Abstract
The tumour necrosis factor family member TNF-related apoptosis-inducing ligand (TRAIL) selectively induces apoptosis in a variety of cancer cells through the activation of death receptors 4 (DR4) and 5 (DR5) and is considered a promising anticancer therapeutic agent. As apoptosis seems to occur primarily via only one of the two death receptors in many cancer cells, the introduction of DR selectivity is thought to create more potent TRAIL agonists with superior therapeutic properties. By use of a computer-aided structure-based design followed by rational combination of mutations, we obtained variants that signal exclusively via DR4. Besides an enhanced selectivity, these TRAIL-DR4 agonists show superior affinity to DR4, and a high apoptosis-inducing activity against several TRAIL-sensitive and -resistant cancer cell lines in vitro. Intriguingly, combined treatment of the DR4-selective variant and a DR5-selective TRAIL variant in cancer cell lines signalling by both death receptors leads to a significant increase in activity when compared with wild-type rhTRAIL or each single rhTRAIL variant. Our results suggest that TRAIL induced apoptosis via high-affinity and rapid-selective homotrimerization of each DR represent an important step towards an efficient cancer treatment.
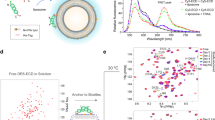
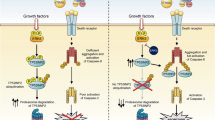
Similar content being viewed by others
Main
Tumour necrosis factor (TNF)-related apoptosis-inducing ligand (TRAIL) is a promiscuous member of the TNF superfamily capable of binding to five different receptors. TRAIL induces apoptosis via the death receptors (DRs) 4 and 5,1, 2 followed by recruitment of the Fas-associated death domain.3, 4, 5 Formation of the death-inducing signalling complex (DISC) is completed by binding of procaspase-8 and -10, leading to their cleavage and activation, that in turn activate downstream caspases (e.g. caspase-3), resulting in apoptosis.6, 7, 8 TRAIL binds also to three decoy receptors, DcR1, DcR2 and OPG.9 Decoy receptors lack a functional death domain, and consequently they do not induce apoptosis and can prevent TRAIL binding to death receptors.2, 9, 10, 11
TRAIL is generating considerable interest as a possible anticancer therapeutic agent because of its selective activation of apoptosis in cancer cells.12, 13 The formation of heterotrimeric non-apoptotic TRAIL–receptor complexes7, 14, 15 and the receptor binding promiscuity of TRAIL endorsed the concept that DR selectivity may allow the creation of more potent TRAIL apoptosis inducers.16, 17, 18, 19 DR5 was described as contributing more than DR4 to the overall apoptotic activity of TRAIL in apoptosis signalling of cancer cells.16, 20, 21 Recently, DR4 has gained increased importance, with the realization that certain cancer cells depend mainly on this receptor for apoptosis via TRAIL.17, 18, 22 Whereas potent DR5-selective variants were created previously,16, 19 no enhancement of apoptosis by selective activation of DR4 has been accomplished by using TRAIL variants. A previous generation of DR4-selective variants designed by us18 or by others17 were either equally active when compared with rhTRAILWT or largely inactive.16, 17
Here we report the design of highly active, DR4-specific TRAIL variants by using computational protein design and a rational combination strategy. This resulted in the first DR4-selective variants that not only have an enhanced selectivity to DR4, but also a significant increase in apoptosis-inducing activity in many cancer cells in vitro. Both receptor selectivity and increased affinity for DR4 appear to contribute to the more efficient apoptosis induction by these TRAIL variants.
These DR4-selective variants and the previously described DR5-selective variants19 allowed us to establish the contribution of DRs in apoptosis signalling of different cancer cells. Cell lines signalling via both DRs showed a large increase in efficacy upon combined treatment with selective TRAIL variants when compared with rhTRAILWT. Our results show these novel DR4-specific variants alone or in combination with a DR5-specific variant as a powerful strategy for targeting DR4/DR5-responsive cancer cells.
Results
Computational design of variants using FoldX
A refined model of the DR4 receptor was created using FoldX. All residues in the TRAIL–receptor interface were considered. A set of predicted energetic values for TRAIL–receptor complex formation was obtained and compared with the TRAILWT values. FoldX proposed the following mutations to favour DR4 selectivity: R149I, R149M, R149N, R149K, S159R, Q193H, Q193K, N199R, K201R, K204D, K204E, K204L, K204Y, K212R, S215D, S215E, S215H, S215K, K251D, K251E and K251Q. Mutations at position 218 were constructed before,18 and were therefore not considered.
DR4-selectivity screening
RhTRAIL variants were expressed and purified as described before,19, 23 and variants were characterized by surface plasmon resonance (SPR). Apoptosis-induction assays were performed in cell lines mediating apoptosis primarily via DR4 or DR5. Mutations at positions 149, 159, 199, 201 and 215 showed increased selectivity to DR4 as predicted by FoldX (Supplementary Figure S1), whereas mutations at positions 193, 204, 212 and 251 did not show the expected changes. Interestingly, positions 199, 201 and 215 have been found before in DR4 variants selected by phage display.16 Variants showing an increase in binding ratio of DR4 over DR5 (DR4/DR5) of more than fourfold as measured by pre-steady state SPR (Supplementary Table 1), and/or presenting a more than threefold increased activity on DR4-mediated cell lines, and showing inactivity in DR5-mediated cell lines were chosen for further studies. R149I, S159R and S215D were selected.
Rational combination of single variants
Several amino-acid substitutions resulted in high DR4/DR5 binding ratios relative to rhTRAILWT, thus favouring DR4 binding. The characteristics of these single mutants were used for rational design of nine combination variants, containing between two and six mutations each. A single mutant (G131R) constructed previously23 showing increased affinity to both DRs and a double mutant (N199R; K201H), showing a decrease in affinity to DR5 have been combined before (to be published elsewhere), and were therefore incorporated in the new combination variants. Mutation S159R was incorporated in all the combination variants, showing a high affinity ratio change (Supplementary Table 1). Mutants displaying similar or higher binding affinities to DR4, together with a lowered affinity for DR5, were characterized. Mutations that resulted in more than 50-fold increased DR4/DR5 binding ratio compared with rhTRAILWT as measured by SPR were selected. Two combination variants showing the highest affinity increase to DR4 and affinity decrease for DR5 were selected for further studies. These variants were named 4C7 (G131R/R149I/S159R/N199R/K201H/S215D) and 4C9 (G131R/R149I/S159R/S215D).
Death receptor selectivity of 4C7 and 4C9
Binding of rhTRAILWT and variants 4C7 and 4C9 to DRs using SPR showed an increased affinity to DR4-Ig with a 9–10-fold lower dissociation constant (KD) when compared with rhTRAILWT (Table 1). In contrast, 4C7 and 4C9 showed a much lower affinity to DR5-Ig with regard to rhTRAILWT with a 40- and 230-fold increase in dissociation constant to DR5-Ig for 4C9 and 4C7, respectively (Table 1). DR4-Ig competed with a 5–7-fold increased effect for the variants 4C7 and 4C9 when compared with rhTRAILWT (Figure 1a). Inversely, DR5-Ig was less capable to compete for 4C7 and 4C9 binding to immobilized DR4-Ig (Figure 1b). A 29-fold increased binding of 4C9 to immobilized DR4-Ig was observed compared with rhTRAILWT when competing with DR5-Ig, indicating a preference to bind to DR4. 4C7 was even less sensitive to competition by DR5-Ig, with 80% binding efficiency for DR4 even when competing with 5000 ng/ml of DR5-Ig. These results indicate that the variants 4C7 and 4C9 bind highly preferentially to DR4.
Competitive inhibition by TRAIL death receptors for rhTRAILWT and variants 4C7 and 4C9, binding to immobilized DR4-Ig, using soluble DR4-Ig as a competitor (a) and soluble DR5-Ig as a competitor (b), in the presence of 100 ng/ml of rhTRAILWT or variants and 0–5000 ng/ml per well of DR4- or DR5-Ig. Binding of rhTRAILWT and variants was measured relative to the wells containing no soluble receptor
Decoy receptors DcR1-Ig and DcR2-Ig were found to equally compete for binding to rhTRAILWT 4C7 and 4C9 (Supplementary Figure S2). Interestingly, another combination variant (4C6: R149I/S159R/S215D) showed not only lowered binding to DR5, but also decreased binding affinity to both decoy receptors in solution (Supplementary Figure S2).
Biological activity of DR4-selective variants
Comparison between single variants and combination variants in BJAB cell lines showed that 4C7 displays a greatly improved capability to induce cell death when compared with rhTRAILWT, to the single variants and to a combination of three mutations (Supplementary Figure S3). To confirm the selectivity of 4C7 and 4C9 in vitro, we tested their activity in different cell lines mediating apoptosis via different DRs. Analysis of cell death induction in Burkitt's lymphoma BJAB cells inducible by both DR4 and DR5 (BJABWT) and BJAB cells deficient in DR5 (BJABDR5 DEF)24 showed that 4C7 and 4C9 do not require DR5 to deliver similar levels of apoptosis in both cell lines. In addition, 4C7 and 4C9 showed a significant increase in cell death induction in BJAB cells when compared with rhTRAILWT in all concentrations measured (Figure 2a and b).
Death receptor 4 selectivity of variants in several cell lines. Cell death induction by wild-type TRAIL and variants on (a) BJAB cell line mediating apoptosis via both DR4 and DR5 receptors (BJABWT), (b) BJAB DR5-deficient cell line (BJABDR5 DEF) mediating apoptosis via DR4, (c) DR5-mediated Jurkat leukaemia and (d) DR5-mediated A2780 ovarian carcinoma cells. BJAB cells were treated with rhTRAILWT and variants for 24 h. The percentage of cell death was calculated relative to control wells containing no ligand. Jurkat leukaemia cells and A2780 ovarian carcinoma cancer cells were treated with the indicated concentrations of rhTRAILWT and variants for 24 h, after which induction of cell death was measured by Annexin V staining. The results are mean values±S.E.M. (n=3)
Cell assays in DR5-mediating A2780 and Jurkat cells showed no apoptosis-inducing activity of the variants (<10%) at almost all concentrations measured (Figure 2c and d). These results support that 4C7 and 4C9 induce apoptosis essentially via DR4. Furthermore, enhanced activity of TRAIL variants 4C7 and 4C9 was shown on a number of colon adenocarcinoma cells sensitive to TRAIL-induced apoptosis when compared with rhTRAILWT, including Colo205, SW948, HCT-15 and DLD-1 (Figure 3a–d).
Several cancer cell lines undergo apoptosis efficiently via TRAIL DR4-selective variants. (a) Colo205, (b) SW948, (c) DLD-1 and (d) HCT-15 colon carcinoma cell lines were incubated with different concentrations of rhTRAILWT and variants 4C7 and 4C9 for 24 h, and assayed for cell viability. The percentage of viable cells was calculated relative to control wells containing no ligand. The results are mean values±S.E.M. (n=3). Death receptor 4-selective variants induce higher levels of apoptosis in MCF-7 human breast adenocarcinoma (e) and PANC-1 human pancreatic carcinoma cells (f). Cells were treated with rhTRAILWT and variants 4C7 and 4C9 for 24 h and cell death was measured by Annexin V staining. The results are mean values±S.E.M. (n=3)
A reduction of 3–5-fold in cell viability of Colo205 by variants 4C7 and 4C9 when compared with rhTRAILWT was observed (Figure 3a). The SW948 cell line showed a strong decrease in cell viability upon treatment with the variants (Figure 3b). Moreover, an impressive decrease in cell viability has been obtained in HCT-15 and DLD-1 cell lines by 4C7 and 4C9 when compared with rhTRAILWT (up to 88% cell death) (Figure 3c and d). Our results indicate that these cell lines can be triggered very efficiently to undergo apoptosis by these novel DR4-specific variants.
We then tested whether selective activation of DR4 would have an impact on cancer cell lines less sensitive to TRAIL-induced apoptosis. Remarkably, our variants 4C7 and 4C9 showed a significant increase in cell killing in breast cancer MCF-7 and pancreatic cancer PANC-1 cell lines (Figure 3e and f). Both cell lines express DR4 and DR5 on their surface (Supplementary Figure S4). Furthermore, the variants were tested for activity on normal cells, namely fibroblasts. No significant activity for rhTRAILWT or variants 4C7 and 4C9 could be observed, suggesting that the increased apoptotic activity on cancer cells had no effect on normal cells (Supplementary Figure S5).
Real-time analysis of caspase activation
The variant 4C7 was subject of study by analysing caspase activation using FRET. Activation of the initiator caspase-8/10 was followed by using a probe based on CFP and YFP (Venus),25 interconnected by a linker containing the preferred cleavage motif for caspase-8 (IETD).26, 27 Owing to its high signal/noise ratio, this probe allows the efficient detection of low caspase activities.25 The 4C7 variant-induced profile of intracellular IETDase activity shows increased cleavage of the CFP-IETD-Venus probe in ovarian carcinoma OVCAR-3 cells at much earlier time points when compared with rhTRAILWT (Figure 4). Cleavage by activated initiator caspases could be observed within 15–20 min after 4C7 addition. More significant differences between rhTRAILWT and 4C7 occur after 50 min of incubation with 50 and 100 ng/ml (Figure 4a and b). Effector caspases have also been shown to cleave IETD recognition sites,27 and consequently a more rapid increase observed after cleavage by activated caspase-8 is most likely owing to the activation of effector caspases during apoptosis or elevated caspase-8 activity, resulting from feedback via caspase-6. Analysis of downstream activation of effector caspase-3/7 based on DEVDase activity using a probe containing CFP and YFP indicates that the rate of cleavage remains low at initial time points, and raises rapidly after 75 min over a 30–40 min period for 4C7 (Figure 4c and d). The rate of DEVDase cleavage for rhTRAILWT was substantially lower. These results indicate a much faster and robust initiator and effector caspase activation by variant 4C7 compared with rhTRAILWT in living cells.
TRAIL-induced IETDase and DEVDase profiles of rhTRAILWT and variant 4C7 in OVCAR-3 ovarian carcinoma cell line. Cells were treated with two different concentrations of rhTRAILWT and 4C7, (a) 50 ng/ml and (b) 100 ng/ml. IETDase activity profile displayed FRET cleavage and consequent increase in CFP/YFP emission ratio is plotted. (c) DEVDase activities displaying FRET cleavage upon addition of 100 ng/ml of rhTRAILWT and variant 4C7. The curves in (a)–(c) represent the analysis of 10 different OVCAR-3 cells, following the TRAIL treatment. Error bars indicate the standard deviations for three single curve profiles using a similar number of cells. (d) CFP/YFP ratio images of a group of OVCAR-3 cells expressing the DEVD FRET probe treated with 100 ng/ml of rhTRAILWT and 4C7 variant
Combination of death receptor-selective variants
In order to establish receptor activity in different cell lines, we tested the new DR4-selective variants or our DR5-selective mutant (D269H/E195R).19 Most cell lines tested undergo apoptosis primarily via one of the DRs; however, some cells did show sensitivity via both DRs. We reasoned that selective activation of both DRs by TRAIL variants may lead to an additive effect in cell death induction and that targeting both DRs separately may be of therapeutic relevance in cancer cell lines. OVCAR-3 cells express both DRs, with almost no detectable levels of decoy receptors (Supplementary Figure S4). Both 4C7 (ED50=6.61±1.3 ng/ml) and 4C9 (ED50=7.86±1.53 ng/ml) caused a substantial decrease in viability when compared with rhTRAILWT (ED50=46.8±1.1 ng/ml), with the DR4-selective variants reaching cell death levels of 88 and 83%, whereas rhTRAILWT was only capable of 50% cell death induction at the highest concentration measured (250 ng/ml) (Figure 5a). Similarly, activation of DR5 by D269H/E195R in this cell line resulted in increased apoptosis induction, with a nearly sevenfold increase in cell death compared with rhTRAILWT. Interestingly, the combination of equimolar concentrations of DR4 and DR5 TRAIL-selective variants resulted in an even lower cell survival at all concentrations measured, when compared with the single treatment with either only rhTRAILWT or selective variants. At the highest concentration used, the combination of selective variants was capable of killing almost 95% of cells upon 24 h incubation. In the Colo205 cell line, the cooperation between DR4- and DR5-selective induction was less pronounced, even though at high concentrations higher levels of cell death could be shown (up to 94%). At these concentrations, also a difference with D269H/E195R could be observed (Figure 5b). In the colon carcinoma CL-34 cell line, both 4C7- and DR5-specific variant showed higher activities than rhTRAILWT at almost all concentrations and when combined induced higher killing levels compared with single treatments, especially at high concentrations (Figure 5c).
DR4- and DR5-specific variants cooperate to induce high apoptosis in different cell lines. (a) OVCAR-3 ovarian carcinoma cell line, (b) Colo205 colon carcinoma and (c) CL-34 colon carcinoma were treated with rhTRAILWT or DR4- and DR5-specific variants (0.9–250 ng/ml) alone and in combination for 24 h and assayed for cell viability. The percentage of viable cells was calculated relative to control wells containing no ligand. The results are mean values±S.E.M. (n=3)
Clonogenic survival assays
To determine the effect of these variants on long-term survival of cancer cells, we performed clonogenic assays (Figure 6). Both DR4- and DR5-selective variants significantly reduced clonogenic growth of OVCAR-3 cells when compared with rhTRAILWT (Figure 6a and b). The combination of the receptor-specific variants results in an increased ability to kill OVCAR-3 cells.
Clonogenic survival and TRAIL-DISC immunoprecipitation. (a and b) Effect of variants of rhTRAIL on the clonogenic survival of ovarian carcinoma OVCAR-3 cells. One hundred (a) and two hundred (b) OVCAR-3 cells were seeded as single cells, and treated with 5 ng/ml of rhTRAILWT or variants for 24 h. After recovering for 14 days, colony formation was determined. (c) Colo205 cells were treated with 250 ng/ml flag-tag rhTRAILWT (WT) or flag-tag rhTRAIL 4C9 (4C9) or left untreated (control) for 15 min. The cells were harvested and the ligated DISC complexes immunoprecipitated using an anti-FLAG antibody. The presence of DR4, DR5 and caspase-8 in the DISC was detected using western blotting. *A nonspecific band detected by the DR5 and caspase-8 antibody
TRAIL-DISC immunoprecipitation
Analysis of the components of the death receptor (DR) DISC formed upon treatment with flag-tag rhTRAILWT and flag-tag rhTRAIL 4C9 indicate the ligation of DR4 and DR5 by flag-tag rhTRAILWT. Only DR4 but not DR5 could be pulled down in the cells treated with flag-tag rhTRAIL 4C9 (Figure 6c), further showing the DR selectivity of these variants. In addition, the activation of procaspase-8 within the 4C9-mediated DISC was more pronounced than in the rhTRAILWT-induced DISC, indicating a more efficient activation of pro-caspase-8 by the DR4-selective variant.
Discussion
TRAIL binds to five different receptors, of which only two are capable of inducing apoptosis, DR4 and DR5. It is generally accepted that decoy receptors can inhibit TRAIL-induced apoptosis because they can bind TRAIL, but lack a functional death domain to form the DISC.2, 9, 11 However, the clarity of this decoy concept has been blurred with the observation that presence of decoy receptors not necessarily translates in protection against apoptosis.21, 28 In a recent study, DcR2 has been reported to heteromerize with DR5 to form inactive complexes.14, 15 Heteromerization of death receptors DR4 and DR5 may also lead to inactive complexes.7, 15 In a large number of cancer cell lines, differential contributions of the DRs have been reported,29 but it is not yet clear whether they differ in their function in the apoptotic signalling. In light of these studies, the introduction of receptor selectivity can contribute to the elucidation of the function of each death receptor. Such variants are expected to have increased in vivo activity and are candidates for new anticancer therapies.
Computational design based on the crystal structure of the TRAIL–DR5 complex allowed the successful introduction of DR5 selectivity by two-point mutations.19 The highly selective variant D269H/E195R was shown to be very efficient to induce apoptosis in an ovarian carcinoma xenograft mouse model.30 For DR4, a highly refined homology model was made, using data from mutational analysis.18, 19, 23 A set of 21 single mutation TRAIL variants predicted to show DR4-selective behaviour were designed, and experimentally characterized. Two of the positions within this set were also reported from a phage display selection study.16 All the 21 mutants were characterized for their ability to bind receptors and to induce apoptosis. Several mutations showed lower affinity for DR5 and higher affinity for DR4. In most cases, these effects could be well explained upon visual inspection of the mutant models.
A representative example is given by the mutation S159R that has a large beneficial effect on the DR4/DR5 binding ratio (Supplementary Table 1). In the WT–TRAIL–DR5 complex, Arg-115 of DR5 forms a salt-bridge with Glu-155 and a hydrogen bond with His-161 of TRAIL (Figure 7b). The hydroxyl group of Ser-159 donates a hydrogen bond to the carboxyl side chain group of Glu-155. Upon mutation of Ser-159 to Arg, the conformation of the side chain of Arg-115 of DR5 is slightly changed and the hydrogen bond interactions with His-161 are destroyed (Figure 7a).
Structural impressions of the area around 159 for S159R and rhTRAILWT as determined by FoldX: (a and b) TRAIL-DR5 and (c and d) TRAIL-DR4. The subunits of TRAIL are depicted in lime and receptors in green. The template selected was 1D4V, the structure at 2.2 Å resolution and of monomeric human TRAIL in complex with the ectodomain of DR5 receptor. The homotrimer was generated using the protein quaternary structure server from the EBI (http://pqs.ebi.ac.uk), having the symmetry coordinates in the PDB file
The hydrogen bond interaction of Arg-115 with His-161 is lost and the salt-bridge with Glu-155 is somewhat weakened. This results in an overall loss of interaction energy for the S159R mutation (ΔΔGinteraction: +1 kcal/mol). On the other hand, in the TRAIL–DR4 complex the equivalent residue of Arg-115 is Pro-115 and, as a consequence, no direct interactions are observed between Glu-155, Ser-159 and His-161 of TRAIL and DR4 receptor residues (Figure 7d). Mutation S159R allows Arg-159 to create a hydrogen bond with the backbone oxygen atom of Cys-113 of DR4 (Figure 7c). As no interactions are destroyed with respect to the WT situation and one new interaction is established, there is a net gain in interaction energy (ΔΔGinteraction: −1.4 kcal/mol). The loss in interaction energy between TRAIL and DR5 on the one hand, and a gain in interaction energy between TRAIL and DR4 upon Ser to Arg mutation on the other hand, explain the observed increase in DR4 binding specificity.
As TRAIL was reported to have a lower affinity for DR4 than to DR5,31 a single mutation seems not sufficient to produce a variant with high affinity and selectivity for DR4. Single mutations were therefore combined based on observed biochemical and biological properties. Two criteria were used for the selection: (1) at least fourfold increase in DR4/DR5 binding ratio based on pre-steady state SPR measurements and (2) at least threefold increased biological activity on DR4-sensitive cancer cells. Nine combinations of the selected mutations were made and most new variants showed affinity improvement to DR4 and largely reduced affinity to DR5. From these, two variants were chosen for further characterization based on their biochemical and apoptosis-inducing characteristics: 4C7 (carrying mutations G131R/N199R/K201H/R149I/S159R/S215D) and 4C9 (carrying G131R/R149I/S159R/S215D). Characterization of the affinities of these two proteins for DR4 and DR5 showed that they have not only a decreased affinity for DR5, but also an increased affinity for DR4 (Table 1). The selective behaviour of these variants was also shown by immunoprecipitation experiments using Colo205 cells, where only DR4 but not DR5 could be detected upon treatment with a flag-tagged version of 4C9 (Figure 6c).
Variants 4C7 and 4C9 showed a remarkable capacity to induce apoptosis in several cancer cells in vitro, including a panel of human colon adenocarcinoma (Colo205, SW948, DLD-1, HCT-15 and CL-34), Burkitt's lymphoma (BJAB), pancreatic carcinoma (PANC-1), the human ovarian carcinoma OVCAR-3 and human breast adenocarcinoma MCF-7 cancer cell line. The colon carcinoma cell lines showed values ranging from three- to 10-fold increased cell death induction by 4C7 and 4C9 when compared with rhTRAILWT (Figure 3). The lack of activity on DR5-mediated A2780 and Jurkat cells and the comparison of activity between BJAB and BJAB DR5-deficient cell line with rhTRAILWT clearly confirmed the DR4 selectivity of the variants 4C7 and 4C9 (Figure 2). The increased activity of the variants was also evident in TRAIL-resistant MCF-7 and PANC-1 cells (Figure 3). The higher affinity and selectivity of our variants for DR4 (Table 1 and Figure 1) seems to correlate to their higher cellular activity in cell lines sensitive via the DR4 receptor.
To further analyse the activity of the variant 4C7, we determined caspase activation in OVCAR-3 cells using FRET and fluorescent probes (Figure 4). A much faster and stronger onset of caspase activation could be observed for 4C7 in comparison to rhTRAILWT caspase-8/10 activation as seen by the IETDase probe cleavage that started already at very early time points with a nearly eightfold faster activation when compared with rhTRAILWT (Figure 4a and b). Downstream effector caspase activation based on DEVDase activity using a probe containing CFP and YFP indicated a significant cleavage rate starting at 75 min over a 30–40 min period for 4C7 (Figure 4c and d). DEVDase cleavage for rhTRAILWT was substantially slower in the time points taken. These results allow us to conclude that variant 4C7 induces a much faster and robust initiator and effector caspase activation than rhTRAILWT in the OVCAR-3 cell line, and underlines the efficacy of the DR4-specific variant.
The high efficacy of our new DR4-selective variants in addition to the earlier published DR5 variant D269H/E195R19 inspired us to test whether their combination would lead to even higher apoptotic activity. Combination treatment using the DR4-selective variants with the DR5-selective variant enhanced apoptosis-inducing activity when compared with the responses obtained for the single treatments (Figure 5). Analysis of response curves using different concentrations of DR-specific variants in combination indicates a significant additive effect on cell death induction on the ovarian carcinoma OVCAR-3 cell line. A similar effect was observed in colon carcinoma cell lines Colo205 and CL-34 at high concentrations of the two TRAIL variants. Even though part of the effect can be explained by the higher affinities and higher association rate constants of these variants for their cognate receptor, these results also point to the inability of these variants to form inactive heteromeric receptor complexes between their target DR and the other receptors, as was shown for rhTRAILWT. Furthermore, our DR4- and DR5-selective variants strongly inhibited clonogenicity of OVCAR-3 ovarian carcinoma cells (Figure 6a and b), showing the enhanced efficacy of these variants compared with rhTRAILWT in reducing long-term survival of cancer cells.
In summary, the newly constructed DR4-specific variants lead to the formation of high-affinity homotrimeric TRAIL–DR4 complexes. These TRAIL variants alone or in combination with our DR5 agonist may prove to be a promising treatment for a range of different cancers.
Materials and Methods
Modelling and computational design of variants using FoldX
Currently, only the crystal structure of TRAIL alone or in complex with DR5 is known. A homology model of DR4 was built based on the coordinates of TRAIL and DR5, from the template TRAIL in complex with the ectodomain of DR5, pdb: ID 1D4V,32 using the high sequence identity between DR5 and DR4 (>50%), with no insertions or deletions between both death receptors. Binding effects of mutations situated in the TRAIL–receptor interface18, 23 allowed the construction of a refined homology model for the TRAIL–DR4 complex using the protein design capabilities of FoldX.33, 34 These coordinates were used to construct the homotrimer TRAIL using the protein quaternary structure server from EBI (http://pqs.ebi.ac.uk). Single mutants of TRAIL were predicted by FoldX. Interaction energies were calculated as the sum of the energies of the individual receptor and ligand subunits and subtracting them from the global energy of the complex. A detailed description of the protein design algorithm FoldX (version 3.0) is available elsewhere in http://foldx.crg.es.
Construction, expression and purification of rhTRAIL variants
Mutants were constructed by PCR reaction using the megaprimer mutagenesis method. The polymerase used was Phusion polymerase supplied by Finnzymes (Espoo, Finland). Introduction of mutations was confirmed by DNA sequencing. RhTRAILWT and variants were cloned into pET15b (Novagen, Madison, WI, USA) using the BamHI and NcoI sites and transformed into Escherichia coli BL21 (DE3). Homotrimeric TRAIL proteins were expressed and purified as described before.19, 23 Flag-tag rhTRAILWT and flag-tag rhTRAIL 4C9 were constructed by introducing the sequence encoding a flag-tag N-terminally of the rhTRAIL (aa 114–281) sequence in pET15b. Analytical gel filtration, dynamic light scattering and non-reducing gel electrophoresis confirmed that rhTRAILWT and variants were stable trimeric molecules and did not form higher-order molecular weight aggregates.
Receptor binding by surface plasmon resonance and competitive ELISA
Binding experiments were performed using a surface plasmon resonance-based Biacore 3000 (GE Healthcare, Eindhoven, The Netherlands). Research grade CM4 sensor chips, N-hydroxysuccimide, N-ethyl-N′-(3-diethylaminopropyl) carbodiimide, ethanolamine hydrochloride and standard buffers, for example, HBS-N, were purchased from the manufacturer (GE Healthcare). Immobilization of Staphylococcal protein A (Sigma, Zwijndrecht, The Netherlands) on the sensor surface of a CM4 sensor chip was performed following a standard amine coupling procedure. Protein A was coated at a level of ∼1000 response units. DR4-Ig and DR5-Ig receptors (R&D Systems, Minneapolis, MN, USA) were captured at high flow rate to low densities (5–20 RU) resulting in binding of a trimeric TRAIL molecule to only one receptor molecule and allowing global fitting of the data to a 1 : 1 Langmuir model.35 A 100 μl aliquot of rhTRAILWT and variants was injected at concentrations ranging from 1 to 250 nM at 50 μl/min and at 37°C using HBS-N supplemented with 0.005% surfactant P20 as running and sample buffer. Binding of ligands to the receptors was monitored in real time. Between cycles, the protein A/sensor surface was regenerated using 10 mM glycine, pH 1.7 and a contact time of 25 s.
For competitive ELISA, Nunc maxisorb plates were coated for 2 h with DR4-Ig (100 ng per well) in 0.1 M sodium carbonate/bicarbonate buffer (pH 8.6) and the remaining binding places subsequently blocked with 2% BSA for 1 h. After washing six times with Tris-buffered saline/0.5% Tween-20 (TBST) (pH 7.5), serial dilutions of soluble DR4-, DR5-, DcR1- or DcR2-Ig (0–5000 ng/ml) and rhTRAILWT or mutants (100 ng/ml) in PBS (pH 7.4) previously incubated for 1.5 h at room temperature were added to the wells and incubated for 1 h. After washing, a 1 : 200 dilution of anti-TRAIL antibody (R&D Systems) was added and incubated for 1 h at room temperature, and, after washing with TBST, subsequently incubated with a 1 : 25 000 dilution of a horse radish peroxidase-conjugated swine anti-goat antibody. Finally, a 100 μl of one-step Turbo TMB solution (Pierce, Rockford, IL, USA) was added. The reaction was quenched with 100 μl of 1 M sulphuric acid and the absorbance measured at 450 nm on a microplate reader (Thermo Labsystems, Breda, The Netherlands). Binding of rhTRAIL or variants to immobilized DR4-Ig with 0 ng per well of the soluble receptors was taken as 100%, and binding at other concentrations of soluble receptors was calculated relative to this value.
Reagents and cell line treatment
Cell culture reagents were purchased from Gibco (Invitrogen, Breda, The Netherlands). Cycloheximide was from Sigma-Aldrich (Zwijndrecht, The Netherlands). Colon adenocarcinoma cell lines Colo205, DLD-1 and HCT-15, Burkitt's lymphoma (BJAB), Jurkat T-cell leukaemia and PANC-1 human pancreatic carcinoma cell lines were maintained in RPMI 1640 medium, supplemented with 10% FBS, 50 U/ml penicillin, 5 mg/ml streptomycin, 2 mM L-glutamine and 1 mM sodium pyruvate in a humidified incubator at 37°C and 5% CO2 environment. A2780 ovarian carcinoma, MFC-7 human breast adenocarcinoma and OVCAR-3 ovarian carcinoma were cultured in DMEM, with 10% FBS, 50 U/ml penicillin, 5 mg/ml streptomycin and glutamine. Colon carcinoma SW948 cells were cultured in Leibovitz L15-RPMI 1640 (1 : 1) enriched with 10% FBS, 0.05 M pyruvate, 0.1 M glutamine and 0.025% β-mercaptoethanol at 37°C in a humidified atmosphere with 5% CO2. All cells were seeded at 50% confluence 24 h before treatment.
TRAIL receptor expression in cell lines
Cells were removed from culture dishes, harvested by centrifugation and washed twice with 1% BSA in PBS. Cells were incubated with 1 : 100 dilution of primary antibodies DR4 and DR5: neutralizing mouse monoclonal antibodies (Alexis, San Diego, CA, USA); DcR1 and DcR2: neutralizing goat polyclonal antibodies (R&D Systems) in 1% BSA in PBS for 40 min on ice. After two washes with 1% BSA in PBS, cells were resuspended in 1 : 50 dilution of FITC-labelled secondary antibody and incubated for 40 min on ice. Excess secondary antibody was removed by washing first in 1% BSA in PBS and then PBS. Cells were fixed in 1% formaldehyde/PBS before analysis by flow cytometry (FacsCalibur, Beckton Dickinson, San Jose, CA, USA).
Cell viability and apoptosis assays
To determine sensitivity to rhTRAIL and variants, cells were plated in 96-well plates, allowed to adhere for 24 h and then treated with rhTRAIL in various concentrations, ranging from 1 to 250 ng/ml. After 24 h incubation, the cells were subjected to an MTS viability assay (Promega, Leiden, The Netherlands) following the manufacturer's protocol. Cell viability was determined after 1–2 h of incubation by measuring the absorption at 490 nm on a microplate reader (Thermo Labsystems). Apoptosis induction was measured by Annexin V staining and quantified by flow cytometric analysis as described before.23
Live cell microscopy: caspase activation in living cells using FRET
Time lapse images were collected using a Leica SP2 AOBS CLSM microscope equipped with an environmental chamber at 37°C and 5% CO2 and × 10 or × 20 objective magnification. Images were collected every minute. OVCAR-3 ovarian carcinoma cell line expressing pSCAT8 producing CFP-IETD-Venus25 and pDEVD containing CFP-DEVD-YFP36 were derived by transfection with Fugene 6 (Roche, Almere, The Netherlands) according to the manufacturer's protocol. Plasmids pSCAT8 and pDEVD were a kind gift from Dr. Markus Rehm. Ratio images were obtained using the Image J software and available custom plugins by calculating the ratio from the obtained CFP and YFP images corrected for background. Signals were normalized by subtracting the minimum value across all time points from each single time-course experiment.
Clonogenic survival assay
The clonogenic ability of ovarian carcinoma cells (OVCAR-3) was determined by plating the cells at the appropriate dilutions in DMEM medium, treated for 24 h with rhTRAILWT and variants and the colonies were counted after 2 weeks. Briefly, 100 or 200 cells were plated out in 94 mm Petri dishes (Greiner) in 10 ml DMEM medium containing 10% fetal bovine serum (Invitrogen), penicillin (10 U/ml) and streptomycin (10 μg/ml), and incubated for 16 h at 37°C and 5% CO2. The cells were treated with 5 ng/ml of rhTRAILWT or variants for 24 h. After treatment, the medium was replaced and cells incubated for 2 weeks at 37°C and 5% CO2. The colonies were fixed and stained using 0.1% Brilliant blue (Sigma), 50% methanol and 10% acetic acid, and destained with wash buffer (10% methanol and 7% acetic acid). Colonies containing more than 50 cells were scored.
TRAIL-DISC immunoprecipitation
The anti-FLAG (M2 antibody, Sigma) was covalently conjugated to epoxy-coated Dynabeads (Invitrogen) following the manufacturer's instructions (5 mg antibody per 1 mg bead). Colo205 cells (approximately 50 mg per sample) were treated with 250 ng/ml flag-tag rhTRAILWT or flag-tag rhTRAIL 4C9 for 15 min. Cells were harvested by scraping in the medium and centrifugation. After washing with PBS, the cell pellet was weighted and the cells lysed with nine volumes of Extraction buffer A (Dynabeads Co-immunoprecipitation kit, Invitrogen) supplemented with 50 mM NaCl and incubated on ice for 15 min. The lysates were cleared by centrifugation and mixed with 1.5 mg of antibody-coupled Dynabeads. The lysates were rotated with the beads for 3 h at 4°C, after which the supernatant was separated from the beads, the beads washed and the DISC complex eluted by following the manufacturer's instructions. The immunoprecipitate was denatured in Laemmli buffer before loading on gels and analysis of the DISC components.
Abbreviations
- TRAIL:
-
tumour necrosis factor-related apoptosis-inducing ligand
- DRs:
-
death receptors
- DR4:
-
death receptor 4
- DR5:
-
death receptor 5
- SPR:
-
surface plasmon resonance
References
Pan G, O'Rourke K, Chinnaiyan AM, Gentz R, Ebner R, Ni J et al. The receptor for the cytotoxic ligand TRAIL. Science 1997; 276: 111–113.
Sheridan JP, Marsters SA, Pitti RM, Gurney A, Skubatch M, Baldwin D et al. Control of TRAIL-induced apoptosis by a family of signaling and decoy receptors. Science 1997; 277: 818–821.
Chaudhary PM, Eby M, Jasmin A, Bookwalter A, Murray J, Hood L . Death receptor 5, a new member of the TNFR family, and DR4 induce FADD-dependent apoptosis and activate the NF-kappaB pathway. Immunity 1997; 7: 821–830.
Kuang AA, Diehl GE, Zhang J, Winoto A . FADD is required for DR4- and DR5-mediated apoptosis: lack of trail-induced apoptosis in FADD-deficient mouse embryonic fibroblasts. J Biol Chem 2000; 275: 25065–25068.
Schneider P, Thome M, Burns K, Bodmer JL, Hofmann K, Kataoka T et al. TRAIL receptors 1 (DR4) and 2 (DR5) signal FADD-dependent apoptosis and activate NF-kappaB. Immunity 1997; 7: 831–836.
Bodmer JL, Holler N, Reynard S, Vinciguerra P, Schneider P, Juo P et al. TRAIL receptor-2 signals apoptosis through FADD and caspase-8. Nat Cell Biol 2000; 2: 241–243.
Kischkel FC, Lawrence DA, Chuntharapai A, Schow P, Kim KJ, Ashkenazi A . Apo2L/TRAIL-dependent recruitment of endogenous FADD and caspase-8 to death receptors 4 and 5. Immunity 2000; 12: 611–620.
Sprick MR, Weigand MA, Rieser E, Rauch CT, Juo P, Blenis J et al. FADD/MORT1 and caspase-8 are recruited to TRAIL receptors 1 and 2 and are essential for apoptosis mediated by TRAIL receptor 2. Immunity 2000; 12: 599–609.
LeBlanc HN, Ashkenazi A . Apo2L/TRAIL and its death and decoy receptors. Cell Death Differ 2003; 10: 66–75.
Kimberley FC, Screaton GR . Following a TRAIL: update on a ligand and its five receptors. Cell Res 2004; 14: 359–372.
Pan G, Ni J, Wei YF, Yu G, Gentz R, Dixit VM . An antagonist decoy receptor and a death domain-containing receptor for TRAIL. Science 1997; 277: 815–818.
Ashkenazi A, Pai RC, Fong S, Leung S, Lawrence DA, Marsters SA et al. Safety and antitumor activity of recombinant soluble Apo2 ligand. J Clin Invest 1999; 104: 155–162.
Lawrence D, Shahrokh Z, Marsters S, Achilles K, Shih D, Mounho B et al. Differential hepatocyte toxicity of recombinant Apo2L/TRAIL versions. Nat Med 2001; 7: 383–385.
Clancy L, Mruk K, Archer K, Woelfel M, Mongkolsapaya J, Screaton G et al. Preligand assembly domain-mediated ligand-independent association between TRAIL receptor 4 (TR4) and TR2 regulates TRAIL-induced apoptosis. Proc Natl Acad Sci USA 2005; 102: 18099–18104.
Merino D, Lalaoui N, Morizot A, Schneider P, Solary E, Micheau O . Differential inhibition of TRAIL-mediated DR5-DISC formation by decoy receptors 1 and 2. Mol Cell Biol 2006; 26: 7046–7055.
Kelley RF, Totpal K, Lindstrom SH, Mathieu M, Billeci K, Deforge L et al. Receptor-selective mutants of apoptosis-inducing ligand 2/tumor necrosis factor-related apoptosis-inducing ligand reveal a greater contribution of death receptor (DR) 5 than DR4 to apoptosis signaling. J Biol Chem 2005; 280: 2205–2212.
MacFarlane M, Kohlhaas SL, Sutcliffe MJ, Dyer MJ, Cohen GM . TRAIL receptor-selective mutants signal to apoptosis via TRAIL-R1 in primary lymphoid malignancies. Cancer Res 2005; 65: 11265–11270.
Tur V, van der Sloot AM, Reis CR, Szegezdi E, Cool RH, Samali A et al. DR4-selective tumor necrosis factor-related apoptosis-inducing ligand (TRAIL) variants obtained by structure-based design. J Biol Chem 2008; 283: 20560–20568.
van der Sloot AM, Tur V, Szegezdi E, Mullally MM, Cool RH, Samali A et al. Designed tumor necrosis factor-related apoptosis-inducing ligand variants initiating apoptosis exclusively via the DR5 receptor. Proc Natl Acad Sci USA 2006; 103: 8634–8639.
Almasan A, Ashkenazi A . Apo2L/TRAIL: apoptosis signaling, biology, and potential for cancer therapy. Cytokine Growth Factor Rev 2003; 14: 337–348.
Ichikawa K, Liu W, Zhao L, Wang Z, Liu D, Ohtsuka T et al. Tumoricidal activity of a novel anti-human DR5 monoclonal antibody without hepatocyte cytotoxicity. Nat Med 2001; 7: 954–960.
MacFarlane M, Inoue S, Kohlhaas SL, Majid A, Harper N, Kennedy DB et al. Chronic lymphocytic leukemic cells exhibit apoptotic signaling via TRAIL-R1. Cell Death Differ 2005; 12: 773–782.
Reis CR, van der Sloot AM, Szegezdi E, Natoni A, Tur V, Cool RH et al. Enhancement of antitumor properties of rhTRAIL by affinity increase toward its death receptors. Biochemistry 2009; 48: 2180–2191.
Thomas LR, Henson A, Reed JC, Salsbury FR, Thorburn A . Direct binding of Fas-associated death domain (FADD) to the tumor necrosis factor-related apoptosis-inducing ligand receptor DR5 is regulated by the death effector domain of FADD. J Biol Chem 2004; 279: 32780–32785.
Hellwig CT, Kohler BF, Lehtivarjo AK, Dussmann H, Courtney MJ, Prehn JH et al. Real time analysis of tumor necrosis factor-related apoptosis-inducing ligand/cycloheximide-induced caspase activities during apoptosis initiation. J Biol Chem 2008; 283: 21676–21685.
Garcia-Calvo M, Peterson EP, Leiting B, Ruel R, Nicholson DW, Thornberry NA . Inhibition of human caspases by peptide-based and macromolecular inhibitors. J Biol Chem 1998; 273: 32608–32613.
McStay GP, Salvesen GS, Green DR . Overlapping cleavage motif selectivity of caspases: implications for analysis of apoptotic pathways. Cell Death Differ 2008; 15: 322–331.
Zhang XD, Franco A, Myers K, Gray C, Nguyen T, Hersey P . Relation of TNF-related apoptosis-inducing ligand (TRAIL) receptor and FLICE-inhibitory protein expression to TRAIL-induced apoptosis of melanoma. Cancer Res 1999; 59: 2747–2753.
Mahalingam D, Szegezdi E, Keane M, Jong S, Samali A . TRAIL receptor signalling and modulation: Are we on the right TRAIL? Cancer Treat Rev 2009; 35: 280–288.
Duiker EW, de Vries EG, Mahalingam D, Meersma GJ, Boersma-van Ek W, Hollema H et al. Enhanced antitumor efficacy of a DR5-specific TRAIL variant over recombinant human TRAIL in a bioluminescent ovarian cancer xenograft model. Clin Cancer Res 2009; 15: 2048–2057.
Truneh A, Sharma S, Silverman C, Khandekar S, Reddy MP, Deen KC et al. Temperature-sensitive differential affinity of TRAIL for its receptors. DR5 is the highest affinity receptor. J Biol Chem 2000; 275: 23319–23325.
Mongkolsapaya J, Grimes JM, Chen N, Xu XN, Stuart DI, Jones EY et al. Structure of the TRAIL–DR5 complex reveals mechanisms conferring specificity in apoptotic initiation. Nat Struct Biol 1999; 6: 1048–1053.
Guerois R, Nielsen JE, Serrano L . Predicting changes in the stability of proteins and protein complexes: a study of more than 1000 mutations. J Mol Biol 2002; 320: 369–387.
Schymkowitz JW, Rousseau F, Martins IC, Ferkinghoff-Borg J, Stricher F, Serrano L . Prediction of water and metal binding sites and their affinities by using the Fold-X force field. Proc Natl Acad Sci USA 2005; 102: 10147–10152.
Reis CR, van Assen AH, Quax WJ, Cool RH . Unraveling the binding mechanism of trivalent tumor necrosis factor ligands and their receptors. Mol Cell Proteomics [Epub ahead of print 17 September 2010].
Rehm M, Dussmann H, Janicke RU, Tavare JM, Kogel D, Prehn JH . Single-cell fluorescence resonance energy transfer analysis demonstrates that caspase activation during apoptosis is a rapid process. Role of caspase-3. J Biol Chem 2002; 277: 24506–24514.
Acknowledgements
We would like to thank Dr. Markus Rehm for providing us the plasmids pSCAT8 and pDEVD for FRET-based experiments; Dr. Andrew Thorburn for kindly providing the BJAB cell lines; and Dr. Steven de Jong and Bodvael Pennarun for providing the SW948 cell line, Pim Weiniger for helpful discussions and technical assistance and Ulrike Schnell for critical reading. This research was partly funded by European Union Fifth Framework Program Grant QLK3-CT-2001-00498 and Sixth Framework Program Grant LSH-2005-2.2.0-2, and by the Dutch Technology Foundation STW, applied science division of NWO and the technology program of the Ministry of Economic Affairs.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Wim J Quax, Luis Serrano and Afshin Samali are founding members of TRISKEL Therapeutics.
Additional information
Edited by R Johnstone
Supplementary Information accompanies the paper on Cell Death and Disease website
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Reis, C., van der Sloot, A., Natoni, A. et al. Rapid and efficient cancer cell killing mediated by high-affinity death receptor homotrimerizing TRAIL variants. Cell Death Dis 1, e83 (2010). https://doi.org/10.1038/cddis.2010.61
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2010.61
Keywords
This article is cited by
-
Hypertonicity-enforced BCL-2 addiction unleashes the cytotoxic potential of death receptors
Oncogene (2018)
-
N-glycosylation of mouse TRAIL-R and human TRAIL-R1 enhances TRAIL-induced death
Cell Death & Differentiation (2017)
-
Onto better TRAILs for cancer treatment
Cell Death & Differentiation (2016)
-
Decoy receptors block TRAIL sensitivity at a supracellular level: the role of stromal cells in controlling tumour TRAIL sensitivity
Oncogene (2016)
-
TRAIL-receptor preferences in pancreatic cancer cells revisited: Both TRAIL-R1 and TRAIL-R2 have a licence to kill
BMC Cancer (2015)