Abstract
MicroRNAs (miRNAs) with tumor-suppressor potential might have therapeutic applications in multiple myeloma (MM) through the modulation of still undiscovered molecular pathways. Here, we investigated the effects of enforced expression of miR-29b on the apoptotic occurrence in MM and highlighted its role in the context of a new transcriptional loop that is finely tuned by the proteasome inhibitor bortezomib. In details, in vitro growth inhibition and apoptosis of MM cells was induced by either transient expression of synthetic miR-29b or its stable lentivirus-enforced expression. We identified Sp1, a transcription factor endowed with oncogenic activity, as a negative regulator of miR-29b expression in MM cells. Since Sp1 expression and functions are regulated via the 26S proteasome, we investigated the effects of bortezomib on miR-29b-Sp1 loop, showing that miR-29b levels were indeed upregulated by the drug. At the same time, the bortezomib/miR-29b combination produced significant pro-apoptotic effects. We also demonstrated that the PI3K/AKT pathway plays a major role in the regulation of miR-29b-Sp1 loop and induction of apoptosis in MM cells. Finally, MM xenografts constitutively expressing miR-29b showed significant reduction of their tumorigenic potential. Our findings indicate that miR-29b is involved in a regulatory loop amenable of pharmacologic intervention and modulates the anti-MM activity of bortezomib in MM cells.
Similar content being viewed by others
Main
MicroRNAs (miRNAs) are a class of naturally occurring small non-coding RNA molecules of 19–25 nucleotides that negatively regulate gene expression at post-transcriptional level by base pairing with the 3′ untranslated region (UTR) of their target mRNAs. It is believed that if this base pairing is perfect, the mRNA is cleaved and degraded.1 However, translational repression by imperfect pairing is the prevalent activity of human miRNAs.2
Since a single miRNA is able to bind to and consequently regulate approximately up to 100 different transcripts,3, 4 miRNAs truly represent master regulators of gene expression and influence virtually all cell activities and events, including cell proliferation, cell death, differentiation, metabolism, infection and cancer.5 Deregulation of miRNA expression commonly occurs in human cancer and is involved in cancer initiation and progression;6, 7 miRNA expression profiles have prognostic implications,8 while targeting the miRNA network exerts anti-tumor effects in vivo,9 indicating a great therapeutic potential of miRNAs for cancer treatment.10 Noteworthy, miRNAs that are deleted or downregulated in malignancies generally play a tumor-suppressor function, whereas those that are upregulated act as transforming oncogenes.11, 12 However, the correlation between expression levels and oncogenetic/suppressor function remains still unclear.13
Multiple myeloma (MM) is an hematologic malignancy characterized by the accumulation of tumor plasma cells (PCs) in the bone marrow (BM), which causes bone destruction and marrow failure.14 Although defined histologically as a single entity, MM encompasses a plethora of disease subsets characterized by distinct genomic abnormalities, which possibly mark different pathogenetic pathways and in most cases retain prognostic value. Such molecular events, beside reprogramming the cell proliferation capacities, finely tune the interaction between malignant PCs and BM stromal cells (BMSCs) within the human BM microenvironment (huBMM).15, 16 The huBMM plays a critical role in promoting MM cell growth, survival and development of drug resistance; moreover, this cell-to-cell interaction provides a framework for the design of novel therapeutic strategies.17 In the last years, the huge advancement in the knowledge of MM pathogenesis allowed some drugs, such as thalidomide,18 bortezomib19 and lenalidomide,20 to reach the clinical side, with significant improvement in terms of disease control and long-term survival.21 However, while many other novel agents are presently in advanced phase of preclinical investigation,22, 23, 24, 25 MM is still an incurable disease and novel therapeutic strategies are urgently needed.
To date, few studies have investigated the expression of miRNAs in MM, mostly by comparing the miRNA expression profile of BM-derived PCs of healthy donors with that of the PCs from subjects with monoclonal gammopaty of undetermined significance (MGUS) or MM patients; from these studies, it clearly emerges that several miRNAs are deregulated in primary or established MM cell lines and are involved in key roles in regulatory networks controlling proliferation and/or survival.26, 27, 28 Among miRNAs frequently deregulated in human cancer, miR-29b is of great interest due to its potential for therapeutic applications. miR-29b belongs to a family whose members are located at two loci on different chromosomes: miR-29b1/29a at 7q32, and miR-29b2/c at 1q23.29 An aggressive clinical behavior of B-cell chronic leukemia correlates with reduced levels of miR-29b and miR-181 and increased expression of their common target, the B- and T-cell malignancy specific oncogene, TCL1.30 Several additional miR-29b targets, including the anti-apoptotic protein Mcl-1,31, 32 the phosphatidyl inositol-3 kinase,33 the DNA methyltransferase 3A and B,34 the extracellular matrix proteins, collagens35 and cell-cycle regulators,36 may contribute to the malignant phenotype and metastatic progression.29 Therefore, it is conceivable that the enforced expression of miR-29b in cancer cells may interfere with multiple oncogenic mechanisms, including tumor-suppressor promoter methylation,34 extracellular matrix remodeling35 and anti-apoptotic signaling.31 Moreover, it is of note that miR-29b/miR-29a are downregulated by the activation of multiple growth and survival promoting signaling such as c-myc, Hedgehog and NF-κB pathways,37 while miR-29b overexpression induces apoptosis in vitro31, 32 and exerts anti-tumor effects in vivo in acute myeloid leukemia and rabdomyosarcoma.36, 38
Based on these premises, we investigated the in vitro and in vivo effects of synthetic miR-29b on MM cell proliferation and apoptosis and explored on the molecular pathways that underline or counteract miR-29b function. Finally, we investigated the role of bortezomib in upregulating miR-29b levels thus antagonizing the escape mechanisms from miR-29b induced growth inhibition and apoptosis in MM cells.
Results
Expression of miR-29b in primary CD138+ cells and established MM cell lines
We evaluated by high-density microarrays miRNA profiling the expression of miR-29b in primary CD138+ cells from intramedullary MM (n=38) or PC leukemia (PCL; n=2) patients. As shown in Figure 1a, we found wide variation of miR-29b expression with 55% (21/38) of MM and 100% (2/2) of PCL patients bearing low miR-29b (<100 relative expression levels – REL – in arbitrary units referred to the median value).
miR-29b expression in MM primary samples and cell lines. (a) miR-29b expression levels obtained by Agilent-miRNA profiling of 723 human and 76 human/viral miRNAs. Histogram bars represent miR-29b expression values in 38 MM and 2 PCL patients, normalized by aroma.light-package for Bioconductor. (b) Quantitative RT–PCR analysis of miR-29b using total RNA from 10 MM cell lines. Raw Ct values were normalized to RNU44 housekeeping snoRNA and expressed as ΔΔCt values referenced to miR-29b levels in PBMC. Values represent mean of three different experiments±S.E. (c) Quantitative RT–PCR of miR-29b in NCI-H929 cells co-cultured with MM patient-derived BMSCs and then immunopurified by immunomagnetic sorting with anti-CD138 beads. Raw Ct values were normalized to RNU44 housekeeping snoRNA and expressed as ΔΔCt values calculated using the comparative cross threshold method. miR-29b levels in NCI-H929 harvested after 6 h of co-culture were set as internal reference. Data are the average of two independent experiments performed in triplicate. P-values were obtained using two-tailed t test. *P<0.01
In addition, we evaluated miR-29b expression by quantitative RT–PCR (qRT–PCR) in a panel of 10 MM cell lines, including OPM1, INA-6, OPM2, NCI-H929, MM1S, LR5, DOX6, SKMM1, KMS12-PE, and RPMI-8226. Heterogeneous levels of miR-29b were detected, suggesting again a deregulated activity (Figure 1b).
Moreover, to study the influence of the huBMM on miR-29b expression, serum-starved NCI-H929 cells were cultured adherent to patient-derived BMSCs, and the expression of miR-29b was analyzed by qRT–PCR at 6, 12, and 24 h in MM cells. Importantly, adhesion of NCI-H929 cells to BMSCs induced a decrease of miR-29b expression after 12 h (Figure 1c), suggesting that low miR-29b might indeed be associated with the supportive effect exerted by huBMM on MM cell growth and survival. Similar results were obtained with RPMI-8226 cells (Supplementary Figure 1).
Synthetic miR-29b reduces cell growth and induces apoptosis in MM cell lines in vitro
On the basis of the previous data, we studied the in vitro activity of miR-29b against different MM cell lines including IL-6-dependent (INA-6) and IL-6-independent (RPMI-8226, OPM1, NCI-H929, SKMM1) MM cells. Cells were electroporated with synthetic miR-29b or scrambled (NC) oligonucleotides and viable cell numbers were determined by trypan blue exclusion assay at different time points (Figure 2; transfection procedures are reported in Supplementary materials and methods). To confirm the successful transfection, qRT–PCR analysis of miR-29b was performed on MM cells 24 h later (Supplementary Figure 2). As shown in Figures 2a–e, overexpression of miR-29b resulted in a significant inhibition of cell growth of MM cell lines starting 48 h after electroporation. Moreover, this effect was independent of IL-6 since it occurred in all cell lines. Interestingly, growth inhibition was associated to upregulation of the cell-cycle inhibitors p21 and p27, both observed 24 h after cell transfection with synthetic miR-29b (Figure 2f; for western blotting procedures, see Supplementary materials and methods).
miR-29b-enforced expression triggers anti-proliferative and pro-apoptotic effects in MM cells. Cell growth curves of (a) RPMI-8226, (b) SKMM1, (c) OPM1, (d) INA-6, and (e) NCI-H929 transfected with synthetic miR-29b (miR-29b) or scrambled oligonucleotides (NC). Averaged values of three independent experiments are plotted including±S.D. P-values were obtained using two-tailed t-test *P<0.01; •P<0.05. (f) Immunoblot of p21WAF1 and p27KIP1 in SKMM1 and NCI-H929 cells 24 h after transfection with synthetic miR-29b or scrambled oligonucleotides (NC). Colony formation assay of SKMM1 (g) and NCI-H929 (h) after transfection with synthetic miR-29b or scrambled oligonucleotides (NC). Six hours after transfection, cells were counted and seeded in triplicate at 200 cells/ml in 18-well plates containing methylcellulose-based medium. After 2 weeks, colony formation was scored by counting colonies including >100 cells. Data indicate the percentage of colonies in the miR-29b methylcellulose cultures with respect to control±S.E. for three independent experiments. *P<0.01. (i) Annexin V-staining of SKMM1 and NCI-H929 cells 48 h after transfection with synthetic miR-29b or scrambled oligonucleotides (NC). The percentage of Annexin-V-positive cells is reported. Data are the average of three independent experiments±S.D. *P<0.01; •P<0.05. (j) Caspase 3/7 activity assay was performed in SKMM1 and NCI-H929 cells 48 h after transfection with synthetic miR-29b or scrambled oligonucleotides (NC). Transfected cells were grown in 96-well microplates and luminescence was measured using the Caspase 3/7 Glo assay (Promega). Results are expressed as fold increase of caspase activity with respect to NC-transfected cells, of at least three independent experiments. *P<0.01. (k) Immunoblot of cleaved caspase 3 and survivin, 48 h after transfection of SKMM1 and NCI-H929 cells with synthetic miR-29b or scrambled oligonucleotides (NC). Loading control was performed using GAPDH
We also found that transfection of synthetic miR-29b in SKMM1 and NCI-H929 MM cells significantly inhibited colony formation in methylcellulose cultures (55 and 35% reduction in colonies, respectively; Figures 2g and h).
Moreover, we evaluated the effects of stably enforced expression of the miR-29b gene carried by lentiviral vectors in MM cells. SKMM1, U266, and RPMI-8226 cell lines were transduced by a lentiviral miR-29b expression vector and subsequently selected by Zeocin (see Supplementary Materials and methods and Supplementary Figure 3A). We found that stable expression of miR-29b strongly reduced cell survival in a time-dependent manner, as demonstrated by MTS assay (Supplementary Figure 3B–D).
We next examined, by Annexin V/7AAD assay, whether apoptotic events occurred in cells transfected with miR-29b. As shown in Figure 2i, ectopic expression of miR-29b triggered apoptosis at 48 h in both SKMM1 and NCI-H929 cells (about twofold increase). Importantly, we found that apoptotic events induced by miR-29b were associated with reduced survivin levels and with caspase 3 activation, as revealed by increased caspase 3/7 activity using an enzymatic assay (Figure 2j) and by enhanced caspase 3 cleavage and reduced survivin expression in western blotting experiments (Figure 2k). Conversely, transfection of miR-29b mimics did not activate caspase 8 in this cell system (data not shown).
Taken together, these results indicate that miR-29b is a negative regulator of MM cell growth and inducer of apoptosis.
miR-29b regulates CDK6 and MCL-1 in MM cells
miR-29b is known to exert anti-proliferative and pro-apoptotic effects in several human cancers by targeting genes involved in cell-cycle progression and/or apoptosis inhibition.31, 33, 36 Based on this consideration, we focused on MCL-1 and CDK6, which are overexpressed in MM and play a pivotal role in apoptosis and cell-cycle regulation, respectively. Enforced expression of miR-29b in SKMM1 cells resulted in a significant downregulation of both MCL-1 and CDK6 mRNAs (Figure 3a) and proteins (Figure 3b), 24 h after cell transfection. As shown in Figure 3b, transfection of miR-29b also resulted in decreased Rb Ser780 phosphorylation. Thus, miR-29b indirectly affected the Rb phosphorylation status, likely by targeting CDK6 that is known to regulate Rb phosphorylation.39 These results were also reproduced in the NCI-H929 cell line: transfection with miR-29b decreased CDK6 and MCL-1 at both mRNA (Figure 3c) and protein levels (Figure 3d). Densitometric analysis of blots is reported in Supplementary Figure 4. Conversely, inhibition of miR-29b in SKMM1 cells by means of antagomirs (Figure 3e) resulted in upregulation of both CDK6 and MCL-1 mRNAs (Figure 3f).
miR-29b expression in MM cells downregulates CDK6 and MCL-1 levels. Quantitative RT–PCR of MCL-1 and CDK6 24 h after transfection with synthetic miR-29b or scrambled oligonucleotides (NC) in SKMM1 (a) and NCI-H929 cells (c). The results are shown as average mRNA expression after normalization with GAPDH and ΔΔCt calculations. Data represent the average of three independent experiments±S.D. Immunoblot of MCL-1, CDK6, and Ser780-Rb phosphorylated 24 h after transfection with synthetic miR-29b or scrambled oligonucleotides (NC) in SKMM1 (b) and NCI-H929 cells (d). The protein loading control was performed using GAPDH. (e) Quantitative RT–PCR of miR-29b in SKMM1 cells after transduction with a lentivirus carrying the empty vector or antagomiR-29b (anti-miR-29b). Raw Ct values were normalized to RNU44 housekeeping snoRNA and expressed as ΔΔCt values. (f) Quantitative RT–PCR of MCL-1 and CDK6 in SKMM1 cells transduced with the empty vector (indicated as vector) or antagomiR-29b (anti-29b). The results are shown as average mRNA expression after normalization with GAPDH and ΔΔCt calculations. *P<0.001; •P<0.05
miR-29b exerts anti-MM activity in vivo
Effect of miR-29b on tumorigenic potential of MM cells was evaluated in vivo. NCI-H929 cells were transduced with a lentivirus expressing miR-29b (pMIF-miR-29b) or the empty vector (pMIF). These cells were inoculated in a cohort of 12 SCID mice, and tumor formation was monitored. As shown in Figure 4a, enforced expression of miR-29b reduced the growth of MM xenografts compared with the control group (P<0.0001), with an average size of tumors of 617±60 S.D. versus 385±60 S.D. at day 23. This effect resulted into a significant survival advantage for mice engrafted with miR-29b-trasduced MM cells (Figure 4b). Quantitative analysis carried out in excised tumors demonstrated a more than fivefold increase of miR-29b expression in pMIF-miR-29b-transduced tumors compared with controls (Figure 4c) that was associated to increased in vivo apoptosis and reduced proliferation, as shown by IHC staining of excised tumors (Figure 4d). A downregulation of CDK6 and MCL-1 proteins was also observed in lysates from retrieved xenografts (Supplementary Figure 5).
miR-29b-enforced expression has anti-tumor activity in a MM mouse model. (a) In vivo tumor formation of NCI-H929 xenografts stably expressing miR-29b after lentivirus transduction with the empty vector (pMIF) or miR-29b (pMIF-miR-29b). Transduced cells were subcutaneously inoculated in the interscapular area of mice and tumor volumes were measured by an electronic caliper in two dimensions every 2–3 days starting the appearance of a palpable tumor mass. The picture inserted in the graph shows the in vivo detection of the tumor mass in a representative pMIF or pMIF-miR-29b NCI-H929-xenografted mouse using IVIS LUMINA II Imaging System (Caliper Life sciences). *P<0.0001. (b) Survival curves (Kaplan–Meier) show prolongation of survival of miR-29b (PMIF-miR-29b) xenografted animals compared with the empty vector (PMIF) control (log-rank test, P<0.05). (c) Quantitative RT–PCR analysis of miR-29b using total RNA from retrieved NCI-H929 xenografts. Raw Ct values were normalized to RNU44 housekeeping snoRNA and expressed as ΔΔCt values. Values represent mean of three different experiments±S.D. (d) Histology (H&E, original magnification, × 40), Caspase 3 and Ki-67 immunohistochemical staining (original magnification, × 20) of NCI-H929 tumor xenografts
All together, these findings indicate that miR-29b exerts in vivo anti-MM activity mediated by apoptosis occurrence, providing a strong rationale for clinical development of miR-29b.
Evidence of a miR-29b-Sp1 regulatory loop in MM cells
Among miR-29b validated targets, we focused our attention on Sp1, a transcription factor that regulates the expression of cell-cycle, differentiation, and apoptosis-related genes. In fact, we have recently shown that high Sp1 activity is a feature of MM cells and that it can be therapeutically targeted.40
We investigated the role of miR-29b on Sp1 mRNA regulation by transfecting into MM cells a luciferase reporter vector that contains the 3′UTR region of Sp1 predicted to interact with miR-29b. A marked reduction in the Luciferase/Renilla ratio was seen in cells transfected with Sp1 construct together with synthetic miR-29b but not with the control (scrambled oligonucleotides; Figure 5a). Moreover, transfection of miR-29b mimics into SKMM1 or NCI-H929 cells decreased Sp1 expression at both mRNA (Figure 5b) and protein levels (Figure 5c). Conversely, inhibition of miR-29b by miR-29b-antagomiR transduction resulted in increased Sp1 mRNA levels (Figure 5d), demonstrating that Sp1 is regulated by miR-29b in MM cells.
Regulatory feedback between miR-29b and its target Sp1. (a) Dual-luciferase assay of SKMM1 and NCI-H929 cells co-transfected with firefly luciferase constructs containing the 3′UTR of Sp1 (nts 2503–5200) and miR-29b or scrambled oligonucleotides (NC) as indicated. The firefly luciferase activity was normalized to renilla luciferase activity. The data are shown as relative luciferase activity of miR-29b-transfected cells as compared with the control (NC) of a total of eight experiments from two independent transfections. (b) Quantitative RT–PCR of Sp1 24 h after transfection with synthetic miR-29b or scrambled oligonucleotides (NC) of SKMM1 and NCI-H929 cells. The results are shown as average mRNA expression after normalization with GAPDH and ΔΔCt calculations. (c) Immunoblot of Sp1 24 h after transfection with synthetic miR-29b or scrambled oligonucleotides in SKMM1 and NCI-H929 cells. The protein loading control was performed using GAPDH. (d) Quantitative RT–PCR of Sp1 in SKMM1 cells transduced with the empty vector or antagomiR-29b (anti-miR-29b). The results are shown as average mRNA expression after normalization with GAPDH and ΔΔCt calculations. (e) Dual-luciferase assay of SKMM1 and NCI-H929 cells transfected with a firefly luciferase construct containing the miR-29a/b1 promoter together with an Sp1 expression construct or pBABE empty vector as control. *P<0.001. (f) Quantitative RT–PCR of miR-29b in SKMM1 and NCI-H929 cells 24 h after transfection with the empty vector (pBABE) or the Sp1-expression vector. Raw Ct values were normalized to RNU44 housekeeping snoRNA and expressed as ΔΔCt values. (g) Quantitative RT–PCR of miR-29b in SKMM1 cells stably expressing two different shRNAs against Sp1 (shSp1#1 and shSp1#2) or a scrambled sequence (SCR). The immunoblot shows the levels of Sp1 after silencing by Sp1-specific shRNAs. GAPDH was used as loading control. (h) Quantitative RT–PCR of miR-29b in SKMM1 cells treated with 160 nM mithramycin-A or vehicle (DMSO) for 15 h. *P<0.001; •P<0.05
Since miRNAs are frequently involved in feedback loops where they target factors that negatively regulate their own expression,41 we investigated whether Sp1 was able to affect miR-29b expression in MM cells. To address this question, we evaluated if Sp1 was able to transcriptionally target miR-29b. Consistently with Sp1-dependent transcriptional repression of miR-29a/b1 promoter, we found that enforced expression of Sp1 (Supplementary Figure 6) decreased promoter activity in SKMM1 and NCI-H929 cells by about 70 and 60%, respectively (Figure 5e); as a consequence, Sp1 overexpression was associated with reduced miR-29b levels (Figure 5f). Moreover, we performed loss-of-function experiments in SKMM1 cells by means of shRNAs against Sp1. As shown in Figure 5g, silencing of Sp1 increased miR-29b levels.
The role of Sp1 in miR-29b regulation was additionally elucidated by treating SKMM1 cells with mithramycin-A, a previously reported Sp1 inhibitor.42 Again, exposure to mithramycin-A led to decrease in Sp1 protein expression (not shown) and increased miR-29b levels (Figure 5h).
miR-29b expression is regulated by bortezomib
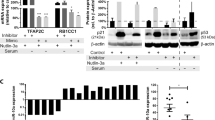
Sp1 expression and functions are, at least in part, regulated via the 26S proteasome, a common pathway controlling the degradation of a plethora of other survival factors.43 Some authors reported that the 26S proteasome inhibitor bortezomib interferes with Sp1/NFkB activity.44 To evaluate the potential regulatory role of bortezomib on the miR-29b-Sp1 loop, NCI-H929 cells were treated for 24 h with vehicle alone or bortezomib 2, 5, and 10 nM. Bortezomib treatment led to a dose-dependent increase in apoptotic cells (Figure 6a) together with downregulation of Sp1 (Figure 6b), and upregulation of miR-29b (Figure 6c). Notably, miR-29b upregulation by bortezomib was further demonstrated in SKMM1 and U266 cells (Figure 6d). A relevant finding was that primary CD138+ cells immunopurified from peripheral blood of a PCL patient also demonstrated upregulation of miR-29b 24 h after treatment with bortezomib (1.3 mg/m2) as compared with PCs before treatment (Figure 6e).
Bortezomib modulates miR-29b expression levels in MM cells. (a) Annexin V-staining of NCI-H929 cells 24 h after treatment with vehicle or bortezomib (indicated as bort) 2, 5, or 10 nM. •P<0.05. (b) Immunoblot of Sp1 in NCI-H929 cells treated for 24 h with vehicle or bortezomib (indicated as bort) 5, 10, or 20 nM. GAPDH was used as loading control. Quantitative RT–PCR of miR-29b in (c) NCI-H929 cells treated with vehicle or 2, 5, or 10 nM bortezomib for 24 h or (d) SKMM1and U266 treated with 10 nM bortezomib or vehicle for 24 h. Raw Ct values were normalized to RNU44 housekeeping snoRNA and expressed as ΔΔCt. *P<0.01; •P<0.05. (e) Quantitative RT–PCR of miR-29b in CD138+ cells immunopurified from peripheral blood of a PCL patient, 24 h after systemic treatment with 1.3 mg/kg bortezomib. Normalization was performed using RNU44 snoRNA and ΔΔCt calculations. *P<0.01. (f) Quantitative RT–PCR of miR-29b in NCI-H929 treated with vehicle (DMSO) or 5 nM bortezomib (indicated as bort) or with 5 nM bortezomib and 20 μM Z-IETD-FMK for 24 h. *P<0.01; •P<0.05
Previous reports showed that bortezomib promotes Sp1 downregulation in MM cells through caspase 8 activation.45 To evaluate whether modulation of caspase 8 activity could also affect miR-29b levels, NCI-H929 cells were treated with bortezomib alone or in combination with caspase 8 inhibitor Z-IETD-FMK. We found that caspase 8 inhibition abrogated bortezomib-induced miR-29b upregulation (Figure 6f).
miR-29b potentiates the anti-MM activity of bortezomib
To evaluate if Sp-1/miR-29b loop has a role in controlling the apoptotic process in MM cells, we investigated the contribution of miR-29b in the response of MM cells to bortezomib. Indeed, enforced expression of miR-29b strongly increased bortezomib-induced growth inhibition (Figure 7a) and apoptosis (Figures 7b and c) in SKMM1 cells. This effect was not limited to SKMM1 cells, as miR-29b mimics significantly enhanced bortezomib effects in NCI-H929 (Figure 7d–f). As a further proof that miR-29b potentiates bortezomib sensitivity in MM cells, we antagonized miR-29b in SKMM1 cells by stably expressing antagomiR-29b through lentivirus transduction: as expected, antagomiR-29b-expressing cells showed a proliferative advantage and were less susceptible to bortezomib-induced growth inhibition (Figure 7g) and apoptosis (Figure 7h) as compared with controls. These findings strongly suggest an exciting translational potential and open a novel scenario for miR-29b mimics in the treatment of MM.
miR-29b-enforced expression enhances bortezomib effects on MM cells. (a) SKMM1 cells were transfected with synthetic mir-29b or scrambled oligonucleotides (NC) and then treated with 5 nM bortezomib (indicated as bort) or vehicle for 48 h. Viable cells were determined by trypan blue exclusion assay and data are expressed as percentage of viable cells (trypan blue negative) in synthetic miR-29b transfected with respect to NC-transfected cells, in three independent experiments performed in triplicate. (b) Apoptosis analysis of SKMM1 cells transfected with synthetic mirR-29b or NC and then treated with vehicle or 5 nM bortezomib for 48 h. Results show the percentage of Annexin-V-positive or Annexin V/7AAD-positive cells (cells in early and late apoptosis, respectively). *P<0.05. (c) Caspase 3/7 activity assay was performed in SKMM1 cells transfected with synthetic mir-29b or NC and then treated with vehicle or 5 nM bortezomib for 48 h. Transfected cells were grown in 96-well microplates and luminescence was measured using the Caspase 3/7 Glo assay (Promega). Results are expressed as fold increase of caspase activity with respect to NC-transfected cells, of at least three independent experiments. (d) NCI-H929 cells were transfected with synthetic mir-29b or NC and then treated with 5 nM bortezomib or vehicle for 24 h. Viable cells were determined by trypan blue exclusion assay and data are represented as percentage of viable cells (trypan blue negative) in synthetic mir-29b transfected with respect to NC-transfected cells in three independent experiments performed in triplicate. (e) Caspase 3–7 activity assay performed in NCI-H929 similarly to SKMM1 cells. (f) NCI-H929 cells transfected with synthetic miR-29b or NC and analyzed for early and late apoptosis by Annexin V/7AAD staining. *P<0.05. (g) SKMM1 cells were transduced with the empty vector or the antagomiR-29b and then treated with 5 nM bortezomib for 48 h. Viable cells were determined by trypan blue exclusion assay and data are expressed as percentage of viable cells (trypan blue negative) in antagomiR-29b with respect to empty vector transduced cells in three independent experiments performed in triplicate. (h) Caspase 3/7 activity assay performed on SKMM1 cells transduced with the empty vector or antagomiR-29b and treated with 5 nM bortezomib for 48 h
PI3K/AKT regulates miR-29b-Sp1 loop in MM cells
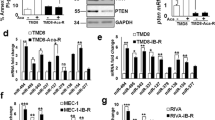
Finally, we explored whether the PI3K/AKT pathway, which plays a pivotal role in the regulation of MM cell proliferation, survival and drug resistance,46, 47 could regulate miR-29b-Sp1 loop. We found that overexpression of a dominant active AKT construct in NCI-H929 cells dramatically reduced miR-29b levels and concomitantly led to Sp1 upregulation (Figure 8a; transfection procedures are reported in Supplementary materials and methods). Moreover, as a further proof of the negative regulation exerted by PI3K/AKT on miR-29b expression, inhibition of PI3K by LY294002 increased miR-29b and reduced Sp1 protein levels. Functionally, overexpression of AKT was able to suppress miR-29b-dependent increase of caspase 3/7 activity (Figure 8c), whereas treatment with LY294002 potentiated miR-29b-dependent apoptosis (Figure 8d). Importantly, miR-29b was demonstrated to be a negative regulator of PI3K/AKT pathway, since transfection of synthetic miR-29b mimics resulted in reduced AKT phosphorylation at S473 together with reduced phosphorylation of the AKT substrate glycogen synthase kinase-3β (GSK-3β) at S9 and upregulation of PTEN protein levels. Moreover, we provided preliminary evidence that miR-29b mimics transfection represses signals mediated by mTOR, as assessed by reduced phosphorylation of 4E-BP1 at S65 (Figure 8e). Altogether, these results delineate a novel druggable microcircuitry in MM cells (an explanatory cartoon is included in Supplementary Figure 7).
Reciprocal regulation between AKT activity and miR-29b expression in MM cells. (a) NCI-H929 cells were electroporated with the pcDNA.3.1-HA-myr-AKT-dominant active construct or the empty vector pcDNA3.1 (indicated as vector; transfection procedures are reported in Supplementary materials and methods) and 48 h later analyzed for miR-29b expression levels by quantitative RT–PCR. Normalization was performed using RNU44 snoRNA and ΔΔCt calculations. Immunoblots show protein levels of transfected AKT (HA tag) and Sp1 in transfected cells; GAPDH was used as loading control. *P<0.01. (b) Quantitative RT–PCR of miR-29b in NCI-H929 cells treated with 20 μM LY294002 or vehicle (DMSO) for 48 h. Immunoblot shows Sp1 and phosphorylated AKT at S473 protein levels in LY294002-treated cells. GAPDH was used as loading control. •P<0.05. (c) Caspase 3/7 activity assay was performed in NCI-H929 cells transfected with synthetic mir-29b or NC together with pcDNA.3.1-HA-myr-AKT dominant active construct or the same amount of the empty vector pcDNA3.1 (indicated as vector). Transfected cells were grown in 96-well microplates and luminescence was measured after 48 h using the Caspase 3/7 Glo assay (Promega). *P<0.01. Results are expressed as fold increase of caspase activity with respect to NC-transfected cells, of at least three independent experiments. (d) Caspase 3/7 activity assay was performed in NCI-H929 cells transfected with synthetic mir-29b or NC and then treated with 20 μM LY294002 or vehicle for 48 h. (e) (Left panel) Immunoblot analysis of AKT, phosphorylated AKT at S473, phosphorylated GSK-3β at S9, GSK-3β, 4E-BP1 phosphorylated at S65 and PTEN in NCI-H929 cells, 24 h after transfection with synthetic mir-29b or NC; GAPDH was used as loading control. (Right panel) AKT activity was determined as reported in Supplementary material and methods, 24 h after transfection of NCI-H929 cells with synthetic miR-29b or NC; α-tubulin was used as loading control
Discussion
In this report, we demonstrate that the enforced expression of miR-29b in MM cell lines inhibits cell growth and triggers apoptosis in vitro. Importantly, the in vivo ability of miR-29b to induce apoptosis was also demonstrated in a murine model of human MM. Our experimental platform was based on MM xenografts bearing cells with constitutive expression of miR-29b after lentiviral transduction. Our findings represent a clear evidence that miR-29b may efficiently prevent the tumor formation in a preclinical in vivo model of MM.
To assess the mechanisms of apoptosis induction by miR-29b, we studied the effects of synthetic miR-29b mimics on the expression of the transcription factor Sp1. In fact, miR-29b, as other miRNAs, is able to downregulate genes involved in proliferation and inhibition of apoptosis, and this property can explain the growth-inhibitory and pro-apoptotic effects triggered by miR-29b in MM cells. Moreover, we recently showed that increased Sp1 activity promotes MM cell growth and survival and its inhibition has therapeutic value in preclinical models of MM.40 Indeed, here we found that miR-29b negatively regulates Sp1 expression in MM cells. The 3′UTR of Sp1 contains four putative miR-29b-binding sites as predicted by Pictar48 and two revealed by TargetScan 4.2 softwares.49 We validated miR-29b regulatory activity by luciferase reporter assay as well as by Sp1 mRNA and protein expression analysis, confirming that miR-29b is able to downregulate Sp1.
Recent studies have also characterized the mechanisms involved in miR-29b regulation. Specifically, several cis elements in the proximal region of miR-29a/b1 promoter have been identified that would account, at least in part, for the transcriptional regulation.37 Consistently with previous findings showing Sp1-binding sites on miR-29b promoter,44 we found that Sp1, which is a miR-29b target, negatively regulates miR-29b: in fact, Sp1 overexpression leads to miR-29b transcriptional inhibition, whereas Sp1 silencing by shRNAs or pharmacologic inhibition by mithramycin-A increase miR-29b levels. Therefore, our results demonstrate that a miR-29b-Sp1 feedback regulatory loop indeed occurs in MM cells, representing a target amenable of therapeutic intervention.
It is well known that Sp1 levels are under proteasome control, and proteasome inhibitors have been proven to decrease Sp1 levels in leukemic cells.44, 50 We have found that inhibition of cell proliferation and induction of apoptosis by the proteasome inhibitor bortezomib, at concentrations that are achievable in MM patients in vivo,51 are associated with Sp1 downregulation and miR-29b upregulation (Figure 6). Importantly, peripheral blood primary CD138+ cells collected from a PCL patient 24 h after bortezomib treatment showed upregulation of miR-29b similar to that observed in MM cell lines. This in vivo observation on PCL is currently under extended analysis in a large cohort of patients. Noteworthy, the clinical efficacy of the proteasome inhibitor bortezomib has been shown in the treatment of patients with refractory and relapsed MM and other tumors.52 An intriguing finding of the present work was that bortezomib regulates the anti-MM effects of miR-29b: in fact, bortezomib increased synthetic miR-29b mimics-induced cytotoxicity on MM cells, whereas miR-29b inhibition dampened bortezomib-induced apoptosis. These results underscore the relevance of miR-29b, and likely of its downstream targets, to apoptosis induction by bortezomib, providing novel information on the unsolved issue of unraveled bortezomib targets.53 Our results could therefore contribute to the understanding of the intracellular functions of miR-29b and of the biochemical perturbations underlying the apoptotic stimuli induced by miR-29b, implementing previous observations on the in vitro anti-myeloma activity of this miRNA.32
Notably, our findings highlight a major role for PI3K/AKT pathway in regulating miR-29b-Sp1 loop. In fact AKT overexpression led to miR-29b downregulation and Sp1 upregulation, whereas pharmacologic inhibition of PI3K by LY294002 had the opposite effect. Conversely, miR-29b behaves as a negative regulator of PI3K/AKT pathway by reducing AKT phosphorylation and its kinase activity on downstream AKT substrate GSK-3β. At the same time, transfection of a constitutively active AKT suppresses caspase 3/7 activity induced by miR-29b. All together, these findings demonstrate that PI3K/AKT is a negative regulator of miR-29b whose proapototic activity involves functional downregulation of AKT. These results highlight a novel exciting scenario and provide new formal evidence for a major role of PI3K/AKT in down-modulating miR-29b activity. Our findings of a preminent role of the survival PI3K/AKT pathway in antagonizing miR-29b activity complement its function of apoptosis inducer also by MCL-1 downregulation, providing a wide scenario of MM apoptosis derangement by miR-29b downregulation in MM.
In conclusions, our investigation demonstrates a relevant in vitro and in vivo anti-MM apoptotic activity of miR-29b and adds novel insights into pathways regulating miRNAs-induced apoptosis, providing the biological rationale for the use of synthetic miR-29b mimics, alone or in combination with bortezomib, as novel therapeutic strategy in MM.
Materials and Methods
Cell lines, primary cells, and drugs
Peripheral blood mononuclear cells (PBMCs), BM mononuclear cells (BMMNCs), and primary MM cells from MM patient BM aspirates, following informed consent and University Magna Graecia IRB approval, were isolated using Ficoll-Hypaque density gradient sedimentation. MM patient cells were separated from BM samples by antibody-mediated positive selection using anti-CD138 magnetic activated cell separation microbeads (Miltenyi Biotech, Gladbach, Germany).
MM cell lines were cultured in RPMI-1640 (Gibco, Life Technologies, Carlsbad, CA, USA) supplemented with 10% fetal bovine serum (Lonza Group, Basel, Switzerland). DOX-6 and LR-5 cell lines were kindly provided by Dr. Eduard Thomson (University of Texas Medical Branch, Galveston, TX, USA); the IL-6-dependent MM cell Line INA-6 (kindly provided from Dr. Renate Burger, University of Erlangen-Nuernberg, Erlangen, Germany) was cultured in the presence of recombinant-human IL-6 as previously described.54 BMSCs were established as previously described.55 Methylcellulose colony assays were performed in RPMI-1640 medium containing 1.1% methyl-cellulose (STEMCELL Technologies Inc., Vancouver, Canada) and 20% FBS. Colony formation was scored at day 14 after plating 1000 cells (SKMM1) or 500 cells (NCI-H929) in 24-well plates containing 1 ml of methylcellulose medium. Mithramycin-A was purchased from Sigma-Aldrich (St Louis, MO, USA) and dissolved in DMSO. Clinical grade bortezomib was purchased from Millennium Pharmaceuticals (Takeda, Cambridge, MA, USA) and dissolved in sodium chloride saline solution at 1 mg/ml. Caspase 8 inhibitor Z-IETD-FMK and PI3K inhibitor LY294002 were purchased from Santa Cruz Biotechnology (Santa Cruz, CA, USA) and Cell Signaling (Danvers, MA, USA), respectively.
miRNA expression profiling of CD138+ cells from MM patients
Whole genome miRNAs profiling of purified (≥90%) PCS from BM aspirates of 40 untreated patients (38 MMs and 2 PCLs) was reported in our previous work.56 Patients have been stratified according to the presence of the recurrent IGH chromosomal translocations and cyclins D expression (TC classification; see Supplementary material and methods for details). The microarray consist of 60-mer DNA probes synthesized in situ, which represent 723 human and 76 human viral miRNAs from the Sanger database v 10.1. (Wellcome Trust Sanger Institute, Hinxton, Cambridge, UK). miRNA data are available at GEO accession number GSE17498.
Virus generation and infection of MM cells
The human Sp1 (NM_138473) mission shRNA set (Sigma) and the mission nontarget control transduction virus (SHC002V; Sigma) were used to generate lentiviral particles in HEK 293T packaging cells as previously described;57 two rounds of transduction of MM cells in the presence of 8 μg/ml of polybrene (Sigma) were performed. Two days after transduction, transduced cells were selected with 1 μg/ml Puromycin (Sigma).
To generate Sp1-encoding retrovirus, HEK 293T cells were co-transfected with 10 μg of pBABEpuro-Sp1 (kindly provided by Dr. Phyllis Dennery, University of Pennsylvania), 10 μg of pCMV-VSVG, and 4 μg of PEQ-PAM and collected and used to transduce MM cells. Transduced cells underwent three rounds of infection (8 h each round) and were selected in medium containing 1 μg/ml puromycin.
To obtain cells stably expressing miR-29b, we used a feline immunodeficiency lentivirus-based construct (System Biosciences, Mountain View, CA, USA), consisting of the stem loop structure of miR-29b and 200 bp of upstream and downstream flanking genomic sequence cloned into the pMIF-cGFPZeo-miR plasmid. Packaging of the miR-29b constructs in pseudoviral particles was performed in 293Ta cells using the Lenti-Pac FIV Expression Packaging Kit (FPK-LvTR-20), according to the manufacturer’s instructions (Genecopoeia, Rockville, MD, USA). After transfection of 293Ta cells, supernatants containing miR-29b lentivirus were collected at 8-h intervals, filtered, and used for two rounds of transduction of SKMM1, U266 and RPMI-8226 cells (1 × 106) in the presence of 8 μg/ml of polybrene (Sigma-Aldrich). Two days after transduction, selection with 200 μg/ml Zeocin for 3 days was performed to achieve almost 100% transduced cells. Empty lentivirus was used as a control for the experiments.
To obtain MM cells stably expressing antagomiR-29b, we used the lentiviral vector miRZip-29b anti-miR-29b construct (System Biosciences); lentiviral particles production and transduction were performed according to the above indicated protocols. Selection was performed in medium containing 1 μg/ml puromycin.
Quantitative real-time amplification of miRNAs and mRNAs
Total RNA from MM cells was prepared with the TRIzol Reagent (Invitrogen, Life Technologies, Carlsbad, CA, USA) according to the manufacturer’s instructions. The single-tube TaqMan miRNA assays (Assay Id 000413, Applied Biosystems, Life Technologies) was used to detect and quantify mature miR-29b according to the manufacturer’s instructions, by the use of iQ5 multicolor detection system (Bio-Rad, Berkeley, CA, USA). miR-29b expression was normalized on RNU44 (Applied Biosystems, Assay Id 001094).
For mRNA dosage studies, oligo-dT-primed cDNA was obtained using the High Capacity cDNA Reverse Transcription Kit (Applied Biosystems) and then used as template to quantify CDK6 (Hs01026371_m1), MCL-1 (Hs01050896_m1) and Sp1 (Hs00412720_m1) levels by TaqMan assay (Applied Biosystems); normalization was performed with GAPDH (Hs03929097_g1). Comparative real-time polymerase chain reaction (RT–PCR) was performed in triplicate, including no-template controls. Relative expression was calculated using the comparative cross threshold (Ct) method.58
Luciferase reporter experiments
The pGL3 Luciferase plasmid containing the miR-29a/b1 promoter (nts –1530 +165) was a kind gift of Dr. J Mott (University of Nebraska Medical Center). Transfections were performed by electroporation and renilla luciferase from pCMV-RL was included to normalize expression of firefly luciferase. The 3′UTR segments containing the target sites for miR-29b from the Sp1 gene corresponding to the sites of interaction with miR-29b, were cloned in pEZX-MT01 vector and purchased from Genecopoeia (product Id HmiT017640b-MT01). MM cells were electroporated as above described using 5 μg of the firefly luciferase reporter; for each plate, 100 nM of the synthetic miR-29b or miR-NC were used. Firefly and Renilla luciferase activities were measured consecutively using the dual-luciferase assay kit (Promega Corporation, Madison, WI, USA) 24 and 48 h after transfection. Data are expressed as luminescence from firefly luciferase divided by luminescence from renilla luciferase.
Animals and in vivo model of human MM
Male CB-17 severe combined immunodeficient (SCID) mice (6- to 8-weeks old; Harlan Laboratories, Inc., Indianapolis, IN, USA) were housed and monitored in our Animal Research Facility. All experimental procedures and protocols had been approved by the Institutional Ethical Committee (Magna Graecia University) and conducted according to the protocols approved by the National Directorate of Veterinary Services (Italy). In accordance with the institutional guidelines, mice were killed when their tumors reached 2 cm in diameter or in the event of paralysis or major compromise in their quality of life, to prevent unnecessary suffering. For our study, 3 × 106 NCI-H929 cells stably expressing miR-29b by lentivirus-based construct, the pMIF-miR-29b plasmid, or transduced with empty vector (pMIF) as control, were injected sc into the flank of 10 SCID mice. Tumor sizes were measured weekly in two dimensions using a caliper, and volume was calculated using the formula: V=0.5 × a × b2, where a and b are the long and short diameter of the tumor, respectively, until the tumor was palpable.54 In this model, tumors were palpable approximately 2 weeks after injection.
In vivo detection of the tumor mass in xenografted mice was performed using IVIS LUMINA II Imaging System (Caliper Life sciences, Hopkinton, MA, USA). Then, tumor sizes were measured every 2 days until the day of first mouse kill. Tumors were retrieved from animals and placed in 10% formalin for histology or in RNAlater (Ambion, Carlsbad, CA, USA) for RNA isolation.
Histology and immunohistochemistry
Retrieved tumors from animals were immediately fixed in 4% buffered formaldehyde for 24 h at 4 °C, washed, dehydrated, and embedded in paraffin. For the light microscopy analysis, sections were cut (4 μm), mounted on poly-lysine slides, and stained with H&E. For immunohistochemistry stainings, 2-μm thick sections were in a Bond Max Automated Immunohistochemistry according to the following protocol. First, tissues were deparaffinized and pretreated with the Epitope Retrieval Solution 2 (EDTA-buffer pH8.8) at 98 °C for 20 min After washing steps, peroxidase blocking was carried out for 10 min using the Bond Polymer Refine. Tissues were again washed and then incubated with the primary antibody directed against Ki-67 (Dako, Milano, Italy, clone: MIB-1; 1 : 150) or capase-3 (Novocastra, Newcastle upon Tyne, UK, clone: JHM62; 1 : 500). Subsequently, tissues were incubated with polymer for 10 min and developed with DAB-Chromogen for 10 min Slides were counterstained with hematoxylin.
Statistical analysis
Student’s t-test, two-tailed and log-rank test were used to evaluate statistical significance by GraphPad software (GraphPad Software, Inc., La Jolla, CA, USA, www.graphpad.com). Graphs were obtained using SigmaPlot version 11.0 (Systat Software, Inc., San Jose, CA, USA).
Accession codes
Abbreviations
- 4E-BP1:
-
eukaryotic initiation factor 4E-binding protein 1
- 7AAD:
-
7-aminoactinomycin D
- BM:
-
bone marrow
- BMMNCs:
-
bone marrow mononuclear cells
- BMSCs:
-
bone marrow stromal cells
- CDK6:
-
cyclin-dependent kinase 6
- GSK:
-
glycogen synthase kinase
- H&E:
-
hematoxylin and eosin
- huBMM:
-
human bone marrow microenvironment
- IL-6:
-
interleukin-6
- MCL-1:
-
myeloid cell leukemia sequence 1
- MM:
-
multiple myeloma
- MGUS:
-
monoclonal gammopaty of undetermined significance
- mTOR:
-
mammalian target of rapamycin
- PBMNCs:
-
peripheral blood mononuclear cells
- PCs:
-
plasma cells
- PCL:
-
plasma cell leukemia
- PI3K:
-
phosphotidyl inositol 3-kinase
- PTEN:
-
phosphatase and tensin homolog
- REL:
-
relative expression level
- Rb:
-
retinoblastoma protein
- SC:
-
subcutaneous
- SD:
-
standard deviation
- Sp1:
-
specificity protein 1
- UTR:
-
untranslated region
References
Pattanayak D, Agarwal S, Sumathi S, Chakrabarti SK, Naik PS, Khurana SM . Small but mighty RNA-mediated interference in plants. Indian J Exp Biol 2005; 43: 7–24.
Hutvagner G, Small RNA . asymmetry in RNAi: function in RISC assembly and gene regulation. FEBS Lett 2005; 579: 5850–5857.
Lim LP, Lau NC, Garrett-Engele P, Grimson A, Schelter JM, Castle J et al. Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 2005; 433: 769–773.
Miranda KC, Huynh T, Tay Y, Ang YS, Tam WL, Thomson AM et al. A pattern-based method for the identification of MicroRNA binding sites and their corresponding heteroduplexes. Cell 2006; 126: 1203–1217.
Miska EA . How microRNAs control cell division, differentiation and death. Curr Opin Genet Dev 2005; 15: 563–568.
Fabbri M, Croce CM, Calin GA . MicroRNAs in the ontogeny of leukemias and lymphomas. Leuk Lymphoma 2009; 50: 160–170.
Visone R, Croce CM . MiRNAs and cancer. Am J Pathol 2009; 174: 1131–1138.
Calin GA, Croce CM . MicroRNA signatures in human cancers. Nat Rev Cancer 2006; 6: 857–866.
Krutzfeldt J, Rajewsky N, Braich R, Rajeev KG, Tuschl T, Manoharan M et al. Silencing of microRNAs in vivo with ‘antagomirs’. Nature 2005; 438: 685–689.
Czech MP . MicroRNAs as therapeutic targets. N Engl J Med 2006; 354: 1194–1195.
Croce CM . Causes and consequences of microRNA dysregulation in cancer. Nat Rev Genet 2009; 10: 704–714.
Esquela-Kerscher A, Slack FJ . Oncomirs—microRNAs with a role in cancer. Nat Rev Cancer 2006; 6: 259–269.
Wiggins JF, Ruffino L, Kelnar K, Omotola M, Patrawala L, Brown D et al. Development of a lung cancer therapeutic based on the tumor suppressor microRNA-34. Cancer Res 2010; 70: 5923–5930.
Anderson KC, Alsina M, Bensinger W, Biermann JS, Chanan-Khan A, Cohen AD et al. Multiple myeloma. J Natl Compr Canc Netw 2011; 9: 1146–1183.
Anderson KC, Carrasco RD . Pathogenesis of myeloma. Annu Rev Pathol 2011; 6: 249–274.
Rossi M, Di Martino MT, Morelli E, Leotta M, Rizzo A, Grimaldi A et al. Molecular targets for the treatment of multiple myeloma. Curr Cancer Drug Targets 2012; 12: 757–767.
Hideshima T, Anderson KC . Molecular mechanisms of novel therapeutic approaches for multiple myeloma. Nat Rev Cancer 2002; 2: 927–937.
Barlogie B, Zangari M, Spencer T, Fassas A, Anaissie E, Badros A et al. Thalidomide in the management of multiple myeloma. Semin Hematol 2001; 38: 250–259.
Richardson PG, Sonneveld P, Schuster MW, Irwin D, Stadtmauer EA, Facon T et al. Bortezomib or high-dose dexamethasone for relapsed multiple myeloma. N Engl J Med 2005; 352: 2487–2498.
Barlogie B . Thalidomide and CC-5013 in multiple myeloma: the University of Arkansas experience. Semin Hematol 2003; 40: 33–38.
Jemal A, Tiwari RC, Murray T, Ghafoor A, Samuels A, Ward E et al. Cancer statistics 2004 CA Cancer J Clin 2004; 54: 8–29.
Fulciniti M, Hideshima T, Vermot-Desroches C, Pozzi S, Nanjappa P, Shen Z et al. A high-affinity fully human anti-IL-6 mAb, 1339, for the treatment of multiple myeloma. Clin Cancer Res 2009; 15: 7144–7152.
Fulciniti M, Tassone P, Hideshima T, Vallet S, Nanjappa P, Ettenberg SA et al. Anti-DKK1 mAb (BHQ880) as a potential therapeutic agent for multiple myeloma. Blood 2009; 114: 371–379.
Shammas MA, Koley H, Bertheau RC, Neri P, Fulciniti M, Tassone P et al. Telomerase inhibitor GRN163L inhibits myeloma cell growth in vitro and in vivo. Leukemia 2008; 22: 1410–1418.
Tassone P, Neri P, Burger R, Savino R, Shammas M, Catley L et al. Combination therapy with interleukin-6 receptor superantagonist Sant7 and dexamethasone induces antitumor effects in a novel SCID-hu in vivo model of human multiple myeloma. Clin Cancer Res 2005; 11: 4251–4258.
Di Martino MT, Leone E, Amodio N, Foresta U, Lionetti M, Pitari MR et al. Synthetic miR-34a mimics as a novel therapeutic agent for Multiple Myeloma: in vitro and in vivo evidence. Clin Cancer Res 2012; 18: 6260–6270.
Lionetti M, Agnelli L, Lombardi L, Tassone P, Neri A . MicroRNAs in the pathobiology of multiple myeloma. Curr Cancer Drug Targets 2012; 12: 823–837.
Tagliaferri P, Rossi M, Di Martino MT, Amodio N, Leone E, Gulla A et al. Promises and challenges of MicroRNA-based treatment of multiple myeloma. Curr Cancer Drug Targets 2012; 12: 838–846.
Le Beau MM, Espinosa R, Davis EM, Eisenbart JD, Larson RA, Green ED . Cytogenetic and molecular delineation of a region of chromosome 7 commonly deleted in malignant myeloid diseases. Blood 1996; 88: 1930–1935.
Pekarsky Y, Santanam U, Cimmino A, Palamarchuk A, Efanov A, Maximov V et al. Tcl1 expression in chronic lymphocytic leukemia is regulated by miR-29 and miR-181. Cancer Res 2006; 66: 11590–11593.
Mott JL, Kobayashi S, Bronk SF, Gores GJ . mir-29 regulates Mcl-1 protein expression and apoptosis. Oncogene 2007; 26: 6133–6140.
Zhang YK, Wang H, Leng Y, Li ZL, Yang YF, Xiao FJ et al. Overexpression of microRNA-29b induces apoptosis of multiple myeloma cells through down regulating Mcl-1. Biochem Biophys Res Commun 2011; 414: 233–239.
Park SY, Lee JH, Ha M, Nam JW, Kim VN . miR-29 miRNAs activate p53 by targeting p85 alpha and CDC42. Nat Struct Mol Biol 2009; 16: 23–29.
Fabbri M, Garzon R, Cimmino A, Liu Z, Zanesi N, Callegari E et al. MicroRNA-29 family reverts aberrant methylation in lung cancer by targeting DNA methyltransferases 3A and 3B. Proc Natl Acad Sci USA 2007; 104: 15805–15810.
Sengupta S, den Boon JA, Chen IH, Newton MA, Stanhope SA, Cheng YJ et al. MicroRNA 29c is down-regulated in nasopharyngeal carcinomas, up-regulating mRNAs encoding extracellular matrix proteins. Proc Natl Acad Sci USA 2008; 105: 5874–5878.
Garzon R, Heaphy CE, Havelange V, Fabbri M, Volinia S, Tsao T et al. MicroRNA 29b functions in acute myeloid leukemia. Blood 2009; 114: 5331–5341.
Mott JL, Kurita S, Cazanave SC, Bronk SF, Werneburg NW, Fernandez-Zapico ME . Transcriptional suppression of mir-29b-1/mir-29a promoter by c-Myc, hedgehog, and NF-kappaB. J Cell Biochem 2010; 110: 1155–1164.
Wang H, Garzon R, Sun H, Ladner KJ, Singh R, Dahlman J et al. NF-kappaB-YY1-miR-29 regulatory circuitry in skeletal myogenesis and rhabdomyosarcoma. Cancer Cell 2008; 14: 369–381.
Zheng L, Lee WH . The retinoblastoma gene: a prototypic and multifunctional tumor suppressor. Exp Cell Res 2001; 264: 2–18.
Fulciniti M, Amin S, Nanjappa P, Rodig S, Prabhala R, Li C et al. Significant biological role of sp1 transactivation in multiple myeloma. Clin Cancer Res 2011; 17: 6500–6509.
Tsang J, Zhu J, van Oudenaarden A . MicroRNA-mediated feedback and feedforward loops are recurrent network motifs in mammals. Mol Cell 2007; 26: 753–767.
Ray R, Snyder RC, Thomas S, Koller CA, Miller DM . Mithramycin blocks protein binding and function of the SV40 early promoter. J Clin Invest 1989; 83: 2003–2007.
Karin M, Yamamoto Y, Wang QM . The IKK NF-kappa B system: a treasure trove for drug development. Nat Rev Drug Discov 2004; 3: 17–26.
Liu S, Wu LC, Pang J, Santhanam R, Schwind S, Wu YZ et al. Sp1/NFkappaB/HDAC/miR-29b regulatory network in KIT-driven myeloid leukemia. Cancer Cell 2010; 17: 333–347.
Kikuchi J, Wada T, Shimizu R, Izumi T, Akutsu M, Mitsunaga K et al. Histone deacetylases are critical targets of bortezomib-induced cytotoxicity in multiple myeloma. Blood 2010; 116: 406–417.
Hideshima T, Chauhan D, Richardson P, Anderson KC . Identification and validation of novel therapeutic targets for multiple myeloma. J Clin Oncol 2005; 23: 6345–6350.
Hsu J, Shi Y, Krajewski S, Renner S, Fisher M, Reed JC et al. The AKT kinase is activated in multiple myeloma tumor cells. Blood 2001; 98: 2853–2855.
Krek A, Grun D, Poy MN, Wolf R, Rosenberg L, Epstein EJ et al. Combinatorial microRNA target predictions. Nat Genet 2005; 37: 495–500.
Lewis BP, Shih IH, Jones-Rhoades MW, Bartel DP, Burge CB . Prediction of mammalian microRNA targets. Cell 2003; 115: 787–798.
Liu S, Liu Z, Xie Z, Pang J, Yu J, Lehmann E et al. Bortezomib induces DNA hypomethylation and silenced gene transcription by interfering with Sp1/NF-kappaB-dependent DNA methyltransferase activity in acute myeloid leukemia. Blood 2008; 111: 2364–2373.
Quinn DI, Nemunaitis J, Fuloria J, Britten CD, Gabrail N, Yee L et al. Effect of the cytochrome P450 2C19 inhibitor omeprazole on the pharmacokinetics and safety profile of bortezomib in patients with advanced solid tumours, non-Hodgkin's lymphoma or multiple myeloma. Clin Pharmacokinet 2009; 48: 199–209.
Molineaux SM . Molecular pathways: targeting proteasomal protein degradation in cancer. Clin Cancer Res 2012; 18: 15–20.
Hideshima T, Ikeda H, Chauhan D, Okawa Y, Raje N, Podar K et al. Bortezomib induces canonical nuclear factor-kappaB activation in multiple myeloma cells. Blood 2009; 114: 1046–1052.
Tassone P, Di Martino MT, Ventura M, Pietragalla A, Cucinotto I, Calimeri T et al. Loss of BRCA1 function increases the antitumor activity of cisplatin against human breast cancer xenografts in vivo. Cancer Biol Ther 2009; 8: 648–653.
Blotta S, Jakubikova J, Calimeri T, Roccaro AM, Amodio N, Azab AK et al. Canonical and non canonical Hedgehog pathway in the pathogenesis of multiple myeloma. Blood 2012 ; [e-pub ahead of print].
Lionetti M, Agnelli L, Mosca L, Fabris S, Andronache A, Todoerti K et al. Integrative high-resolution microarray analysis of human myeloma cell lines reveals deregulated miRNA expression associated with allelic imbalances and gene expression profiles. Genes Chromosomes Cancer 2009; 48: 521–531.
Amodio N, Scrima M, Palaia L, Salman AN, Quintiero A, Franco R et al. Oncogenic role of the E3 ubiquitin ligase NEDD4-1, a PTEN negative regulator, in non-small-cell lung carcinomas. Am J Pathol 2010; 177: 2622–2634.
Livak KJ, Schmittgen TD . Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 2001; 25: 402–408.
Acknowledgements
This work was supported by funds of Italian Association for Cancer Research (AIRC), PI: PT. ‘Special Program Molecular Clinical Oncology—5 per mille’ n. 9980, 2010/15.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by P Salomoni
Supplementary Information accompanies the paper on Cell Death and Disease website
Supplementary information
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Amodio, N., Di Martino, M., Foresta, U. et al. miR-29b sensitizes multiple myeloma cells to bortezomib-induced apoptosis through the activation of a feedback loop with the transcription factor Sp1. Cell Death Dis 3, e436 (2012). https://doi.org/10.1038/cddis.2012.175
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2012.175
Keywords
This article is cited by
-
miRNA-seq and clinical evaluation in multiple myeloma: miR-181a overexpression predicts short-term disease progression and poor post-treatment outcome
British Journal of Cancer (2022)
-
Elevation of miR-302b prevents multiple myeloma cell growth and bone destruction by blocking DKK1 secretion
Cancer Cell International (2021)
-
Roles of miRNA dysregulation in the pathogenesis of multiple myeloma
Cancer Gene Therapy (2021)
-
Ectopic expression of BIRC5-targeting miR-101-3p overcomes bone marrow stroma-mediated drug resistance in multiple myeloma cells
BMC Cancer (2019)
-
miR-22 suppresses DNA ligase III addiction in multiple myeloma
Leukemia (2019)