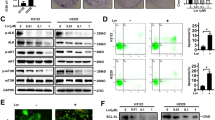
As shown in Figure 5a, we observed that PF-4708671 significantly increased either mitomycin C or the combination-induced apoptosis (PARP cleavage, hallmark of apoptosis), indicating S6K1 negatively regulated the combination treatment-induced apoptosis and S6K1 inactivation in the combination treatment contributed to the induction of apoptosis. Similar results were obtained by flow cytometry assay (Figure 5b). Interestingly, phosphorylation of Bad was also decreased by the treatment of PF-4708671, indicating that the S6K1/Bad pathway was involved in the combination treatment (Figure 5a).
Role of S6K1 in the combination of mitomycin C and rapamycin-induced apoptosis. (a) LS174T cells were pretreated with 10 μM PF-4708671 for 30 min followed by mitomycin C (5 μg/ml) and/or rapamycin (10 μg/ml) for 24 h. PARP cleavage and p-Bad/Bad were detected by immunoblotting. Actin was used to confirm the equal amount of proteins loaded in each lane. (b) After treatment, cells were stained with Annexin V and PI. Apoptosis was detected by the flow cytometric assay. (c) LS174T cells were transiently transfected with a sham plasmid, S6K1 WT-HA, S6K1 E389 DeltaCT-HA and S6K1 F5A-HA; 48 h later, cells were treated with mitomycin C (5 μg/ml) for 24 h. PARP, HA, and p-Bad/Bad were detected by immunoblotting. Actin was shown as an internal control. (d) Cell viability was analyzed by MTS assay. **P<0.01 compared with the sham group; ##P<0.01 compared with the S6K WT group. (e) LS174T cells were transiently transfected with S6K1 WT-HA, S6K1 E389 DeltaCT-HA and S6K1 F5A-HA; 48 h later, cells were treated with mitomycin C (5 μg/ml) and/or rapamycin (10 μg/ml) for 24 h. PARP, HA, p-S6K1/S6K1 and p-Bad/Bad were detected by immunoblotting. Actin was shown as an internal standard. (f) LS174T cells were transiently transfected with S6K1 WT-HA, S6K1 E389 DeltaCT-HA and S6K1 F5A-HA; 48 h later, cells were treated with mitomycin C (5 μg/ml) and/or rapamycin (10 μg/ml) for 24 h. Cell viability was analyzed by MTS assay. Error bars represent S.D. from triplicate experiments. The asterisk (*) represents a statistically significant difference between the two groups (P<0.05). (g) LS174T cells were transfected with control siRNA or Bad siRNA. After 48 h, cells were treated with mitomycin C (5 μg/ml) and/or rapamycin (10 μg/ml). Cell viability was analyzed by MTS assay 24 h later. The levels of Bad were detected by immunoblotting 72 h after transfection. Actin was used as a loading control. The asterisk (**) represents a statistically significant difference compared with the control siRNA group (P<0.01)
The activity of S6K1 is regulated by the phosphorylation events of several sites within the catalytic domain, linker and pseudosubstrate domains.27 Thr389 appears to most closely correlate with S6K1 activity.21 We transiently transfected LS174T with a sham plasmid, S6K1 WT-HA, S6K1 E389 DeltaCT-HA (C-terminal deletion and replacing Thr389 with glutamic acid, S6K1 active form) and S6K1 F5A-HA (replacing phenylalanine residue number 5 with alanine, S6K1 inactive form);28 48 h later, cells were treated with mitomycin C (5 μg/ml) for 24 h. We observed that S6K1 WT and E389 DeltaCT-HA prevented mitomycin-induced PARP cleavage compared with the sham group; S6K1 F5A-HA restored mitomycin-induced PARP cleavage (Figure 5c). As shown in Figure 5d, S6K1 WT and E389 DeltaCT-HA decreased mitomycin C-induced cell death compared with the sham group (P<0.01); S6K1 F5A-HA enhanced mitomycin C-induced cell death compared with S6K1 WT (P<0.01).
Then, LS174T cells were transiently transfected with S6K1 WT-HA, S6K1 E389 DeltaCT-HA and S6K1 F5A-HA and treated with mitomycin C (5 μg/ml) and/or rapamycin (10 μg/ml) for 24 h. We observed that PARP cleavage was inhibited with the active form S6K1 E389 DeltaCT-HA, whereas there was no difference with the inactive form S6K1 F5A-HA compared with S6K1 WT-HA in the combination treatment, indicating active S6K1 negatively regulated the combination of mitomycin C and rapamycin-induced apoptosis (Figure 5e). The expression of HA was almost equal among the three different plasmids. We also noticed that the phospho-p70 S6 Kinase (Thr389) antibody did not recognize S6K1 E389 DeltaCT-HA, and the level of p-S6K1 (Thr389) was decreased in S6K1 F5A-HA. Interestingly, we observed that the combination treatment in S6K1 E389 DeltaCT-HA failed to dephosphorylate Bad, which indicated consistently that S6K1 contributed to the phosphorylation of Bad and S6K1/Bad played an important role in the combination of mitomycin C and rapamycin-induced apoptosis. Similar results were obtained in the survival assay with the three different S6K1 plasmids in Figure 5f.
To confirm the role of Bad in the combination of mitomycin C and rapamycin-induced cell death, LS174T cells were transfected with control siRNA or Bad siRNA (Figure 5g). After 48 h, cells were treated with mitomycin C (5 μg/ml) and/or rapamycin (10 μg/ml) for 24 h. Cell viability was analyzed by MTS assay. We observed that knockdown of Bad protected the combination of mitomycin C and rapamycin-induced cell death (P<0.01), which confirmed that Bad was at least partially involved in mitomycin C and rapamycin-induced cell death.
Effect of mitomycin C and rapamycin on the growth of LS174T intraperitoneal tumors
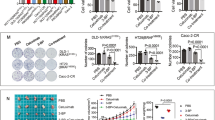
Finally, in vivo studies were performed to examine the effect of the combination treatment of mitomycin C and rapamycin on growth of LS174T intraperitoneal tumors. Figures 6a and b show that rapamycin alone induced a significant decrease in tumor growth compared with the control group (P<0.01), which was consistent with the report that rapamycin exposure caused very slight inhibition of cell growth in standard monolayer cultures, but dramatically inhibited the growth of in vivo tumors,29 which underscores the therapeutic potential of rapamycin in peritoneal tumors. Mitomycin C alone caused a statistically significant decrease of tumor growth (P<0.01). In particular, the combination of mitomycin C and rapamycin was significantly more effective at inhibiting tumor growth compared with single treatment (P<0.01). Hematoxylin and eosin (H&E) and CEA (used as a tumor marker) staining showed that all the sections were in the tumor tissue (Figure 6c). TUNEL assay confirmed more apoptotic death in tumor tissues in the combination treatment group compared with sham group and the single treatment group (Figure 6c). However, rapamycin alone did not induce significant apoptotic cell death but a decrease in Ki67 expression, which indicated that its inhibition of tumor growth was due to the inhibition of cell proliferation.
Effect of mitomycin C and rapamycin on the growth of LS174T intraperitoneal tumors. (a) Nude mice were intraperitoneally inoculated with 5 × 105 LS174T-luc cells/mouse on day 0. Four days after tumor inoculation, all the tumor-bearing mice were treated with either PBS alone (sham) or rapamycin alone at a dose of (0.25 mg/kg body weight) or treated with mitomycin C (1.5 mg/kg body weight) alone or treated with combination of mitomycin C and rapamycin every other day from day 4 to day 28. Mice were imaged using the IVIS Imaging System Series 200. (b) Line graph illustrating the luciferase activity (photons/second) in LS174T tumor-bearing mice treated with PBS alone, rapamycin alone or mitomycin C alone or combination of rapamycin with mitomycin C from week 1 to week 4. ** represents a statistically significant difference compared with the control group. ## represents a statistically significant difference compared with the single treatment group. (c) Tumor tissues were harvested at day 28 and subjected to H&E staining, immunohistochemistry staining with anti-CEA and Ki67 primary antibodies and TUNEL assay. Cell nuclei were stained with DAPI. Representative images are shown (magnification × 400)
Discussion
Recently, it has been reported that shortened survival is associated with activation of the phosphoinositide-3-kinase and mTOR signaling pathways in malignant peritoneal mesothelioma patients.30 Phase I study of everolimus and mitomycin C for patients with metastatic esophagogastric adenocarcinoma showed safety and encouraging signs of antitumor activity.31, 32 These evidences indicate that the combination of mitomycin C and rapamycin deserves further investigation and evaluation for PC. Our aim here is to study the efficacy and investigate the mechanism of mitomycin C and rapamycin on peritoneal cancer cells in vitro and in vivo. In this study, we observed that the DNA-damaging drug mitomycin C in combination with mTOR inhibitor rapamycin synergistically induced cytotoxicity and apoptosis.
The serine/threonine kinase mTOR affects its downstream effector S6K1 and the binding protein of eukaryotic translation initiation factor 4E (4EBP1) mainly through direct phosphorylation to control translation and autophagy. Moreover, mTOR also has intense apoptosis-controlling effects.33, 34 mTOR has been reported to have both pro- and anti-apoptotic functions,33, 34, 35 depending on mTOR level, the involvement of other genes, drug action, cell types and some unknown factors.34, 36
Recent studies show rapamycin inhibited mTOR complex-1 (mTORC1), but not mTOR complex-2 (mTORC2), which may limit its antitumor agents.37 Akt is another key component of the mTOR axis that involves complicated regulatory networks. Inhibition of mTORC1 induces feedback Akt activation, which decreased the anticancer efficacy.38 However, in other circumstances rapamycin decreased Akt phosphorylation and activity.39 In this study, p-Akt is increased slightly in the treatment of mitomycin C, whereas rapamycin did not significantly change its activity (Figures 2a and b), indicating that Akt played a minor role in the combination of mitomycin C and rapamycin in LS174T and LS180 cells.
S6K1, a dual pathway kinase, controls cell survival and cell death through different mechanisms: (1) phosphorylation of ribosomal subunit S6, which regulates cell growth by inducing protein synthesis components;14 and (2) binding of S6K1 to mitochondrial membranes, which phosphorylates the pro-apoptotic molecule Bad on serine 136 and inactivates Bad function.40 Overexpression of S6K1 in brain tumors has been reported to predict patients’ survival.41 In this study, combination of mitomycin C and rapamycin C potently decreased the activity of S6K1, mainly due to the effect of rapamycin. Mitomycin C alone did not have that effect; however, the combination treatment dramatically decreased the activity of S6K1. From our observations, it is possible that S6K1 could be used as a biomarker to predict the response to the treatment of mitomycin C and rapamycin. Of note, mitomycin C but not rapamycin induced sustained JNK activation. JNK is also well known to promote Bax translocation to mitochondria.42, 43 The question of how JNK and S6K1 interplay and induce synergistic effect awaits further investigation.
Bad is a BH3 domain-only pro-apoptotic member of the Bcl-2 family that regulates cell death and survival through posttranslational modification.22 Data from siRNA knockdown experiments demonstrated that Bad played an important role as a pro-apoptotic protein in mitomycin C and rapamycin-induced cell death. Bad phosphorylated on three serine residues (S112, S136 and S155) is reported to be the inactive moiety sequestered in the cytosol and then embedded within 14-3-3 consensus binding sites, disrupting the binding of Bad to Bcl-xL or Bcl-2 to promote survival.23, 44, 45 Both Akt/PKB and S6K1 were shown to phosphorylate Bad, which inactivated the pro-apoptotic function of Bad.41, 46 Bad phosphorylation was reported to determine ovarian cancer chemosensitivity and patient survival47 and could possibly serve as a biomarker to predict the response to the treatment of mitomycin C and rapamycin. In our current study, the combination of mitomycin C and rapamycin inactivated not only Akt but also S6K1, led to dephosphorylated Bad, and resulted in dissociation of Bcl-xL from pro-apoptotic protein Bak, inducing mitochondria-dependent apoptosis. LS174T is a Bax null cell line25 and a good model to study the role of Bak. We observed Bak oligomerization was sufficient to induce mitochondria-dependent apoptosis by the combination of mitomycin C and rapamycin that was consistent with other reports.46, 48
Taken together, we presented here a novel combination treatment of chemotherapeutic agent mitomycin C and mTOR inhibitor rapamycin that induced synergistic apoptosis through the S6K1–Bad–Bak pathway in PC. Mitomycin C and rapamycin are both commonly used FDA-approved drugs and could be considered for PC patients in clinics. Understanding the mechanisms involved in this combination treatment is important not only to predict and interpret the responses but also to enhance the efficacy of this combination.
Materials and Methods
Cell cultures and transfection
LS174T and LS180 are human colon adenocarcinoma cell lines and were purchased from American Type Culture Collection (ATCC, Manassas, VA, USA) and cultured in Eagle’s Minimum Essential Medium (ATCC, Rockville, MD, USA) and 10% fetal bovine serum at 37°C and 5% CO2. For transient transfection, cells were transfected with Lipofectamine 2000 (Life Technologies, Gaithersburg, MD, USA), and were treated 48 h after transfection.
Reagents and antibodies
Mitomycin C was from Santa Cruz Biotechnology (Santa Cruz, CA, USA). Rapamycin and protease inhibitor cocktail were obtained from Sigma-Aldrich (St Louis, MO, USA). S6K1WT-HA, S6K1E389DeltaCT-HA and S6K1F5A were purchased from Addgene (Cambridge, MA, USA). Anti-HA, anti-phospho-Akt, anti-Akt, anti-CEA, anti-phospho-JNK, anti-JNK, anti-Bak, anti-caspase 8, anti-caspase 9, anti-caspase 3, anti-phospho-p70 S6 kinase (Thr389), anti-p70 S6 kinase, anti-phospho-Bad (Ser136), anti-Bad, anti-COX-IV and anti-PARP antibody were from Cell Signaling (Danvers, MA, USA). Anti-actin antibody was from Sigma-Aldrich. Anti-cytochrome c antibody was from BD PharMingen (San Jose, CA, USA). Anti-Ki67 was purchased from Dako (Carpinteria, CA, USA).
MTS [3-(4,5-dimethylthiazol-2-yl)-5-(3-carboxymethoxyphenyl)-2-(4-sulfophenyl)-2H-tetrazolium] assays
MTS studies were carried out using the Promega CellTiter 96 AQueous One Solution Cell Proliferation Assay (Promega, Madison, WI, USA). CX-1 cells were grown in tissue culture-coated 96-well plates and treated as described in Results. Cells were then treated with the MTS/phenazine methosulfate solution for 1 h at 37°C. Absorbance at 490 nm was determined using an enzyme-linked immunosorbent assay plate reader.
Annexin V binding
Cells were harvested by trypsinization, washed in a serum-free medium, and suspended in binding buffer (Annexin V-FITC Staining Kit, PharMingen). This cell suspension was stained with mouse anti-human Annexin V antibody and PI and immediately analyzed by flow cytometry.
Immunoblot analysis
Cells were lysed with Laemmli lysis buffer and boiled for 10 min. The protein content was measured with BCA Protein Assay Reagent (Thermo Scientific, Hudson, NH, USA), separated by SDS-PAGE and electrophoretically transferred to a nitrocellulose membrane. The nitrocellulose membrane was blocked with 5% nonfat dry milk in PBS-Tween-20 (0.1%, v/v) for 1 h and incubated with primary antibody at room temperature for 2 h. Horseradish peroxidase conjugated anti-rabbit or anti-mouse IgG was used as the secondary antibody. Immunoreactive protein was visualized by the chemiluminescence protocol. To ensure equal protein loading, each membrane was stripped and reprobed with anti-actin antibody to normalize for differences in protein loading. Densitometry analysis was performed with Gel-pro analyzer program (Media Cybernetics, Silver Spring, MD, USA).
Immunoprecipitation
Briefly, cells were lysed in CHAPS lysis buffer with protease inhibitor cocktail (Calbiochem, San Diego, CA, USA). Approximately, 0.5–1 mg of lysate was incubated with 1.5 μg of anti-Bad/Bcl-xL antibody or rabbit IgG (Santa Cruz) at 4°C overnight, followed by the addition of protein G PLUS-agarose beads (Santa Cruz) and rotation at room temperature for 2 h, followed by immunoblot analysis.
JC-1 mitochondrial membrane potential assay
JC-1 dye was used to monitor mitochondrial transmembrane potential (ΔΨm). In the undamaged mitochondria, the aggregated dye appears as red fluorescence, whereas in the apoptotic cell with altered ΔΨm, the dye remains as monomers in the cytoplasm with diffuse green fluorescence. The red/green fluorescence ratio is dependent on ΔΨm. Cells were then stained with JC-1 mitochondrial membrane potential detection kit (Invitrogen, Carlsbad, CA, USA) for 10 min and analyzed by flow cytometry. Fluorescence intensity was measured with the Accuri C6 flow cytometer (Accuri Cytometers, Inc, San Jose, MI, USA). Results were analyzed with VenturiOne software (Applied Cytometry, Inc, Plano, TX, USA).
Bak oligomerization
Cells were pelleted and resuspended in HB buffer. The cell suspension was homogenized, and spun at 1000 × g for 15 min at 4°C. The supernatant was transferred and spun at 10 000 × g for 15 min at 4°C to pellet mitochondria. Aliquots of isolated mitochondrial fractions and cytosolic fractions were cross-linked with 1 mM dithiobis (Pierce, Rockford, IL, USA). Samples were subjected to SDS-PAGE under non-denaturing conditions followed by immunoblotting for Bak.
Confocal microscope
Cells were stained with 300 nM MitoTracker (Invitrogen). Cells were washed three times with 0.5% BSA in PBS, followed by fixation in 4% paraformaldehyde for 15 min. Bak was stained with anti-Bak antibody. Nuclei were stained with DAPI (Cell Signaling). Slides were mounted and visualized in 0.4-μm sections using an OlympusFluoview 1000 confocal microscope and the companion software FV10-ASW2.1 under a × 63 oil immersion objective.
Measurement of cytochrome c release
To determine the release of cytochrome c from the mitochondria, mitochondrial and cytosolic fractions were prepared using the Mitochondrial Fractionation Kit (Active Motif, Carlsbad, CA, USA) following company instructions and reagents included in the kit. Cytosolic fractions were subjected to SDS-PAGE gel electrophoresis and analyzed by immunoblotting using anti-cytochrome c antibody.
Knockdown of Bad with siRNA oligomers
To generate Bad-knockdown LS174T cells, cells were transfected with 200 nM of siRNA Bad or control siRNA from Sigma-Aldrich, using Lipofectamine 2000 (Invitrogen). Target sequences for preparing siRNAs of human Bad were as follows: 5′-AAGAAGGGACUUCCUCGCCCG-3′ (sense strand) and 5′-CGGGCGAGGAAGUCCCUUCUU-3′ (complement strand). Expression levels were determined by immunoblot analysis.
Animal model
Nude mice were obtained from Taconic (Germantown, NY, USA) at 6 weeks old. Stably transduced LS174T cells expressing the firefly luciferase gene were generated by lentiviral transfection of the pGL4 Luciferase Reporter Vector (Promega) and selected with puromycin. Mice were intraperitoneally challenged with 5 × 105 LS174T luciferase cells. On the fourth day, the mice were examined using the IVIS image system for bioluminescent activity indicating basal tumor loading. Briefly, the mice were injected with the luciferase substrate luciferin (150 mg/kg i.p.; GoldBio, St. Louis, MO, USA), and anesthetized with 2% isoflurane. Bioluminescence images were acquired 5 min after luciferin administration on an IVIS200 system (Perkin Elmer, Waltham, MA, USA). Bioluminescence signal was quantified using the LivingImage software (Perkin Elmer). Mice with the same level of bioluminescence were divided into four groups (five per group) and subjected to different treatments, including control, rapamycin (0.25 mg/kg body weight, p.o.) alone, mitomycin C (1.5 mg/kg body weight, i.p.) alone or in combination. Each group of mice was treated every other day. Signs of discomfort, moribund and weight loss were not observed during the combinatorial treatment in tumor bearing and control mice. The tumor load, represented by bioluminescence signal determined using the IVIS bioluminescence Imaging System was checked from week 1 to week 4. Mice were fed ad libitum and maintained in environments with controlled temperature of 22–24°C and 12 h light and dark cycles. All animal experiments were carried out at the University of Pittsburgh in accordance with the Guide for the Care and Use of Laboratory Animals.
TUNEL assay
The TUNEL method was used to detect apoptotic cells. An in situ cell apoptosis detection kit (Trevigen, Gaithersburg, MD, USA) was used. The staining was performed according to the manufacturer’s instructions. Tissue sections in the vehicle group were stained and served as negative controls. Briefly, sections of paraffin-embedded tissues were deparaffinized, and then washed with TBS buffer and permeabilized with proteinase K. DNA strand breaks were then end-labeled with terminal transferase, and the labeled DNA was visualized by fluorescence microscopy.
Immunohistochemical staining
Tumor tissues recovered from each group were fixed in 10% neutral buffered formalin and embedded in paraffin. Sections were cut 4 μm thick and then were stained either by standard H&E staining or immunohistochemical staining with mAbs specific for human CEA and Ki67, and were visualized with the Envision+ dual link system (Dako) following the manufacturer’s instructions. Pictures were taken using an Olympus BH2 microscope (Olympus, Tokyo, Japan) with a camera (Coolsnap3.3; Photometrics, Tucson, AZ, USA). Tissue sections of inoculation sites were examined under an Olympus BX71 microscope (Olympus, Lake Success, NY, USA) equipped with Hoffman objective lenses ( × 40; Diagnostic Instruments, Sterling Heights, MI, USA) connected to a SPOT One Digital Camera (Diagnostic Instruments). Images were then processed using Adobe Photoshop software (Adobe, San Jose, CA, USA).
Statistical analysis
Statistical analysis was carried out using Graphpad InStat 3 software (GraphPad Software, Inc, San Diego, CA, USA). Data showing comparisons between two groups were assessed using the Student’s t-test. Comparisons among more than two groups were done using ANOVA with the appropriate post hoc testing. Statistical significance is marked with asterisks (*P<0.05 and ** or ##P<0.01).
Abbreviations
- BAD:
-
Bcl-2-associated death promoter
- BAK:
-
Bcl-2 homologous antagonist/killer
- BAX:
-
Bcl-2-associated X protein
- CEA:
-
carcinoembryonic antigen
- JNK:
-
c-Jun NH2-terminal kinase
- mTOR:
-
mammalian target of rapamycin
- PARP:
-
poly (ADP-ribose) polymerase
- PAGE:
-
polyacrylamide gel electrophoresis
- SDS:
-
sodium dodecyl sulfate
- S6K1:
-
p70 S6 ribosomal kinase
- PC:
-
peritoneal carcinomatosis
References
Seidl C, Essler M . Radioimmunotherapy for peritoneal cancers. Immunotherapy 2013; 5: 395–405.
Knorr C, Reingruber B, Meyer T, Hohenberger W, Stremmel C . Peritoneal carcinomatosis of colorectal cancer: incidence, prognosis, and treatment modalities. Int J Colorectal Dis 2004; 19: 181–187.
Ceelen WP . Current management of peritoneal carcinomatosis from colorectal cancer. Minerva Chir 2013; 68: 77–86.
Pirnia F, Schneider E, Betticher DC, Borner MM . Mitomycin C induces apoptosis and caspase-8 and -9 processing through a caspase-3 and Fas-independent pathway. Cell Death Differ 2002; 9: 905–914.
Wu KY, Wang HZ, Hong SJ . Mechanism of mitomycin-induced apoptosis in cultured corneal endothelial cells. Mol Vis 2008; 14: 1705–1712.
Van der Speeten K, Stuart OA, Chang D, Mahteme H, Sugarbaker PH . Changes induced by surgical and clinical factors in the pharmacology of intraperitoneal mitomycin C in 145 patients with peritoneal carcinomatosis. Cancer Chemother Pharmacol 2011; 68: 147–156.
Cohen MS, Al-Kasspooles MF, Williamson SK, Henry D, Broward M, Roby KF . Combination intraperitoneal chemotherapy is superior to mitomycin C or oxaliplatin for colorectal carcinomatosis in vivo. Ann Surg Oncol 2010; 17: 296–303.
Kornek GV, Schuell B, Laengle F, Gruenberger T, Penz M, Karall K et al. Mitomycin C in combination with capecitabine or biweekly high-dose gemcitabine in patients with advanced biliary tract cancer: a randomised phase II trial. Ann Oncol 2004; 15: 478–483.
Saif MW, Kaley K, Brennan M, Garcon MC, Rodriguez G . Mitomycin-C and capecitabine (MIXE) as salvage treatment in patients with refractory metastatic colorectal cancer: a retrospective study. Anticancer Res 2013; 33: 2743–2746.
Guertin DA, Sabatini DM . An expanding role for mTOR in cancer. Trends Mol Med 2005; 11: 353–361.
Tee AR, Proud CG . Staurosporine inhibits phosphorylation of translational regulators linked to mTOR. Cell Death Differ 2001; 8: 841–849.
Tee AR, Proud CG . DNA-damaging agents cause inactivation of translational regulators linked to mTOR signalling. Oncogene 2000; 19: 3021–3031.
Ramanathan A, Schreiber SL . Direct control of mitochondrial function by mTOR. Proc Natl Acad Sci USA 2009; 106: 22229–22232.
Choo AY, Yoon SO, Kim SG, Roux PP, Blenis J . Rapamycin differentially inhibits S6Ks and 4E-BP1 to mediate cell-type-specific repression of mRNA translation. Proc Natl Acad Sci USA 2008; 105: 17414–17419.
Fasolo A, Sessa C . mTOR inhibitors in the treatment of cancer. Expert Opin Investig Drugs 2008; 17: 1717–1734.
Faivre S, Kroemer G, Raymond E . Current development of mTOR inhibitors as anticancer agents. Nat Rev Drug Discov 2006; 5: 671–688.
Dhanasekaran DN, Reddy EP . JNK signaling in apoptosis. Oncogene 2008; 27: 6245–6251.
Tandon P, Gallo CA, Khatri S, Barger JF, Yepiskoposyan H, Plas DR . Requirement for ribosomal protein S6 kinase 1 to mediate glycolysis and apoptosis resistance induced by Pten deficiency. Proc Natl Acad Sci USA 2011; 108: 2361–2365.
Dudek H, Datta SR, Franke TF, Birnbaum MJ, Yao R, Cooper GM et al. Regulation of neuronal survival by the serine-threonine protein kinase Akt. Science 1997; 275: 661–665.
Downward J . Mechanisms and consequences of activation of protein kinase B/Akt. Curr Opin Cell Biol 1998; 10: 262–267.
Pearson RB, Dennis PB, Han JW, Williamson NA, Kozma SC, Wettenhall RE et al. The principal target of rapamycin-induced p70s6k inactivation is a novel phosphorylation site within a conserved hydrophobic domain. Embo J 1995; 14: 5279–5287.
Danial NN . BAD: undertaker by night, candyman by day. Oncogene 2008; 27 (Suppl 1): S53–S70.
Yang E, Zha J, Jockel J, Boise LH, Thompson CB, Korsmeyer SJ . Bad, a heterodimeric partner for Bcl-XL and Bcl-2, displaces Bax and promotes cell death. Cell 1995; 80: 285–291.
Wei MC, Zong WX, Cheng EH, Lindsten T, Panoutsakopoulou V, Ross AJ et al. Proapoptotic BAX and BAK: a requisite gateway to mitochondrial dysfunction and death. Science 2001; 292: 727–730.
Carethers JM, Pham TT . Mutations of transforming growth factor beta 1 type II receptor, BAX, and insulin-like growth factor II receptor genes in microsatellite unstable cell lines. In Vivo 2000; 14: 13–20.
Pearce LR, Alton GR, Richter DT, Kath JC, Lingardo L, Chapman J et al. Characterization of PF-4708671, a novel and highly specific inhibitor of p70 ribosomal S6 kinase (S6K1). Biochem J 2010; 431: 245–255.
Dennis PB, Pullen N, Kozma SC, Thomas G . The principal rapamycin-sensitive p70(s6k) phosphorylation sites, T-229 and T-389, are differentially regulated by rapamycin-insensitive kinase kinases. Mol Cell Biol 1996; 16: 6242–6251.
Schalm SS, Blenis J . Identification of a conserved motif required for mTOR signaling. Curr Biol 2002; 12: 632–639.
Liu M, Howes A, Lesperance J, Stallcup WB, Hauser CA, Kadoya K et al. Antitumor activity of rapamycin in a transgenic mouse model of ErbB2-dependent human breast cancer. Cancer Res 2005; 65: 5325–5336.
Varghese S, Chen Z, Bartlett DL, Pingpank JF, Libutti SK, Steinberg SM et al. Activation of the phosphoinositide-3-kinase and mammalian target of rapamycin signaling pathways are associated with shortened survival in patients with malignant peritoneal mesothelioma. Cancer 2011; 117: 361–371.
Wagner OJ, Inglin RA, Bisch-Knaden S, Mettler D, Borner M, Candinas D et al. Sirolimus and intraoperative hyperthermic peritoneal chemoperfusion with mitomycin-C do not impair healing of bowel anastomoses. Transpl Int 2008; 21: 554–563.
Werner D, Atmaca A, Pauligk C, Pustowka A, Jager E, Al-Batran SE . Phase I study of everolimus and mitomycin C for patients with metastatic esophagogastric adenocarcinoma. Cancer Med 2013; 2: 325–333.
Shang YC, Chong ZZ, Wang S, Maiese K . Prevention of beta-amyloid degeneration of microglia by erythropoietin depends on Wnt1, the PI 3-K/mTOR pathway, Bad, and Bcl-xL. Aging (Albany NY) 2012; 4: 187–201.
Wang Y, Hu Z, Liu Z, Chen R, Peng H, Guo J et al. MTOR inhibition attenuates DNA damage and apoptosis through autophagy-mediated suppression of CREB1. Autophagy 2013; 9: 2069–2086.
Chen L, Xu B, Liu L, Luo Y, Yin J, Zhou H et al. Hydrogen peroxide inhibits mTOR signaling by activation of AMPKalpha leading to apoptosis of neuronal cells. Lab Invest 2010; 90: 762–773.
Maiese K, Chong ZZ, Shang YC, Wang S . Targeting disease through novel pathways of apoptosis and autophagy. Expert Opin Ther Targets 2012; 16: 1203–1214.
Shor B, Gibbons JJ, Abraham RT, Yu K . Targeting mTOR globally in cancer: thinking beyond rapamycin. Cell Cycle 2009; 8: 3831–3837.
Wang X, Sun SY . Enhancing mTOR-targeted cancer therapy. Expert Opin Ther Targets 2009; 13: 1193–1203.
Dormond O, Madsen JC, Briscoe DM . The effects of mTOR-Akt interactions on anti-apoptotic signaling in vascular endothelial cells. J Biol Chem 2007; 282: 23679–23686.
Harada H, Andersen JS, Mann M, Terada N, Korsmeyer SJ . p70S6 kinase signals cell survival as well as growth, inactivating the pro-apoptotic molecule BAD. Proc Natl Acad Sci USA 2001; 98: 9666–9670.
Ismail HM . Overexpression of s6 kinase 1 in brain tumours is associated with induction of hypoxia-responsive genes and predicts patients’ survival. J Oncol 2012; 2012: 416927.
Tsuruta F, Sunayama J, Mori Y, Hattori S, Shimizu S, Tsujimoto Y et al. JNK promotes Bax translocation to mitochondria through phosphorylation of 14-3-3 proteins. EMBO J 2004; 23: 1889–1899.
Song X, Kim SY, Lee YJ . The role of Bcl-xL in synergistic induction of apoptosis by mapatumumab and oxaliplatin in combination with hyperthermia on human colon cancer. Mol Cancer Res 2012; 10: 1567–1579.
Taghiyev AF, Guseva NV, Harada H, Knudson CM, Rokhlin OW, Cohen MB . Overexpression of BAD potentiates sensitivity to tumor necrosis factor-related apoptosis-inducing ligand treatment in the prostatic carcinoma cell line LNCaP. Mol Cancer Res 2003; 1: 500–507.
Masters SC, Yang H, Datta SR, Greenberg ME, Fu H . 14-3-3 inhibits Bad-induced cell death through interaction with serine-136. Mol Pharmacol 2001; 60: 1325–1331.
Aluvila S, Mandal T, Hustedt E, Fajer P, Choe JY, Oh KJ . Organization of the mitochondrial apoptotic BAK pore: oligomerization of the BAK homodimers. J Biol Chem 2013; 11: 11.
Marchion DC, Cottrill HM, Xiong Y, Chen N, Bicaku E, Fulp WJ et al. BAD phosphorylation determines ovarian cancer chemosensitivity and patient survival. Clin Cancer Res 2011; 17: 6356–6366.
Pecot CV, Rupaimoole R, Yang D, Akbani R, Ivan C, Lu C et al. Tumour angiogenesis regulation by the miR-200 family. Nat Commun 2013; 4: 2427.
Acknowledgements
This work was supported by the NCI grant fund (CA140554). This project used the UPCI Core Facility and was supported in part by the award P30CA047904.
Author contributions
Conceived and designed the experiments, and wrote the paper: XS and YJL. Performed the experiments: XS, A-KD, and S-YK. Analyzed the data: XS, HAC, and YJL.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by R Aqeilan
Rights and permissions
Cell Death and Disease is an open-access journal published by Nature Publishing Group. This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Song, X., Dilly, AK., Kim, SY. et al. Rapamycin-enhanced mitomycin C-induced apoptotic death is mediated through the S6K1–Bad–Bak pathway in peritoneal carcinomatosis. Cell Death Dis 5, e1281 (2014). https://doi.org/10.1038/cddis.2014.242
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2014.242
This article is cited by
-
8-Hydroxyquinoline a natural chelating agent from Streptomyces spp. inhibits A549 lung cancer cell lines via BCL2/STAT3 regulating pathways
World Journal of Microbiology and Biotechnology (2022)
-
mTOR is a fine tuning molecule in CDK inhibitors-induced distinct cell death mechanisms via PI3K/AKT/mTOR signaling axis in prostate cancer cells
Apoptosis (2016)