Abstract
Our previous studies have shown that microRNA-383 (miR-383) expression is downregulated in the testes of infertile men with maturation arrest (MA). However, the underlying mechanisms of miR-383 involved in the pathogenesis of MA remain unknown. In this study, we showed that downregulation of miR-383 was associated with hyperactive proliferation of germ cells in patients with mixed patterns of MA. Overexpression of miR-383 in NT2 (testicular embryonal carcinoma) cells resulted in suppression of proliferation, G1-phase arrest and induction of apoptosis, whereas silencing of miR-383 reversed these effects. The effects of miR-383 were mediated through targeting a tumor suppressor, interferon regulatory factor-1 (IRF1), and miR-383 was negatively correlated with IRF1 protein expression in vivo. miR-383 inhibited IRF1 by affecting its mRNA stability, which subsequently reduced the levels of the targets of IRF1, namely cyclin D1, CDK2 and p21. Downregulation of IRF1 or cyclin D1, but not that of CDK2, enhanced miR-383-mediated effects, whereas silencing of p21 partially inhibited the effects of miR-383. Moreover, miR-383 downregulated CDK4 by increasing proteasome-dependent degradation of CDK4, which in turn resulted in an inhibition of phosphorylated retinoblastoma protein (pRb) phosphorylation. These results suggest that miR-383 functions as a negative regulator of proliferation by targeting IRF1, in part, through inactivation of the pRb pathway. Abnormal testicular miR-383 expression may potentiate the connections between male infertility and testicular germ cell tumor.
Similar content being viewed by others
Main
Infertility affects 10–15% of couples who are trying to conceive, and half of the cases are due to male factors.1 Despite several decades of efforts to elucidate the etiological mechanism of male infertility, most cases are idiopathic. Furthermore, patients with unexplained nonobstructive azoospermia (NOA), especially men with maturation arrest (MA, a cause of NOA), usually have a significantly lower sperm retrieval rate and a clinical pregnancy rate.2 Therefore, elucidating the underlying pathogenesis of MA may help to improve treatment outcomes in these patients.
Spermatogenesis is an elaborate process involving the development of spermatogonial stem cells to highly differentiated spermatozoon, including the stages of actively proliferating spermatogonia and meiotic spermatocytes. Some histological studies have suggested the existence of accelerated germ cell apoptosis3 and decreased mitotic activity of spermatogonia4 in infertile men with MA at the spermatocyte stage. However, a recent study has shown normal active spermatogonia in testes arrested at the pachytene stage.5 In these cases, spermatogenic failure may be due to meiotic impairment, such as impaired chromosome synapsis and decreased meiotic recombination.6, 7 These histological findings demonstrate various patterns of spermatogonial proliferation in men with MA. However, the molecular mechanisms regulating germ cell proliferation remain poorly understood.
A significant feature of mammalian spermatogenesis is that numerous mRNAs in differentiating germ cells undergo posttranscriptional and translational regulation.8 microRNAs (miRNAs), a family of small noncoding RNAs (typically 19–23 nt), post-transcriptionally regulate gene expression through base pair binding to 3′-untranslated region (3′ UTR) sequences of target mRNAs.9 Several studies have indicated that miRNAs could have an important role during spermatogenesis. For example, the loss of Dicer1 (an RNase III enzyme required for miRNA processing) in primordial germ cells and spermatogonia resulted in defective proliferation;10 although in Dicer-deleted testis, spermatogenesis was arrested at an early stage during proliferation and/or early differentiation,10, 11 miR-372 and miR-373 have been implicated as oncogenes in testicular germ cell tumors (TGCTs).12 However, little is known about the mechanisms of miRNAs in the regulation of mammalian spermatogenesis.
Recently, we have identified many deregulated miRNAs in testicular tissues of NOA patients with MA.13 miR-383, the expression of which is restricted to the spermatogonia and early spermatocytes, exhibited a significant downregulation of gene expression in MA patients.13 These findings suggest that miR-383 may act on early stages of spermatogenesis and thus may regulate germ cell proliferation or death. In this study, miR-383 was functionally characterized in the testicular embryonal carcinoma cell line NTERA-2 (NT2) and during spermatogenesis by identifying its target genes.
Results
Hyperactive proliferation of germ cells in patients with mixed patterns of MA
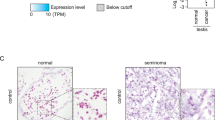
Germ cell proliferation and apoptosis were determined with immunohistochemical expression of proliferating cell nuclear antigen (PCNA) and in situ DNA 3′-end labeling (terminal deoxy-nucleotidyl transferase-mediated digoxigenin-dUTP nick-end labeling, TUNEL) in the testes of infertile men with mixed patterns of MA, respectively. As shown in Figure 1a, positive TUNEL staining was detected predominantly in spermatogonia and spermatocytes. Compared with normal controls, the rate of germ cell apoptosis in testicular biopsy specimens obtained from MA patients was slightly upregulated (Table 1). However, the difference between the groups did not reach statistical significance (P=0.21). Unexpectedly, most of the seminiferous tubules in all seven of MA patients exhibited high immunoreactivity for PCNA in the nuclei of spermatogonia and primary spermatocytes (Figure 1b and Table 2). Furthermore, real-time PCR results also suggested the upregulation of PCNA mRNA in some patients with MA (Figure 1c), although specimens obtained from men with the Sertoli-cell-only (SCO) syndrome showed no PCNA-immunopositive staining (Figure 1b). These results indicate that some human spermatogenic failure could be due to hyperactive proliferation of germ cells.
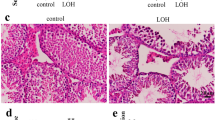
Downregulation of miR-383 expression is associated with hyperactive proliferation of germ cells in MA patients. (a) Apoptotic germ cells in testicular specimens of normal controls and MA patients were identified by the TUNEL assay. Arrowheads indicate TUNEL-positive cells. (b) Proliferating germ cells were determined by immunohistochemical staining for PCNA. Scale bar: 13 μm. (c) The expression of PCNA mRNA in the testes of normal controls (N1) and MA patients (A1–A5) was examined by real-time PCR in triplicate. (d) The state of Sertoli cell differentiation was assessed by immunohistochemical staining for CK-18. (e) Localization of miR-383 in the testes of normal controls and MA patients by LNA-based miRNA in situ hybridization. Representative hematoxylin and eosin staining, hybridization signals (purple) for miR-383 and negative controls are shown. (f and g) Expressions of mature miR-383 (panel f) and pri-miR-383 (panel g) in the testes of normal controls (N1) and MA patients (A1–A5) were examined by real-time PCR in triplicate. (h) A significant positive correlation was found between the levels of pri-miR-383 and mature miR-383 (panel d, r=0.8802, P<0.05) Scale bar: 50 μm. SPG, spermatogonia; PS, primary spermatocyte; Sp, spermatid. The Ct value for the control samples was arbitrarily set at 1. Data were expressed as mean±S.E.M. (n=3). **P<0.01, compared with normal controls
It is reported that spermatogenic impairment is associated with the presence of immature Sertoli cells.14 We then examined the immunohistochemical expression of cytokeratin-18 (CK-18, a marker of immature or undifferentiated Sertoli cells) in testicular tissues of infertile men with mixed patterns of MA. Histological analysis revealed remarkable expression of CK-18 in SCO tubules (Figure 1d). Only the residual expression of the marker was demonstrated in adjacent tubules showing arrest of spermatogenesis at the level of spermatogonia or primary spermatocytes, as well as normal spermatogenesis (Figure 1d). These results suggest that in patients with mixed patterns of MA, spermatogenic arrest is intrinsic to the germ line without affecting Sertoli cell differentiation.
Downregulation of miR-383 was associated with enhanced proliferative activity of germ cells in MA patients
The expression of miR-383 in testicular biopsy specimens was performed by in situ hybridization (ISH) and real-time PCR. In seminiferous tubules with normal spermatogenesis, the miR-383 expression was predominantly found in spermatogonia and primary spermatocytes (Figure 1e). Consistent with the miRNA microarray results,13 miR-383 was decreased significantly in testicular specimens with MA (Figure 1e). In addition, real-time PCR results also revealed a significant downregulation of miR-383 expression in testes obtained from all five NOA patients compared with normal controls (Figure 1f). This downregulation may be exclusive for these patients, as miR-383 was not altered in infertile patients with hypospermatogenesis (Supplementary Figure 1). These data indicate that the downregulation of miR-383 is associated with increased spermatogonial proliferation of MA patients.
To determine whether the downregulation of miR-383 occurs at the level of transcription, the expression of primary miR-383 gene transcripts (pri-miR-383) in the same NOA patients was examined. Four out of five NOA patients (A2–A5) with downregulated mature miR-383 expression also showed a significant decrease in pri-miR-383 levels (Figure 1g). A significant correlation between the levels of pri-miR-383 and mature miR-383 in these four cases was identified (r=0.8802, P<0.05; Figure 1h), indicating that the miR-383 expression is mainly regulated at the transcriptional level. However, one case (A1) displayed much higher levels of pri-miR-383 than did normal controls (Figure 1g), suggesting an additional layer of post-transcriptional regulation of miR-383 expression.
The effects of miR-383 on NT2 cell survival were evaluated. Overexpression of miR-383 by transfecting the synthetic miR-383 mimic into NT2 cells not only significantly decelerated cell proliferation (Figure 2a) but also induced G1 cell-cycle arrest (Figure 2b) and apoptosis (Figure 2c). Conversely, inhibition of endogenous miR-383 with antisense oligonucleotides resulted in a significant elevation in cell proliferation (Figure 2d), a decrease in cell population in the G1 phase (Figure 2e) and suppression of basal apoptosis (Figure 2f).
miR-383 functions as a negative regulator of proliferation in NT2 cells. (a and d) miR-383 repressed cell proliferation. Cell viability was measured by the cell counting kit-8 assay after transfection with 25, 50 or 100 nM mimic negative control (Mimic NC) or miR-383 mimic (panel a) or with 150 nM inhibitor negative control (Inhibitor NC) or miR-383 inhibitor (panel d). (b and e) miR-383 induced G1-phase arrest. Cell-cycle profiles were analyzed using flow cytometry on cells transfected either with 150 nM miR-383 mimic (panel b) or with 150 nM miR-383 inhibitor (panel e). (c and f) miR-383 promoted cell apoptosis. The apoptosis rate was measured using flow cytometry with Annexin V and PI staining after transfection either with 100 nM miR-383 mimic (panel c), or with 150 nM miR-383 inhibitor (panel f). Data are presented as means±S.E.M. (n=3). *P<0.05, **P<0.01, compared with negative controls
IRF1 is a direct target of miR-383
A potential target of miR-383, interferon regulatory factor-1 (IRF1), is predicted by the TargetScan database. The 3′ UTR of IRF1 mRNA harbors a putative miR-383-binding site, which is evolutionarily conserved in vertebrates (Figure 3a). To determine whether IRF1 is a true target of miR-383, NT2 cells were transfected with miR-383 mimic/control or miR-383 inhibitor/control. Both IRF1 protein (Figures 3b and c) and IRF1 mRNA (Figure 3d) expressions were significantly decreased in miR-383-transfected NT2 cells than in control cells, whereas knockdown of miR-383 increased IRF1 protein expression.
IRF1 is a direct target of miR-383. (a) Putative-binding sites for human (hsa) miR-383 and mouse (mmu) miR-383 are predicted in the 3′ UTR of IRF1 mRNA. The underlined nucleotides indicate the mutated bases of predicted miR-383-binding sites. MT IRF1–3′ UTR, Mutation-type IRF1 3′ UTR. (b–d) miR-383 downregulated the IRF1 protein (panels b and c) and mRNA (panel d) expression levels. mRNA and protein expressions of IRF1 were determined by real-time PCR and western blotting, respectively. Representative western blot bands (panel b) and statistical summary (panel c) are shown. Relative band intensities were calculated by densitometry and normalized to the loading control (GAPDH). (e) miR-383 targeted 3′ UTR of IRF1. Luciferase reporters containing either miR-383 complementary sites from wild-type IRF1 3′ UTR (WT IRF1-3′ UTR) or mutation-type IRF1 3′ UTR (MT IRF1-3′ UTR) were cotransfected with the indicated oligonucleotides into 293T cells. Luciferase activity was measured 30 h after transfection. The value from cells cotransfected with Mimic NC and a plasmid expressing WT IRF1-3′ UTR was arbitrarily set at 1. Data were presented as means±S.E.M. for at least three independent experiments. *P<0.05, **P<0.01, compared with negative controls
To validate whether the IRF1 gene is a direct target of miR-383, we constructed Renilla luciferase reporters containing either the wild-type (WT) full-length 3′ UTR or the mutant forms of seeding sites. Cotransfection of the miR-383 mimic and the reporters into 293T cells resulted in an ∼45% decrease in luciferase activity, whereas inhibition of miR-383 expression increased the reporter activity (Figure 3e). On the other hand, mutation of seed sequences abolished the silencing effects of miR-383 (Figures 3a and e). Taken together, these results demonstrate that miR-383 downregulates IRF1 directly by decreasing mRNA stability.
To verify whether IRF1 induced phenotypes associated with miR-383 function, we specifically silenced IRF1 (si-IRF1) in NT2 cells by using RNA interference (RNAi). As expected, knockdown of IRF1 (Figure 4a) led to a significantly decreased cell proliferation in a dose-dependent manner (Figure 4b), accumulation of cells in the G1 phase (Figure 4c) and induction of apoptosis (Figure 4d). Furthermore, compared with NT2 cells transfected with the miR-383 mimic, cotransfection of si-IRF1 and the miR-383 mimic into cells resulted in a much more decreased cell proliferation (Figure 4e), a more increased G1-phase population (Figure 4f) and a higher apoptotic rate (Figure 4g). These results demonstrate that IRF1, at least in part, mediates the effects of miR-383 on cell proliferation, cell-cycle regulation and apoptosis.
Silencing of endogenous IRF1 enhances miR-383-mediated effects on cell cycle and apoptosis in NT2 cells. (a) The efficacy of IRF1 siRNA was evaluated by western blotting after 48 h transfection with either siRNA negative control (si.NC) or IRF1 siRNA (si-IRF1). (b–d) Knockdown of IRF1 inhibited cell proliferation (panel b), induced G1-phase arrest (panel c) and activated apoptosis (panel d). Cell viability (panel b) was measured by the cell counting kit-8 assay after transfection with 5, 10, 20, 30, 50, 100 nM si.NC or si-IRF1. Proportions of cells in the G1, S, and G2/M phases of the cell cycle (panel c) and apoptotic rates (panel d) were determined using flow cytometry. (e–g) Silencing of IRF1 enhanced miR-383-induced proliferation suppression (panel e), G1-phase arrest (panel f) and apoptosis (panel g). Cell viability, cell-cycle profiles and apoptotic rates were analyzed after cotransfection with 100 nM miR-383 mimic and 30 nM si-IRF1 into NT2 cells. (h) IRF1 was induced rapidly by serum. NT2 cells were growth arrested by serum starvation for 24 h and then harvested for western blot analysis at the indicated time points after serum stimulation. Untreated cells were also included as a control (Exp). (i) Overexpression of miR-383 or silencing of IRF1 inhibited serum-inducible IRF1 protein expression. Cells were transfected with the indicated oligonucleotides after serum starvation for 6 h and growth arrested for a further 18 h. For western blot analysis, cells were harvested 2 h after serum stimulation. SFM, serum-free medium. Error bars indicate S.E.M. (n=3). *P<0.05, **P<0.01, ***P<0.001, compared with negative controls
As described in this study, IRF1 might exhibit a pro-mitogenic role in NT2 cells. However, in general, IRF1 had an anti-mitogenic role in most other cell lines.15 To test this hypothesis, NT2 cells were growth arrested by serum starvation. As shown in Figure 4h, the IRF1 protein was markedly elevated within 2 h after serum stimulation of growth-arrested NT2 cells, just the time point when the IRF1 protein rapidly declined in NIH3T3 cells.16 Meanwhile, both miR-383 mimic and si-IRF1 inhibited serum-inducible IRF1 protein expression (Figure 4i). These observations suggest that, besides its function as an anti-mitogenic factor, IRF1 also exhibits a pro-mitogenic role in NT2 cells.
miR-383 expression correlates negatively with the IRF1 protein in vivo
To further validate the regulatory relationship between miR-383 and its target IRF1 in vivo, expressions of miR-383 and IRF1 protein were identified in the normal human/mouse testis and in human infertile testis. First, the developmental changes of miR-383 and IRF1 protein expression in the mouse testis were surveyed by real-time PCR and western blot analysis, respectively. The miR-383 expression was high at birth, then gradually decreased until 14 days postpartum (dpp) (except for an increase at 8 dpp), and became stable from 3 weeks onwards (Figure 5a). ISH analysis showed that miR-383 was mainly expressed in spermatogonia and primary spermatocytes, which was consistent with the results obtained in the human testis (Figure 5b). The IRF1 expression was increased soon after birth, peaked at 5 dpp and then gradually declined (Figures 5c and d). Immunocytochemical localization demonstrated that IRF1 was predominantly expressed in the cytoplasm of spermatogonia (Figure 5e). However, a nuclear accumulation of IRF1 was observed at 5 dpp (Figure 5e). Furthermore, a significant negative correlation between the expression of miR-383 and IRF1 at 0–8 dpp was found (r=−0.9861, P<0.05; Figure 5f).
IRF1 is negatively correlated with miR-383 in vivo. (a) Real-time PCR analysis of miR-383 expression in the mouse testes on days postpartum (dpp) 0, 2, 5, 8, 14, 21, 35, 49 and 63. The bars represent data obtained from at least three mice, expressed as mean±S.E.M. of miR-383 expression levels normalized to day 0 control. (b) Localization of miR-383 in the mouse testes at 30 dpp using LNA-based miRNA in situ hybridization. Scale bar: 50 μm. (c and d) western blot analysis of IRF1 protein expression in the mouse testes on indicated dpp. Representative images (panel c) and statistical summary (panel d) are shown. Relative band intensities were calculated by densitometry and normalized to the loading control (GAPDH). Data were shown as mean±S.E.M. from at least three mice. (e) Immunohistochemical staining for IRF1 in the mouse testes during postnatal development (0, 5, 36 dpp). Scale bar: 13 μm. (f) A significant negative correlation was found between mature miR-383 and IRF1 protein expressions in the mouse testes at 0–8 dpp (r=−0.9861, P<0.05). (g) Immunohistochemical staining for IRF1 in testicular biopsy specimens of normal controls and MA patients. Scale bar: 50 μm. SC, Sertoli cell; SPG, spermatogonia; PS, primary spermatocyte; Sp, spermatid
Second, the IRF1 expression in human testicular specimens of two controls and seven MA patients was analyzed by immunohistochemistry. As showed in Figure 5g and Table 2, IRF1 was also detected in a very small proportion of spermatogonia and restricted to the cytoplasm in controls. However, all MA patients but one (case A2) showed significantly higher signal intensity in the nucleus of both spermatogonia and primary spermatocytes, or in either of them (case A7 for spermatogonia, case A3 for primary spermatocytes). Meanwhile, four out of the five cases of testicular tissues that had reduced miR-383 levels, showed an elevated IRF1 expression (Table 2). The other two cases were not examined for miR-383 levels as we did not have enough tissues to analyze. These observations corroborate the negative correlation between the expression of miR-383 and IRF1 in MA patients.
miR-383 represses IRF1-targeted cell cycle-related genes
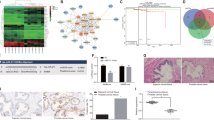
It is well known that the transcription factor IRF1 exerts multiple effects by activating a set of target genes, depending on the cell type and on the specific stimuli.17 We next examined the effects of IRF1/miR-383 on the expressions of known IRF1 target genes involved in the regulation of cell apoptosis, cell cycle and DNA repair. Our results revealed that mRNA (Figure 6a) and protein (Figure 6b) levels of three cell cycle-related genes, namely Cyclin D1,18 CDK219 and p21,20 were all lower in NT2 cells transfected with si-IRF1 than in cells transfected with negative controls (NCs). Similarly, the miR-383 mimic clearly decreased mRNA and protein levels of these three genes (Figures 6c and d), whereas knockdown of miR-383 induced their mRNA and protein expressions (Figures 6c and d).
miR-383 regulates the expression of cyclin D1, CDK2 and p21 at the transcriptional level. (a–d) Effects of si-IRF1 and miR-383 mimic/inhibitor on mRNA (panel a for si-IRF1, panel c for miR-383) and protein (panel b for si-IRF1, panel d for miR-383) expression levels of IRF1 targets in NT2 cells were determined by RT-PCR and western blotting, respectively
We were then interested in identifying a major gene responsible for miR-383-mediated proliferation control in NT2 cells. Protein expressions of cyclin D1, CDK2 and p21 were analyzed after serum stimulation of growth-arrested cells. As shown in Figure 7a, mitogen stimulation markedly induced cyclin D1 and p21 expression, but CDK2 levels remained constant. Only cyclin D1 was inhibited by the miR-383 mimic or si-IRF1. In addition, although si-cyclin D1, si-CDK2 and si-p21 (Figure 7b) all resulted in inhibition of proliferation (Supplementary Figures 2a and b), only knockdown of cyclin D1 caused G1-phase arrest in NT2 cells (Supplementary Figures 2c and d). Furthermore, overexpression of cyclin D1 partly rescued miR-383-induced inhibition of proliferation (Figure 7c). These results suggested that cyclin D1 might be the major gene responsible for miR-383-mediated antiproliferative effects in NT2 cells.
The pRb pathway mediated miR-383-induced phenotype in NT2 cells. (a) Cells were treated as in Figure 4i. For western blotting analysis of the indicated proteins, cells were harvested 2 h after serum stimulation of growth-arrested cells. (b) The efficacy of siRNAs for cyclin D1, CDK2 and p21 was evaluated by western blot. (c) Cyclin D1 partly rescued NT2 cells from growth inhibition by miR-383. Twenty-four hours after transfection with 100 nM miR-383 mimic, cells were further transfected with 0.1 μg cyclin D1 expression vector (pcDNA3.1/cyclin D1) for 24 h and then subjected to cell viability assay. (d and e) Effects of knockdown of cyclin D1 or p21 on miR-383-induced G1-phase arrest (panel d) and apoptosis (panel e). NT2 cells were cotransfected with the miR-383 mimic and the indicated siRNAs. (f) Immunoblot analysis of phosphorylation of pRb using an antibody specific for phosphorylated Ser249/Thr252 (P-pRb). Cells were treated as in Figure 4i. For western blotting analysis, cells were harvested 24 h after serum stimulation of growth-arrested cells. (g and h) The protein level of CDK4 was determined by western blotting 24 h after transfection with the indicated oligonucleotides (panel g) or cotreatment of the miR-383 mimic with 20 μM of MG-132 or DMSO (panel h). SFM, serum-free medium. Error bars indicate S.E.M. (n=3). *P<0.05, **P<0.01, compared with negative controls
Interestingly, contrary to the role of cyclin D1 in NT2 cells, p21 promoted G1/S transition (Supplementary Figures 2c and d). In addition, knockdown of p21 partly inhibited miR-383-induced G1-phase arrest, but si-cyclin D1 dramatically enhanced the effects of miR-383 (Figure 7d). Similarly, silencing of cyclin D1 and silencing of p21 resulted in a slight increase and decrease in apoptosis rates, respectively (Figure 7e).
miR-383 inhibits pRb phosphorylation and CDK4 protein levels
The assembly of cyclin D1–CDK4–p21 complex is required for phosphorylated retinoblastoma protein (pRb)-mediated G1/S transition.21 As described above, miR-383 represses the expression of two members of this complex, cyclin D1 and p21, which leads us to ask whether miR-383 has a role in regulating the activity of cyclin D1–CDK4–p21 complex. We first examined the effects of miR-383 on pRb phosphorylation using specific antibodies recognizing pRb phosphorylated on Ser249/Thr252. Phosphorylations of Ser249/Thr252 in vitro are preferentially catalyzed by cyclin D1–CDK4 complex.22 The results revealed that both miR-383 mimic and si-IRF1 inhibited serum-inducible pRb phosphorylation (Figure 7f).
Second, immunofluorescence showed that neither the miR-383 mimic nor si-IRF1 affected the nuclear localization of cyclin D1 and CDK4 in NT2 cells (Supplementary Figures 3a–c). However, although silencing of IRF1 did not affect CDK4 protein levels (Figure 7g), the miR-383 mimic repressed CDK4 expression (Figure 7g) without a significant decrease in CDK4 mRNA levels (Supplementary Figure 4a). We then explored the possibility that CDK4 might also be a target of miR-383. However, luciferase assay showed that the miR-383 mimic did not inhibit luciferase activity of a reporter vector containing the full-length CDK4 3′ UTR (Supplementary Figure 4b).
As miR-383 affects neither CDK4 transcription nor CDK4 mRNA stability, we speculate whether miR-383 induces degradation of CDK4 through a proteasome-dependent pathway. As shown in Figure 7h, treatment with the proteasome inhibitor MG-132 restored CDK4 expression that was downregulated by miR-383. These results confirm that miR-383 suppresses CDK4 protein expression by promoting proteasome degradation of CDK4.
Widespread upregulation of P-pRb expression in MA patients
To further understand the role of the above-mentioned miR-383-pRb pathway in spermatogenesis, the expression of cyclin D1, p21, CDK2, CDK4 and phosphorylated pRb (P-pRb) was analyzed in testicular specimens of MA patients. The immunohistochemical expression patterns of these proteins are summarized in Table 2. In germ cells in normal controls, expressions of cyclin D1, p21 and CDK2 were nearly undetectable, whereas both diffuse immunostaining for P-pRb and distinct staining pattern for CDK4 were detected in the nucleus of spermatogonia, spermatocytes and round spermatids (Figure 8). Although nuclear localization of staining for all these proteins was higher in some seminiferous tubules of MA patients characterized by the downregulation of mature miR-383 (Table 2), only enhanced nuclear staining for P-pRb was identified in all MA patients studied. For example, one case (A1) showed enhanced nuclear staining for P-pRb, but could not detect the expression of cyclin D1 in the nucleus (Table 2); in particular, one case (A6) with downregulated miR-383 expression and upregulated IRF1 protein expression in the testis, exhibited a markedly enhanced nuclear staining for all these proteins (Figure 8). These results indicate that dysregulation of the miR-383-mediated pRb pathways may contribute to spermatogenic failure in MA patients.
Discussion
The completion of spermatogenesis depends on not only the proliferative activity of spermatogonia but also the loss of germ cells during meiosis and spermiogenesis.4 In this study, we found that downregulation of miR-383 and upregulated IRF1 expressions are associated with increased mitotic activity of germ cells in patients with mixed patterns of MA. In addition, the miR-383-IRF1-pRb pathways are identified in the regulation of NT2 cell survivals. We provide, for the first time, mechanistic insights into the role of miR-383 in spermatogenesis and in NT2 cell proliferation.
We have reported that miR-224 can promote TGF-β1-induced germ cell proliferation,23 and that miR-383 might regulate germ cell proliferation/death at early stages of spermatogenesis.13 In this study, miR-383 acts as a negative regulator of proliferation in NT2 cells; in addition, downregulation of miR-383 in germ cells was observed in all our MA patients. These results indicate that miR-383 could contribute to spermatogenic arrest caused by the dysregulation of germ cell proliferation. Furthermore, miR-383 was also expressed at high levels in primary spermatocytes from normal human testes, suggesting a role for miR-383 in meiosis. We found that both CDK2 and CDK4 were regulated by miR-383 in NT2 cells, whereas CDK4 and CDK2 were colocalized with RPA (a marker for early meiotic recombination nodules) and MLH1 (a marker for late recombination nodules) on the mouse synaptonemal complex, respectively.24 Whether miR-383 is involved in meiotic recombination in human germ cells requires further investigation.
miRNAs are known to control diverse biological processes through the regulation of target genes. In this study, we demonstrate that IRF1 is a downstream target of miR-383 and mediates miR-383-induced antiproliferative effects in vitro. An inverse correlation between miR-383 and IRF1 protein levels was also identified in the testes of 80% (4/5) of MA patients. IRF1 has been identified as a tumor suppressor and tumor susceptibility gene. It can inhibit cell growth, cause G1-phase cell-cycle arrest and promote apoptosis.15, 25 However, in this study, we suggest a pro-mitogenic role for IRF1 in NT2 cells. In addition, MA patients with reduced levels of miR-383 and high proliferative activity of germ cells showed upregulation of IRF1. These results indicate that IRF1 may also act as a survival factor for germ cells both in vitro and in vivo. This novel role has also been demonstrated in vivo by Chapman et al.,26 who observed that the loss of IRF1 in mice resulted in accelerated apoptosis of epithelial cells leading to premature involution after weaning.
Over the past few years, several studies have focused on the role of IRF1 in the immune system and in cancer.15, 17, 25, 27 However, a function for IRF1 in gametogenesis has not yet been investigated. In this study, IRF1 expression was gradually increased after birth and reached a maximum at 5 dpp, at which time spermatogonia start to undergo sequential mitotic divisions before the initiation of meiosis.28 IRF1 levels were also markedly elevated soon after serum stimulation of growth-arrested NT2 cells. On the contrary, miR-383 was decreased after birth and reached the lowest levels at 14 dpp, at which time testicular cell proliferation and expansion occur at the highest level.28 Furthermore, a significant negative correlation between the expression levels of mature miR-383 and IRF1 protein was observed in the mouse testis from 0 to 8 dpp, further indicating that IRF1 is a target of miR-383. These results suggest that the pro-mitogenic role of IRF1 may be limited to only the spermatogonial stage.
Notably, we also found that miR-383 influenced the expression of proteins required for the G1/S transition phase, the cyclin D1–CDK4–p21 complex and CDK2. Moreover, the downregulation of miR-383 expression is accompanied by enhanced nuclear staining for P-pRb in all MA patients. In the early G1 phase, the cyclin D1–CDK4 complex partially activates pRb by phosphorylation, resulting in the inhibition of pRb function; in the late G1 phase, CDK2 complexed with cyclin E further phosphorylates pRb to complete its inactivation and allows the cell to enter the S phase.29, 30 Phosphorylations of pRb mediated by the cyclin D1–CDK4 complex and CDK2 are required for cell-cycle re-entry.31 Our data indicate that defection in the miR-383-pRb pathway may be a cause for proliferative dysfunction of germ cells in MA patients.
Cyclin D1 has an important role in G1/S phase transition.32 In this study, we found that among the targets of IRF1, cyclin D1 is a major regulator responsible for miR-383-mediated cell-cycle effects on NT2 cells. Although no cyclin D1 expression is detected in adult human male germ cells in the current study, as also noticed by other researchers,33, 34 nuclear cyclin D1 expression was observed in germ cells of 75% (3/4) MA patients. Ikumasa et al.35 also found that 44.4% (4/9) of MA patients showed a cyclin D1 expression in germ cells. Although miR-383-mediated effects were partially dependent on p21 in vitro, p21 was rarely expressed in the testes of both MA patients and normal controls, indicating a p21-independent pathway mediating spermatogenesis. However, in contrast to the expression of cyclin D1 and p21, CDK4 expression was observed in the testis of all MA patients and normal controls. Combined with the observation that adult germ cells proliferate through a cyclin D1-independent pathway,33 our results suggest that the effect of miR-383 on pRb phosphorylation in the human testis may be mainly mediated by the activated CDK4 expression.
The existence of the widespread expression of P-pRb in the testes of MA patients indicates a status of ‘physiological absence of pRb’ in infertile men. The loss of pRb is highly related to the occurrence of TGCTs.36, 37 Recent reports have shown strong evidence that infertile men are nearly three times more likely to develop TGCTs than are those who are fertile;38 however, the mechanisms linking infertility and TGCT remain uncertain.39 Whether the activated P-pRb expression accompanied by downregulation of miR-383 in infertile testes drives the association between infertility and TGCT is warranted.
In summary, we demonstrate that miR-383 functions as a negative regulator of cell proliferation by directly targeting IRF1. Defective miR-383-pRb pathways are associated with spermatogeneic arrest. The mechanistic studies regarding the role of miRNAs in spermatogenesis will provide new clues to treating male infertility or, on the opposite, designing new male contraceptives. In addition, our results also propose a potential pathway connecting male infertility with TGCT.
Materials and Methods
Mouse and human testicular samples
ICR mice were obtained from the Animal Center, University of Science and Technology of China (USTC, Hefei, China). All animal experiments were conducted following the guidelines for the care and use of laboratory animals from the USTC.
Testicular biopsy specimens obtained from 10 patients (aged 20–35 years) with NOA and from 2 control subjects (aged 28–69 years) were obtained from the First Affiliated Hospital of Anhui Medical University (Hefei, China). Each testicular specimen was divided into several portions for different experiments. Some tissues were too small to complete all experiments. The conditions for specimen use are summarized in Table 2. According to standard pathological criteria, eight patients with mixed pattern of MA, one patient with hypospermatogenesis and one patient with SCO syndrome were recruited for this study. The mixed pattern of MA is characterized by various spermatogenic impairments in adjacent seminiferous tubules showing both SCO and spermatogenic arrest at the level of spermatogonia or primary spermatocytes. As normal controls, the optimal study population would consist of volunteers of known fertility, but difficulties in acquiring testicular materials make this impractical. Instead, testicular samples were obtained from patients undergoing orchiectomy for prostate carcinoma before chemotherapy. None of them had a history of meiotic defects or infertility, and histological examination revealed normal spermatogenesis. All patients gave informed consent, and this study received ethical approval from the Institutional Review Boards of the USTC and the Anhui Medical University.
Vectors
The psiCHECK-2 dual luciferase reporter vector was kindly provided by Biliang Zhang (Guangzhou Institute of Biomedicine and Health, Chinese Academy of Sciences, China), and the pcDNA3.1 vector was kindly provided by Mian Wu (USTC, China). The cyclin D1 expression vector was constructed by cloning human cyclin D1 cDNAs into the pcDNA3.1 vector at the BamHI and XhoI sites. For construction of luciferase reporter plasmids, WT IRF1 3′ UTR was obtained by amplifying a 790-bp 3′ UTR fragment of IRF1 harboring the miR-383-binding site predicted by the TargetScan (http://www.targetscan.org/), whereas mutated IRF1 3′ UTR was generated by PCR-based site-directed mutagenesis. WT and mutated IRF1 3′ UTR, as well as full-length CDK4 3′ UTR sequences were inserted into the XhoI and NotI sites of the psiCHECK-2 reporter vector immediately downstream of the stop codon of an SV40 promotor-driven Renilla luciferase gene. Primer sequences are listed in Supplementary Table 1.
Oligonucleotides
siRNA duplex homologs in sequence with the miRNA-383 mimic and miR-383 inhibitor, as well as scramble NCs, were synthesized and purified by Shanghai Gene-Pharma Co. (Shanghai, China). The miR-383 inhibitor is a single-stranded 2-O-methyl (2-OMe)-modified oligoribonucleotide fragment exactly antisense to miR-383. siRNAs (si-IRF1, si-cyclin D1, si-CDK2, si-p21) were obtained from Santa Cruz Biotechnology Inc. (Santa Cruz, CA, USA). For ISH assay, 5′ end digoxigenin-labeled locked nucleic acid (LNA)-modified DNA oligonucleotides complementary to mature miR-383 (LNA miR-383) and scramble NCs (LNA scrambled) were supplied by Exiqon A/S (Vedbaek, Denmark). Oligonucleotide sequences are provided in Supplementary Table 2.
Histological analysis, immunohistochemistry and ISH
Tissues were dissected out, fixed with 4% paraformaldehyde, embedded in paraffin wax and sectioned at 4 μm. The sections were then used for histological evaluation, immunohistochemical detection of protein expression and TUNEL assay. For histological evaluation, sections were stained with hematoxylin and eosin using standard techniques.
For protein immunolocalization in human testes, sections were microwave heated in 10 mM sodium citrate buffer, pH 6.0 for 10 min after deparaffinization and were then treated with 3% H2O2 in phosphate-buffered saline (PBS) containing 0.1% Triton X-100 to inhibit endogenous peroxidase. After blocking nonspecific-binding sites with 10% normal donkey serum (Jackson ImmunoResearch Labs Inc., West Grove, PA, USA), sections were then incubated with primary antibodies overnight at 4°C and secondary antibodies for 2 h at room temperature, respectively. Immunosignals were visualized using streptavidin–peroxidase and 3, 3′-diaminobenzidine (Maixin Bio, Fuzhou, China). The primary and secondary antibodies used in this study are described in Supplementary Tables 3 and 4, respectively.
For the detection of apoptosis in tissues, sections were subjected to biotin–streptavidin–horseradish peroxidase labeling of DNA strand breaks by TUNEL, using the DeadEnd Colorimetric TUNEL System (Promega, Madison, WI, USA) according to the manufacturer's protocol. As a NC, the TdT enzyme was omitted in parallel reactions.
ISH was performed on 10-μm frozen tissue sections using LNA-modified DNA probes, as described previously.13 The probe sequences are listed in Supplementary Table 2. In brief, 10 μm testis biopsy sections obtained from normal controls and NOA patients were fixed with 4% paraformaldehyde for 15 min at room temperature. To block endogenous alkaline phosphatase activity, slides were immersed and stirred gently in 0.1 M ethanolamine and 2.5% acetic anhydride for 10 min, followed by treatment with 5 μg/ml proteinase K for 3 min after extensive washing with PBS. Prehybridizations were performed for 6 h in a hybridization oven at a temperature between 21 and 23°C below the reported melting temperature of the LNAs (57°C) with 700 μl prehybridization buffer (50% formamide, 5 × SSC, 5 × Denhardt’s, 200 μg/ml yeast RNA, 500 μg/ml salmon sperm DNA, 2% Roche blocking reagents (Roche, Basel, Switzerland) and DEPC-treated water). A 1 pmol probe was added to 150 μl denaturizing hybridization buffer (50% formamide, 5 × SSC, 5 × Denhardt’s, 200 μg/ml yeast RNA, 500 μg/ml salmon sperm DNA, 2% Roche blocking reagents, 0.25% CHAPS, 0.1% Tween and DEPC-treated water). After denatured at 80°C for 5 min, hybridization occurred overnight at the prehybridization temperature-covered glass coverslips. To remove the coverslips, slides were soaked in pre-warmed 60°C 5 × SSC. After incubation in 0.2 × SSC at 60°C for 1 h, sections were washed in B1 solution (0.1 M Tris pH 7.5/0.15 M NaCl) at room temperature for 10 min. After blocking for 1 h in 20% sheep serum (Santa Cruz Biotechnology Inc.) diluted with B1 solution, the sections were incubated overnight at 4°C 10% sheep serum containing anti-Digoxigenin-AP FAB fragments (Roche; 1:250). After washing three times for 5 min in the B1 solution at room temperature and equilibrated for 10 min in B3 solution (0.1 M Tris pH 9.5/0.1 M NaCl/50 mM MgCl2), the sections were stained with NBT/BCIP (Roche) overnight at room temperature. When each probe had yielded a strong signal, or until the NCs began to show background label, reactions were stopped by washing with PBS. Signals were visualized under a standard light microscopy.
Cell culture and transfection
NTERA-2 (NT2) and HEK 293T cells were grown in Dulbecco's modified Eagle's medium supplemented with 10% fetal bovine serum (Life Technologies Inc., Grand Island, NY, USA) and 1% antibiotics (100 Units/ml penicillin and 100 μg/ml streptomycin; Life Technologies Inc.) and cultured at 37°C with 5% CO2. Transfection of 293T cells was performed by Lipofectamine 2000 Reagent (Invitrogen, Carlsbad, CA, USA). Lipofectamine RNAiMAX (Invitrogen) and Fugene HD (Roche) were used for the transfection of oligonucleotides and plasmids into NT2 cells, respectively. The transfection procedure was performed according to the manufacturer's instructions. Unless otherwise indicated, cells were transfected with 30 nM siRNAs, 100 nM miR-383 mimic or 150 nM miR-383 inhibitor. For serum stimulation experiments, NT2 cells were growth arrested for 24 h in serum-free medium and DMEM containing 10% FBS was then added. Cells were harvested at the indicated time points.
RT-PCR and real-time PCR
Total RNA was extracted using the SV Total RNA Isolation System (Promega), and semi-quantitative reverse transcriptase (RT)-PCR was performed using the PrimeScript One-Step RT-PCR Kit Version 2 (TaKaRa Bio Inc., Otsu, Japan). Relative quantification in real-time PCR was performed on ABI Step One System (Applied Biosystems, Foster City, CA, USA), using the SYBR Premix EX Taq kit (TaKaRa Bio Inc.), following the manufacturer's protocols. For pri-miRNA real-time PCR analysis, primers were designed to sit down ∼170 bp 5′ and 3′ to the predicted miRNA stem loop precursor, as described by Cai et al.,40 and the downstream primer was used for RT reaction as well. mRNA levels were normalized to human β-actin mRNA or mouse GAPDH mRNA. Primer sequences for RT-PCR and real-time PCR are provided in Supplementary Tables 5 and 6, respectively.
For real-time PCR detection of miR-383, small RNAs were isolated using the mirPremier miRNA isolation kit (Sigma-Aldrich, St. Louis, MO, USA), and the miR-383 expression was quantified using TaqMan miRNA assays (Applied Biosystems) according to the manufacturer’ instructions. U6 snRNA (Applied Biosystems) was used for normalization. The threshold cycle (Ct) is defined as the fractional cycle number at which the fluorescence passes the fixed threshold. Each PCR reaction was performed in triplicate.
Western blotting
Cells were lysed in RIPA buffer (50 mM Tris-HCl, pH 7.4, 150 mM NaCl, 1% Triton X-100, 1% sodium dodecyl sulfate, 1% sodium deoxycholate, 1 mM EDTA) together with complete EDTA-free protease inhibitor cocktail (Roche), 1 mM phenylmethylsulfonyl fluoride and phosphatase inhibitors (5 mM sodium orthovanadate). Tissue lysates were prepared using the KeyGen total protein extraction kit (KeyGEN, Nanjing, China) according to the manufacturer's instructions. Protein lysates were resolved by SDS-PAGE, transferred to Hybond enhanced chemiluminescence (ECL) Nitrocellulose membrane (Amersham Biosciences, Freiburg, Germany), immunoblotted with antibodies and visualized by ECL (Kodak, Rochester, NY, USA). Protein levels were normalized to GAPDH and quantified using the Scion Image program 4.0.3.2 (Scion Corporation, Frederick, MD, USA). The proteasome inhibitor MG-132 was purchased from Sigma-Aldrich. The primary and secondary antibodies used for immunoblotting are listed in Supplementary Tables 3 and 4, respectively.
Cell proliferation assay
Cell proliferation was measured using cell counting kit-8 (CCK-8) (Dojindo Laboratories, Kumamoto, Japan). NT2 cells were plated at 3–5 × 103 cells/well in 96-well plates with 5 replicate wells for each condition. Forty-eight hours after transfection, 10 μl of CCK-8 solution was added to the medium and then incubated at 37°C for 2 h. Cell numbers were measured by measuring the absorbance at 450 nm using a 96-well format plate reader (ELx800 Universal Microplate Reader; Biotek Instrument Inc., Highland Park, VT, USA).
Flow cytometric analysis
Cell-cycle regulation and apoptosis were analyzed using the FACScalibur flow cytometer (BD Biosciences, San Jose, CA, USA). Data analysis was carried out using WinMDI 2.9 software (The Scripps Research Institute, La Jolla, CA, USA). For cell-cycle analysis, cells were seeded in 6-well or 12-well plates at 60–70% confluency. Twenty-four hours after transfection with the indicated oligonucleotides, NT2 cells were harvested, washed in PBS and fixed overnight in 70% ethanol. The next day, cells were collected, washed in PBS, incubated at 37°C for 10 min with PBS containing 100 μg/ml RNase A and 25 μg/ml propidium iodide (PI) (Sigma-Aldrich) and analyzed by flow cytometry. For cell apoptosis analysis, cells were seeded in 6-well or 12-well plates at 30–50% confluency. Cells were harvested at 72 h after transfection with 80 nM indicated siRNAs, 100 nM miR-383 mimic or 150 nM miR-383 inhibitor, stained using the Annexin V FITC Apoptosis Detection Kit (Beijing Biosea Biotechnology Co., Ltd, Beijing, China) and PI staining solution, and analyzed by flow cytometry.
Luciferase reporter assay
293T cells were cotransfected with either 20 pmol miR-383 mimic or 30 pmol miR-383 inhibitor and 200 ng psiCHECK-2 vectors in 24-well plates with three replicate wells for each condition. Luciferase activity was measured 30 h after transfection using the Dual Luciferase Reporter Assay System (Promega). Renilla luciferase activity was normalized to Firefly luciferase activity for each transfected well. All experiments were performed in triplicate.
Immunofluorescence analysis
See Supplementary Methods for details.
Statistics
Student's t-test was applied to examine the differences among variables. Data were shown as mean±S.E.M. unless otherwise indicated. The relationships between pri-miR-383 and mature miR-383 expressions in human testes, and those between mature miR-383 and IRF1 protein expressions in the mouse testes were evaluated using Pearson's correlation analysis. P-values <0.05 were considered to be statistically significant.
Abbreviations
- CCK-8:
-
cell counting kit-8
- CK-18:
-
cytokeratin-18
- Ct:
-
threshold cycle
- dpp:
-
days postpartum
- IRF1:
-
interferon regulatory factor-1
- ISH:
-
in situ hybridization
- LNA:
-
locked nucleic acid
- MA:
-
maturation arrest
- miRNA:
-
microRNA
- MT:
-
mutation type
- NC:
-
negative control
- NOA:
-
nonobstructive azoospermia
- NT2:
-
NTERA-2
- PCNA:
-
proliferating cell nuclear antigen
- PI:
-
propidium iodide
- P-pRb:
-
phosphorylated retinoblastoma protein
- pri-miRNA:
-
primary microRNA
- SCO:
-
Sertoli-cell-only
- UTR:
-
untranslated region
- WT:
-
wild type
References
Matzuk MM, Lamb DJ . Genetic dissection of mammalian fertility pathways. Nat Cell Biol 2002; 4 (Suppl): s41–s49.
Hung AJ, King P, Schlegel PN . Uniform testicular maturation arrest: a unique subset of men with nonobstructive azoospermia. J Urol 2007; 178: 608–612; discussion 612.
Lin WW, Lamb DJ, Lipshultz LI, Kim ED . Demonstration of testicular apoptosis in human male infertility states using a DNA laddering technique. Int Urol Nephrol 1999; 31: 361–370.
Steger K, Aleithe I, Behre H, Bergmann M . The proliferation of spermatogonia in normal and pathological human seminiferous epithelium: an immunohistochemical study using monoclonal antibodies against Ki-67 protein and proliferating cell nuclear antigen. Mol Hum Reprod 1998; 4: 227–233.
Bar-Shira Maymon B, Yogev L, Yavetz H, Lifschitz-Mercer B, Schreiber L, Kleiman SE et al. Spermatogonial proliferation patterns in men with azoospermia of different etiologies. Fertil Steril 2003; 80: 1175–1180.
Gonsalves J, Sun F, Schlegel PN, Turek PJ, Hopps CV, Greene C et al. Defective recombination in infertile men. Hum Mol Genet 2004; 13: 2875–2883.
Sun F, Kozak G, Scott S, Trpkov K, Ko E, Mikhaail-Philips M et al. Meiotic defects in a man with non-obstructive azoospermia: case report. Hum Reprod 2004; 19: 1770–1773.
Braun RE . Post-transcriptional control of gene expression during spermatogenesis. Semin Cell Dev Biol 1998; 9: 483–489.
Bartel DP . MicroRNAs: target recognition and regulatory functions. Cell 2009; 136: 215–233.
Hayashi K, Chuva de Sousa Lopes SM, Kaneda M, Tang F, Hajkova P, Lao K et al. MicroRNA biogenesis is required for mouse primordial germ cell development and spermatogenesis. PLoS One 2008; 3: e1738.
Maatouk DM, Loveland KL, McManus MT, Moore K, Harfe BD . Dicer1 is required for differentiation of the mouse male germline. Biol Reprod 2008; 79: 696–703.
Voorhoeve PM, le Sage C, Schrier M, Gillis AJ, Stoop H, Nagel R et al. A genetic screen implicates miRNA-372 and miRNA-373 as oncogenes in testicular germ cell tumors. Cell 2006; 124: 1169–1181.
Lian J, Zhang X, Tian H, Liang N, Wang Y, Liang C et al. Altered microRNA expression in patients with non-obstructive azoospermia. Reprod Biol Endocrinol 2009; 7: 13.
Bergmann M, Kliesch S . The distribution pattern of cytokeratin and vimentin immunoreactivity in testicular biopsies of infertile men. Anat Embryol (Berl) 1994; 190: 515–520.
Kroger A, Koster M, Schroeder K, Hauser H, Mueller PP . Activities of IRF-1. J Interferon Cytokine Res 2002; 22: 5–14.
Harada H, Kitagawa M, Tanaka N, Yamamoto H, Harada K, Ishihara M et al. Anti-oncogenic and oncogenic potentials of interferon regulatory factors-1 and -2. Science 1993; 259: 971–974.
Romeo G, Fiorucci G, Chiantore MV, Percario ZA, Vannucchi S, Affabris E . IRF-1 as a negative regulator of cell proliferation. J Interferon Cytokine Res 2002; 22: 39–47.
Kroger A, Stirnweiss A, Pulverer JE, Klages K, Grashoff M, Reimann J et al. Tumor suppression by IFN regulatory factor-1 is mediated by transcriptional down-regulation of cyclin D1. Cancer Res 2007; 67: 2972–2981.
Xie RL, Gupta S, Miele A, Shiffman D, Stein JL, Stein GS et al. The tumor suppressor interferon regulatory factor 1 interferes with SP1 activation to repress the human CDK2 promoter. J Biol Chem 2003; 278: 26589–26596.
Tanaka N, Ishihara M, Lamphier MS, Nozawa H, Matsuyama T, Mak TW et al. Cooperation of the tumour suppressors IRF-1 and p53 in response to DNA damage. Nature 1996; 382: 816–818.
LaBaer J, Garrett MD, Stevenson LF, Slingerland JM, Sandhu C, Chou HS et al. New functional activities for the p21 family of CDK inhibitors. Genes Dev 1997; 11: 847–862.
Brown KA, Roberts RL, Arteaga CL, Law BK . Transforming growth factor-beta induces Cdk2 relocalization to the cytoplasm coincident with dephosphorylation of retinoblastoma tumor suppressor protein. Breast Cancer Res 2004; 6: R130–R139.
Yao G, Yin M, Lian J, Tian H, Liu L, Li X et al. MicroRNA-224 is involved in transforming growth factor-beta-mediated mouse granulosa cell proliferation and granulosa cell function by targeting Smad4. Mol Endocrinol 2010; 24: 540–551.
Ashley T, Walpita D, de Rooij DG . Localization of two mammalian cyclin dependent kinases during mammalian meiosis. J Cell Sci 2001; 114: 685–693.
Tanaka N, Ishihara M, Taniguchi T . Suppression of c-myc or fosB-induced cell transformation by the transcription factor IRF-1. Cancer Lett 1994; 83: 191–196.
Chapman RS, Duff EK, Lourenco PC, Tonner E, Flint DJ, Clarke AR et al. A novel role for IRF-1 as a suppressor of apoptosis. Oncogene 2000; 19: 6386–6391.
Matsuyama T, Kimura T, Kitagawa M, Pfeffer K, Kawakami T, Watanabe N et al. Targeted disruption of IRF-1 or IRF-2 results in abnormal type I IFN gene induction and aberrant lymphocyte development. Cell 1993; 75: 83–97.
Jarvis S, Elliott DJ, Morgan D, Winston R, Readhead C . Molecular markers for the assessment of postnatal male germ cell development in the mouse. Hum Reprod 2005; 20: 108–116.
Weinberg RA . The retinoblastoma protein and cell cycle control. Cell 1995; 81: 323–330.
Sherr CJ . G1 phase progression: cycling on cue. Cell 1994; 79: 551–555.
Guo J, Sheng G, Warner BW . Epidermal growth factor-induced rapid retinoblastoma phosphorylation at Ser780 and Ser795 is mediated by ERK1/2 in small intestine epithelial cells. J Biol Chem 2005; 280: 35992–35998.
Fu M, Wang C, Li Z, Sakamaki T, Pestell RG . Minireview: cyclin D1: normal and abnormal functions. Endocrinology 2004; 145: 5439–5447.
Bartkova J, Rajpert-de Meyts E, Skakkebaek NE, Bartek J . D-type cyclins in adult human testis and testicular cancer: relation to cell type, proliferation, differentiation, and malignancy. J Pathol 1999; 187: 573–581.
Houldsworth J, Reuter V, Bosl GJ, Chaganti RS . Aberrant expression of cyclin D2 is an early event in human male germ cell tumorigenesis. Cell Growth Differ 1997; 8: 293–299.
Ikumasa TXZ, Yoshihiro K, Nobufumi U . The expression of cyclin D1 in the human testicular tissue. I. Immunohistochemical detection of cyclin D1 in the testicular tissue of infertile patients. Nishinihon J Urol 2001; 63: 79–82.
Strohmeyer T, Reissmann P, Cordon-Cardo C, Hartmann M, Ackermann R, Slamon D . Correlation between retinoblastoma gene expression and differentiation in human testicular tumors. Proc Natl Acad Sci USA 1991; 88: 6662–6666.
Bartkova J, Lukas C, Sorensen CS, Meyts ER, Skakkebaek NE, Lukas J et al. Deregulation of the RB pathway in human testicular germ cell tumours. J Pathol 2003; 200: 149–156.
Walsh TJ, Croughan MS, Schembri M, Chan JM, Turek PJ . Increased risk of testicular germ cell cancer among infertile men. Arch Intern Med 2009; 169: 351–356.
Hotaling JM, Walsh TJ . Male infertility: a risk factor for testicular cancer. Nat Rev Urol 2009; 6: 550–556.
Cai X, Hagedorn CH, Cullen BR . Human microRNAs are processed from capped, polyadenylated transcripts that can also function as mRNAs. RNA 2004; 10: 1957–1966.
Acknowledgements
We thank Dr. Mian Wu for the HEK293T cell line. This work was supported by the following grants (to FS): the National Basic Research Program of China (2009CB941700 and 2007CB947400) and the National Natural Science Foundation of China (30770811 and 30971091).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by D Bano
Supplementary Information accompanies the paper on Cell Death and Disease website
Supplementary information
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Lian, J., Tian, H., Liu, L. et al. Downregulation of microRNA-383 is associated with male infertility and promotes testicular embryonal carcinoma cell proliferation by targeting IRF1. Cell Death Dis 1, e94 (2010). https://doi.org/10.1038/cddis.2010.70
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2010.70
Keywords
This article is cited by
-
miRNA-383-5p Regulated Migration and Invasion of Tumor Cells by Inhibiting NCKAP1 Expression in Gastric Cancer
Biochemical Genetics (2024)
-
MicroRNA-383 located in frequently deleted chromosomal locus 8p22 regulates CD44 in prostate cancer
Oncogene (2017)
-
The Value of MiR-383, an Intronic MiRNA, as a Diagnostic and Prognostic Biomarker in Intestinal-Type Gastric Cancer
Biochemical Genetics (2017)
-
SMAR1 binds to T(C/G) repeat and inhibits tumor progression by regulating miR-371-373 cluster
Scientific Reports (2016)
-
miR-383 inhibits hepatocellular carcinoma cell proliferation via targeting APRIL
Tumor Biology (2016)