Abstract
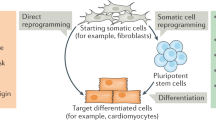
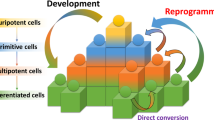
Cell fate decision is a critical step during physiological development when embryonic stem cells commit to either becoming adult stem cells or somatic cells. Recent advances in reprogramming demonstrate that a similar set of transcription factors (TFs), which are important for maintaining the pluripotent state of stem cells, can also reprogram somatic cells to induced pluripotent stem cells (iPSCs). In addition, trans-differentiation, which entails the use of different sets of defined factors, whereby one type of somatic cell can be directly converted into another and even to cell types from different germ layers has become a parallel widely used approach for switching cell fate. All these progresses have provided powerful tools to manipulate cells for basic science and therapeutic purposes. Besides protein-based factors, non-coding RNAs (ncRNAs), particularly microRNAs and long ncRNAs, are also involved in cell fate determination, including maintaining self-renewal of pluripotent stem cells and directing cell lineage. Targeting specific ncRNAs represents an alternative promising approach to optimize cell-based disease modeling and regenerative therapy. Here we focus on recent advances of ncRNAs in cell fate decision, including ncRNA-induced iPSCs and lineage conversion. We also discuss some underlying mechanisms and implications in molecular pathogenesis of human diseases.
Similar content being viewed by others
Facts
-
1
Certain non-coding RNAs (ncRNAs) such as some microRNAs (miRNAs) or long ncRNAs (lncRNAs) are critically involved in induced pluripotency or trans-differentiation.
-
2
Depending on cellular context and species, different miRNAs or lncRNAs can have opposite roles during cell fate transition process.
-
3
ncRNAs regulate various signaling pathways that are critical for cell fate determination.
-
4
Studies of ncRNAs on cell fate decision are very important for understanding pathogenesis of genetic diseases and clinical applications, but it is only emerging and more interesting questions are being raised and await answers.
Open Questions
-
1
How to improve transfection efficiency of ncRNAs into host cells and how to maintain the effective dose to convert cell fate.
-
2
How to avoid activation or repression of unwanted targets to initiate lineage-specific programs, and how to activate or shut down certain pathways to obtain temporal gene expression signatures amenable to unique cell fate.
-
3
How to evaluate potential safety issues caused by ectopic overexpression of ncRNAs such as the off-target effect, cell-type specificity and dose dependency, especially in the context of clinical applications.
ncRNAs
ncRNAs consist of various RNA species that are not translated and evolutionarily conserved among organisms. One single ncRNA may control hundreds of genes. Based on their length, these regulatory ncRNAs can be further divided into short ncRNAs including small interfering RNAs, miRNAs and PIWI-interacting RNAs, intermediate ncRNAs like small nucleolar RNAs, and lncRNAs. Of particular interest, we will give some brief background of miRNAs and lncRNAs. Mature miRNAs are a group of short ncRNAs with approximate 20 nucleotides that target specific mRNA motifs, which may be located within the coding regions or untranslated regions (UTRs). Most miRNAs target hundreds of genes and mainly repress post-transcriptional protein expression. Processing miRNAs require specific factors including Drosha and Dicer, deletion of which abolishes miRNA maturation and critical for embryonic development.1 LncRNAs are a group of ncRNAs >200 nucleotides and widely distributed,1 among which long intergenic ncRNAs represent one particular class located within intergenic regions in the genome with a specific chromatin signature, usually lysine methylation on histones.2, 3 They contain some characteristics of mRNA, including 5’ capping and splicing, but have no peptide-encoding open reading frames.4, 5 LncRNAs are able to regulate protein expression at transcriptional or post-transcriptional level by targeting modifiers to a specific genomic position or working as an enhancer. Below we summarized recent studies that focused on elucidating the essential roles of miRNAs (Table 1) and lncRNAs (Table 2) in somatic cell reprogramming and trans-differentiation.
ncRNAs and Reprogramming
miRNAs and reprogramming
The idea that miRNAs might be involved in pluripotency of stem cells came from the initial discovery of embryonic stem cell (ESC)-specific miRNAs.6, 7 Follow-up studies demonstrated that a subset of these miRNAs have essential roles in cell cycle regulation and self-renewal of ESCs.1, 8, 9, 10, 11 Of note, transcription factors (TFs) could be completely replaced by certain ESC-specific miRNAs to efficiently reprogram human or mouse somatic cells to induced pluripotent stem cells (iPSCs).11 The role of miRNAs in reprogramming is indispensable and irreplaceable because the same set of reprogramming factors fail to reprogram somatic cells when certain miRNA expression is defective.12 One good example of ESC-specific miRNAs is the miR-302 family that is shown to drive the initiation of a pluripotent state.13, 14, 15, 16, 17
Many signaling pathways have been implicated in mediating miRNA-induced reprogramming, including those involved in mesenchymal–epithelial transition (MET), cell cycle regulation, epigenetic modification, and others like nuclear factor kappa-light-chain-enhancer of activated B cell (NF- κB) and transforming growth factor (TGF)-β pathways18 (Figure 1). The miR-302/367 family are able to facilitate pluripotency by regulating all above pathways, which may explain why miR302/367 are sufficient to induce somatic cell reprogramming.14, 15, 17, 19, 20 MET has been proposed to be required for the initial phase of reprogramming and thus could be a preferred target for various miRNAs.21, 22 Some of those miRNAs regulate TGF-β signaling, leading to increased E-cadherin expression, a hallmark of epithelial cells. For example, miR-205 and miR-200 are induced at the initial stage of reprogramming and promote MET in a bone morphogenic protein (BMP, TGF-β superfamily member)-dependent manner, likely through inhibiting Zeb1 and Zeb2, two transcriptional repressors for E-cadherin expression.22, 23 miR-93 and miR-106b regulate TGF-β receptor 2 during MET process.24 miRNAs in the miR-290 cluster share a similar seeding sequence to activate NF-κB signaling pathway.24, 25 p53, a well-studied tumor-suppressor gene and whose activation has been known to be a roadblock for reprogramming, is proved to be a good target for reprogramming-inducing miRNAs.26, 27, 28, 29, 30, 31 miR-138 directly targets 3′-UTR of p53 mRNA and significantly increases reprogramming efficiency.32 In addition to regulate specific pathways mentioned above, miRNAs could modify global gene expression profile by controlling epigenetic factors to induce pluripotency.15, 19 miR-302 represses at least four different epigenetic regulators including lysine-specific histone demethylase 1 and 2, and methyl-CpG-binding proteins 1 and 2, which in turn leads to global demethylation and activation of pluripotency-associated genes.15, 19 Another set of tissue-specific miRNAs, including miR-21, miR29a, miR-34 and miR-199a-3p, have a suppressive role during reprogramming16, 33, 34, 35 (Figure 1). Such miRNAs use various strategies to inhibit reprogramming. miR-21 and miR-29a target pluripotent factors involved in p53 and Erk1/2 pathways to build tissue-specific barriers; miR-34 and miR-199a-3p repress proliferation;33, 34, 35 miR-34 and miR-199a-3p are also involved in p53-associated inhibition of reprogramming in synergy with p21, another p53 downstream effector.27, 33, 34
Scheme describing how ncRNAs modulate induction of somatic cells to iPSCs. Multiple mechanisms are involved: (I) activating pluripotency-associated TFs; (II) activating MET in the context of iPSC formation; (III) promoting cell cycle progression and/or inhibiting apoptosis; and (IV) modulating chromatin-modifying enzymes to affect epigenetic reprogramming of somatic cells. Of note, conventional TFs are able to activate ncRNAs targeting various signaling pathways to facilitate reprogramming and PcG components block the transcription of tissue-specific ncRNAs by co-occupying their promoters with TFs
LncRNAs and reprogramming
LncRNAs represent another group of ncRNAs that are involved in cell fate decision. Loss of function studies demonstrate that lncRNAs regulate genetic and epigenetic activities primarily in a trans-manner at transcription level, a mechanism that differs from siRNA/miRNA pathway.36, 37 The first direct evidence of lncRNA in reprogramming came from the Rinn lab who demonstrated that lincRNA-regulator of reprogramming bears the ability to modulate reprogramming.38 Another example is Xist, a marker of X-chromosome inactivation (XCI) and identified as a molecular signature of human iPSCs.39 Xist-deficient iPSCs exhibit increased expression of some X-linked oncogenes, abnormal growth rates and deficient differentiation potential relative to normal iPSCs.39 LncRNAs may serve as a good benchmark to evaluate certain aspects of stem cell quality. In addition, both Xist lncRNA and its target polycomb repressive complex 2 (PRC2) are required for XCI.40 A positive feedback loop is identified between lncRNAs and TFs in ESCs, probably through epigenetic activation.2, 36, 37, 41, 42, 43 AK028326 and AK141205 are two lncRNAs that were identified as direct targets of the key pluripotent factors Oct4 and Nanog in mouse ESCs, respectively. Although a direct connection between lncRNAs and reprogramming is still missing, owing to the critical role of Oct4 in pluripotency and ES lineage-specific differentiation, these results strongly imply a role of these lncRNAs in controlling stem cell fate.41
ncRNAs and Trans-Differentiation
Trans-differentiation refers to direct conversion of one somatic cell type into another. This approach could avoid the induced pluripotent state that bears perceivable higher oncogenic potential than somatic cells and directly generate patient-specific progenitors or somatic cells for disease modeling and personalized regenerative therapy.44 The capability of TFs to trigger trans-differentiation was initially unveiled by Davis et al.45 More recently, the Wernig group46 demonstrated that mesoderm cells (e.g., fibroblasts) can be directly converted into functional neurons. During the process of trans-differentiation into neurons or their precursors, combinations of multiple TFs have been used.47, 48, 49, 50, 51, 52, 53 Besides TFs, ncRNAs are also involved in trans-differentiation, either alone or in combination with TFs. As recently summarized by Shenoy and Blelloch,54 miRNAs seem to inhibit lineage suppressors to lower the threshold for commitment. One example is miR-124, when combined with MYT1L and BRN2 or miR-9/9*, is able to convert human fibroblasts to functional neurons.55, 56 However, at this stage it is not clear how miRNAs manage to activate neuronal-specific pathways. It will be logical to hypothesize that miR-124 and miR-9/9* target components of chromatin-remodeling complexes, such as BAF53a, PTBP-1 and components of the repressor element-1 silencing transcription factor (REST) complex, which in turn remodels chromatin structure and turns on the neuron-specific epigenetic switch. Yoo et al.56 demonstrated that miR-124 and miR-9/9* suppress fibroblast-expressing BAF53a and activated neurogenesis-essential BAF53b (a 53KD subunit of BRG1/brm-associated factor complex), serving as a potential explanation for miRNA-induced neuronal commitment.54
Although direct evidence for a role of lncRNAs in trans-differentiation is yet-to-be established, lncRNAs have been shown to be critical for regulating the expression of Malat1, Gomafu, Neat1 and RMST, factors that are involved neurogenesis and neural cell fate specification.57, 58, 59, 60 Some lncRNAs physically associate with neural TFs such as REST or epigenetic modulators PRC2, implying that lncRNAs may critically regulate neural trans-differentiation.58, 59, 60
Although miRNAs and lncRNAs possess distinct regulatory mechanisms, they can interplay with each other during cell fate determination.61 Certain lncRNAs share the same imprinted genomic region with miRNAs, as miRNAs are also mapped in non-coding genomic regions and sometimes share genomic regions with lncRNAs.3, 62, 63 Several studies identified active Dlk1-Dio3 region as a marker to distinguish iPSCs with full pluripotency from those that are partially reprogrammed.64, 65, 66 Liu et al.63 demonstrated that miRNA cluster transcribed from active Dlk1-Dio3 region may in turn attenuate imprinting and promote expression of genes and lncRNAs located within the Dlk1-Dio3 region in an epigenetic-dependent pattern by physically targeting constituent parts of PRC2. Therefore, transcription of these lncRNAs is predicted to be under the control of the neighbor miRNAs in fully pluripotent stem cells.63 Meanwhile, miRNAs could be transcriptionally adjusted by lncRNAs in corresponding region.67 Epigenetic changes including DNA and histone modifications may serve as a switch of reciprocal regulations between miRNAs and lncRNAs.3, 63, 68
Perspectives
The conversion of terminally differentiated cells to iPSCs or to other lineages entails dramatic transformations of epigenetic remodeling and gene expression, which was initially examined and validated by studying protein-based factors. Accumulating evidence indicates that ncRNAs target diverse cellular processes including epigenetic modifiers, key TFs, MET, as well as cell cycle regulators. Thus, the combined effect of ncRNAs could regulate cell fate decision in a similar way, if no more efficient than protein factors (Figure 2). Compared with protein-mediated reprogramming or lineage conversions, ncRNAs, especially miRNAs, are more easily introduced into primary cells relative to protein-coding vectors or in vitro recombinant proteins. They are also easier to be degraded and diluted in cells within several passages. In principle, serial transfection of small ncRNAs together with protein-encoding mRNAs can effectively alter cell fate with minimal toxic effect and alterations in genomic DNA. In addition, ncRNA-mediated cell fate switching appears to be more efficient. The Morrisey team reported that miR302/367 can induce iPSCs generation approximately two-fold more efficiently than standard protein-based reprogramming.13 This high efficiency may be partially explained by a coordinated action of more targeting effectors of miRNAs compared with protein factors. Despite these advantages, there are still many barriers before using ncRNAs in basic and therapeutic applications. These barriers include: (1) how to improve the transfection efficiency of miRNAs into host cells, and how to maintain their sustained cellular concentrations; (2) how to avoid activation or repression of unwanted targets of specific ncRNAs to initiate lineage-specific programs, and how to timely activate or shut down certain ncRNAs pathways to obtain spatio-temporal gene expression signatures amenable to unique cell fate; and (3) how to evaluate potential safety issues caused by ectopic overexpression of ncRNAs including off-target effects, cell-type-specific effects and dose-dependent effects, factors to be especially considered in the context of clinical applications. Addressing all these questions will significantly help to advance the development of optimal strategies for basic and therapeutic studies.
Roles of ncRNAs in dedifferentiation and trans-differentiation. Like protein-coding factors, ncRNAs could promote dedifferentiation of somatic cells (e.g., fibroblasts) to iPSCs and other groups of ncRNAs could induce iPSCs to certain functional cells such as neural stem cells or neurons. ncRNAs could also directly convert one type of somatic cells to another type
Increasing evidence has linked ncRNAs disregulation to human diseases.69, 70 Particularly, genetic defects in ncRNAs are a common hallmark of human diseases like cancer and neurological disorders. Yin et al. found that depletion of one class of small nucleolar long ncRNAs is functionally related to Prader–Willi syndrome.71 Such disease-causing disregulation of ncRNAs will provide superior opportunities for studying disease pathophysiology and serve as targets of intervention for therapeutic purposes. Recent progress in iPSC-based gene targeting has established worldwide platforms to study the cellular and molecular mechanisms involved in various hereditary diseases. These platforms could be extended to manipulate expression of ncRNAs in patient-derived iPSCs.72, 73 Correction of disease-causing events related to dysregulation of ncRNAs may provide alternative strategies for cell replacement therapies of genetic diseases.73
Abbreviations
- ncRNAs:
-
non-coding RNAs
- miRNAs:
-
microRNAs
- lncRNAs:
-
long ncRNAs
- lincRNAs:
-
long intergenic non-coding RNAs
- MET:
-
mesenchymal–epithelial transition
- TGF-β:
-
transforming growth factor-β
- NF-κB:
-
nuclear factor kappa-light-chain-enhancer of activated B cells
- TFs:
-
transcription factors
- ESC:
-
embryonic stem cell
- iPSCs:
-
induced pluripotent stem cells
- UTRs:
-
untranslated regions
- BMP:
-
bone morphogenic protein
- XCI:
-
X-chromosome inactivation
- PRC2:
-
polycomb repressive complex 2
- REST:
-
repressor element-1 silencing transcription factor
- lincRNA-RoR:
-
lincRNA-regulator of reprogramming
- BAF53b:
-
a 53KD subunit of BRG1/brm-associated factor complex
- OSKM:
-
the combination of Oct4, Sox2, Klf4 and c-Myc
References
Pauli A, Rinn JL, Schier AF . Non-coding RNAs as regulators of embryogenesis. Nat Rev Genet 2011; 12: 136–149.
Guttman M, Amit I, Garber M, French C, Lin MF, Feldser D et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 2009; 458: 223–227.
Benetatos L, Hatzimichael E, Londin E, Vartholomatos G, Loher P, Rigoutsos I et al. The microRNAs within the DLK1-DIO3 genomic region: involvement in disease pathogenesis. Cell Mol Life Sci 2012.
Consortium TF, Carninci P, Kasukawa T, Katayama S, Gough J, Frith MC et al. The transcriptional landscape of the mammalian genome. Science 2005; 309: 1559–1563.
Kampa D, Cheng J, Kapranov P, Yamanaka M, Brubaker S, Cawley S et al. Novel RNAs identified from an in-depth analysis of the transcriptome of human chromosomes 21 and 22. Genome Res 2004; 14: 331–342.
Houbaviy HB, Murray MF, Sharp PA . Embryonic stem cell-specific MicroRNAs. Dev Cell 2003; 5: 351–358.
Suh M-R, Lee Y, Kim JY, Kim S-K, Moon S-H, Lee JY et al. Human embryonic stem cells express a unique set of microRNAs. Dev Biol 2004; 270: 488–498.
Qi J, Yu J-Y, Shcherbata HR, Mathieu J, Wang AJ, Seal S et al. microRNAs regulate human embryonic stem cell division. Cell Cycle 2009; 8: 3729–3741.
Shcherbata HR, Hatfield S, Ward EJ, Reynolds S, Fischer KA, Ruohola-Baker H . The microRNA pathway plays a regulatory role in stem cell division. Cell Cycle 2006; 5: 172–175.
Murchison EP, Partridge JF, Tam OH, Cheloufi S, Hannon GJ . Characterization of Dicer-deficient murine embryonic stem cells. Proc Natl Acad Sci USA 2005; 102: 12135–12140.
Judson RL, Babiarz JE, Venere M, Blelloch R . Embryonic stem cell-specific microRNAs promote induced pluripotency. Nat Biotechnol 2009; 27: 459–461.
Kim B-M, Thier M-C, Oh S, Sherwood R, Kanellopoulou C, Edenhofer F et al. MicroRNAs are indispensable for reprogramming mouse embryonic fibroblasts into induced stem cell-like cells. PLoS One 2012; 7: e39239.
Anokye-Danso F, Trivedi CM, Juhr D, Gupta M, Cui Z, Tian Y et al. Highly efficient miRNA-mediated reprogramming of mouse and human somatic cells to pluripotency. Cell Stem Cell 2011; 8: 376–388.
Kuo CH, Deng JH, Deng Q, Ying SY . A novel role of miR-302/367 in reprogramming. Biochem Biophys Res Commun 2012; 417: 11–16.
Lin S-L, Chang DC, Chang-Lin S, Lin C-H, Wu DTS, Chen DT et al. Mir-302 reprograms human skin cancer cells into a pluripotent ES-cell-like state. RNA 2008; 14: 2115–2124.
Marson A, Levine SS, Cole MF, Frampton GM, Brambrink T, Johnstone S et al. Connecting microRNA genes to the core transcriptional regulatory circuitry of embryonic stem cells. Cell 2008; 134: 521–533.
Subramanyam D, Lamouille S, Judson RL, Liu JY, Bucay N, Derynck R et al. Multiple targets of miR-302 and miR-372 promote reprogramming of human fibroblasts to induced pluripotent stem cells. Nat Biotechnol 2011; 29: 443–448.
Onder TT, Daley GQ . microRNAs become macro players in somatic cell reprogramming. Genome Med 2011; 3: 40.
Lin S-L, Chang DC, Lin C-H, Ying S-Y, Leu D, Wu DTS . Regulation of somatic cell reprogramming through inducible mir-302 expression. Nucleic Acids Res 2011; 39: 1054–1065.
Lipchina I, Elkabetz Y, Hafner M, Sheridan R, Mihailovic A, Tuschl T et al. Genome-wide identification of microRNA targets in human ES cells reveals a role for miR-302 in modulating BMP response. Genes Dev 2011; 25: 2173–2186.
Li R, Liang J, Ni S, Zhou T, Qing X, Li H et al. A mesenchymal-to-epithelial transition initiates and is required for the nuclear reprogramming of mouse fibroblasts. Cell Stem Cell 2010; 7: 51–63.
Samavarchi-Tehrani P, Golipour A, David L, Sung HK, Beyer TA, Datti A et al. Functional genomics reveals a BMP-driven mesenchymal-to-epithelial transition in the initiation of somatic cell reprogramming. Cell Stem Cell 2010; 7: 64–77.
Gregory PA, Bert AG, Paterson EL, Barry SC, Tsykin A, Farshid G et al. The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nat Cell Biol 2008; 10: 593–601.
Li Z, Yang CS, Nakashima K, Rana TM . Small RNA-mediated regulation of iPS cell generation. EMBO J 2011; 30: 823–834.
Lüningschrör P, Stöcker B, Kaltschmidt B, Kaltschmidt C . miR-290 cluster modulates pluripotency by repressing canonical NF-κB signaling. Stem Cells 2012; 30: 655–664.
Banito A, Rashid ST, Acosta JC, Li S, Pereira CF, Geti I et al. Senescence impairs successful reprogramming to pluripotent stem cells. Genes Dev 2009; 23: 2134–2139.
Hong H, Takahashi K, Ichisaka T, Aoi T, Kanagawa O, Nakagawa M et al. Suppression of induced pluripotent stem cell generation by the p53–p21 pathway. Nature 2009; 460: 1132–1135.
Kawamura T, Suzuki J, Wang YV, Menendez S, Morera LB, Raya A et al. Linking the p53 tumour suppressor pathway to somatic cell reprogramming. Nature 2009; 460: 1140–1144.
Liu Y, Hoya-Arias R, Nimer SD . The role of p53 in limiting somatic cell reprogramming. Cell Res 2009; 19: 1227–1228.
Marion RM, Strati K, Li H, Murga M, Blanco R, Ortega S et al. A p53-mediated DNA damage response limits reprogramming to ensure iPS cell genomic integrity. Nature 2009; 460: 1149–1153.
Utikal J, Polo JM, Stadtfeld M, Maherali N, Kulalert W, Walsh RM et al. Immortalization eliminates a roadblock during cellular reprogramming into iPS cells. Nature 2009; 460: 1145–1148.
Ye D, Wang G, Liu Y, Huang W, Wu M, Zhu S et al. MiR-138 promotes induced pluripotent stem cell generation through the regulation of the p53 signaling. Stem Cells 2012; 30: 1645–1654.
Choi YJ, Lin CP, Ho JJ, He X, Okada N, Bu P et al. miR-34 miRNAs provide a barrier for somatic cell reprogramming. Nat Cell Biol 2011; 13: 1353–1360.
Wang J, He Q, Han C, Gu H, Jin L, Li Q et al. p53-facilitated miR-199a-3p regulates somatic cell reprogramming. Stem Cells 2012; 30: 1405–1413.
Yang CS, Li Z, Rana TM . microRNAs modulate iPS cell generation. RNA 2011; 17: 1451–1460.
Au PCK, Zhu Q-H, Dennis ES, Wang M-B . Long non-coding RNA-mediated mechanisms independent of the RNAi pathway in animals and plants. RNA Biol 2011; 8: 404–414.
Guttman M, Donaghey J, Carey BW, Garber M, Grenier JK, Munson G et al. lincRNAs act in the circuitry controlling pluripotency and differentiation. Nature 2011; 477: 295–300.
Loewer S, Cabili MN, Guttman M, Loh Y-H, Thomas K, Park IH et al. Large intergenic non-coding RNA-RoR modulates reprogramming of human induced pluripotent stem cells. Nat Genet 2010; 42: 1113–1117.
Anguera Montserrat C, Sadreyev R, Zhang Z, Szanto A, Payer B, Sheridan Steven D et al. Molecular signatures of human induced pluripotent stem cells highlight sex differences and cancer genes. Cell Stem Cell 2012; 11: 75–90.
Zhao J, Sun BK, Erwin JA, Song J-J, Lee JT . Polycomb proteins targeted by a short repeat RNA to the mouse X chromosome. Science 2008; 322: 750–756.
Sheik Mohamed J, Gaughwin PM, Lim B, Robson P, Lipovich L . Conserved long noncoding RNAs transcriptionally regulated by Oct4 and Nanog modulate pluripotency in mouse embryonic stem cells. RNA 2010; 16: 324–337.
Dinger ME, Amaral PP, Mercer TR, Pang KC, Bruce SJ, Gardiner BB et al. Long noncoding RNAs in mouse embryonic stem cell pluripotency and differentiation. Genome Res 2008; 18: 1433–1445.
Tsai M-C, Manor O, Wan Y, Mosammaparast N, Wang JK, Lan F et al. Long noncoding RNA as modular scaffold of histone modification complexes. Science 2010; 329: 689–693.
Pfisterer U, Wood J, Nihlberg K, Hallgren O, Bjermer L, Thorsson GW et al. Efficient induction of functional neurons from adult human fibroblasts. Cell Cycle 2011; 10: 3311–3316.
Davis RL, Weintraub H, Lassar AB . Expression of a single transfected cDNA converts fibroblasts to myoblasts. Cell 1987; 51: 987–1000.
Vierbuchen T, Ostermeier A, Pang ZP, Kokubu Y, Südhof TC, Wernig M . Direct conversion of fibroblasts to functional neurons by defined factors. Nature 2010; 463: 1035–1041.
Caiazzo M, Dell/'Anno MT, Dvoretskova E, Lazarevic D, Taverna S, Leo D et al. Direct generation of functional dopaminergic neurons from mouse and human fibroblasts. Nature 2011; 476: 224–227.
Lujan E, Chanda S, Ahlenius H, Südhof TC, Wernig M . Direct conversion of mouse fibroblasts to self-renewing, tripotent neural precursor cells. Proc Natl Acad Sci USA 2012; 109: 2527–2532.
Pfisterer U, Kirkeby A, Torper O, Wood J, Nelander J, Dufour A et al. Direct conversion of human fibroblasts to dopaminergic neurons. Proc Natl Acad Sci USA 2011; 108: 10343–10348.
Qiang L, Fujita R, Yamashita T, Angulo S, Rhinn H, Rhee D et al. Directed conversion of Alzheimer's disease patient skin fibroblasts into functional neurons. Cell 2011; 146: 359–371.
Ring Karen L, Tong Leslie M, Balestra Maureen E, Javier R, Andrews-Zwilling Y, Li G et al. Direct reprogramming of mouse and human fibroblasts into multipotent neural stem cells with a single factor. Cell Stem Cell 2012; 11: 100–109.
Son Esther Y, Ichida Justin K, Wainger Brian J, Toma Jeremy S, Rafuse Victor F, Woolf Clifford J et al. Conversion of mouse and human fibroblasts into functional spinal motor neurons. Cell Stem Cell 2011; 9: 205–218.
Thier M, Wörsdörfer P, Lakes Yenal B, Gorris R, Herms S, Opitz T et al. Direct conversion of fibroblasts into stably expandable neural stem cells. Cell Stem Cell 2012; 10: 473–479.
Shenoy A, Blelloch R . microRNA induced transdifferentiation. F1000 Biol Rep 2012; 4: 3.
Ambasudhan R, Talantova M, Coleman R, Yuan X, Zhu S, Lipton Stuart A et al. Direct reprogramming of adult human fibroblasts to functional neurons under defined conditions. Cell Stem Cell 2011; 9: 113–118.
Yoo AS, Sun AX, Li L, Shcheglovitov A, Portmann T, Li Y et al. MicroRNA-mediated conversion of human fibroblasts to neurons. Nature 2011; 476: 228–231.
Bond AM, VanGompel MJW, Sametsky EA, Clark MF, Savage JC, Disterhoft JF et al. Balanced gene regulation by an embryonic brain ncRNA is critical for adult hippocampal GABA circuitry. Nat Neurosci 2009; 12: 1020–1027.
Johnson R, CH-L Teh, Jia H, Vanisri RR, Pandey T, Lu Z-H et al. Regulation of neural macroRNAs by the transcriptional repressor REST. RNA 2009; 15: 85–96.
Mercer T, Qureshi I, Gokhan S, Dinger M, Li G, Mattick J et al. Long noncoding RNAs in neuronal-glial fate specification and oligodendrocyte lineage maturation. BMC Neurosci 2010; 11: 14.
Ng SY, Johnson R, Stanton LW . Human long non-coding RNAs promote pluripotency and neuronal differentiation by association with chromatin modifiers and transcription factors. EMBO J 2012; 31: 522–533.
Salmena L, Poliseno L, Tay Y, Kats L, Pandolfi PP . A ceRNA hypothesis: the Rosetta Stone of a hidden RNA language? Cell 2011; 146: 353–358.
Carthew RW, Sontheimer EJ . Origins and mechanisms of miRNAs and siRNAs. Cell 2009; 136: 642–655.
Liu L, Luo GZ, Yang W, Zhao X, Zheng Q, Lv Z et al. Activation of the imprinted Dlk1-Dio3 region correlates with pluripotency levels of mouse stem cells. J Biol Chem 2010; 285: 19483–19490.
Stadtfeld M, Apostolou E, Akutsu H, Fukuda A, Follett P, Natesan S et al. Aberrant silencing of imprinted genes on chromosome 12qF1 in mouse induced pluripotent stem cells. Nature 2010; 465: 175–181.
Stadtfeld M, Apostolou E, Ferrari F, Choi J, Walsh RM, Chen T et al. Ascorbic acid prevents loss of Dlk1-Dio3 imprinting and facilitates generation of all-iPS cell mice from terminally differentiated B cells. Nat Genet 2012; 44: 398–405 S391-392.
Zhang W, Guan D, Qu J, Liu GH . Non-viral iPSCs: a safe way for therapy? Protein Cell 2012; 3: 241–245.
Augoff K, McCue B, Plow EF, Sossey-Alaoui K . miR-31 and its host gene lncRNA LOC554202 are regulated by promoter hypermethylation in triple-negative breast cancer. Mol Cancer 2012; 11: 5.
Wu SC, Kallin EM, Zhang Y . Role of H3K27 methylation in the regulation of lncRNA expression. Cell Res 2010; 20: 1109–1116.
Esteller M . Non-coding RNAs in human disease. Nat Rev Genet 2011; 12: 861–874.
Mendell Joshua T, Olson Eric N . MicroRNAs in stress signaling and human disease. Cell 2012; 148: 1172–1187.
Yin QF, Yang L, Zhang Y, Xiang JF, Wu YW, Carmichael GG et al. Long noncoding RNAs with snoRNA ends. Mol Cell 2012; 48: 219–230.
Liu GH, Suzuki K, Qu J, Sancho-Martinez I, Yi F, Li M et al. Targeted gene correction of laminopathy-associated LMNA mutations in patient-specific iPSCs. Cell Stem Cell 2011; 8: 688–694.
Pan H, Zhang W, Liu GH . Find and replace: editing human genome in pluripotent stem cells. Protein Cell 2011; 2: 950–956.
Lee SI, Lee BR, Hwang YS, Lee HC, Rengaraj D, Song G et al. MicroRNA-mediated posttranscriptional regulation is required for maintaining undifferentiated properties of blastoderm and primordial germ cells in chickens. Proc Natl Acad Sci USA 2011; 108: 10426–10431.
Miyoshi N, Ishii H, Nagano H, Haraguchi N, Dewi DL, Kano Y et al. Reprogramming of mouse and human cells to pluripotency using mature microRNAs. Cell Stem Cell 2011; 8: 633–638.
Tabuchi T, Satoh M, Itoh T, Nakamura M . MicroRNA-34a regulates the longevity-associated protein SIRT1 in coronary artery disease: effect of statins on SIRT1 and microRNA-34a expression. Clin Sci 2012; 123: 11.
Xu N, Papagiannakopoulos T, Pan G, Thomson JA, Kosik KS . MicroRNA-145 regulates OCT4, SOX2, and KLF4 and represses pluripotency in human embryonic stem cells. Cell 2009; 137: 647–658.
Bin G, Jiarong Z, Shihao W, Xiuli S, Cheng X, Liangbiao C et al. Aire promotes the self-renewal of embryonic stem cells through Lin28. Stem Cells Dev 2012; 21: 2878–2890.
Acknowledgements
We are grateful to P Schwarz, M Schwarz and A Goebl for critical reading of the manuscript. G-HL was supported by the Thousand Young Talents program of China, the National Laboratory of Biomacromolecules, the Strategic Priority Research Program of the Chinese Academy of Sciences (XDA01020312), and NSFC (81271266, 31222039 and 31201111). WZ was supported by a grant CA158055 from the National Institutes of Health, USA and a Startup Funding from the Department of Pathology, University of Iowa. JCIB was supported by TERCEL-ISCIII-MINECO, CIBER, Fundacion Cellex, the Glenn Foundation, G Harold and Leila Y Mathers Charitable Foundation, Sanofi, the Leona M and Harry B Helmsley Charitable Trust and the Ellison Medical Foundation.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by A Stephanou
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Guan, D., Zhang, W., Zhang, W. et al. Switching cell fate, ncRNAs coming to play. Cell Death Dis 4, e464 (2013). https://doi.org/10.1038/cddis.2012.196
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2012.196
Keywords
This article is cited by
-
Ensemble Deep Learning Based on Multi-level Information Enhancement and Greedy Fuzzy Decision for Plant miRNA–lncRNA Interaction Prediction
Interdisciplinary Sciences: Computational Life Sciences (2021)
-
Vigor of survival determinism: subtle evolutionary gradualism interspersed with robust phylogenetic leaping
Theory in Biosciences (2017)
-
Chromatin modifiers and remodellers: regulators of cellular differentiation
Nature Reviews Genetics (2014)
-
A Helm model for microRNA regulation in cell fate decision and conversion
Science China Life Sciences (2013)