Abstract
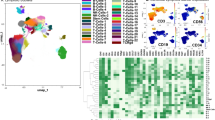
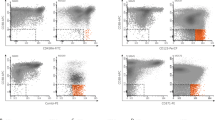
Relapse of Ph chromosome-positive ALL (Ph+ALL) results from the persistence of leukemia-propagating cells (LPCs). In Ph+ALL, a xenograft assay recently determined that LPCs are enriched in the CD34+CD38−CD58− fraction. Therefore, the prognostic significance of LPCs in Ph+ALL subjects after allogeneic hematopoietic SCT (allo-HSCT) was investigated. A total of 80 consecutive adults with Ph+ALL who underwent allo-HSCT were eligible. A multi-parameter flow cytometry analysis examining CD58–FITC/CD10–PE/ CD19–APC–Cy7/CD34–PerCP/CD45–Vioblue/ CD38–APC on gated leukemia BM blasts was performed at diagnosis. Based on the original blast phenotypes, subjects were stratified into the CD34+CD38−CD58−group (N=15) and other phenotype group (N=65). During minimal residual disease monitoring, significantly higher levels of BCR/ABL transcripts were detected in subjects in the CD34+CD38−CD58− group than in other phenotype group, especially at 3 months post HSCT. In addition, CD34+CD38−CD58−LPCs are directly correlated with a higher 3-year cumulative incidence of relapse (CIR) and worse leukemia-free survival (LFS) and OS. Multivariate analyses indicated that presence of CD34+CD38−CD58− LPCs at diagnosis, and BCR–ABL reduction at 3 months post HSCT were independent risk factors for relapse, LFS and OS. Our data suggest that presence of CD34+CD38−CD58− LPCs at diagnosis allows rapid identification of high-risk patients for relapse after allo-HSCT.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Fielding AK . How I treat Philadelphia chromosome-positive acute lymphoblastic leukemia. Blood 2010; 116: 3409–3417.
Pui CH, Robison LL, Look AT . Acute lymphoblastic leukaemia. Lancet 2008; 371: 1030–1043.
Chen H, Liu KY, Xu LP, Liu DH, Chen YH, Zhao XY et al. Administration of imatinib after allogeneic hematopoietic stem cell transplantation may improve disease-free survival for patients with Philadelphia chromosome-positive acute lymphobla stic leukemia. J Hematol Oncol 2012; 5: 29.
Fielding AK, Rowe JM, Buck G, Foroni L, Gerrard G, Litzow MR et al. UKALLXII/ECOG2993: addition of imatinib to a standard treatment regimen enhances long-term outcomes in Philadelphia positive acute lymphoblastic leukemia. Blood 2014; 123: 843–850.
Mizuta S, Matsuo K, Nishiwaki S, Imai K, Kanamori H, Ohashi K et al. Pretransplant administration of imatinib for allo-HSCT in patients with BCR-ABL-positive acute lymphoblastic leukemia. Blood 2014; 123: 2325–2332.
Valent P, Bonnet D, De Maria R, Lapidot T, Copland M, Melo JV et al. Cancer stem cell definitions and terminology: the devil is in the details. Nat Rev Cancer 2012; 12: 767–775.
Lapidot T, Sirard C, Vormoor J, Murdoch B, Hoang T, Caceres-Cortes J et al. A cell initiating human acute myeloid leukaemia after transplantation into SCID mice. Nature 1994; 367: 645–648.
Kong Y, Yoshida S, Saito Y, Doi T, Nagatoshi Y, Fukata M et al. CD34+CD38+CD19+ as well as CD34+CD38-CD19+ cells are leukemia-initiating cells with self-renewal capacity in human B-precursor ALL. Leukemia 2008; 22: 1207–1213.
Eppert K, Takenaka K, Lechman ER, Waldron L, Nilsson B, van Galen P et al. Stem cell gene expression programs influence clinical outcome in human leukemia. Nat Med 2011; 17: 1086–1093.
Gerber JM, Smith BD, Ngwang B, Zhang H, Vala MS, Morsberger L et al. A clinically relevant population of leukemic CD34(+)CD38(−) cells in acute myeloid leukemia. Blood 2012; 119: 3571–3577.
van Rhenen A, Feller N, Kelder A, Westra AH, Rombouts E, Zweegman S et al. High stem cell frequency in acute myeloid leukemia at diagnosis predicts high minimal residual disease and poor survival. Clin Cancer Res 2005; 11: 6520–6527.
Gentles AJ, Plevritis SK, Majeti R, Alizadeh AA . Association of a leukemic stem cell gene expression signature with clinical outcomes in acute myeloid leukemia. JAMA 2010; 304: 2706–2715.
Kong Y, Chang YJ, Liu YR, Wang YZ, Jiang Q, Jiang H et al. CD34+CD38−CD58− cells are leukemia-propagating cells in Philadelphia chromosome-positive acute lymphoblastic leukemia. Leukemia 2014 (epub ahead of print 25 July 2014; doi:10.1038/leu.2014.228).
Baumgarth N, Roederer M . A practical approach to multicolor flow cytometry for immunophenotyping. J Immunol Methods 2000; 243: 77–97.
Bayer J, Grunwald D, Lambert C, Mayol JF, Maynadie M . Thematic workshop on fluorescence compensation settings in multicolor flow cytometry. Clin Cytom 2007; 72: 8–13.
Bene MC, Castoldi G, Knapp W, Ludwig WD, Matutes E, Orfao A et al. Proposals for the immunological classification of acute leukemias. Leukemia 1995; 9: 1783–1786.
Xiao-Jun H, Lan-Ping X, Kai-Yan L, Dai-Hong L, Yu W, Huan C et al. Partially matched related donor transplantation can achieve outcomes comparable with unrelated donor transplantation for patients with hematologic malignancies. Clin Cancer Res 2009; 15: 4777–4783.
Huang XJ, Zhu HH, Chang YJ, Xu LP, Liu DH, Zhang XH et al. The superiority of haploidentical related stem cell transplantation over chemotherapy alone as postremission treatment for patients with intermediate- or high-risk acute myeloid leukemia in first complete remission. Blood 2012; 119: 5584–5590.
Yan CH, Liu DH, Liu KY, Xu LP, Liu YR, Chen H et al. Risk stratification-directed donor lymphocyte infusion could reduce relapse of standard-risk acute leukemia patients after allogeneic hematopoietic stem cell transplantation. Blood 2012; 119: 3256–3262.
Wang Y, Wu DP, Liu QF, Qin YZ, Wang JB, Xu LP et al. RUNX1/RUNX1T1-based MRD-monitoring early after allogeneic transplantation rather than c-KIT mutations in adult t(8;21) AML allows further risk stratification. Blood, (e-pub ahead of print 31 July 2014; pii: blood-2014-03-563403).
Qin YZ, Liu YR, Zhu HH, Li JL, Ruan GR, Zhang Y et al. Different kinetic patterns of BCR-ABL transcript levels in imatinib-treated chronic myeloid leukemia patients after achieving complete cytogenetic response. Int J Lab Hematol 2008; 30: 317–323.
Beillard E, Pallisgaard N, van der Velden VH, Bi W, Dee R, van der Schoot E et al. Evaluation of candidate control genes for diagnosis and residual disease detection in leukemic patients using 'real-time' quantitative reverse-transcriptase polymerase chain reaction (RQ-PCR) - a Europe against cancer program. Leukemia 2003; 17: 2474–2486.
Gabert J, Beillard E, van der Velden VH, Bi W, Grimwade D, Pallisgaard N et al. Standardization and quality control studies of 'real-time' quantitative reverse transcriptase polymerase chain reaction of fusion gene transcripts for residual disease detection in leukemia-a Europe Against Cancer program. Leukemia 2003; 17: 2318–2357.
Kong Y, Chang YJ, Wang YZ, Chen YH, Han W, Wang Y et al. Association of an impaired bone marrow microenvironment with secondary poor graft function after allogeneic hematopoietic stem cell transplantation. Biol Blood Marrow Transplant 2013; 19: 1465–1473.
Preudhomme C, Henic N, Cazin B, Lai JL, Bertheas MF, Vanrumbeke M et al. Good correlation between RT-PCR analysis and relapse in Philadelphia (Ph1)-positive acute lymphoblastic leukemia (ALL). Leukemia 1997; 11: 294–298.
Pane F, Cimino G, Izzo B, Camera A, Vitale A, Quintarelli C et al. Significant reduction of the hybrid BCR/ABL transcripts after induction and consolidation therapy is a powerful predictor of treatment response in adult Philadelphia-positive acute lymphoblastic leukemia. Leukemia 2005; 19: 628–635.
Lee S, Kim YJ, Chung NG, Lim J, Lee DG, Kim HJ et al. The extent of minimal residual disease reduction after the first 4-week imatinib therapy determines outcome of allogeneic stem cell transplantation in adults with Philadelphia chromosome-positive acute lymphoblastic leukemia. Cancer 2009; 115: 561–570.
Pfeifer H, Wassmann B, Bethge W, Dengler J, Bornhauser M, Stadler M et al. Randomized comparison of prophylactic and minimal residual disease-triggered imatinib after allogeneic stem cell transplantation for BCR-ABL1-positive acute lymphoblastic leukemia. Leukemia 2013; 27: 1254–1262.
Wassmann B, Pfeifer H, Stadler M, Bornhauser M, Bug G, Scheuring UJ et al. Early molecular response to posttransplantation imatinib determines outcome in MRD+ Philadelphia-positive acute lymphoblastic leukemia (Ph+ALL). Blood 2005; 106: 458–463.
Yanada M, Sugiura I, Takeuchi J, Akiyama H, Maruta A, Ueda Y et al. Prospective monitoring of BCR-ABL1 transcript levels in patients with Philadelphia chromosome-positive acute lymphoblastic leukaemia undergoing imatinib-combined chemotherapy. Bri J Haematol 2008; 143: 503–510.
Copland M, Hamilton A, Elrick LJ, Baird JW, Allan EK, Jordanides N et al. Dasatinib (BMS-354825) targets an earlier progenitor population than imatinib in primary CML but does not eliminate the quiescent fraction. Blood 2006; 107: 4532–4539.
Saito Y, Kitamura H, Hijikata A, Tomizawa-Murasawa M, Tanaka S, Takagi S et al. Identification of therapeutic targets for quiescent, chemotherapy-resistant human leukemia stem cells. Sci Transl Med 2010; 2: 17ra9.
Talpaz M, Shah NP, Kantarjian H, Donato N, Nicoll J, Paquette R et al. Dasatinib in imatinib-resistant Philadelphia chromosome-positive leukemias. N Engl J Med 2006; 354: 2531–2541.
Cortes JE, Kantarjian H, Shah NP, Bixby D, Mauro MJ, Flinn I et al. Ponatinib in refractory Philadelphia chromosome-positive leukemias. N Engl J Med 2012; 367: 2075–2088.
Savani BN, Srinivasan R, Espinoza-Delgado I, Dorrance C, Takahashi Y, Igarashi T et al. Treatment of relapsed blast-phase Philadelphia-chromosome-positive leukaemia after non-myeloablative stem-cell transplantation with donor lymphocytes and imatinib. Lancet Oncol 2005; 6: 809–812.
Visani G, Martinelli G, Piccaluga P, Tosi P, Amabile M, Pastano R et al. Alpha-interferon improves survival and remission duration in P-190BCR-ABL positive adult acute lymphoblastic leukemia. Leukemia 2000; 14: 22–27.
Wassmann B, Scheuring U, Pfeifer H, Binckebanck A, Kabisch A, Lubbert M et al. Efficacy and safety of imatinib mesylate (Glivec) in combination with interferon-alpha (IFN-alpha) in Philadelphia chromosome-positive acute lymphoblastic leukemia (Ph+ALL). Leukemia 2003; 17: 1919–1924.
Martinelli G, Iacobucci I, Storlazzi CT, Vignetti M, Paoloni F, Cilloni D et al. IKZF1 (Ikaros) deletions in BCR-ABL1-positive acute lymphoblastic leukemia are associated with short disease-free survival and high rate of cumulative incidence of relapse: a GIMEMA AL WP report. J Clin Oncol 2009; 27: 5202–5207.
Mullighan CG, Miller CB, Radtke I, Phillips LA, Dalton J, Ma J et al. BCR-ABL1 lymphoblastic leukaemia is characterized by the deletion of Ikaros. Nature 2008; 453: 110–114.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (grant no. 81370638 & 81230013), the Beijing Municipal Science and Technology Program, the National Clinical Priority Specialty and the Peking University People’s Hospital Research and Development Funds (grant no. RDB2012-23). American Journal Experts (www.journalexperts.com) offered editorial assistance to the authors during the preparation of this manuscript. We thank all of the core facilities at the Peking University Institute of Hematology for sample collection.
Author Contributions
X-JH designed the study and supervised the analyses and manuscript preparation. YK performed the research, analyzed and interpreted the data, performed statistical analyses and wrote the manuscript. All other authors participated in the collection of patient data. All the authors agreed to submit the final manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Kong, Y., Xu, LP., Liu, YR. et al. Presence of CD34+CD38−CD58− leukemia-propagating cells at diagnosis identifies patients at high risk of relapse with Ph chromosome-positive ALL after allo-hematopoietic SCT. Bone Marrow Transplant 50, 348–353 (2015). https://doi.org/10.1038/bmt.2014.274
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bmt.2014.274
This article is cited by
-
Leukemia-propagating cells demonstrate distinctive gene expression profiles compared with other cell fractions from patients with de novo Philadelphia chromosome-positive ALL
Annals of Hematology (2018)
-
Ruxolitinib/nilotinib cotreatment inhibits leukemia-propagating cells in Philadelphia chromosome-positive ALL
Journal of Translational Medicine (2017)