Abstract
Huntington's disease (HD) is a complex and severe disorder characterized by the gradual and the progressive loss of neurons, predominantly in the striatum, which leads to the typical motor and cognitive impairments associated with this pathology. HD is caused by a highly polymorphic CAG trinucleotide repeat expansion in the exon-1 of the gene encoding for huntingtin protein. Since the first discovery of the huntingtin gene, investigations with a consistent number of in-vitro and in-vivo models have provided insights into the toxic events related to the expression of the mutant protein. In this review, we will summarize the progress made in characterizing the signaling pathways that contribute to neuronal degeneration in HD. We will highlight the age-dependent loss of proteostasis that is primarily responsible for the formation of aggregates observed in HD patients. The most promising molecular targets for the development of pharmacological interventions will also be discussed.
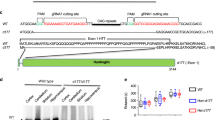
Similar content being viewed by others
Main
Huntington's disease (HD) is an inherited autosomal dominant neurodegenerative disorder characterized by adult-onset of motor dysfunctions, psychiatric disturbances and intellectual decline.1 As revealed by postmortem analysis of tissues from HD patients, the neuropathological changes are predominantly detected in the striatum, although marked alterations have also been observed in other areas of the brain, including the cerebellar cortex, thalamus and cerebellum.2, 3 HD is associated with an unstable CAG expansion in the huntingtin gene (HTT) on chromosome 4. In humans, the exon-1 of HTT gene normally contains between 6 and 35 CAG repeats, whereas in patients affected by HD more than 40 trinucleotides have been described.4 In most cases, an intermediate number (36–40) of CAG repeats leads to a slower progression of the pathology as a result of the incomplete penetrance of the mutant allele. Importantly, the onset and severity of the pathology is directly correlated with the number of CAG repeats, although the actual function of the trinucleotide stretch remains unknown.5, 6 As reported by recent findings, the length of the CAG repeats might be relevant in the translation of the HTT mRNA transcript, as a result of binding with a ribosome-containing complex7 (Krauss S., unpublished data). The HTT gene encodes for an approximately 350 kDa protein composed of several subdomains. At the N-terminus, the polyglutamine (polyQ) stretch encoded by the CAG repeats functions as potential membrane association signal.8 In mammals, the polyQ-containing domain is followed by a polyproline sequence that stabilizes protein conformation. The N-terminal portion of HTT is followed by three main clusters of HEAT repeats, which are essential for the binding with interacting proteins. In addition to these motifs, HTT contains a range of consensus sites for posttranslational modifications, including proteolytic cleavage, phosphorylation and sumoylation. Within cells, HTT has been detected in the nucleus, mitochondria, Golgi and endoplasmic reticulum and can be found in the neuronal body, dendrites and synapses.9, 10 At the molecular level, there is evidence that HTT can interact with a variety of proteins, including some transcriptional factors, synaptic complexes, plasma membrane and cytoskeleton proteins.11 HTT is ubiquitously expressed during embryonic development and at high levels in testis and in mature postmitotic neurons in adult human brain.12
Although the physiological role of HTT has not been fully defined, analysis of transgenic mice with a targeted deletion of the Htt gene has demonstrated its role in mammalian development. Complete suppression of Htt expression in mice leads to embryonic lethality as a result of increased apoptosis,13, 14 while heterozygous knockout animals exhibit severe cognitive deficits as a consequence of increased neuronal loss in the subthalamic nucleus of the basal ganglia.13 Similarly, postnatal neuronal-specific inactivation of Htt is accompanied by progressive apoptotic neuronal degeneration,15 which suggests an essential function of the protein in the neuronal maintenance and activity. The antiapoptotic effect is likely due to the both inhibition of caspase-3 activity by its direct binding16 as well as to the activation of prosurvival pathways controlled by the serine/threonine kinase Akt.17 This pattern strongly supports the idea that HD pathogenesis results from a combination of increased gain-of-function of the mutant HTT together with the decreased wild-type HTT physiological function. This physiological function may be related to the N-terminal polyglutamine region, as it can form polar zipper structure able to bind transcription factors.18 Importantly, the physiological role of the polyQ-repeated expansion in higher organisms has been recently explored in mice carrying only seven CAG repeats within the murine Htt gene. These animals revealed subtle memory and learning deficits, with an altered energy status caused by changes in mitochondrial function.19 In a knock-in mouse model for HD, overexpression of the full-length Htt lacking the polyQ specifically stimulates the catabolic process of autophagy, significantly reduces mutant Htt-containing aggregates and, as a result, extends the lifespan in comparison with HD mice.20 Taken together, this evidence suggests the presence of an evolutionary positive selection favouring the expansion of the repetitive element as modulator of the protein activity itself.
HD is characterized by protein aggregates that accumulate within cells in a manner similar to that seen in various forms of spinocerebellar ataxia, as well as in other neurodegenerative disorders such as Alzheimer's disease (AD) and Parkinson's disease (PD). In human patients affected by HD, immunohistochemical analyses of postmortem brain tissue has demonstrated the presence of intracellular inclusions,21 which are mainly associated with the selective loss of medium spiny neurons in the striatum.22 These aggregates are enriched in truncated polyglutamine containing-fragments generated by several proteases, however the precise mechanisms responsible for the toxicity of these proteolytic products remain elusive.20, 23, 24, 25 Even though some mouse models expressing N-terminal truncated mutant HTT exhibit abnormal behavioral and neurological phenotypes,26 other transgenic lines present widespread intracellular inclusion formation without any functional neuronal deficits. For example, the ‘shortstop’ is a mouse line expressing the first two exons of HTT with an expanded CAG repeat. In these transgenic mice, there is no evident neurodegenerative phenotype, and neurons are less susceptible to excitotoxic cell death compared with other HD mouse models.27 Thus, as the full-length HTT is indispensable for manifestation of neuropathology clearly analogous to human HD,28, 29 the deposition of proteolytic products is not sufficient to initiate a toxic cycle leading to extensive neuronal damage in the striatum. In AD30, 31, 32 and in PD,33 (reviewed in refs Douglas and Dillin34; McCormack and DiMonte35) inclusions do trigger neurotoxicity. In HD, in a limited number of conditions, intracellular aggregates can also sequester toxic soluble fragment and therefore have beneficial effect.36 Nevertheless, the majority of evidences indicates that any mechanism promoting maintenance of the correct protein folding conformation or that enhances the clearance of huntingtin-containing aggregates represents a powerful therapeutic approach in HD.37, 38 In the next section, some of the key molecular mechanisms that influence proteostasis will be outlined and their relevance in the progression of HD will be discussed.
Proteolytic Cleavage of HTT
HTT is susceptible to proteolysis by a number of proteases (Figure 1). Historically, HTT was initially identified as a caspase substrate and it was the first example of a protein associated with a neurodegenerative disorder cleaved during apoptosis.39 Caspases are highly conserved cysteine-aspartic proteases associated primarily with apoptotic cell death and essential for the processing of a large number of substrates.40 Proteolytic fragments processed by caspases are detectable in brains of HD patients and HD mice before the loss of neurons in the striatum,41 with the cleavage efficiency dependent on the polyQ tract length.39 Blocking HTT cleavage by site-directed mutagenesis or by pharmacological approaches reduces cytotoxicity in cultured cells.42 In line with these findings, mice overexpressing a caspase-6 non-cleavable mutant HTT have milder neuropathological defects and are protected against excitotoxic stimulation compared with mice carrying the cleavable mutant HTT.43 This strongly suggests that caspase-dependent proteolytic cleavage of the aberrant protein might be a key step in the toxic events during HD, and that HTT functions as prosurvival factor.
Representative intracellular events in neurons expressing mutant HTT. In HD, processing of mutant HTT by caspases, calpains and MMPs facilitates the formation of intracellular aggregates, which are mainly degraded by autophagy. Failure in the clearance of HTT proteolytic fragments eventually results in excessive cytosolic Ca2+ concentration and organelle dysfunctions
HTT is also a substrate of calcium-activated proteases, that is, calpains. Calpains belong to the family of cysteine proteases typically activated by the elevation of intracellular Ca2+ levels, either in response to plasma membrane depolarization or in response to Ca2+ release from the intracellular stores.44 In mice overexpressing mutant HTT, increased glutamate release from afferent neurons enhances NMDA-R activity. This leads to an intracellular Ca2+ increase and therefore activation of calpains, which in turn cleave the HTT protein into a series of proteolytic products45 that promote NMDA-R-mediated excitotoxicity.46 Moreover, calpains can modulate HTT homeostasis via the catabolic process of autophagy. As shown by recent RNAi and chemical compound screenings in cultured cells, inhibition of calpains likely stimulates the lysosome-mediated degradation of intracellular aggregates.47, 48 Another RNAi screening study has also shown that small HTT fragments can be generated by the proteolytic activity of some matrix metalloproteinases (MMPs).48 The activation of the MMPs and the resulting cleavage of HTT were confirmed in samples from HD mouse models. Reduced MMP activity, especially MMP-10 and MMP-14, correlates with lower amount of proteolytic fragments and, as a result, suppression of neuronal degeneration induced by mutant HTT in cellular model systems as well as in Drosophila.48 Collectively, these findings suggest that protease inhibition might be a beneficial therapeutic approach for HD as it delays the formation of HTT-containing intracellular aggregates.
Autophagy
Autophagy is a cellular catabolic process that seems to have an important role in the pathogenesis of cancer as well as in neurodegenerative disorders.49, 50, 51, 52, 53, 54, 55, 56, 57 The process of autophagy involves the formation of a double-membrane structure (autophagosome) that then encloses a portion of cytosol and delivers its cargo content to the lysosomes for digestion.58, 59, 60 This nonspecific bulk degradation pathway is highly conserved from yeast to mammals. Autophagy occurs at constant low levels in all cells as part of ongoing cellular protein quality control and organelle turnover. However, it also has a primary role in the response to nutrient deprivation as it sustains metabolic functions by providing energy and metabolites to the cells. In different experimental settings, autophagy activation blocks detrimental processes and therefore facilitates cell stress resilience and survival.61, 62, 63, 64, 65 Furthermore, autophagy is one of the primary degradation pathways for various aggregate-prone proteins associated with neurodegenerative diseases.66, 67 As the tight regulation of autophagy is essential for cellular homeostasis, it is not surprising that autophagic dysfunction can cause metabolic stress and cell death68, 69, 70 mainly through apoptosis resulting from mitochondrial deficiency or via cleavage of Atg proteins.71
Among several key regulators of autophagy, the ‘target of rapamycin’ (TOR) senses energy status and the availability of the nutrients within the cell through the upstream class I phosphoinositol 3-kinase (PI3K), the serine/threonine kinase Akt and the 5′-AMP-activated protein kinase (AMPK).72 Inhibition of the TOR complex promotes the recruitment of Beclin-1 and Atg proteins involved in the formation of the mature autophagosome. The modulation of autophagy is therapeutically promising in HD: the inhibition of TOR by rapamycin enhances the clearance of mutant HTT-containing aggregates via the autophagy-lysosome pathway (Figure 1).64, 66 Similarly, drugs that block a rise in intracellular Ca2+, such as L-type Ca2+ channel antagonists, decrease the activity of calpains and result in the indirect activation of autophagy, likely by preventing the degradation of Beclin-1- and Atg-related proteins.73, 74 Although calpain inhibition promotes autophagy in-vitro75 and in zebrafish,76 it still remains to be determined whether it can be effective against HD in in-vivo mammalian models or in clinical settings.
Autophagy induction clearly represents an appealing approach for HD treatment; however, the therapeutic window remains to be determined as mutant HTT has a negative effect on the sequestration of the autophagic cargo. Although the autophagosomes seem to form and fuse to the lysosomes efficiently, there is a failure in the recognition of targeting signals, such as p62 or polyubiquitin, that results in delayed engulfment of cytosolic macromolecules and damaged organelles.77 Several strategies have been suggested to improve the clearance of HTT-containing aggregates by autophagy. One of them is based on the observation that histone deacetylase inhibitors block the polyQ protein-dependent neuronal degeneration in Drosophila.78 In this case, the acetylation of mutant HTT facilitates the recruitment of the protein to the autophagosome and therefore increases the removal of toxic species within the cells.79 More recent evidence demonstrates that HTT-mediated neuronal loss in Drosophila can be suppressed by genetic or pharmacological inhibition of NAD+-dependent class III deacetylases sirtuins.80 As pharmacological manipulation of sirtuins by resveratrol81 has been proposed to activate several pathways, including autophagy, these studies are of particular interest from a potential therapeutic standpoint.82, 83
Ageing Modifiers as Regulators of Proteostasis
Loss of proteostasis is a hallmark of several neurodegenerative disorders such as PD, AD and HD. In all of these disorders, aggregate-prone proteins trigger the formation of insoluble intracellular or extracellular aggregates as a result of environmental stress or metabolic changes. Whether the fibrillar protein aggregates are pathogenic or have protective roles, remains controversial.84, 85, 86 In nematodes and in mice, loss-of-function or decreased insulin/insulin-like growth factor 1 (IGF-1) signaling prevent the proteotoxicity caused by aggregate-prone peptides.87, 88 The insulin/IGF-1 signaling pathway is an evolutionarily conserved process that stimulates cellular growth according to nutrient availability.89, 90 The activation of the receptor leads to the potent activation of the downstream target PI3K and Akt, which coordinates multiple cellular processes such as proliferation, energy metabolism and survival. Together with TOR, Akt integrates the extracellular inputs with the intracellular status and tunes the cellular responses accordingly.
In Caenorhabditis elegans, loss-of-function mutations of the sole insulin/IGF-1 receptor daf-2 extend the lifespan to more than twofold.91 Genetic studies in C. elegans have revealed that the shift of polyQ-containing proteins from the soluble to the aggregate form is time-dependent. Loss-of-function of the PI3K age-1 not only extends the lifespan of nematodes but also significantly delays polyQ aggregation and toxicity.92 These protective effects are determined by increased expression of stress-response genes, such as heat shock proteins under the control of the transcription factors DAF-16 and HSF-1.93 Interestingly, overexpression of full-length, but not of truncated, HTT lowers the expression of plasma IGF-1 levels and, as result, affects body weight in mice.94 A decrease in IGF-1 expression has also been observed in different tissues of HD patients, which indicates that HTT loss-of-function can modulate IGF-1 signaling over time. In primary dissociated neurons expressing mutant HTT, treatment with IGF-1 induces specific activation of Akt and the direct phosphorylation of HTT, which results in a reduced number of HTT-containing intracellular inclusions and therefore neuroprotection.17 Thus, these findings suggest that IGF-1 signaling and HTT can apparently influence each other, although it still remains elusive whether this cross-talk potentiates or prevents detrimental cascades, including apoptosis.95 Modification of proteostasis by the Insulin/IGF-1 signaling pathway is not the only process, which affects HTT homeostasis. Recent screenings in C. elegans identified the evolutionarily conserved protein MOAG-4/SERF1-2 as a modifier of protein aggregation during ageing. Loss-of-function or silencing of MOAG-4 suppress the formation of aggregates in animals carrying mutant huntingtin, α-synuclein or β-amyloid.96 Whether MOAG-4/SERF1-2 and the interplay with other prosurvival pathways are relevant in HD remains to be explored, nevertheless the modulation of proteostasis remains a promising approach for the treatment of neurodegenerative disorders.
Mitochondrial Deficiency, Excitotoxicity and Inflammation
Energetic disturbances in HD is well described by post mortem, in-vitro and in-vivo evidences.11 The high metabolic rate of excitable cells such as neurons makes them strongly reliant upon mitochondrial functions. Mitochondria are highly motile organelles that control dendritic spine formation and synaptic activity by buffering intracellular Ca2+ rise underneath the plasma membrane.97, 98, 99 Mutant HTT has been shown to affect mitochondrial morphology and the bioenergetic status by altering the balance between mitochondrial fusion and fission under the control of the dynamin-related protein 1100, 101 or the interaction with other mitochondria-associated proteins.102 Alterations in mitochondria dynamics are reflected in deficits of the electron transport chain and of cellular respiration. The use of energy-related supplements, such as creatine, has been attempted in some clinical trials in order to correct mitochondrial defects in HD patients.38 As a result of extensive mitochondrial depolarization, neurons exposed to prolonged Ca2+ rise become vulnerable to excitotoxic insults (Figure 1).40, 103, 104 In HD, mutant HTT affects glutamatergic signals as a result of altered neurotransmitter release and activity of the glutamate-ionotropic receptors at the plasma membrane (Figure 1). In addition, aberrant HTT with the expanded polyQ tract inhibits the expression of the transcriptional co-activator PGC-1α, therefore compromising mitochondrial biogenesis and respiration.105 Thus, the combination of the two effects – alteration of Ca2+ influx and diminished capability of Ca2+ clearance by mitochondria – seriously increases the susceptibility of striatal cells expressing mutant HTT to excitotoxic insults. For this reason, agents that can affect glutamatergic signaling (i.e. NMDA receptor antagonists-like memantine) have been undergoing clinical trials.38 Similarly, other downstream targets that affect NMDA signaling and the excitotoxic neuronal demise might have some potential applications for the treatment of HD.106
Mitochondrial dysfunction resulting from Ca2+ overload, prolonged membrane depolarization or impairment of the electron transfer chain is the main source of intracellular reactive oxidative species.107, 108 Under certain circumstances, enhanced production of oxidative stress triggers neuroinflammatory responses by activation of the inflammasome in a cell-autonomous or non-autonomous manner.107 Neuroinflammatory processes are key determinants of neurodegenerative disorders characterized by aggregate-prone proteins, as in the case of PD and AD.109 Although the activation of inflammatory responses can be triggered by a variety of toxic species, the evidence indicates that most of the common neurodegenerative disorders have converging mechanisms that amplify the detrimental cascades. Remarkably, in the majority of the brain pathologies, neuroinflammation is a presymptomatic event and similar patterns have been shown in unrelated pathologies.110, 111 In case of HD, the expression of mutant HTT in glial cells affects the buffering capacity by altering the expression of the glutamate transporters, thus precluding the uptake of glutamate and enhancing neuronal excitotoxicity.112 Inflammation is a critical process that affects neuronal survival during pathological conditions.111 It has been shown that mutant HTT can lower the expression and release of glial chemokine,113 which can be neuroprotective under different circumstances.114 These data add additional complexity to the interaction between neurons and other brain cells. Whether targeting excessive activation of immune responses can be beneficial to HD remains to be determined, although it is tempting to consider it as a feasible possibility.115
Concluding Remarks
The identification of the HTT gene has contributed enormously to our understanding of the multiple pathogenic mechanisms involved in the onset of HD and in the selectively enhanced vulnerability of a subset of neurons to the mutant HTT. As discussed in this review, HD is a monogenic disease that results in a gain-of-function of the mutant form and in the loss-of-functions of the wild-type protein, which together severely compromise cellular homeostasis in a complex manner. To date, there is no cure for HD and most of the treatments available only help to alleviate some of the movement and psychiatric symptoms associated with the pathology. As mutant HTT is not considered to be an ideal pharmacological target due to its myriad biological functions, other biochemical pathways, such as those that prevent the abnormal accumulation of unfolded proteins, represent an encouraging alternative for the treatment of this neurodegenerative disorder. The identification and characterization of additional detrimental processes underlying cellular deficits in HD patients might provide new efficient and beneficial targets for neuroprotective intervention.
Abbreviations
- HD:
-
Huntington's disease
- HTT:
-
huntingtin
- polyQ:
-
polyglutamine
- AD:
-
Alzheimer's disease
- PD:
-
Parkinson's disease
- NMDA-R:
-
N-methyl-D-aspartate receptor
- MMP:
-
matrix metalloproteinases
- TOR:
-
target of rapamycin
- PI3K:
-
phosphoinositol 3-kinase
- IGF-1:
-
insulin-like growth factor 1
References
Martin JB, Gusella JF . Huntington's disease. Pathogenesis and management. N Engl J Med 1986; 315: 1267–1276.
Reiner A, Albin RL, Anderson KD, D’Amato CJ, Penney JB, Young AB . Differential loss of striatal projection neurons in Huntington disease. Proc Natl Acad Sci USA 1988; 85: 5733–5737.
Rosas HD, Koroshetz WJ, Chen YI, Skeuse C, Vangel M, Cudkowicz ME et al. Evidence for more widespread cerebral pathology in early HD: an MRI-based morphometric analysis. Neurology 2003; 60: 1615–1620.
A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington's disease chromosomes. The Huntington's Disease Collaborative Research Group. Cell 1993; 72: 971–983.
Andrew SE, Goldberg YP, Kremer B, Telenius H, Theilmann J, Adam S et al. The relationship between trinucleotide (CAG) repeat length and clinical features of Huntington's disease. Nat Genet 1993; 4: 398–403.
Rubinsztein DC, Barton DE, Davison BC, Ferguson-Smith MA . Analysis of the huntingtin gene reveals a trinucleotide-length polymorphism in the region of the gene that contains two CCG-rich stretches and a correlation between decreased age of onset of Huntington's disease and CAG repeat number. Hum Mol Genet 1993; 2: 1713–1715.
Aranda-Orgilles B, Aigner J, Kunath M, Lurz R, Schneider R, Schweiger S . Active transport of the ubiquitin ligase MID1 along the microtubules is regulated by protein phosphatase 2A. PLoS One 2008; 3: e3507.
Atwal RS, Xia J, Pinchev D, Taylor J, Epand RM, Truant R . Huntingtin has a membrane association signal that can modulate huntingtin aggregation, nuclear entry and toxicity. Hum Mol Genet 2007; 16: 2600–2615.
DiFiglia M, Sapp E, Chase K, Schwarz C, Meloni A, Young C et al. Huntingtin is a cytoplasmic protein associated with vesicles in human and rat brain neurons. Neuron 1995; 14: 1075–1081.
Trottier Y, Devys D, Imbert G, Saudou F, An I, Lutz Y et al. Cellular localization of the Huntington's disease protein and discrimination of the normal and mutated form. Nat Genet 1995; 10: 104–110.
Zuccato C, Valenza M, Cattaneo E . Molecular mechanisms and potential therapeutical targets in Huntington's disease. Physiol Rev 2010; 90: 905–981.
Strong TV, Tagle DA, Valdes JM, Elmer LW, Boehm K, Swaroop M et al. Widespread expression of the human and rat Huntington's disease gene in brain and nonneural tissues. Nat Genet 1993; 5: 259–265.
Nasir J, Floresco SB, O’Kusky JR, Diewert VM, Richman JM, Zeisler J et al. Targeted disruption of the Huntington's disease gene results in embryonic lethality and behavioral and morphological changes in heterozygotes. Cell 1995; 81: 811–823.
Zeitlin S, Liu JP, Chapman DL, Papaioannou VE, Efstratiadis A . Increased apoptosis and early embryonic lethality in mice nullizygous for the Huntington's disease gene homologue. Nat Genet 1995; 11: 155–163.
Dragatsis I, Levine MS, Zeitlin S . Inactivation of Hdh in the brain and testis results in progressive neurodegeneration and sterility in mice. Nat Genet 2000; 26: 300–306.
Zhang Y, Li M, Drozda M, Chen M, Ren S, Mejia Sanchez RO et al. Depletion of wild-type huntingtin in mouse models of neurologic diseases. J Neurochem 2003; 87: 101–106.
Humbert S, Bryson EA, Cordelieres FP, Connors NC, Datta SR, Finkbeiner S et al. The IGF-1/Akt pathway is neuroprotective in Huntington's disease and involves Huntingtin phosphorylation by Akt. Dev Cell 2002; 2: 831–837.
Perutz MF, Johnson T, Suzuki M, Finch JT . Glutamine repeats as polar zippers: their possible role in inherited neurodegenerative diseases. Proc Natl Acad Sci USA 1994; 91: 5355–5358.
Clabough EB, Zeitlin SO . Deletion of the triplet repeat encoding polyglutamine within the mouse Huntington's disease gene results in subtle behavioral/motor phenotypes in vivo and elevated levels of ATP with cellular senescence in vitro. Hum Mol Genet 2006; 15: 607–623.
Zheng S, Clabough EB, Sarkar S, Futter M, Rubinsztein DC, Zeitlin SO . Deletion of the huntingtin polyglutamine stretch enhances neuronal autophagy and longevity in mice. PLoS Genet 2010; 6: e1000838.
DiFiglia M, Sapp E, Chase KO, Davies SW, Bates GP, Vonsattel JP et al. Aggregation of huntingtin in neuronal intranuclear inclusions and dystrophic neurites in brain. Science 1997; 277: 1990–1993.
Martindale D, Hackam A, Wieczorek A, Ellerby L, Wellington C, McCutcheon K et al. Length of huntingtin and its polyglutamine tract influences localization and frequency of intracellular aggregates. Nat Genet 1998; 18: 150–154.
Saudou F, Finkbeiner S, Devys D, Greenberg ME . Huntingtin acts in the nucleus to induce apoptosis but death does not correlate with the formation of intranuclear inclusions. Cell 1998; 95: 55–66.
Thakur AK, Jayaraman M, Mishra R, Thakur M, Chellgren VM, Byeon IJ et al. Polyglutamine disruption of the huntingtin exon 1 N terminus triggers a complex aggregation mechanism. Nat Struct Mol Biol 2009; 16: 380–389.
Zuchner T, Brundin P . Mutant huntingtin can paradoxically protect neurons from death. Cell Death Differ 2008; 15: 435–442.
Davies SW, Turmaine M, Cozens BA, DiFiglia M, Sharp AH, Ross CA et al. Formation of neuronal intranuclear inclusions underlies the neurological dysfunction in mice transgenic for the HD mutation. Cell 1997; 90: 537–548.
Slow EJ, Graham RK, Osmand AP, Devon RS, Lu G, Deng Y et al. Absence of behavioral abnormalities and neurodegeneration in vivo despite widespread neuronal huntingtin inclusions. Proc Natl Acad Sci USA 2005; 102: 11402–11407.
Slow EJ, van Raamsdonk J, Rogers D, Coleman SH, Graham RK, Deng Y et al. Selective striatal neuronal loss in a YAC128 mouse model of Huntington disease. Hum Mol Genet 2003; 12: 1555–1567.
Hodgson JG, Agopyan N, Gutekunst CA, Leavitt BR, LePiane F, Singaraja R et al. A YAC mouse model for Huntington's disease with full-length mutant huntingtin, cytoplasmic toxicity, and selective striatal neurodegeneration. Neuron 1999; 23: 181–192.
Ciavardelli D, Silvestri E, Viscovo AD, Bomba M, Gregorio DD, Moreno M et al. Alterations of brain and cerebellar proteomes linked to Abeta and tau pathology in a female triple-transgenic murine model of Alzheimer's disease. Cell Death Dis 2010; 1: e90.
Lee MH, Lin SR, Chang JY, Schultz L, Heath J, Hsu LJ et al. TGF-beta induces TIAF1 self-aggregation via type II receptor-independent signaling that leads to generation of amyloid beta plaques in Alzheimer's disease. Cell Death Dis 2010; 1: e110.
Sivananthan SN, Lee AW, Goodyer CG, LeBlanc AC . Familial amyloid precursor protein mutants cause caspase-6-dependent but amyloid beta-peptide-independent neuronal degeneration in primary human neuron cultures. Cell Death Dis 2010; 1: e100.
Yacoubian TA, Slone SR, Harrington AJ, Hamamichi S, Schieltz JM, Caldwell KA et al. Differential neuroprotective effects of 14-3-3 proteins in models of Parkinson's disease. Cell Death Dis 2010; 1: e2.
Douglas PM, Dillin A . Protein homeostasis and aging in neurodegeneration. J Cell Biol 2010; 190: 719–729.
McCormack AL, Di Monte DA . Enhanced alpha-synuclein expression in human neurodegenerative diseases: pathogenetic and therapeutic implications. Curr Protein Pept Sci 2009; 10: 476–482.
Bodner RA, Outeiro TF, Altmann S, Maxwell MM, Cho SH, Hyman BT et al. Pharmacological promotion of inclusion formation: a therapeutic approach for Huntington's and Parkinson's diseases. Proc Natl Acad Sci USA 2006; 103: 4246–4251.
Krainc D . Clearance of mutant proteins as a therapeutic target in neurodegenerative diseases. Arch Neurol 2010; 67: 388–392.
Munoz-Sanjuan I, Bates GP . The importance of integrating basic and clinical research toward the development of new therapies for Huntington disease. J Clin Invest 2011; 121: 476–483.
Goldberg YP, Nicholson DW, Rasper DM, Kalchman MA, Koide HB, Graham RK et al. Cleavage of huntingtin by apopain, a proapoptotic cysteine protease, is modulated by the polyglutamine tract. Nat Genet 1996; 13: 442–449.
Orrenius S, Zhivotovsky B, Nicotera P . Regulation of cell death: the calcium-apoptosis link. Nat Rev Mol Cell Biol 2003; 4: 552–565.
Wellington CL, Ellerby LM, Gutekunst CA, Rogers D, Warby S, Graham RK et al. Caspase cleavage of mutant huntingtin precedes neurodegeneration in Huntington's disease. J Neurosci 2002; 22: 7862–7872.
Wellington CL, Singaraja R, Ellerby L, Savill J, Roy S, Leavitt B et al. Inhibiting caspase cleavage of huntingtin reduces toxicity and aggregate formation in neuronal and nonneuronal cells. J Biol Chem 2000; 275: 19831–19838.
Graham RK, Deng Y, Slow EJ, Haigh B, Bissada N, Lu G et al. Cleavage at the caspase-6 site is required for neuronal dysfunction and degeneration due to mutant huntingtin. Cell 2006; 125: 1179–1191.
Goll DE, Thompson VF, Li H, Wei W, Cong J . The calpain system. Physiol Rev 2003; 83: 731–801.
Gafni J, Ellerby LM . Calpain activation in Huntington's disease. J Neurosci 2002; 22: 4842–4849.
Cowan CM, Fan MM, Fan J, Shehadeh J, Zhang LY, Graham RK et al. Polyglutamine-modulated striatal calpain activity in YAC transgenic huntington disease mouse model: impact on NMDA receptor function and toxicity. J Neurosci 2008; 28: 12725–12735.
Williams A, Sarkar S, Cuddon P, Ttofi EK, Saiki S, Siddiqi FH et al. Novel targets for Huntington's disease in an mTOR-independent autophagy pathway. Nat Chem Biol 2008; 4: 295–305.
Miller JP, Holcomb J, Al-Ramahi I, de Haro M, Gafni J, Zhang N et al. Matrix metalloproteinases are modifiers of huntingtin proteolysis and toxicity in Huntington's disease. Neuron 2010; 67: 199–212.
Heidari N, Hicks MA, Harada H . GX15-070 (obatoclax) overcomes glucocorticoid resistance in acute lymphoblastic leukemia through induction of apoptosis and autophagy. Cell Death Dis 2010; 1: e76.
Bennett HL, Fleming JT, O’Prey J, Ryan KM, Leung HY . Androgens modulate autophagy and cell death via regulation of the endoplasmic reticulum chaperone glucose-regulated protein 78/BiP in prostate cancer cells. Cell Death Dis 2010; 1: e72.
Ruela-de-Sousa RR, Fuhler GM, Blom N, Ferreira CV, Aoyama H, Peppelenbosch MP . Cytotoxicity of apigenin on leukemia cell lines: implications for prevention and therapy. Cell Death Dis 2010; 1: e19.
Li Q, Li H, Roughton K, Wang X, Kroemer G, Blomgren K et al. Lithium reduces apoptosis and autophagy after neonatal hypoxia-ischemia. Cell Death Dis 2010; 1: e56.
Liang C . Negative regulation of autophagy. Cell Death Differ 2010; 17: 1807–1815.
Eisenberg-Lerner A, Bialik S, Simon HU, Kimchi A . Life and death partners: apoptosis, autophagy and the cross-talk between them. Cell Death Differ 2009; 16: 966–975.
Takahashi Y, Meyerkord CL, Wang HG . Bif-1/endophilin B1: a candidate for crescent driving force in autophagy. Cell Death Differ 2009; 16: 947–955.
Zhang J, Ney PA . Role of BNIP3 and NIX in cell death, autophagy, and mitophagy. Cell Death Differ 2009; 16: 939–946.
Maiuri MC, Tasdemir E, Criollo A, Morselli E, Vicencio JM, Carnuccio R et al. Control of autophagy by oncogenes and tumor suppressor genes. Cell Death Differ 2009; 16: 87–93.
Levine B, Kroemer G . Autophagy in the pathogenesis of disease. Cell 2008; 132: 27–42.
Noda T, Fujita N, Yoshimori T . The late stages of autophagy: how does the end begin? Cell Death Differ 2009; 16: 984–990.
Longatti A, Tooze SA . Vesicular trafficking and autophagosome formation. Cell Death Differ 2009; 16: 956–965.
Deng L, Feng J, Broaddus RR . The novel estrogen-induced gene EIG121 regulates autophagy and promotes cell survival under stress. Cell Death Dis 2010; 1: e32.
Marino ML, Fais S, Djavaheri-Mergny M, Villa A, Meschini S, Lozupone F et al. Proton pump inhibition induces autophagy as a survival mechanism following oxidative stress in human melanoma cells. Cell Death Dis 2010; 1: e87.
Silver N, Proctor GB, Arno M, Carpenter GH . Activation of mTOR coincides with autophagy during ligation-induced atrophy in the rat submandibular gland. Cell Death Dis 2010; 1: e14.
Ravikumar B, Vacher C, Berger Z, Davies JE, Luo S, Oroz LG et al. Inhibition of mTOR induces autophagy and reduces toxicity of polyglutamine expansions in fly and mouse models of Huntington disease. Nat Genet 2004; 36: 585–595.
Orvedahl A, Levine B . Eating the enemy within: autophagy in infectious diseases. Cell Death Differ 2009; 16: 57–69.
Williams A, Jahreiss L, Sarkar S, Saiki S, Menzies FM, Ravikumar B et al. Aggregate-prone proteins are cleared from the cytosol by autophagy: therapeutic implications. Curr Top Dev Biol 2006; 76: 89–101.
Sarkar S, Ravikumar B, Floto RA, Rubinsztein DC . Rapamycin and mTOR-independent autophagy inducers ameliorate toxicity of polyglutamine-expanded huntingtin and related proteinopathies. Cell Death Differ 2009; 16: 46–56.
Mizushima N, Levine B, Cuervo AM, Klionsky DJ . Autophagy fights disease through cellular self-digestion. Nature 2008; 451: 1069–1075.
Vellai T . Autophagy genes and ageing. Cell Death Differ 2009; 16: 94–102.
Kourtis N, Tavernarakis N . Autophagy and cell death in model organisms. Cell Death Differ 2009; 16: 21–30.
Wirawan E, Vande Walle L, Kersse K, Cornelis S, Claerhout S, Vanoverberghe I et al. Caspase-mediated cleavage of Beclin-1 inactivates Beclin-1-induced autophagy and enhances apoptosis by promoting the release of proapoptotic factors from mitochondria. Cell Death Dis 2010; 1: e18.
Kroemer G, Marino G, Levine B . Autophagy and the integrated stress response. Mol Cell 2010; 40: 280–293.
Russo R, Berliocchi L, Adornetto A, Varano GP, Cavaliere F, Nucci C et al. Calpain-mediated cleavage of Beclin-1 and autophagy deregulation following retinal ischemic injury in vivo. Cell Death Dis 2011; 2: e144.
Yousefi S, Perozzo R, Schmid I, Ziemiecki A, Schaffner T, Scapozza L et al. Calpain-mediated cleavage of Atg5 switches autophagy to apoptosis. Nat Cell Biol 2006; 8: 1124–1132.
Xia HG, Zhang L, Chen G, Zhang T, Liu J, Jin M et al. Control of basal autophagy by calpain1 mediated cleavage of ATG5. Autophagy 2010; 6: 61–66.
He C, Bartholomew CR, Zhou W, Klionsky DJ . Assaying autophagic activity in transgenic GFP-Lc3 and GFP-Gabarap zebrafish embryos. Autophagy 2009; 5: 520–526.
Martinez-Vicente M, Talloczy Z, Wong E, Tang G, Koga H, Kaushik S et al. Cargo recognition failure is responsible for inefficient autophagy in Huntington's disease. Nat Neurosci 2010; 13: 567–576.
Steffan JS, Bodai L, Pallos J, Poelman M, McCampbell A, Apostol BL et al. Histone deacetylase inhibitors arrest polyglutamine-dependent neurodegeneration in Drosophila. Nature 2001; 413: 739–743.
Jeong H, Then F, Melia Jr TJ, Mazzulli JR, Cui L, Savas JN et al. Acetylation targets mutant huntingtin to autophagosomes for degradation. Cell 2009; 137: 60–72.
Pallos J, Bodai L, Lukacsovich T, Purcell JM, Steffan JS, Thompson LM et al. Inhibition of specific HDACs and sirtuins suppresses pathogenesis in a Drosophila model of Huntington's disease. Hum Mol Genet 2008; 17: 3767–3775.
Howitz KT, Bitterman KJ, Cohen HY, Lamming DW, Lavu S, Wood JG et al. Small molecule activators of sirtuins extend Saccharomyces cerevisiae lifespan. Nature 2003; 425: 191–196.
Morselli E, Maiuri MC, Markaki M, Megalou E, Pasparaki A, Palikaras K et al. Caloric restriction and resveratrol promote longevity through the Sirtuin-1-dependent induction of autophagy. Cell Death Dis 2010; 1: e10.
Blagosklonny MV . Linking calorie restriction to longevity through sirtuins and autophagy: any role for TOR. Cell Death Dis 2010; 1: e12.
Haass C, Selkoe DJ . Soluble protein oligomers in neurodegeneration: lessons from the Alzheimer's amyloid beta-peptide. Nat Rev Mol Cell Biol 2007; 8: 101–112.
Caughey B, Lansbury PT . Protofibrils, pores, fibrils, and neurodegeneration: separating the responsible protein aggregates from the innocent bystanders. Annu Rev Neurosci 2003; 26: 267–298.
Goedert M, Spillantini MG . A century of Alzheimer's disease. Science 2006; 314: 777–781.
Cohen E, Bieschke J, Perciavalle RM, Kelly JW, Dillin A . Opposing activities protect against age-onset proteotoxicity. Science 2006; 313: 1604–1610.
Cohen E, Paulsson JF, Blinder P, Burstyn-Cohen T, Du D, Estepa G et al. Reduced IGF-1 signaling delays age-associated proteotoxicity in mice. Cell 2009; 139: 1157–1169.
Cohen E, Dillin A . The insulin paradox: aging, proteotoxicity and neurodegeneration. Nat Rev Neurosci 2008; 9: 759–767.
Bano D, Agostini M, Melino G, Nicotera P . Ageing, neuronal connectivity and brain disorders: an unsolved ripple effect. Mol Neurobiol 2011; 43: 124–130.
Kenyon C, Chang J, Gensch E, Rudner A, Tabtiang R . A C. elegans mutant that lives twice as long as wild type. Nature 1993; 366: 461–464.
Morley JF, Brignull HR, Weyers JJ, Morimoto RI . The threshold for polyglutamine-expansion protein aggregation and cellular toxicity is dynamic and influenced by aging in Caenorhabditis elegans. Proc Natl Acad Sci USA 2002; 99: 10417–10422.
Hsu AL, Murphy CT, Kenyon C . Regulation of aging and age-related disease by DAF-16 and heat-shock factor. Science 2003; 300: 1142–1145.
Pouladi MA, Xie Y, Skotte NH, Ehrnhoefer DE, Graham RK, Kim JE et al. Full-length huntingtin levels modulate body weight by influencing insulin-like growth factor 1 expression. Hum Mol Genet 2010; 19: 1528–1538.
Mitchell GC, Fillinger JL, Sittadjody S, Avila JL, Burd R, Limesand KH . IGF1 activates cell cycle arrest following irradiation by reducing binding of DeltaNp63 to the p21 promoter. Cell Death Dis 2010; 1: e50.
van Ham TJ, Holmberg MA, van der Goot AT, Teuling E, Garcia-Arencibia M, Kim HE et al. Identification of MOAG-4/SERF as a regulator of age-related proteotoxicity. Cell 2010; 142: 601–612.
Giacomello M, Drago I, Pizzo P, Pozzan T . Mitochondrial Ca2+ as a key regulator of cell life and death. Cell Death Differ 2007; 14: 1267–1274.
Li Z, Okamoto K, Hayashi Y, Sheng M . The importance of dendritic mitochondria in the morphogenesis and plasticity of spines and synapses. Cell 2004; 119: 873–887.
Young KW, Bampton ET, Pinon L, Bano D, Nicotera P . Mitochondrial Ca2+ signalling in hippocampal neurons. Cell Calcium 2008; 43: 296–306.
Costa V, Giacomello M, Hudec R, Lopreiato R, Ermak G, Lim D et al. Mitochondrial fission and cristae disruption increase the response of cell models of Huntington's disease to apoptotic stimuli. EMBO Mol Med 2010; 2: 490–503.
Song W, Chen J, Petrilli A, Liot G, Klinglmayr E, Zhou Y et al. Mutant huntingtin binds the mitochondrial fission GTPase dynamin-related protein-1 and increases its enzymatic activity. Nat Med 2011; 17: 377–382.
Sassone J, Colciago C, Marchi P, Ascardi C, Alberti L, Di Pardo A et al. Mutant Huntingtin induces activation of the Bcl-2/adenovirus E1B 19-kDa interacting protein (BNip3). Cell Death Dis 2010; 1: e7.
Bano D, Nicotera P . Ca2+ signals and neuronal death in brain ischemia. Stroke 2007; 38 (2 Suppl): 674–676.
Ankarcrona M, Dypbukt JM, Bonfoco E, Zhivotovsky B, Orrenius S, Lipton SA et al. Glutamate-induced neuronal death: a succession of necrosis or apoptosis depending on mitochondrial function. Neuron 1995; 15: 961–973.
Cui L, Jeong H, Borovecki F, Parkhurst CN, Tanese N, Krainc D . Transcriptional repression of PGC-1alpha by mutant huntingtin leads to mitochondrial dysfunction and neurodegeneration. Cell 2006; 127: 59–69.
Zhu S, Zhang Y, Bai G, Li H . Necrostatin-1 ameliorates symptoms in R6/2 transgenic mouse model of Huntington's disease. Cell Death Dis 2011; 2: e115.
Zhou R, Yazdi AS, Menu P, Tschopp J . A role for mitochondria in NLRP3 inflammasome activation. Nature 2011; 469: 221–225.
Ruiz A, Matute C, Alberdi E . Intracellular Ca2+ release through ryanodine receptors contributes to AMPA receptor-mediated mitochondrial dysfunction and ER stress in oligodendrocytes. Cell Death Dis 2010; 1: e54.
Glass CK, Saijo K, Winner B, Marchetto MC, Gage FH . Mechanisms underlying inflammation in neurodegeneration. Cell 2010; 140: 918–934.
Sapp E, Kegel KB, Aronin N, Hashikawa T, Uchiyama Y, Tohyama K et al. Early and progressive accumulation of reactive microglia in the Huntington disease brain. J Neuropathol Exp Neurol 2001; 60: 161–172.
Rodriguez JJ, Witton J, Olabarria M, Noristani HN, Verkhratsky A . Increase in the density of resting microglia precedes neuritic plaque formation and microglial activation in a transgenic model of Alzheimer's disease. Cell Death Dis 2010; 1: e1.
Shin JY, Fang ZH, Yu ZX, Wang CE, Li SH, Li XJ . Expression of mutant huntingtin in glial cells contributes to neuronal excitotoxicity. J Cell Biol 2005; 171: 1001–1012.
Chou SY, Weng JY, Lai HL, Liao F, Sun SH, Tu PH et al. Expanded-polyglutamine huntingtin protein suppresses the secretion and production of a chemokine (CCL5/RANTES) by astrocytes. J Neurosci 2008; 28: 3277–3290.
Nicolai J, Burbassi S, Rubin J, Meucci O . CXCL12 inhibits expression of the NMDA receptor's NR2B subunit through a histone deacetylase-dependent pathway contributing to neuronal survival. Cell Death Dis 2010; 1: e33.
Yong VW, Rivest S . Taking advantage of the systemic immune system to cure brain diseases. Neuron 2009; 64: 55–60.
Acknowledgements
We thank Dr. Sarah Jewell, Dr. Sybille Krauss, Professor Ina Vorberg and Professor Gerry Melino for their useful comments.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by G Melino
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Bano, D., Zanetti, F., Mende, Y. et al. Neurodegenerative processes in Huntington's disease. Cell Death Dis 2, e228 (2011). https://doi.org/10.1038/cddis.2011.112
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2011.112
Keywords
This article is cited by
-
Interleukin 3 Inhibits Glutamate-Cytotoxicity in Neuroblastoma Cell Line
Neurochemical Research (2024)
-
Huntington Disease Gene Expression Signatures in Blood Compared to Brain of YAC128 Mice as Candidates for Monitoring of Pathology
Molecular Neurobiology (2022)
-
Immune-microbiome interplay and its implications in neurodegenerative disorders
Metabolic Brain Disease (2022)
-
The impact of proteostasis dysfunction secondary to environmental and genetic causes on neurodegenerative diseases progression and potential therapeutic intervention
Environmental Science and Pollution Research (2020)
-
Therapeutic Effect of Agmatine on Neurological Disease: Focus on Ion Channels and Receptors
Neurochemical Research (2019)