Abstract
The anti-tumoral effects of cannabinoids have been described in different tumor systems, including pancreatic adenocarcinoma, but their mechanism of action remains unclear. We used cannabinoids specific for the CB1 (ACPA) and CB2 (GW) receptors and metabolomic analyses to unravel the potential pathways mediating cannabinoid-dependent inhibition of pancreatic cancer cell growth. Panc1 cells treated with cannabinoids show elevated AMPK activation induced by a ROS-dependent increase of AMP/ATP ratio. ROS promote nuclear translocation of GAPDH, which is further amplified by AMPK, thereby attenuating glycolysis. Furthermore, ROS determine the accumulation of NADH, suggestive of a blockage in the respiratory chain, which in turn inhibits the Krebs cycle. Concomitantly, inhibition of Akt/c-Myc pathway leads to decreased activity of both the pyruvate kinase isoform M2 (PKM2), further downregulating glycolysis, and glutamine uptake. Altogether, these alterations of pancreatic cancer cell metabolism mediated by cannabinoids result in a strong induction of autophagy and in the inhibition of cell growth.
Similar content being viewed by others
Main
Cannabinoids are a class of bioactive lipids1, 2, 3 that have a range of interesting activities, including the ability to reduce the growth of tumours such as glioma,4 breast cancer,5 prostate cancer,6 and colon cancer7 in different animal models. They impair tumour progression at different levels, with the most prevalent effects being the inhibition of cell proliferation by apoptosis,8 cell cycle arrest,9 and autophagy.10 Cannabinoids induce autophagy in various types of cancer cell lines, and pharmacological or genetic inhibition of autophagy prevents their antiproliferative action, thus demonstrating that autophagy is important for cannabinoid antineoplastic activity.11 Autophagy is an evolutionarily conserved process in eukaryotes by which cytoplasmic cargo sequestered inside double-membrane vesicles are delivered to the lysosome for degradation.12 This process has the role to rid the cell of intracellular misfolded or long-lived proteins, superfluous or damaged organelles, and invading microorganisms, and also is an adaptive response to provide nutrients and energy on exposure to various stresses.13 In hepatocellular carcinoma cells, cannabinoids can trigger an ER stress-dependent activation of AMP-activated protein kinase (AMPK) that cooperates with the TRIB3-mediated inhibition of the Akt–mTORC1 axis in the stimulation of autophagy-mediated cell death.14 AMPK is a sensor of energy status that responds to the increase of AMP or ADP cellular concentration to maintain cellular energy homeostasis.15 AMPKs appear to exist universally as heterotrimeric complexes comprising catalytic α subunits and regulatory β and γ subunits.15 The α subunits contain a typical serine/threonine kinase domain at the N terminus and is significantly active only when phosphorylated by upstream kinases.15 The γ subunits contain four regulatory adenine nucleotide-binding sites, two of which competitively bind AMP, ADP and ATP, and are the sites via which cellular energy status is sensed.15 The major upstream kinase phosphorylating Thr 172 of the α subunit, and thus activating AMPK, in most mammalian cells is the tumour suppressor kinase LKB1.16, 17, 18 Although LKB1 has to be expressed in mammalian cells for agents that increase the cellular AMP/ATP and ADP/ATP ratios to cause the activation of AMPK,16 it is worth emphasizing that these effects are due to the binding of adenine nucleotides to the γ subunit of AMPK and that the LKB1 complex itself appears to be constitutively active.19 In some cell types, Thr 172 can also be phosphorylated by the Ca2+/calmodulin-dependent protein kinase, CaMKKβ, providing a Ca2+-activated pathway to switch on AMPK.20, 21, 22 Activation by this mechanism can occur in the absence of any change in the adenine nucleotide ratios, although increases in Ca2+ can act synergistically with increases in AMP or ADP.23
Recently, we have demonstrated that cannabinoids and gemcitabine, a nucleoside analogue used in cancer chemotherapy, synergistically inhibit pancreatic adenocarcinoma cell growth by a ROS-mediated autophagy induction.10
Here, to shed light on the molecular mechanisms of autophagy induction by cannabinoids in pancreatic adenocarcinoma cells, we have investigated whether the AMPK has a role in this effect and whether this mechanism is related to the alteration of the energetic metabolism. For this purpose, we performed analysis of autophagy with a mutant of the γ subunit isoform 2 of AMPK unable to bind AMP, metabolomic analyses and determination of phosphorylation, activity or localization of proteins involved in the energetic metabolism or autophagy. We show that cannabinoids induce AMPK-mediated autophagy in pancreatic adenocarcinoma cells through a ROS-dependent increase of the AMP/ATP ratio.
Results
Cannabinoid-induced autophagy depends on AMPK activation
We previously demonstrated that arachidonoyl cyclopropamide (ACPA) or GW405833 (GW), two synthetic cannabinoid ligands specific for CB1 and CB2, respectively, are able to induce ROS-mediated autophagy in pancreatic adenocarcinoma cell lines.10 To more deeply investigate the molecular mechanisms of this effect, we performed in Panc1 cells treated with ACPA or GW kinetic analyses of the Thr172 phosphorylated AMPK (p-AMPK) and of the Thr389 phosphorylated p70S6K (p-p70S6K), HIF-1α and PDHK downstream targets of mTORC1, a known autophagy inhibitor. As shown in Figure 1, after 1 h of treatment with cannabinoids, the increase of AMPK phosphorylation already occurred, with a further extension of the effect up to 24 h (Figure 1a). It is worthy to note that the AMPK total protein levels decreased after 12 and 24-h treatment. The ratio phospho-p70S6K/p70S6K decreased after 1-h treatment and to a larger extent at 12 and 24 h (Figure 1b), although the total protein formed also decreased after treatment with cannabinoids. Instead, HIF-1α (Figure 1c) showed a reduction of the expression starting at 12-h treatment and PDHK (Figure 1d) at 12 and 24 h for GW and ACPA, respectively. These results strongly suggested that AMPK could be involved in autophagy induction by cannabinoids in pancreatic adenocarcinoma cells. To further examine the role of AMPK in this effect, we treated the cells with cannabinoids in the absence or presence of compound C (CC), an AMPK inhibitor,24 and we analysed AMPK phosphorylation as control (Supplementary Figure 1) and the regulation of the autophagy marker phosphoethanolaminated LC3-II (Figure 2a). CC prevented AMPK phosphorylation and autophagy induction by cannabinoids, providing a strong support to the dependence of cannabinoid-induced autophagy on AMPK activation. To rule out the nonspecific effect of CC and to characterize the upstream events of AMPK activation by cannabinoids, we performed transient transfection of Panc1 cells using a plasmid coding for the mutated γ subunit isoform 2 (γ2 of AMPK, which is unable to bind AMP). The results reported in Figures 2b and c show that overexpression of the mutant AMPK (γ2 R531G) significantly decreased both p70S6K phosphorylation inhibition and LC3-II induction by cannabinoids, whereas the wild-type AMPK (γ2wt) had no effect on those cannabinoid activities. These findings indicated that AMPK was involved in cannabinoid-induced autophagy via an AMP-dependent mechanism. To confirm the role of AMP in this event and to verify whether it could depend on ROS production, we analysed the cellular AMP/ATP ratio following GW or ACPA treatment in the absence or presence of the radical scavenger N-acetyl-L-cysteine (NAC). As shown in Figure 2d, the level of AMP/ATP was strongly increased by cannabinoids, while it was similar to the control upon NAC pre-treatment of the cells.
Effect of cannabinoids on key metabolic proteins. Western blot analyses of (a) phospho-AMPK, (b) phospho-p70S6K, (c) HIF-1 alpha, and (d) pyruvate dehydrogenase kinase (PDHK) were performed after treatment of Panc1 cells for 1, 12, and 24 h with 200 μM GW or ACPA. The bands were scanned as digital peaks and the areas of the peaks were calculated in arbitrary units, as described in Materials and Methods. The value of α-tubulin was used as a normalizing factor and quantifications represent the ratio phosphorylated/total protein. Quantification values are the means of three independent experiments (±S.D.). Statistical analysis: *P<0.05, **P<0.01, and ***P<0.001
Role of AMPK in cannabinoid-induced autophagy. Western blot analysis of (a) LC3-II after treatment of Panc1 cells for 12 h with 200 μM GW or ACPA and 20 μM CC, and of (b) p-p70S6K and (c) LC3-II after treatment of Panc1 cells for 12 h with 200 μM GW or ACPA, in the presence or absence of the expression vectors for the AMPK wt or mutant R531G gamma-2 subunit. The bands were scanned as digital peaks and the areas of the peaks were calculated in arbitrary units, as described in Materials and Methods. The value of α-tubulin was used as a normalizing factor. Values are the means of three independent experiments (±S.D.). (d) AMP/ATP ratio was measured after 12 h treatment with 200 μM GW or ACPA, in the absence or presence of a pre-treatment for 1 h with 20 mM NAC, as reported in Materials and Methods. Values are the means of three independent experiments (±S.D.). Statistical analysis: *P<0.05, **P<0.01, and ***P<0.001
Cannabinoids inhibit the glycolytic pathway
To assess whether a restriction of the energetic metabolism by cannabinoids could be responsible for the enhancement of the cellular AMP level, we performed a targeted metabolomic analysis. We determined fold-change variations of the concentration levels of several key metabolites from the glycolitic pathway upon GW or ACPA treatment. Figure 3a shows a significant increase of glyceraldehyde 3-phosphate (G3P) and phosphoenolpyruvate (PEP) and a decrease of lactate (LACT), following a 12-h treatment with GW. Instead, ACPA treatment determined only a G3P increase (Figure 3b). To exclude the possibility that the glycolysis metabolite increase was determined by a higher glucose uptake by the cells, we measured the amount of glucose in the supernatant of treated or untreated cells, and we found that glucose incorporation did not significantly change upon cannabinoid treatment (Figure 3c).
Effect of cannabinoids on glycolysis. Panc1 were treated for 12 h with (a) GW or (b) ACPA and the metabolites analysis was performed as reported in Materials and Methods.FDP, fructose 1,6-bisphosphate; F6P, fructose-6-phosphate; G3P, glyceraldehyde-3-phosphate; G6P, glucose-6-phosphate; LACT, lactate; PEP, phosphoenolpyruvate. (c) Glucose consumption was measured as reported in Materials and Methods. Values are the means of three independent experiments (±S.D.). (d) PKM2 activity was performed after treatment of Panc1 cells for 12 or 24 h with 200 μM GW or ACPA, as reported in Materials and Methods. Values are the means of three independent experiments (±S.D.). Statistical analysis: *P<0.05, **P<0.01, and ***P<0.001
As the glycolytic enzyme glyceraldehyde 3-phosphate dehydrogenase (GAPDH) is known to contribute to the upregulation of autophagy by translocating to the nucleus,25 we analysed the cellular distribution of GAPDH after the treatment with GW or ACPA. Figure 4 shows the confocal images where GAPDH appear to be translocated into the nucleus of the cells after 12-h cannabinoid treatments. This effect was inhibited by NAC or CC, indicating its dependence on ROS and AMPK activation. These data suggested that the increased level of G3P observed in the metabolomic analysis could arise from the diminished presence of GAPDH in the cytosol of cells treated with cannabinoids.
Cannabinoids induce AMPK-dependent GAPDH nuclear translocation. Representative confocal images of GAPDH translocation in Panc1 cell nuclei after 12 h treatment with 200 μM GW or ACPA in the absence or presence of a pre-treatment for 1 h with 20 mM NAC or 20 μM C.C. Values are the means of three independent experiments (±S.D.)
PKM2 is an embryonic isoform of pyruvate kinase (PK) re-expressed in cancer cells, which dephosphorylates PEP to pyruvate, and determines a metabolic advantage for the cell by allowing the availability of phosphometabolites upstream of pyruvate as precursors for cellular syntheses.26 To assess whether the PKM2 activity might be responsible for an increase of PEP level by GW, we analysed the activity of PKM2 on crude protein extracts from cells treated with cannabinoids for 12 or 24 h. Figure 3d shows that PKM2 activity after GW treatment significantly decreased at 12 h and even more at 24 h, whereas it was inhibited by ACPA only after 24-h treatment. These data are consistent with the increased level of PEP observed in the metabolomic analysis after 12 h of treatment with GW. Furthermore, data not shown indicate that PEP did increase after 24-h treatment with ACPA.
Taken together, these findings suggested an impairment of the glycolytic pathway by cannabinoids that correlated with the increase of the AMP/ATP ratio.
Cannabinoids inhibit the Krebs cycle
To further examine the involvement of the energetic metabolism in the induction of AMPK-mediated autophagy by cannabinoids, we analysed the critical metabolites of the Krebs cycle. As shown in Figure 5, only the levels of the α-ketoglutarate (Figures 5a and b) and, to a very large extent, those of NADH (Figure 5c) increased after cannabinoid treatments. These results suggested the occurrence of the inhibition by NADH of the α-ketoglutarate dehydrogenase, the enzyme of the Krebs cycle most sensitive to the levels of that coenzyme, and the impairment of the oxidative phosphorylation in the increased AMP/ATP ratio determined by cannabinoids. Interestingly, NAC was able to significantly reduce the increase of NADH by GW or ACPA (Figure 5c), suggesting the involvement of the oxidative stress in that effect. To verify whether the ROS-dependent NADH increase correlated with the activation of the pentose phosphate pathway and thus with the accumulation of the coenzyme NADPH, known to have a key role in the antioxidant response of the cell, we measured the amounts of the phosphogluconolactone and of the NADPH. The levels of both the compounds remained unchanged after treatment with GW or ACPA, indicating that cannabinoids were unable to activate the pentose phosphate pathway in our experimental conditions (Supplementary Figure 2).
Effect of cannabinoids on Krebs cycle. Panc1 were treated for 12 h with 200 μM (a) GW or (b) ACPA and the metabolites analysis was performed as reported in Materials and Methods.KET, alpha-ketoglutarate; MA, malate; SUCC, succinate. (c) NADH was measured after 12 h treatment with 200 μM GW or ACPA in the presence or absence of a pre-treatment for 1 h with 20 mM NAC, as reported in Materials and Methods. Values are the means of three independent experiments (±S.D.). Statistical analysis: *P<0.05, **P<0.01, and ***P<0.001
Cannabinoids inhibit the anaplerotic flux of the Krebs cycle from glutamine
Glutamine and glucose are the only two molecules catabolized in appreciable quantities in most mammalian cells in culture, to supply the cell with most of the carbon, nitrogen, free energy and reducing equivalents. To assess whether the regulation of glutamine catabolism was involved in the downregulation of the energetic metabolism, we measured the levels of glutamine and glutamate in the culture medium after the treatment of cells with cannabinoids. Figure 6a shows that glutamine incorporation was strongly reduced by both GW and ACPA, whereas glutamate release remained unchanged. The oncogene c-Myc has been described to coordinate the expression of genes involved in glutamine catabolism, including the induction of glutamine transporters.27 Furthermore, it has been shown that Akt induces the upregulation of c-Myc and that Akt suppression inhibits the c-Myc expression.28 We evaluated both Akt phosphorylation on serine 473, which is a marker of activation for this kinase, and c-Myc activity after GW or ACPA treatments. Figures 6b and c show that GW strongly inhibited Akt phosphorylation and c-Myc activity at 12-h treatment and even more at 24 h, while ACPA determined a significant decrease of both the proteins only at 24-h treatment. Altogether, these results suggested that the inhibition of glutamine uptake by cannabinoids could depend, at least for GW, on the repression of glutamine transporters determined by the decrease of c-Myc activity.
Effect of cannabinoids on glutamine metabolism. (a) The analyses of glutamine uptake and glutamate release were performed after treatment of Panc1 cells for 12 h with 200 μM GW or ACPA. (b) Western blot analysis was performed using total protein extracts of Panc1 cells treated for 1, 12 and 24 h with 200 μM GW or ACPA. The bands were scanned as digital peaks and the areas of the peaks were calculated in arbitrary units, as described in Materials and Methods. The value of α-tubulin was used as a normalizing factor and quantifications represent the ratio phosphorylated/total protein. Values are the means of three independent experiments (±S.D.). (c) C-Myc activity was analysed after treatment for 1, 12 and 24 h with 200 μM GW or ACPA. Values are the means of three independent experiments (±S.D.). Statistical analysis: *P<0.05, **P<0.01, and ***P<0.001
Discussion
AMPK has been shown to have a crucial role in the cannabinoid-induced autophagy.14 We previously reported10 that cannabinoids inhibit pancreatic cancer cell proliferation both in vitro and in vivo, and autophagy has been demonstrated to mediate this process10, 11 or to be itself a death mechanism.10 As the antitumour effects of the cannabinoids are beginning to be clinically assessed, following the promising preclinical data, the need to clearly establish the molecular mechanisms of cannabinoid-induced autophagy has become more critical. In this study, we demonstrated for the first time that two synthetic cannabinoids, ACPA and GW, activated AMPK and autophagy in the pancreatic adenocarcinoma cells by increasing the cellular AMP/ATP ratio.
It is well known that AMPK exerts an active role in autophagy by inhibiting mTORC1, the major regulator of protein synthesis and cell growth.29 In our study, we found that the phosphorylation level of p70S6K, a direct target of mTORC1, significantly decreased following cannabinoid treatment and that this event started at a very early stage, well correlating with the increase of the AMPK phosphorylation. Furthermore, we found that the decrease of HIF-1α, a transcription factor that is indirectly regulated by mTORC1 via protein stabilization, started only after 12 h of treatment and, in turn, the decrease of one of its targets, PDHK, mainly after 24 h with both cannabinoids. Taken together, these data strongly suggested that the main site of autophagy regulation by cannabinoid-activated AMPK was mTORC1.
AMPK activation is generally mediated by the increase of the cellular AMP/ATP and ADP/ATP ratios that favours the binding of adenine nucleotides to the γ subunit of AMPK.15 In most of the cell types, this event is followed by Thr 172 phosphorylation by the LKB1 complex, which appears to be constitutively active.15 The Ca2+/calmodulin-dependent protein kinase (CaMKKβ) has been also reported to phosphorylate AMPK at the same site in some cell types.15 However, activation by this mechanism can occur in the absence of any change in adenine nucleotide ratios, although increases in Ca2+ can act synergistically with increases in AMP or ADP.23 Our data demonstrated that in cells overexpressing an AMPK containing a mutation in the γ subunit isoform 2 that renders the enzyme insensitive to increases in AMP or ADP, the autophagy pathway was not activated following cannabinoid treatment. This result indicated that in our experimental system the main mechanism of AMPK activation by cannabinoids depended on the alteration of adenine nucleotide ratios and suggested that LKB1 could be involved in the subsequent phosphorylation of AMPK at Thr 172.
AMPK has been described as an intracellular energy sensor and regulator,30 but is also important in maintaining intracellular homoeostasis during many kinds of stress challenges, such as oxidative stress, which has been shown to induce AMPK.31 In our previous paper, we demonstrated that the cannabinoids ACPA and GW were able to induce oxidative stress in pancreatic adenocarcinoma cell lines, which was crucial for triggering autophagic cell death in combination with gemcitabine.10 Here, we reported that ROS were necessary to increase the AMP/ATP ratio, which in turn mediated the activation of AMPK by cannabinoids leading to autophagy. In agreement with our result, Shrivastava et al.32 have recently shown that cannabidiol induces apoptosis and autophagy by increasing the generation of ROS.32 We found that NADH cellular level strongly increased after treatment of cells with either ACPA or GW and that this increase was inhibited in the presence of the radical scavenger NAC. Furthermore, our previous paper10 and unpublished results demonstrated that ROS generation occurred as early as 30 min after the beginning of cannabinoid treatment. Taken together, these data suggested that ROS production by cannabinoids could impair the electron transport chain leading to NADH accumulation and to oxidative phosphorylation inhibition, which in turn could further increase the level of ROS. Consistent with this hypothesis, many articles have been published describing the oxidative stress as a cause of mitochondrial injury, often occurring at the level of complex I of the respiratory chain by direct oxidation of the proteins belonging to the complex.33 In correlation with the strong increase of the NADH level after cannabinoid treatment, we found a strong increase of the AMP/ATP ratio, which suggested that the oxidative phosphorylation inhibition could play a crucial role in generating the energetic blackout of the cell. Further studies have been planned to clarify this aspect.
Our analyses of key metabolites and enzymes of the glycolysis suggested a general inhibition of the pathway after ACPA or GW treatment. Although glucose uptake by cells remained unchanged, the level of two glycolytic metabolites, G3P and PEP, significantly increased in cells treated with cannabinoids, well correlating with a decrease of the respective enzymes, GAPDH and PKM2.
GAPDH has been recently reported as a key redox-sensitive protein, the activity of which is largely affected by covalent oxidative modifications at its highly reactive Cys152.34 These modifications stimulate nuclear translocation of the enzyme and regulate the fate of the cell,35 often leading to autophagy activation by upregulating the autophagy protein Atg12.34 Furthermore, also AMPK has been shown to stimulate GAPDH translocation into the nuclei.36 Our data showed that both NAC and CC inhibited GAPDH nuclear translocation, demonstrating that this event was mediated by both ROS and AMPK.
PKM2 is the embryonic pyruvate kinase isoform almost universally re-expressed in cancer that promotes aerobic glycolysis.37, 38 It has recently been shown that c-Myc upregulates the transcription of genes involved in the alternative splicing leading to the expression of PKM2.37, 39 Our results showed a strict correlation between PKM2 and c-Myc activities in both ACPA and GW-treated cells, suggesting that cannabinoids determined the downregulation of PKM2 via the inhibition of c-Myc.
c-Myc is a classical oncogene that promotes not only proliferation but also the production of accompanying macromolecules and antioxidants that are required for growth.37 c-Myc increases glutamine uptake by directly inducing the expression of glutamine transporters27 and promotes the expression of the PKM2 isoform.39 Furthermore, c-Myc is upregulated by Akt, the suppression of which inhibits c-Myc expression.28 Our kinetic analyses showed that GW inhibited Akt phosphorylation at 12 h and even more at 24 h, while ACPA inhibited Akt phosphorylation at 24 h treatment. Similar results were obtained on analysing c-Myc activity. These findings suggested the dependence of c-Myc downregulation on the inhibition of Akt phosphorylation by cannabinoids. One of the major regulators of Akt is the phosphatase PTEN, a tumour suppressor mutated in a wide range of human cancers, the activity of which has been found to increase upon endocannabinoid treatment.40
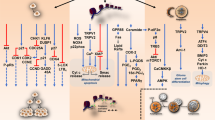
In summary, our results demonstrated for the first time that cannabinoid-dependent autophagy induction in pancreatic adenocarcinoma cells is strictly related to the inhibition of the energetic metabolism, which, in turn, is dependent on the early production of ROS induced by the compounds. In Figure 7, we propose a model of the mechanism of autophagy induction by the cannabinoids ACPA and GW in pancreatic adenocarcinoma cells that is based on the results presented in this paper and on literature data.
Materials and Methods
Materials
ACPA was obtained from Cayman Chemicals (Inalco, Milan, Italy); GW405833 hydrochloride (1-(2,3-dichlorobenzoyl)-2-methyl-3-(2-(1-morpholine)ethyl)-5-methoxyindole) and NAC (N-acetyl-L-cysteine) were obtained from Sigma (Milan, Italy); Compound C (CC) was obtained from Calbiochem. Acetonitrile, formic acid, and HPLC-grade water were purchased from Sigma Aldrich (Milan, Italy). Standards (equal to or greater than 98% chemical purity) D-glucose 6-phosphate, fructose 6-phosphate, D-fructose 1,6 biphosphate, glyceraldehyde 3-phosphate, phosphoenolpyruvic acid, L-lactic acid, α-ketoglutaric acid, L-malic acid, succinic acid, ATP, NADH, NADPH, 6-phosphogluconolactic acid, L-glutamic acid, glutamine, reduced glutathione, and oxidized glutathione were purchased from Sigma Aldrich.
Cell culture
Panc1 cell line was grown in RPMI 1640 supplemented with 2 mM glutamine, 10% FBS, and 50 μg/ml gentamicin sulphate (BioWhittaker, Lonza, Bergamo, Italy), and incubated at 37 °C with 5% CO2.
Immunoblot analysis
Cells (1.2 × 106 cells/dish 10 cm) were treated as reported in the figures and then lysated with RIPA buffer for total protein extract. Protein concentration was measured with the Bradford protein assay reagent (Pierce, Rockford, IL, USA) using bovine serum albumin as a standard. Forty μg of protein extracts was electrophoresed through SDS–polyacrylamide gel and electroblotted onto polyvinylidene difluoride membranes (Millipore, Milan, Italy). Membranes were then incubated for 1 h at room temperature with blocking solution (5% low-fat milk in 100 mM Tris pH 7.5, 0.9% NaCl, 0.1% Tween 20) and probed overnight at 4 °C with the primary monoclonal antibodies (1 : 1000 in blocking solution). Light chain protein-II (LC-3II), AMPK, phospho-AMPK Thr172, p70S6K, phopsho-p70S6K Thr389, Akt, phospho-Akt Ser473, and PDHK were obtained from Cell Signaling, Euroclone, Milan, Italy, HIF-1 alpha antibody from Novus Biologicals, Milan, Italy, and α-tubulin antibody from Oncogene, Cambridge, MA, USA. Horseradish peroxidase-conjugated secondary antibodies IgG (1 : 8000 in blocking solution; Upstate Biotechnology, DBA Italia, Milan, Italy) were used to detect specific proteins. Immunodetection was carried out using chemiluminescent substrates (Amersham Pharmacia Biotech, Euroclone, Milan, Italy) and recorded using HyperfilmECL (Amersham Pharmacia Biotech). The bands for total and phosphorylated proteins were scanned as digital peaks and the areas of the peaks were calculated in arbitrary units using the public domain NIH Image software (http://rsb.info.nih.gov/nih-image/) and then normalized with α-tubulin expression. Quantifications were obtained as fold induction relative to controls and, for phosphorylated proteins, quantifications represent the ratio phosphorylated/total protein.
Metabolite extraction
Metabolomic analyses after treatment for 12 h with 200 μM GW or ACPA and 20 mM NAC were performed as previously reported.41 Cells were prepared following the protocol by Sana et al.,42 with minor modifications as previously reported.41 The sample was resuspended by adding 0.15 ml of ice-cold ultra-pure water (18 MΩ) to lyse cells. The tubes were plunged into dry ice or a circulating bath at −25 °C for 0.5 min and then into a water bath at 37 °C for 0.5 min. To each tube was added 0.6 ml of −20 °C methanol and then 0.45 ml of −20 °C chloroform. The tubes were mixed every 5 min for 30 min. Subsequently, 0.15 ml of ice-cold pH-adjusted ultra-pure water was added to each tube and these were centrifuged at 1000 × g for 1 min at 4 °C, before being transferred to −20 °C for 2–8 h. After thawing, liquid phases were recovered and an equivalent volume of acetonitrile was added to precipitate any residual protein. The tubes were then transferred to a refrigerator (4 °C) for 20 min, centrifuged at 10 000 × g for 10 min at 4 °C and the collected supernatants were dried to obtain visible pellets. Finally, the dried samples were re-suspended in 1 ml of water, 5% formic acid and transferred to glass autosampler vials for LC/MS analysis.
Rapid-resolution reverse-phase HPLC
An Ultimate 3000 Rapid Resolution HPLC system (LC Packings, DIONEX, Sunnyvale, CA, USA) was used to perform metabolite separation. A Dionex Acclaim RSLC 120 C18 column 2.1 mm × 150 mm × 2.2 μm was used to separate the extracted metabolites. A 0–95% linear gradient of solvent A (0.1% formic acid in water) to B (0.1% formic acid in acetonitrile) was employed over 15 min followed by a solvent B hold of 2 min, returning to 100% A in 2 min and a 6-min post-time solvent A hold.
ESI mass spectrometry
Metabolites were directly eluted into a High Capacity ion Trap HCTplus (Bruker-Daltonik, Bremen, Germany). Mass spectra for metabolite extracted samples were acquired in positive ion mode. ESI capillary voltage was set at 3000 V (+) ion mode. The liquid nebulizer was set to 30 psig and the nitrogen drying gas was set to a flow rate of 9 l/min. Dry gas temperature was maintained at 300 °C. Data were stored in centroid mode. Internal reference ions were used to continuously maintain mass accuracy. Data were acquired at a rate of 5 spectra/s with a stored mass range of m/z 50–1500. Data were collected using Bruker Esquire Control (v. 5.3 - build 11) data acquisition software. In MRM analysis, m/z values of interest were isolated, fragmented and monitored (either the parental or the fragment ions) throughout the whole RT range. Validation of HPLC on-line MS-eluted metabolites was performed by comparing transitions fingerprint, upon fragmentation and matching against the standard metabolites through direct infusion with a syringe pump (infusion rate 4 μl/min). Standard curve calibration was performed either on precursor or on fragment ion signals. Double analyses were performed on both parental and product ion species and results were adopted for quantitation. Transitions were monitored to validate each detected metabolite.
Metabolite data elaboration
LC/MS data files were processed by Bruker DataAnalysis 4.0 (build 234) software, Milan, Italy. Files from each run were either analysed as ‘.d’ files or exported as ‘.mzXML’ files, to be further elaborated for spectra alignment, peak picking and quantitation with InSilicos Viewer 1.5.4 (Insilicos LLC; Seattle, WA, USA).
Quantitative analyses of standard compounds were performed on MRM data. Each standard metabolite was run in triplicate, at incremental dilution until LOD and LOQ were reached. The limit of detection for each compound was calculated as the minimum amount injected that gave a detector response higher than three times the signal-to-noise ratio (S/N). Linearity of the observed quantities, slope, intercept and linear correlation values were all calculated via Microsoft Excel (Microsoft, Redmond, WA, USA).
Analysis of metabolite uptake and release
Cells (1.2 × 106/dish 10 cm) were treated for 12 h with 200 μM GW or ACPA. The cell culture medium was collected and analysed with Bioprofile Flex (Nova Biomedical, Waltham, MA, USA).
Immunofluorescence analysis
Cells (1.6 × 104) were grown on coverslips and treated for 12 h with 200 μM GW or ACPA and 20 μM CC or 20 mM NAC. Cells were incubated with rabbit GAPDH antibody (1 : 100) at RT for 90 min and then incubated with Alexa Fluor 488 anti-rabbit IgG antibody (1 : 500) at RT for 60 min. To assess nuclear morphology, cells were incubated with HOECHST for 2 min at RT. Fluorescence was visualized using excitation/emission wavelengths of 488/520 nm (green) and 350/460 nm (blue) for GAPDH and HOECHST, respectively. Cells were examined using TCS-SP5 Leica confocal microscope, Milan, Italy, at × 40 and × 63 magnification.
PKM2 activity assay
One μg of total protein extract obtained from Panc1 cells treated for 12 and 24 h with 200 μM GW or ACPA was used to analyse PKM2 activity as previously reported.43
c-Myc activity assay
Cells (4 × 105 cells/dish 6 cm) were treated for 1, 12 or 24 h with 200 μM GW or ACPA and then were lysed with nuclear extract kit (Active Motif, Vinci Biochem, Florence, Italy) for nuclear and cytosolic protein extracts. Five μg of the nuclear extract was used to measured c-Myc activity through ELISA assay (Active Motif, TransAM, c-Myc).
Transfection experiments
Cells (2.5 × 105 cells/dish 6 cm) were transfected with the pcDNA5/FRT expression vector containing the human AMPK gamma-2 subunit wt or R531G mutated using TransIT-LT1 transfection reagent (Mirus Bologna, Italy) according to the manufacturer’s directions. Cells were incubated for 72 h and then treated with 200 μM GW or ACPA for 12 h, to evaluate the role of AMP production on AMPK induction. Transfection efficiency was assessed by cytofluorimetric analysis and was ∼56%. The expression vectors for the AMPK wt and mutant R531G γ2 subunit were kindly provided by Dr. Hawley (University of Dundee, Scotland, UK).
Statistical analysis
ANOVA (post hoc Bonferroni) and graphical presentations were performed by GraphPad Prism 5. P-values of *P<0.05, **P<0.01, or ***P<0.001 are indicated in the figures.
Abbreviations
- LC-MS:
-
liquid chromatography–mass spectrometry
- AMPK:
-
AMP-activated protein kinase
- ACPA:
-
arachidonoyl cyclopropylamide
- GW:
-
GW405833 hydrochloride
- NAC:
-
N-acetyl-L-cysteine
- CC:
-
Compound C
- GAPDH:
-
glyceraldehyde 3-phosphate dehydrogenase
- PDHK:
-
pyruvate dehydrogenase kinase
- ROS:
-
reactive oxygen species
- PKM2:
-
pyruvate kinase M2
- CaMKKβ:
-
Ca2+/calmodulin-dependent protein kinase
- F6P:
-
fructose-6-phosphate
- G6P:
-
glucose-6-phosphate
- FDP:
-
fructose 1,6-bisphosphate
- G3P:
-
glyceraldehyde-3-phosphate
- PEP:
-
phosphoenolpyruvate
- LACT:
-
lactate
- KET:
-
alpha-ketoglutarate
- SUCC:
-
succinate
- MA:
-
malate
- PPP:
-
pentose phosphate pathway
- PGL:
-
phosphogluconolactone
References
Hermanson DJ, Cannabinoids Marnett LJ . endocannabinoids, and cancer. Cancer Metastasis Rev 2011; 30: 599–612.
Pisanti S, Bifulco M . Endocannabinoid system modulation in cancer biology and therapy. Pharmacol Res 2009; 60: 107–116.
Pisanti S, Malfitano AM, Grimaldi C, Santoro A, Gazzerro P, Laezza C et al. Use of cannabinoid receptor agonists in cancer therapy as palliative and curative agents. Best Pract Res Clin Endocrinol Metab 2009; 23: 117–131.
Velasco G, Carracedo A, Blazquez C, Lorente M, Aguado T, Haro A et al. Cannabinoids and gliomas. Mol Neurobiol 2007; 36: 60–67.
Qamri Z, Preet A, Nasser MW, Bass CE, Leone G, Barsky SH et al. Synthetic cannabinoid receptor agonists inhibit tumor growth and metastasis of breast cancer. Mol Cancer Ther 2009; 8: 3117–3129.
Sarfaraz S, Afaq F, Adhami VM, Mukhtar H . Cannabinoid receptor as a novel target for the treatment of prostate cancer. Cancer Res 2005; 65: 1635–1641.
Proto MC, Gazzerro P, Di Croce L, Santoro A, Malfitano AM, Pisanti S et al. Interaction of endocannabinoid system and steroid hormones in the control of colon cancer cell growth. J Cell Physiol 2012; 227: 250–258.
Guindon J, Hohmann AG . The endocannabinoid system and cancer: therapeutic implication. Br J Pharmacol 2011; 163: 1447–1463.
De Petrocellis L, Melck D, Palmisano A, Bisogno T, Laezza C, Bifulco M et al. The endogenous cannabinoid anandamide inhibits human breast cancer cell proliferation. Proc Natl Acad Sci USA 1998; 95: 8375–8380.
Donadelli M, Dando I, Zaniboni T, Costanzo C, Dalla Pozza E, Scupoli MT et al. Gemcitabine/cannabinoid combination triggers autophagy in pancreatic cancer cells through a ROS-mediated mechanism. Cell Death Dis 2011; 2: e152.
Salazar M, Carracedo A, Salanueva IJ, Hernandez-Tiedra S, Lorente M, Egia A et al. Cannabinoid action induces autophagy-mediated cell death through stimulation of ER stress in human glioma cells. J Clin Invest 2009; 119: 1359–1372.
Yu L, McPhee CK, Zheng L, Mardones GA, Rong Y, Peng J et al. Termination of autophagy and reformation of lysosomes regulated by mTOR. Nature 2010; 465: 942–946.
Levine B . Cell biology: autophagy and cancer. Nature 2007; 446: 745–747.
Vara D, Salazar M, Olea-Herrero N, Guzman M, Velasco G, Diaz-Laviada I . Anti-tumoral action of cannabinoids on hepatocellular carcinoma: role of AMPK-dependent activation of autophagy. Cell Death Differ 2011; 18: 1099–1111.
Hardie DG . AMP-activated protein kinase: an energy sensor that regulates all aspects of cell function. Genes Dev 2011; 25: 1895–1908.
Hawley SA, Boudeau J, Reid JL, Mustard KJ, Udd L, Makela TP et al. Complexes between the LKB1 tumor suppressor, STRAD alpha/beta and MO25 alpha/beta are upstream kinases in the AMP-activated protein kinase cascade. J Biol 2003; 2: 28.
Shaw RJ, Kosmatka M, Bardeesy N, Hurley RL, Witters LA, DePinho RA et al. The tumor suppressor LKB1 kinase directly activates AMP-activated kinase and regulates apoptosis in response to energy stress. Proc Natl Acad Sci USA 2004; 101: 3329–3335.
Woods A, Johnstone SR, Dickerson K, Leiper FC, Fryer LG, Neumann D et al. LKB1 is the upstream kinase in the AMP-activated protein kinase cascade. Curr Biol 2003; 13: 2004–2008.
Sakamoto K, Goransson O, Hardie DG, Alessi DR . Activity of LKB1 and AMPK-related kinases in skeletal muscle: effects of contraction, phenformin, and AICAR. Am J Physiol Endocrinol Metab 2004; 287: E310–E317.
Hawley SA, Pan DA, Mustard KJ, Ross L, Bain J, Edelman AM et al. Calmodulin-dependent protein kinase kinase-beta is an alternative upstream kinase for AMP-activated protein kinase. Cell Metab 2005; 2: 9–19.
Hurley RL, Anderson KA, Franzone JM, Kemp BE, Means AR, Witters LA . The Ca2+/calmodulin-dependent protein kinase kinases are AMP-activated protein kinase kinases. J Biol Chem 2005; 280: 29060–29066.
Woods A, Dickerson K, Heath R, Hong SP, Momcilovic M, Johnstone SR et al. Ca2+/calmodulin-dependent protein kinase kinase-beta acts upstream of AMP-activated protein kinase in mammalian cells. Cell Metab 2005; 2: 21–33.
Fogarty S, Hawley SA, Green KA, Saner N, Mustard KJ, Hardie DG . Calmodulin-dependent protein kinase kinase-beta activates AMPK without forming a stable complex: synergistic effects of Ca2+ and AMP. Biochem J 2010; 426: 109–118.
Zhou G, Myers R, Li Y, Chen Y, Shen X, Fenyk-Melody J et al. Role of AMP-activated protein kinase in mechanism of metformin action. J Clin Invest 2001; 108: 1167–1174.
Colell A, Ricci JE, Tait S, Milasta S, Maurer U, Bouchier-Hayes L et al. GAPDH and autophagy preserve survival after apoptotic cytochrome c release in the absence of caspase activation. Cell 2007; 129: 983–997.
Kroemer G, Pouyssegur J . Tumor cell metabolism: cancer's Achilles' heel. Cancer Cell 2008; 13: 472–482.
Wise DR, DeBerardinis RJ, Mancuso A, Sayed N, Zhang XY, Pfeiffer HK et al. Myc regulates a transcriptional program that stimulates mitochondrial glutaminolysis and leads to glutamine addiction. Proc Natl Acad Sci USA 2008; 105: 18782–18787.
Castellino RC, Durden DL . Mechanisms of disease: the PI3K-Akt-PTEN signaling node--an intercept point for the control of angiogenesis in brain tumors. Nat Clin Pract Neurol 2007; 3: 682–693.
Fleming A, Noda T, Yoshimori T, Rubinsztein DC . Chemical modulators of autophagy as biological probes and potential therapeutics. Nat Chem Biol 2011; 7: 9–17.
Wang S, Song P, Zou MH . AMP-activated protein kinase, stress responses and cardiovascular diseases. Clin Sci (Lond) 2012; 122: 555–573.
Lamberts RR, Onderwater G, Hamdani N, Vreden MJ, Steenhuisen J, Eringa EC et al. Reactive oxygen species-induced stimulation of 5'AMP-activated protein kinase mediates sevoflurane-induced cardioprotection. Circulation 2009; 120: S10–S15.
Shrivastava A, Kuzontkoski PM, Groopman JE, Prasad A . Cannabidiol induces programmed cell death in breast cancer cells by coordinating the cross-talk between apoptosis and autophagy. Mol Cancer Ther 2011; 10: 1161–1172.
Murray J, Taylor SW, Zhang B, Ghosh SS, Capaldi RA . Oxidative damage to mitochondrial complex I due to peroxynitrite: identification of reactive tyrosines by mass spectrometry. J Biol Chem 2003; 278: 37223–37230.
Colell A, Green DR, Ricci JE . Novel roles for GAPDH in cell death and carcinogenesis. Cell Death Differ 2009; 16: 1573–1581.
Dando I, Fiorini C, Pozza ED, Padroni C, Costanzo C, Palmieri M et al. UCP2 inhibition triggers ROS-dependent nuclear translocation of GAPDH and autophagic cell death in pancreatic adenocarcinoma cells. Biochim Biophys Acta 2013; 1833: 672–679.
Kwon HJ, Rhim JH, Jang IS, Kim GE, Park SC, Yeo EJ . Activation of AMP-activated protein kinase stimulates the nuclear localization of glyceraldehyde 3-phosphate dehydrogenase in human diploid fibroblasts. Exp Mol Med 2010; 42: 254–269.
Cairns RA, Harris IS, Mak TW . Regulation of cancer cell metabolism. Nat Rev Cancer 2011; 11: 85–95.
Hitosugi T, Kang S, Vander Heiden MG, Chung TW, Elf S, Lythgoe K et al. Tyrosine phosphorylation inhibits PKM2 to promote the Warburg effect and tumor growth. Sci Signal 2009; 2: ra73.
David CJ, Chen M, Assanah M, Canoll P, Manley JL . HnRNP proteins controlled by c-Myc deregulate pyruvate kinase mRNA splicing in cancer. Nature 2010; 463: 364–368.
Aquila S, Guido C, Santoro A, Perrotta I, Laezza C, Bifulco M et al. Human sperm anatomy: ultrastructural localization of the cannabinoid1 receptor and a potential role of anandamide in sperm survival and acrosome reaction. Anat Rec (Hoboken) 2010; 293: 298–309.
D'Alessandro A, Gevi F, Zolla L . A robust high resolution reversed-phase HPLC strategy to investigate various metabolic species in different biological models. Mol Biosyst 2011; 7: 1024–1032.
Sana TR, Waddell K, Fischer SM . A sample extraction and chromatographic strategy for increasing LC/MS detection coverage of the erythrocyte metabolome. J Chromatogr B Analyt Technol Biomed Life Sci 2008; 871: 314–321.
Christofk HR, Vander Heiden MG, Wu N, Asara JM, Cantley LC . Pyruvate kinase M2 is a phosphotyrosine-binding protein. Nature 2008; 452: 181–186.
Acknowledgements
This work was supported by Associazione Italiana Ricerca Cancro (AIRC), Milan, Italy; Fondazione CariPaRo, Padova, Italy; Progetti di Ricerca di Interesse Nazionale (PRIN, MIUR), Rome, Italy. We thank Prof. Michela Giuliano and Prof. Maurizio Bifulco for their critical reading of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by G Melino
Supplementary Information accompanies this paper on Cell Death and Disease website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Dando, I., Donadelli, M., Costanzo, C. et al. Cannabinoids inhibit energetic metabolism and induce AMPK-dependent autophagy in pancreatic cancer cells. Cell Death Dis 4, e664 (2013). https://doi.org/10.1038/cddis.2013.151
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2013.151