Abstract
The presence of α-synuclein aggregates in the characteristic Lewy body pathology seen in idiopathic Parkinson’s disease (PD), together with α-synuclein gene mutations in familial PD, places α-synuclein at the center of PD pathogenesis. Decreased levels of the chaperone-mediated autophagy (CMA) proteins LAMP-2A and hsc70 in PD brain samples suggests compromised α-synuclein degradation by CMA may underpin the Lewy body pathology. Decreased CMA protein levels were not secondary to the various pathological changes associated with PD, including mitochondrial respiratory chain dysfunction, increased oxidative stress and proteasomal inhibition. However, decreased hsc70 and LAMP-2A protein levels in PD brains were associated with decreases in their respective mRNA levels. MicroRNA (miRNA) deregulation has been reported in PD brains and we have identified eight miRNAs predicted to regulate LAMP-2A or hsc70 expression that were reported to be increased in PD. Using a luciferase reporter assay in SH-SY5Y cells, four and three of these miRNAs significantly decreased luciferase activity expressed upstream of the lamp-2a and hsc70 3′UTR sequences respectively. We confirmed that transfection of these miRNAs also decreased endogenous LAMP-2A and hsc70 protein levels respectively and resulted in significant α-synuclein accumulation. The analysis of PD brains confirmed that six and two of these miRNAs were significantly increased in substantia nigra compacta and amygdala respectively. These data support the hypothesis that decreased CMA caused by miRNA-induced downregulation of CMA proteins plays an important role in the α-synuclein pathology associated with PD, and opens up a new avenue to investigate PD pathogenesis.
Similar content being viewed by others
Main
Parkinson’s disease (PD) is a progressive neurodegenerative disorder characterized pathologically by the loss of dopaminergic neurons in the substantia nigra pars compacta (SNc) and the presence of Lewy bodies (LBs) that contain aggregates of α-synuclein protein.1 The finding of mutations2 and multiplications3 of the α-synuclein gene in familial forms of PD and the genome-wide association studies linking the α-synuclein gene to sporadic PD4 further reinforces the importance of α-synuclein in PD pathogenesis. The accumulation and aggregation of α-synuclein in PD may reflect changes to its synthesis and/or degradation. Although increased α-synuclein gene copy number3 supports the role of increased α-synuclein synthesis in PD, there is increasing evidence that the degradation pathways responsible for α-synuclein turnover may also be compromised in PD.5, 6
Many cytosolic proteins are targeted for degradation in lysosomes by chaperone-mediated autophagy (CMA).7 In this pathway, a pentapeptide motif (KFERQ) present in the protein is recognized by the heat shock cognate protein 70 (hsc70) chaperone and internalized into the lysosome by the membrane receptor lysosomal-associated membrane protein 2A (LAMP-2A).8 α-Synuclein contains a pentapeptide sequence (VKKDQ) demonstrated to target it to the lysosome5 and, in cells, we6 and others9 have confirmed that inhibition of CMA through downregulation of LAMP-2A protein levels leads to α-synuclein accumulation. The relevance of this mechanism to α-synuclein pathology in PD has been emphasized by our recent observation that LAMP-2A and hsc70 proteins were decreased in the SNc and amygdala in PD brains compared with both age-matched controls and brain samples from Alzheimer’s patients.6 Consequently, it is important to understand the mechanism that leads to the decrease in these CMA proteins in PD patients.
A mechanism that has been suggested to play an important role in the regulation of up to a third of coding mRNAs involves the expression of microRNAs (miRNAs), which are single-stranded RNA molecules of between 21 and 23 nucleotides in length.10 Many different miRNAs have been identified and shown to down-regulate protein translation or promote degradation of specific mRNA molecules through association with the RNA-induced silencing complex.11 miRNAs are involved in the normal function of eukaryotic cells and miRNA deregulation has been associated with various diseases.12 A recent expression analysis of 224 miRNAs in PD brains showed significant variation in the level of many miRNAs in comparison with controls,13 suggesting that miRNA deregulation may play a role in PD pathology.
PD is also associated with a range of other biochemical changes in the brain, the most consistent including oxidative stress, mitochondrial dysfunction and decreased proteasomal function. Consequently, we have investigated whether these pathological changes or changes in miRNA levels underpin the decreased levels of CMA proteins observed in PD brains and can cause the α-synuclein accumulation characteristic of PD pathology.
Results
To investigate whether biochemical features associated with PD influenced LAMP-2A or hsc70 levels, normal SH-SY5Y cells were exposed to conditions previously demonstrated to induce oxidative stress (50 and 100 μM paraquat), mitochondrial dysfunction (100, 200 nM rotenone) or proteasome dysfunction (5 and 10 nM epoxomycin), but in the absence of marked cell death (Supplementary Figure 1a).14, 15 None of these conditions significantly affected LAMP-2A or hsc70 protein (Figures 1a and b) or mRNA levels (Figure 1c) in normal SH-SY5Y cells. α-Synuclein protein levels in normal SH-SH5Y cells are not readily detectable, consequently to determine the effect of these stresses on α-synuclein levels a stable cell line overexpressing wild-type (WT) α-synuclein was exposed to optimal concentrations of these agents (5 nM epoxomycin, 100 μM paraquat or 200 nM rotenone) for 48 h. Confirming the previous experiments, LAMP-2A and hsc70 protein levels were unaltered (Figures 1d and e); however all three treatments caused a significant increase in α-synuclein protein levels (Figures 1d and f). This was only associated with increased α-synuclein mRNA levels with epoxomycin treatment (Figure 1f), suggesting UPS inhibition increased α-synuclein transcription. To determine if this increase in transcription was also observed for the endogenous α-synuclein gene, normal SH-SY5Y and SKMEL-28 cells were treated with these agents. There was no effect on endogenous α-synuclein mRNA expression with any of these agents (Supplementary Figure 1b). This suggested that proteasome inhibition influenced the transcriptional regulation of exogenous α-synuclein under the control of the cytomegalovirus (CMV) promoter but did not influence endogenous α-synuclein regulation.
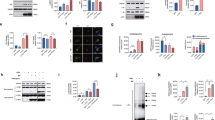
Influence of pathological features associated with PD on LAMP-2A, hsc70 and α-synuclein levels. (a–c) Normal SH-SY5Y cells after 48 h of treatment with different concentrations of epoxomycin (E), paraquat (P), rotenone (R) or untreated (C). (a) Western blot analyses of hsc70, LAMP-2A and actin. (b) Quantitation of the western blot data for LAMP-2A (open bars) and hsc70 (closed bars) protein levels relative to actin and normalized to untreated cells. (c) qPCR analysis of hsc70 and LAMP-2A mRNA relative to actin mRNA and normalized to untreated cells. (d–f) SH-SY5Y cells overexpressing WT α-synuclein after 48 h of exposure to epoxomycin (E, 5 nM), paraquat (P, 100 μM), rotenone (R, 200 nM) or under normal conditions (C). (d) Western blot analyses and (e) quantitation of the western blot data for LAMP-2A (open bars) and hsc70 (closed bars) protein levels relative to actin and normalized to untreated (control) cells. (f) Quantitation of the western blot data for α-synuclein protein relative to actin and mRNA levels relative to actin mRNA; all data normalized to untreated (control) cells. Data expressed as mean±S.E.M. (n=3), and statistical analyses compared with the respective control group, *P<0.05
Using the miRBase Target (Sanger Institute, Cambridge, UK) database, we analyzed the miRNAs evaluated by Kim et al.13 to identify which of the miRNAs expressed in brain were predicted to target the 3′ untranslated region (3′UTR) region of lamp-2a or hsc70 mRNA in humans. Although lamp-2a, lamp-2b and lamp-2c have common exons, they differ in their 3′UTRs and exon 9,16, 17 and therefore will be differentially influenced by specific miRNAs. Four miRNAs strongly predicted to target lamp-2a mRNA (hsa-miR-224; hsa-miR-320a; hsa-miR-373*; and hsa-miR-379), and two predicted to target hsc70 mRNA (hsa-miR-26b and hsa-miR-106a*; Table 1) were reported to be increased in PD brains.13 In addition, we used miRBase to identify which of the miRNAs reported to be expressed in the brain, but not previously shown to be increased in PD,13 gave the highest target prediction for lamp-2a (hsa-miR-21) or hsc70 (hsa-miR-301b).
To determine the ability of these miRNAs to target the 3′UTR sequences of lamp-2a or hsc70, two luciferase reporter constructs were generated in psiCheck2.2 with the 3′UTR sequence for either hsc70 or lamp-2a (Figure 2a). SH-SY5Y cells were transfected with these luciferase constructs and increasing concentrations (0–50 nM) of two different miRNAs predicted to target the 3′UTR of either hsc70 (hsa-miR-106a*) or lamp-2a (hsa-miR-224). After 48 h, the analysis of luciferase activity demonstrated that hsa-miR-106a* and hsa-miR-224 caused dose-dependent decreases in the activity of luciferase with the hsc70 and lamp-2a 3′UTRs, respectively (Figure 2b). However, as a negative control, even at the higher concentrations, these miRNAs had no effect on the luciferase activity linked to the alternative 3′UTR sequences (Figure 2b). Using 10 nM miRNAs to minimize nonspecific effects, four miRNAs (hsa-miR-21*; hsa-miR-224; hsa-miR-373*; and hsa-miR-379) and three miRNAs (hsa-miR-26b; hsa-miR-106a*; and hsa-miR-301b) significantly decreased the luciferase activity linked to lamp2a and hsc70 3′UTRs, respectively (Figure 2c). However, hsa-miR-320a, which was predicted to target the 3′UTR of lamp-2a, did not affect the luciferase activity linked to lamp-2a 3′UTR. As an additional control, the specificity of the miRNAs for the predicted lamp2a or hsc70 3′UTR sequences were confirmed for miRNAs hsa-miR-373*, hsa-miR-379*, hsa-miR-106a* and hsa-miR-301b using the luciferase constructs where the putative recognition sequence was mutated (Supplementary Figure 2a).
Luciferase reporter assays to analyze the influence of miRNAs on the 3′UTR of lamp-2a and hsc70. (a) Schematic representation of the Renilla luciferase reporter constructs in psiCHECK2.2 for lamp-2a and hsc70 3′UTR. (b) The influence of increasing concentrations of hsa-miR-106a* and hsa-miR-224 on Renilla luciferase activity when cotransfected with luciferase-3′UTR hsc70 or luciferase-3′UTR lamp-2a constructs. (c) The effect of cotransfection of the different miRNAs (10 nM) with either luciferase-3′UTR hsc70 or luciferase-3′UTR lamp-2a reporter constructs upon Renilla luciferase activity 48 h after transfection. Data normalized to cells in the absence of miRNA (C). Values are mean±S.E.M. (n=6), statistical analyses compared with the respective control group, *P<0.05, **P<0.01, ***P<0.001
Increasing concentrations (5,10 and 50 nM) of the eight miRNAs under investigation were transfected into normal SH-SY5Y cells and their impact on endogenous LAMP-2A or hsc70 protein levels evaluated. Consistent with the luciferase reporter assays, 7 of the miRNAs resulted in a dose-dependent decrease in either LAMP-2A or hsc70 protein levels at 10 and 50 nM, whereas the remaining miRNA (hsa-miR-320a) had no effect even at 50 nM (Supplementary Figure 2b). For subsequent studies, the 7 effective miRNAs were used at 10 nM, predicted to give a 30–70% decrease in protein levels. After transfection of normal SH-SY5Y cells with the 7 selected miRNAs (10 nM, 72 h), the 3 miRNAs predicted to target hsc70 and the 4 miRNAs predicted to target lamp-2a all decreased the levels of the respective proteins relative to actin (Figures 3a and b), but these were only statistically significant for hsa-miR-106a* and hsa-miR-301 (hsc70 protein) and hsa-miR-224, hsa-miR-373* and hsa-miR-379 (LAMP-2A protein). There were no changes to the levels of lamp-2a or hsc70 mRNA relative to actin mRNA (Supplementary Figure 3a). The impact of these changes on intracellular α-synuclein levels was analyzed using SH-SY5Y cells overexpressing α-synuclein and an extended 9-day protocol. The absence of 3′UTR sequences with the overexpressed α-synuclein excluded any direct influences of the miRNAs on α-synuclein regulation and the analysis after 9 days permitted sufficient time for the influence of decreased CMA proteins and altered α-synuclein turnover rates to be detected. Using this protocol the miRNA-dependent decrease in hsc70 and LAMP-2A protein levels reported in the normal SH-SY5Y cells were replicated in the α-synuclein overexpressing cells (Figures 3c and d). In addition, as a control, the miRNAs predicted to target lamp-2a or hsc70 had no influence on hsc70 or LAMP-2A protein levels respectively, confirming the relative specificity of these miRNAs at these concentrations (Figures 3c and d). α-Synuclein protein levels increased significantly in response to all the miRNAs tested (Figures 3c and d). Although α-synuclein mRNA levels did not change significantly for most of the miRNAs used (Supplementary Figure 3b), hsa-miR-106a* and hsa-miR-301b caused a significant decrease in α-synuclein mRNA levels (Supplementary Figure 3b). These miRNAs also caused a decrease in endogenous α-synuclein mRNA (Supplementary Figure 3a), suggesting they may target α-synuclein transcripts. Indeed hsa-miR-106a* is predicted to target the 3′UTR of α-synuclein with a miRBase Target predicted score of 15.22. The absence of increased α-synuclein transcript levels suggests that upregulation of miRNAs predicted to target specific CMA proteins were sufficient to result in intracellular α-synuclein protein accumulation.
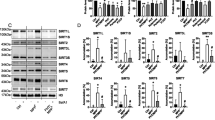
Impact of increased miRNA levels upon LAMP-2A, hsc70 and α-synuclein protein levels. (a and b) Western blot analyses of normal SHSY5Y cells 72 h after transfection with the respective miRNAs (10 nM) and untreated cells (C) for (a) LAMP-2A, hsc70 and actin (b) and their quantitation. (c and d) Western blot analyses of α-synuclein overexpressing cells 9 days after treatment with miRNAs (10 nM) for (c) α-synuclein, LAMP-2A, hsc70 and actin. (d) Quantitation of α-synuclein, LAMP-2A and hsc70 protein levels. Values are relative to actin and normalized to untreated (control) cells. Data expressed as mean±S.E.M. (n=3), statistical analyses compared with the respective control group, *P<0.05
We previously reported a significant decrease in the CMA proteins LAMP-2A and hsc70 in PD SNc and amygdala samples.6 To determine if increased miRNA levels could be responsible for these changes, we analyzed the levels of the miRNAs shown to influence these proteins in control and PD SNc and amygdala samples matched for post-mortem delay (mean (S.D.): control, 4.8 (2.1) h; PD, 4.1 (3.0) h) and age (mean (S.D.): control, 70.2 (6.9) years; PD, 76.6 (3.4) years, Supplementary Table 1).
The levels of the three miRNAs targeting lamp-2a (hsa-miR-21*; hsa-miR-224; and hsa-miR-373*) and the three miRNAs targeting hsc70 (hsa-miR-26b; hsa-miR-106a*; and hsa-miR-301b) were significantly increased in PD SNc relative to actin mRNA levels (Figure 4a). These increases corresponded to a significant decrease in lamp-2a (71%) and hsc70 (78%) mRNA levels (Figure 4c) and a concomitant decrease in LAMP-2A (45%) and hsc70 (51%) protein levels previously reported.6 Similar but milder changes were observed in PD amygdala where there was a significant increase in the two miRNAs targeting lamp-2a (hsa-miR-224 and hsa-miR-373*) and a nonsignificant increase in the two miRNAs targeting hsc70 (hsa-miR-26b and hsa-miR-106a*; Figure 4b). These were associated with a mild decrease in LAMP-2A (36%) and hsc70 (32%) protein levels6 and a mild downregulation of lamp-2a (30%) and hsc70 (10%) mRNA levels (Figure 4c). The changes in miRNA levels were confirmed when the data were analyzed relative to gapdh mRNA (Supplementary Figures 4a and b).
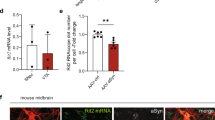
Analysis of PD brain samples and the dose-dependent impact of miRNA-373* upon LAMP-2A. Relative change in miRNAs normalized to actin mRNA levels and compared with control in (a) SNc from PD patients and (b) the amygdala. (c) mRNA levels for lamp-2a, hsc70 and α-synuclein relative to actin mRNA and compared with control values in PD amygdala and SNc. Data expressed as mean±S.E.M., controls (n=5) and PD (n=6). Statistical analyses compared with control group, *P<0.05. (d) The influence of transfecting increasing concentrations of miRNA hsa-miR-373* into normal SH-SY5Y cells on LAMP-2A protein and mRNA levels 72 h after transfection. The upper panel depicts the western blot of LAMP-2A levels relative to actin with increasing miR-373* levels, and the lower panel depicts the relationship between LAMP-2A protein and lamp-2a mRNA levels relative to actin and normalized to untreated cells
In SNc, the decrease in lamp-2a mRNA (71%) exceeded the decrease in protein levels (45%); however, this was not the situation in the amygdala (30 versus 35%). To determine if increasing concentrations of miRNAs could account for this difference, we evaluated the impact of increasing concentrations of hsa-miR-373* on the mRNA and protein levels of LAMP-2A. Relatively low miRNA concentrations (10 nM) reduced LAMP-2A protein levels but had no impact on mRNA levels, whereas higher concentrations (50 and 100 nM) downregulated both LAMP-2A protein and mRNA levels (Figure 4d). These data are in agreement with the high levels of miRNA in the SNc leading to lamp-2a mRNA degradation, whereas in the amygdala the milder increase in miRNA concentrations had less impact on mRNA stability.
We detected a significant decrease in α-synuclein mRNA levels in both the SNc and amygdala samples from PD patients (Figure 4c), in agreement with previous studies.18, 19 Although α-synuclein aggregates accumulate in LBs, we found that the levels of SDS-soluble α-synuclein monomer were not changed in the amygdala (Supplementary Figure 4d).20 This is consistent with the suggestion that α-synuclein monomer levels may be sustained, despite a decrease in transcripts, through a decrease in its degradation via CMA.
Discussion
α-Synuclein aggregation and the formation of LBs are characteristic pathological features of sporadic PD, but the underlying biochemical abnormality responsible remains to be defined. Multiplication of the α-synuclein locus is a rare cause of familial PD but suggests that increased synthesis of α-synuclein is sufficient to cause the disease.3 However, decreased α-synuclein degradation leading to α-synuclein accumulation and aggregation has been the focus for the other patients. There is a consensus that α-synuclein turnover predominantly involves CMA.6, 9, 10 Factors that influence its processing via CMA including various α-synuclein mutations and modifications including oxidative damage and S129 phosphorylation21, 22 can lead to α-synuclein accumulation. We have recently reported a decrease in the CMA proteins LAMP-2A and hsc70 in PD brains,6 further suggesting α-synuclein aggregation may be a consequence of its impaired degradation. It is important to understand the cause of the decreased LAMP-2A and hsc70 protein levels in PD brains to identify where these changes should be placed in the overall disease mechanisms.
Modeling the mitochondrial dysfunction, oxidative stress and UPS dysfunction frequently reported in PD brains23 failed to replicate the decrease in LAMP-2A or hsc70 levels seen in PD brains, although α-synuclein levels were increased. The increased α-synuclein levels observed with oxidative stress and mitochondrial inhibition have been previously reported,24, 25 and normal mRNA levels suggest this is a consequence of impaired α-synuclein degradation. Conversely, epoxomycin treatment led to an increase in the level of α-synuclein, which was associated with a large increase in mRNA. This increase was only seen in cells expressing ectopic α-synuclein under the CMV promoter and not observed in normal SH-SY5Y or melanoma cells that express endogenous α-synuclein. This is in agreement with a previous report demonstrating that epoxomycin enhanced the expression of genes regulated by the CMV promoter26 and therefore was an artifact of the system used. In the absence of changes to LAMP-2A or hsc70 levels with these treatments, the increased α-synuclein levels with rotenone or paraquat were likely to be related to either an inactivation of CMA or an impaired turnover of oxidatively modified or S129 phosphorylated α-synuclein.21, 22 We have previously shown that increased α-synuclein levels per se did not influence LAMP-2A or hsc70 protein levels,6 and we have now confirmed that although increased oxidative stress and mitochondrial dysfunction may interfere with α-synuclein degradation, these are not predicted to lead to the decrease in CMA proteins.
In recent years, the influence of miRNAs on the post-transcriptional regulation of protein levels has received much attention, in particular related to cancer initiation and progression, with studies describing the downregulation of miRNAs in cancer cells.27, 28 More recently, several reports have suggested that miRNA deregulation may play an important role in various neurodegenerative disorders including Alzheimer’s disease29 and spinocerebellar ataxia type 1.30 In PD, significant changes in the miRNA expression profile have been reported with numerous miRNA precursors elevated in the midbrain.13 We have focused on eight miRNAs increased in PD brains that were strongly predicted to target the 3′UTR of either lamp-2a or hsc70. The luciferase reporter constructs confirmed that four and three of these miRNAs effectively targeted lamp-2a and hsc70 3′UTRs, respectively, suggesting they could underpin the decrease in hsc70 and LAMP-2A reported in PD brains.6 We also demonstrated that these miRNAs were able to cause a dose-dependent decrease in endogenous LAMP-2A or hsc70 protein levels in SHSY5Y cells. These decreases were not associated with reduced mRNA levels, suggesting the miRNAs were acting at the level of translational regulation.31 The associated decrease in CMA would account for the decreased turnover and accumulation of α-synuclein under these conditions, an observation further supported by the lack of an increase in α-synuclein mRNA levels. Our results demonstrated that increases in specific miRNA levels, predicted to target lamp-2a or hsc70, were sufficient to induce α-synuclein accumulation in cell models.
The previously published changes to miRNA profiles in PD were limited to a small number of samples,13 and therefore it was important to replicate these changes in a larger cohort of PD brain samples. We confirmed that of the seven miRNAs capable of decreasing either LAMP-2A or hsc70 levels, six miRNAs were markedly increased in PD SNc and more mildly increased in the amygdala. Interestingly, the relative increase in these miRNAs in SNc and amygdala paralleled the relative decreases in hsc70 and LAMP-2A protein levels6 and severity of LB pathology. There were marked decreases in lamp-2a, hsc70 and α-synuclein mRNA levels in PD SNc, which were milder in the amygdala. It has been suggested that miRNAs can act via translational repression or increased mRNA degradation.31 In the cell cultures, we found that as the levels of miR-373* increased, there was an incremental decrease in LAMP-2A protein levels, whereas lamp-2a mRNA levels were only reduced at the higher miRNA concentrations. This may help explain the greater decrease in lamp-2a and hsc70 mRNA levels in PD SNc where the miRNA levels were increased the most.
The decrease in α-synuclein mRNA levels in PD has been previously reported.18, 19 We noted that cells treated with miR-106*a demonstrated a significant increase in α-synuclein protein levels in association with decreased mRNA levels, consistent with the observations seen in PD brain samples. Interestingly, miR-106*a is predicted to target the 3′UTRs of both α-synuclein (miRBase Target score 15.22) and hsc70 (miRBase Target score 17.39), which may explain this observation. Other miRNAs may also contribute to these effects in PD as both miR-7 and miR-153 have been previously demonstrated to modulate α-synuclein expression in cell models32, 33 and the precursors of miR-7 were significantly increased in PD brain samples.13
It is important to understand the cause of this deregulation and if it relates to changes to the synthesis or stability of the miRNAs in PD. The recent identification of a direct interaction between the PD causing G2019S LRRK2 mutant with the miRNA processing enzyme Argonaute2 and its effect upon mRNA and protein levels reinforces the potential importance of this pathway in PD.34
In conclusion, we demonstrated that in two brain regions (SNc and amygdala) associated with PD α-synuclein pathology, there were significant increases in specific miRNAs that were predicted to target LAMP-2A and hsc70 expression. These miRNAs were experimentally shown to decrease LAMP-2A and hsc70 protein levels in cell culture and resulted in significant α-synuclein intracellular accumulation. This supports the hypothesis that elevated miRNA levels in Parkinson’s disease brains will lead to the reported downregulation of LAMP-2A and hsc70 levels and compromised α-synuclein degradation, adding a further dimension to the pathogenesis of PD and LB formation. Modulation of CMA function in PD by miRNA silencing might represent a suitable target for drug intervention to modify the deleterious effects of impaired protein handling.
Materials and Methods
Brain samples
Substantia nigra pars compacta and amygdala samples from controls and patients with PD were obtained from the Navarra Brain Bank (Pamplona, Spain) and used with the consent of the local ethics committee and were part of a previous study.6 Controls had no clinical evidence ante mortem or pathological evidence post mortem of any neurodegenerative disease. Pathological diagnoses of PD were made according to recognized criteria (Queen Square Brain Bank criteria).
Cell cultures
The human SH-SY5Y neuroblastoma cell line (American Type Culture Collection, Manassas, VA, USA) and SH-SY5Y clones constitutively expressing full-length human WT α-synuclein with a C-terminal hemagglutinin tag have been previously described.21 The human melanoma SK-MEL28 cell line (American Type Culture Collection) was cultured in DMEM/F12 media supplemented with 10% FBS and penicillin/streptomycin. Other reagents were obtained from Sigma Aldrich (Dorset, UK) or Merck (Nottingham, UK) unless otherwise stated. Where stated, cells were treated for 48 h with rotenone (100 and 200 nM), paraquat (50 and 100 μM) or epoxomycin (5 and 10 nM).
Western blot analysis
Cell samples were solubilized, separated on NuPAGE Novex 4–12% Bis-Tris Gel (Invitrogen, Paisley, UK) and analyzed by western blot as previously described6 using the following primary antibodies: anti-α-synuclein (1 : 1000; Zymed, Invitrogen); anti-LAMP-2A (1 : 400); anti-hsc70 (1 : 500); and anti-β-actin (1 : 2000; all from Abcam, Cambridge, UK). Horseradish peroxidase-conjugated anti-mouse or anti-rabbit IgG secondary antibodies (DAKO, Ely, UK), were detected using ECL Plus Reagents and Hyperfilm ECL (GE Healthcare, Little Chalfont, UK). Films were scanned and signals in the linear range were quantified using DigiDoc software (Alpha Innotech, San Leandro, CA, USA) and normalized to β-actin levels.
Luciferase reporter constructs
Luciferase targets were produced by PCR amplification of the 3′UTRs of lamp-2a and hsc70 from genomic DNA from human embryonic kidney cells and ligating them into psiCheck2.2 (kind gift from Dr. Marc Weinberg, University of Witwatersrand, Johannesburg, South Africa) downstream of Renilla luciferase (Figure 2a) between XhoI and NotI sites as previously described.35 The relevant NCBI accession numbers used to design appropriate PCR primers were NM_006597.4 (HSPA8) and NM_002294.2 (LAMP2). The primers used for PCR were (XhoI hspa8 3′UTR F) CATCTCGAGCCAACCAAGTGTAGATGTAG; (NotI hspa8 3′UTR R) CATGCGGCCGCATTGCATTTTCCACTTACAATAC; (XhoI lamp2a 3′UTR F) CATCTCGAGAGCAATTTTAGAATCTGCAACC; (NotI lamp2a 3′UTR R) CATGCGGCCGCCATCTTGAAGAAACAAGAGC. Mutations to the 3′UTR sequences of the lamp2a and hsc70 luciferase constructs were generated using Phusion Site-Directed Mutagenesis Kit (Thermo Scientific, Loughborough, UK) and the following forward primers (mutations in upper case); tcccagcatttCAagatcagtctt (miRNA 373*); gcaatgttttaaggAAtatcttaagaagccc (miRNA 379); ggaaggaaataacaAAgcactttataaacac (miRNA 106*a, miRNA 301b).
Treatment of cells with miRNAs
Cells were transfected using HiPerfect Transfection reagent (Qiagen, Hilden, Germany) with miRNAs (Dharmacon, Epsom, UK) listed in Table 1, up to 50 nM for 72 h to determine their optimal concentrations. For long-term treatments, cells were transfected with 10 nM miRNAs at 0, 3 and 6 days.
Luciferase assay
SH-SY5Y cells were transfected with 1 μg of luciferase-3′UTR lamp-2a or luciferase-3′UTR hsc70 plasmids and 1 μl of x-tremeGENE HP transfection reagent (Roche, Burgess Hill, UK) as per the manufacturer’s instructions in 24-well plates with 105 cells per well. At 4 h after transfection, cells were treated with the miRNAs as described above. Luciferase activity was measure after 48 h with Dual-Glo Luciferase Assay System (Promega, Southampton, UK) using a luminometer (Synergy HT, Bio-Tek, Potton, UK).
Cell survival analysis
Cells were seeded in a 96-well plate and treated with the toxin or miRNAs. Viable cell numbers were determined using the Celltiter Blue kit (Promega) and expressed as a percentage of the control for each cell line.
Quantitative PCR
Total RNA was harvested using the RNeasy kit for cells or miRNeasy kit for brain tissue (Qiagen) as per the manufacturer’s protocol. Only samples with a low post-mortem delay (<11 h) were used and sample quality was assessed by RNA Integrity Number (RIN).36 Reverse transcription (RT) was performed with qSCRIP Reverse Transcriptase kit (Primer Design, Southampton, UK) or TaqMan MicroRNA RT kit (Applied Biosystems, Carlsbad, CA, USA) as per the manufacturer’s instructions. qPCR experiments were performed on a StepOne Real-Time PCR system (Applied Biosystems) using Precision qPCR Mastermix or TaqMan PCR (Applied Biosystems). TaqMan primers for the analysis of the various miRNAs were obtained from Applied Biosystems, and primers for the mRNA analysis are listed in Supplementary Table 2. Values were calculated using the standard ΔΔCt method.
Statistical analysis
Statistical analyses of the data were performed using SPSS, program 16.0 (IBM, North Harbour Portsmouth, UK), using the nonparametric Kruskal–Wallis test followed by the Mann–Whitney U-test.
Abbreviations
- hsc70:
-
heat shock cognate protein 70
- LAMP-2A:
-
lysosomal-associated membrane protein 2A
- PD:
-
Parkinson’s disease
- CMA:
-
chaperone-mediated autophagy
- miRNA:
-
microRNA
- LB:
-
Lewy body
- CMV:
-
cytomegalovirus
- 3′UTR:
-
3′ untranslated region
References
Spillantini MG, Schmidt ML, Lee VM, Trojanowski JQ, Jakes R, Goedert M . Alpha-synuclein in Lewy bodies. Nature 1997; 388: 839–840.
Polymeropoulos MH, Lavedan C, Leroy E, Ide SE, Dehejia A, Dutra A et al. Mutation in the alpha-synuclein gene identified in families with Parkinson’s disease. Science 1997; 276: 2045–2047.
Singleton AB, Farrer M, Johnson J, Singleton A, Hague S, Kachergus J et al. Alpha-synuclein locus triplication causes Parkinson’s disease. Science 2003; 302: 841.
Nalls MA, Plagnol V, Hernandez DG, Sharma M, Sheerin UM, Saad M et al. Imputation of sequence variants for identification of genetic risk for Parkinson’s disease: a meta-analysis of genome wide association studies. Lancet 2011; 377: 641–649.
Cuervo AM, Stefanis L, Fredenburg R, Lansbury PT, Sulzer D . Impaired degradation of mutant alpha-synuclein by chaperone-mediated autophagy. Science 2004; 305: 1292–1295.
Alvarez-Erviti L, Rodriguez-Oroz MC, Cooper JM, Caballero C, Ferrer I, Obeso JA et al. Chaperone-mediated autophagy markers in Parkinson disease brains. Arch Neurol 2010; 67: 1464–1472.
Dice JF . Peptide sequences that target cytosolic proteins for lysosomal proteolysis. Trends Biochem Sci 1990; 15: 305–309.
Majeski AE, Dice JF . Mechanisms of chaperone-mediated autophagy. Int J Biochem Cell Biol 2004; 36: 2435–2444.
Vogiatzi T, Xilouri M, Vekrellis K, Stefanis L . Wild type alpha-synuclein is degraded by chaperone-mediated autophagy and macroautophagy in neuronal cells. J Biol Chem 2008; 283: 23542–23556.
Lewis BP, Burge CB, Bartel DP . Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 2005; 120: 15–20.
Hutvágner G, Zamore PD . A microRNA in a multiple-turnover RNAi enzyme complex. Science 2002; 297: 2056–2060.
Yang B, Lin H, Xiao J, Lu Y, Luo X, Li B et al. The muscle-specific microRNA miR-1 regulates cardiac arrhythmogenic potential by targeting GJA1 and KCNJ2. Nat Med 2007; 13: 486–491.
Kim J, Inoue K, Ishii J, Vanti WB, Voronov SV, Murchison E et al. microRNA feedback circuit in midbrain dopamine neurons. Science 2007; 317: 1220–1224.
Bradley JL, Homayoun S, Hart PE, Schapira AH, Cooper JM . Role of oxidative damage in Friedreich’s ataxia. Neurochem Res 2004; 29: 561–567.
Hartley A, Stone JM, Heron C, Cooper JM, Schapira AH . Complex I inhibitors induce dose-dependent apoptosis in PC12 cells: relevance to Parkinson’s disease. J Neurochem 1994; 63: 1987–1990.
Konecki DS, Foetisch K, Zimmer KP, Schlotter M, Lichter-Konecki U . An alternatively spliced form of the human lysosome-associated membrane protein-2 gene is expressed in a tissue-specific manner. Biochem Biophys Res Commun 1995; 215: 757–767.
Gough NR, Fambrough DM . Different steady state subcellular distributions of the three splice variants of lysosome-associated membrane protein LAMP-2 are determined largely by the COOH-terminal amino acid residue. J Cell Biol 1997; 137: 1161–1169.
Neystat M, Lynch T, Przedborski S, Kholodilov N, Rzhetskaya M, Burke RE . Alpha-synuclein expression in substantia nigra and cortex in Parkinson’s disease. Mov Disord 1999; 14: 417–422.
Kingsbury AE, Daniel SE, Sangha H, Eisen S, Lees AJ, Foster OJ . Alteration in alpha-synuclein mRNA expression in Parkinson’s disease. Mov Disord 2004; 19: 162–170.
Tong J, Wong H, Guttman M, Ang LC, Forno LS, Shimadzu M et al. Brain alpha-synuclein accumulation in multiple system atrophy, Parkinson’s disease and progressive supranuclear palsy: a comparative investigation. Brain 2010; 133: 172–188.
Chau KY, Ching HL, Schapira AH, Cooper JM . Relationship between alpha synuclein phosphorylation, proteasomal inhibition and cell death: relevance to Parkinson’s disease pathogenesis. J Neurochem 2009; 110: 1005–1013.
Martinez-Vicente M, Talloczy Z, Kaushik S, Massey AC, Mazzulli J, Mosharov EV et al. Dopamine-modified alpha-synuclein blocks chaperone-mediated autophagy. J Clin Invest 2008; 118: 777–788.
Shulman JM, De Jager PF, Feany MB . Parkinson’s disease: genetics and pathogenesis. Annu Rev Pathol Mech Dis 2011; 6: 193–222.
Sherer TB, Betarbet R, Stout AK, Lund S, Baptista M, Panov AV et al. An in vitro model of Parkinson’s disease: linking mitochondrial impairment to altered alpha-synuclein metabolism and oxidative damage. J Neurosci 2002; 22: 7006–7015.
Manning-Bog AB, McCormack AL, Li J, Uversky VN, Fink AL, Di Monte DA . The herbicide paraquat causes up-regulation and aggregation of alpha-synuclein in mice: paraquat and alpha-synuclein. J Biol Chem 2002; 277: 1641–1644.
Biasini E, Fioriti L, Ceglia I, Invernizzi R, Bertoli A, Chiesa R et al. Proteasome inhibition and aggregation in Parkinson’s disease: a comparative study in untransfected and transfected cells. J Neurochem 2004; 88: 545–553.
Yanaihara N, Caplen N, Bowman E, Seike M, Kumamoto K, Yi M et al. Unique microRNA molecular profiles in lung cancer diagnosis and prognosis. Cancer Cell 2006; 9: 189–198.
Melo SA, Ropero S, Moutinho C, Aaltonen LA, Yamamoto H, Calin GA et al. A TARBP2 mutation in human cancer impairs microRNA processing and DICER1 function. Nat Genet 2009; 41: 365–370.
Hébert SS, Horré K, Nicolaï L, Hébert SS, Horré K, Nicolaï L et al. Loss of microRNA cluster miR-29a/b-1 in sporadic Alzheimer’s disease correlates with increased BACE1/beta-secretase expression. Proc Natl Acad Sci USA 2008; 105: 6415–6420.
Lee Y, Samaco RC, Gatchel JR, Thaller C, Orr HT, Zoghbi HY . miR-19, miR-101 and miR-130 co-regulate ATXN1 levels to potentially modulate SCA1 pathogenesis. Nat Neurosci 2008; 11: 1137–1139.
Pasquinelli AE . MicroRNAs and their targets: recognition, regulation and an emerging reciprocal relationship. Nat Rev 2012; 13: 271–282.
Junn E, Lee KW, Jeong BS, Chan TW, Im JY, Mouradian MM . Repression of alpha-synuclein expression and toxicity by microRNA-7. Proc Natl Acad Sci USA 2009; 1060: 13052–13057.
Doxakis E . Post-transcriptional regulation of alpha-synuclein expression by mir-7 and mir-153. J Biol Chem 2010; 285: 12726–12734.
Gehrke S, Imai Y, Sokol N, Lu B . Pathogenic LRRK2 negatively regulates microRNA-mediated translational repression. Nature 2010; 466: 637–641.
Ely A, Naidoo T, Mufamadi S, Crowther C, Arbuthnot P . Expressed anti-HBV primary microRNA shuttles inhibit viral replication efficiently in vitro and in vivo. Mol Ther 2008; 16: 1105–1112.
Schroeder A, Mueller O, Stocker S, Salowsky R, Leiber M, Gassmann M et al. The RIN: an RNA integrity number for assigning integrity values to RNA measurements. BMC Mol Biol 2006; 7: 3.
Acknowledgements
We are grateful to Dr. Zudaire, Dr. Agorreta, and Dr. Tuñon (Pamplona) for helpful discussions. This work was supported by the Brain Research Trust (to JMC and LA-E), Parkinson’s UK (to JMC and LA-E), the Wellcome Trust/MRC Joint Call in Neurodegeneration award (WT089698) to the UK Parkinson’s Disease Consortium (to JMC and AHVS).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Dr. JA Obeso has served on the Advisory Board of TEVA Pharmaceutica in the past 3 years and once for TEVA Neuroscience (USA) in 2012 and has received honorarium for lecturing in meetings organized by GSK, Lundbeck-TEVA and UCB in Spain. He was funded by the Spanish Science and Education Ministry and European Union (REPLACES). Maria C Rodriguez-Oroz has received payment for lectures, as well as travel and accommodation to attend scientific meetings, from UCB and Lundbeck. She has received research funding from national and regional government bodies in Spain. Professor Schapira and Drs. Alvarez-Erviti, Seow and Cooper declare no conflict of interest.
Additional information
Edited by D Bano
Supplementary Information accompanies this paper on Cell Death and Disease website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Alvarez-Erviti, L., Seow, Y., Schapira, A. et al. Influence of microRNA deregulation on chaperone-mediated autophagy and α-synuclein pathology in Parkinson's disease. Cell Death Dis 4, e545 (2013). https://doi.org/10.1038/cddis.2013.73
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2013.73
Keywords
This article is cited by
-
The role of exosomes in pathogenesis and the therapeutic efficacy of mesenchymal stem cell-derived exosomes against Parkinson’s disease
Neurological Sciences (2023)
-
Lebensverlängernde Substanzen und Lebensmittel für ein gesundes Altern
ProCare (2022)
-
Chaperone-mediated Autophagy Regulates Cell Growth by Targeting SMAD3 in Glioma
Neuroscience Bulletin (2022)
-
Down-Regulation of ID2-AS1 Alleviates the Neuronal Injury Induced by 1-Methy1-4-Phenylpyridinium in Human Neuroblastoma Cell Line SH-SY5Y Cells Through Regulating miR-199a-5p/IFNAR1/JAK2/STAT1 Axis
Neurochemical Research (2021)
-
Autophagy and disease: unanswered questions
Cell Death & Differentiation (2020)