Abstract
Background:
In Australia, more money is spent on skin cancer than any other malignancy. Despite this, the mortality rate of melanoma, the deadliest form, has steadily increased over the past 50 years. Diagnostic imprecision and a lack of complimentary molecular biomarkers are partially responsible for this lack of progress.
Methods:
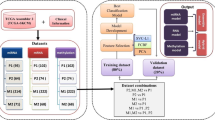
Whole-microRNAome profiling was performed on plasma samples from 32 patients with histologically confirmed melanoma and 16 normal controls. A classification algorithm was trained on these data and independently validated on multiple previously published microRNA data sets, representing (i) melanoma patient- and normal-blood, (ii) melanoma and nevi biopsy tissue, and (iii) cell lines and purified exosomes.
Results:
38 circulating microRNAs had biologically and statistically significant differences between melanoma and normal plasma samples (MEL38). A support vector machine algorithm, trained on these markers, showed strong independent classification accuracy (AUC 0.79–0.94). A majority of MEL38 genes have been previously associated with melanoma and are known regulators of angiogenesis, metastasis, tumour suppression, and treatment resistance.
Conclusions:
MEL38 exhibits disease state specificity and robustness to platform and specimen-type variation. It has potential to become an objective diagnostic biomarker and improve the precision and accuracy of melanoma detection and monitoring.
Similar content being viewed by others
Main
The age standardised mortality rates of four of the five most common cancers in Australia (i.e., prostate – first; breast – second; colorectal – third; lung – fifth) are currently at their lowest levels since government records began. In stark contrast, the mortality rate of the fourth most common cancer, melanoma, has consistently increased and is now >200% higher than it was in the 1960s (Australian Institute of Health and Welfare, 2017) (Supplementary Figure 1). In 2017, melanoma kills at least three people per day, resulting in an annual death toll higher than from motor vehicle accidents. Despite this lack of progress, the Australian Government spends an estimated $900 million on skin cancer each year (melanoma $200 m, non-melanoma $700 m), which is more than it spends on any other cancer type (Fransen et al, 2012; Elliott TM et al, 2017). Recent advances in immunotherapy for late stage melanoma patients, which can cost upwards of $200 000 per patient per year, are likely to add to this figure (Luftos and Wilslow, 2015). As melanoma is easily and inexpensively cured if diagnosed early, this combination of increasing mortality and ballooning expenditure suggest that new thinking is needed on how skin cancer, particularly melanoma, is diagnosed and managed.
In order to determine the pathological status of a suspicious skin lesion, visual examination, dermoscopy, biopsy, and histopathological examination are the methods currently employed. Pigmented lesions can be difficult to diagnose and subjective to the pathologist viewing them, especially in the case of in situ or borderline. A recent study of pathologists’ diagnostic accuracy concluded that up to one in six melanomas may be misdiagnosed due to an inter-pathologist variation of up to 45%. This study also showed that 33% of skin lesion biopsies receive a different diagnosis when reviewed by the same pathologist 8 or more months apart. The authors conclude that diagnosis of skin lesions ranging from benign to invasive melanoma are nether accurate nor reproducible. The development of molecular tools to compliment visual assessments is suggessted (Elmore et al, 2017). In addition to the risk of under and overtreatment caused by diagnostic imprecision, there is significant economic impact. In the United States alone, it is estimated that this diagnostic imprecision adds US$673 million to the cost of melanoma management. (Bhattacharya et al, 2017; Elmore et al, 2017).
MicroRNAs (miRNAs) are a class of small regulatory molecules with unique physical properties that make them strong candidates for diagnostic or prognostic assay development (Keller et al, 2011). These non-protein-coding RNAs are present both within and external to melanoma cells and have strong tissue and disease specificity. Extracellular miRNAs are secreted by cells in exosomes; virus-sized particles that facilitate inter-cellular exchange of molecular information. While all cells release exosomes into their microenvironment, those from cancer cells have a higher concentration of miRNAs compared to normal cells (Tomasetti et al, 2017). MicroRNA-rich melanoma exosomes have a functional role in tumour invasion, by stimulating cancer-activated fibroblasts at the primary tumour site and also triggering events at distant sites to facilitate the growth of metastases (Peinado et al, 2012).
The first comprehensive study of circulating miRNAs in melanoma patients was performed in 2010 (Leidinger et al, 2010). Whole-blood samples from 35 individuals with melanoma of various clinical stages were compared with normal controls, resulting in a set of 51 differentially expressed miRNAs. While this study demonstrated the diagnostic potential of these molecules, the work was performed using a custom miRNA detection platform not readily available outside of the research setting and the genes identified were not evaluated for association with other disease types, limiting the clinical utility of the work.
An extensive review of circulating miRNA studies for melanoma diagnosis was performed by Carpi et al (2016), identifying over 40 publications on the topic. The authors conclude that for miRNA technology to be useful, clinical practice for melanoma four areas need to be addressed, namely (i) the lack of reproducibility between studies, (ii) the wide variety of evaluation techniques, (iii) individual cancer variation, and (iv) prospective trials validation. The robust level of scientific consensus as to the suitability of circulating miRNAs as melanoma biomarkers was also noted.
While there are well defined challenges in developing a novel cancer biomarker, there is a clear need for additional methods of detecting the presence of malignant melanoma, particularly for high risk individuals, in which up to 50% of all melanomas occur (Williams et al, 2011). The goal of this study was to perform miRNA analysis using readily available, high throughput laboratory methods and to develop a diagnostic biomarker for cutaneous melanoma in order to complement existing methods of detection. The NanoString system is highly suited for non-invasive detection of miRNAs in plasma, where RNA quantities are often miniscule. It is an amplification-free based method of gene expression analysis that requires lower amounts of starting materials and has lower cost compared with microarray or sequencing-based methods (Chatterjee et al, 2015). A robust biomarker may also be useful in the post-treatment setting, where objective and non-invasive methods for detecting melanoma recurrence and monitoring response to novel therapies are also needed in order to improve patient outcomes.
Patients and methods
Specimen collection and microRNA profiling
To create a database of circulating miRNA expression profiles of individuals with and without melanoma, 0.5–2 ml of cell-free plasma was obtained from 16 healthy controls and 32 Caucasian individuals at the time of their diagnosis with cutaneous melanoma (Cureline Inc., CA, USA and Folio Biosciences, OH, USA). The institutional ethics board of each hospital, in collaboration with Cureline and Folio Biosciences, approved the use of the tissue material. Written informed consent was obtained from each patient.
All melanoma cases were confirmed by histopathological examination before the corresponding plasma sample was processed further. Details of the 48 individuals used are provided in Table 1, including tumour depth (T), nodal involvement (N), and the presence or absence of metastases (M). Using current American Joint Committee on Cancer staging guidelines; 4, 18, 4, and 4 melanoma patients involved in this section of the study had stage I, II, III, and IV disease at the time of specimen collection, respectively.
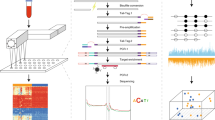
Cell-free miRNA purification was performed by Canopy Biosciences (St Louis, MO, USA) using the QIAGEN miRNeasy Serum/Plasma Kit (cat no./ID: 217184), incorporating the miRNeasy Serum/Plasma Spike-In Control (cat no./ID: 219610) as per manufacturer recommendations. Total RNA was eluted in 100 μl water and concentrated to 20 μl. Three microlitres was used for profiling on the multiplexed nCounter microRNA Assay V3 from NanoString Technologies (Seattle, WA, USA). The assay detects 800 human miRNAs curated from miRBase Version 21 (Kozomara and Griffiths-Jones, 2014).
Purified RNA samples were prepared for detection and quantification by ligating a specific DNA tag (miR-tag) onto the 3′ end of each mature miRNA per the manufacturer’s instructions. Excess tags were removed by restriction digestion at 37 °C. Hybridisations were carried out by combining 5 μl of each miRNA-miR-Tag sample with 20 μl of nCounter Reporter probes in hybridisation buffer and 5 μl of nCounter Capture probes at 65 °C for 16–20 h.
Excess probes were removed using a two-step magnetic bead-based purification on the nCounter Prep Station. Levels of specific target molecules were quantified using the nCounter Digital Analyzer by counting individual fluorescent barcodes. Assay quality control was performed using nSolver version 3 using a combination of metrics including percent field of view registration, minimum binding density and positive control linearity, with manufacturer recommended thresholds for miRNA analysis.
Data analysis
Raw gene counts were adjusted for background noise by negative control adjustment and normalised to spike-in genes cel-miR-254 and osa-miR-414 using Nanostring nSolver V3.0 (Nanostring Inc, WA, USA). To prepare the data for statistical comparison between normal and melanoma classes, genes with no or low detection rates across all samples were excluded and variance modelling at the observational level (voom) transformation was then applied (Law et al, 2014). This method estimates the mean-variance relationship of the log-counts, generates a precision weight for each observation and enters these into the limma empirical Bayes analysis pipeline for downstream gene selection.
For count-based data such as Nanostring or RNA-seq, voom transformation has been shown to produce the most accurate assessment of true differential expression, with the lowest false discovery rate (Ritchie et al, 2015). The empirical Bayes method of gene expression analysis uses a pooled estimate of sample variance and gives a stable assessment of expression level variance between and within classes. This is important when the number of samples measurements is smaller than the number of genes measured in each.
Classification models were constructed using linear support vector machine (SVM) algorithm developed by Vapnik et al (1995). The SVM predictor is a linear function of voom-transformed count data that best separates the data subject to penalty costs on the number of specimens misclassified. Statistical analysis and algorithm development were performed using R 3.4 (R Core Team, 2014), Bioconductor 3.5, Minitab 17.1 and Medcalc 17.6 (2010; Schoonjans et al, 1995; Gentleman et al, 2004).
Independent validation
To further explore the performance of the novel miRNA signature identified, multiple independent data sets were downloaded from public gene expression data repositories (Table 2). In total, these data sets represent 473 unique melanoma patients, normal control individuals or cell-line models, with the clinical or technical details of each as originally published. An downside to this approach is that due to the evolving nature of mirBase (Kozomara & Griffiths-Jones, 2014), there is an incomplete overlap in the miRNA content of the Nanostring, Agilent, Affymetrix, and custom platforms used to generate the data sets used in this study. In addition to varying probe content, the method of miRNA isolation/enrichment, detection chemistry and data normalisation methods associated with each technology are unique and can lead to challenges in comparing data between experiments(Kolbert et al, 2013).
Results
Identification of circulating microRNAs differentially expressed between melanoma patients and normal controls
Gene selection and functional annotation
Circulating miRNA gene expression profiles of melanoma patients and normal control donors were compared using voom and limma (Law et al, 2014), after probes corresponding to non-expressed genes in a majority of samples were excluded. Those miRNAs with a limma FDR of <0.01 and an absolute fold-change >2.0 were selected (n=38). Supplementary Figure 2 shows the relationship between log FDR and fold change for each gene before (a) and after (b) filtering and gene selection. The 38 genes differentially expressed between melanoma and normal donor expression profiles are hereon referred to as MEL38 and shown in Table 3.
Literature searches revealed that 22 of the miRNAs in the MEL38 signature (58%) have been previously associated with melanoma, predominantly in studies of cell lines or skin biopsies rather than blood, serum or plasma. Notably, two of the 38 genes (hsa-miR-1537-3p and hsa-miR-181b-5p) have only been previously identified in studies that used next-generation sequencing methods, demonstrating the comparable level of sensitivity of the Nanostring platform (Kolbert et al, 2013).
The reported biological or molecular functions of each gene in MEL38 can be distilled into four categories. These are angiogenesis and inflammation (n=2), cancer cell invasion and metastasis (n=14), immune system and treatment resistance (n=11), and tumour suppression or oncogene regulation (n=8), with a number of genes having dual functions. A summary of the reported functions or biological roles of each member of MEL38 and relevant references is provided in Supplementary Table 3.
Gene set analysis was also performed using the miRNA Enrichment Analysis and Annotation Tool (miEAA; https://ccb-compute2.cs.uni-saarland.de/mieaa_tool/), which uses a database of miRNA predicted or validated target gene binding sites to identify significantly enriched functional or pathway gene sets (Backes et al, 2016). Among the biologically relevant and statistically significant pathways targeted by MEL38 are melanogenesis (hsa04916), T-cell activation (P00053) and both RAS (P04393) and MAPK (WP422) oncogene activation, as shown in Supplementary Table 1.
A classification algorithm to predict disease state from circulating microRNAs
In order to demonstrate the potential of the MEL38 signature to predict the melanoma or normal (non-melanoma) status of a plasma sample, a support vector machine algorithm was trained on the information present in the Nanostring plasma miRNA discovery series (i.e., 38 genes × 48 samples) using leave-one-out cross validation (Ben-Dor et al, 2000). Gene re-selection was not performed as limma method used for their selection strictly controls for false discoveries and adjusts for multiple testing bias. As expected from a discovery series, the MEL38 SVM showed a high degree of accuracy as shown in Figure 1A, with clear separation of scores based on disease status apparent.
Training and independent validation series results. (A) Training series support vector machine (SVM): MEL38 scores generated from circulating microRNA profiles of normal control individuals and melanoma patients with stage I–IV disease. (B) A 28-gene subset of MEL38 applied to circulating microRNA profiles generated from blood collected from normal controls and melanoma patients with stage I–IV disease (n=57). The 28-gene subset SVM was partially retrained with leave-one-out cross validation to and applied to the peripheral blood microRNA independent validation series. 3C Independent validation of the microRNA signature of melanoma in. (C) Solid line: ROC analysis of the SVM classifier as trained on the 48-sample Nanostring discovery series. AUC=0.79, P<0.001. Dotted line: ROC analysis of the SVM classifier partially retrained on the 57 sample independent validation series using the 28 out of 38 genes available. AUC=0.94, P<0.001.
Independent validation of MEL38 using peripheral blood microRNA profiles of individuals with or without melanoma
To independently validate the significance of the circulating miRNAs identified from the discovery series of forty eight Nanostring whole-miRNAome profiles, additional genomic data sets were sourced from the NCBI Gene Expression Omnibus (GEO) and EBI ArrayExpress. To date, no other group has published a multi-sample plasma ‘microRNome’ profiling study of melanoma. As such, the most suitable validation data set identified was GEO ID GSE20994, which is a series of 57 miRNA profiles generated from peripheral blood of melanoma patients (n=35) and normal controls (n=22). These data were generated using a custom miRNA oligonucleotide microarray, based on version 13.0 of the Sanger MirBase; (febit Homo Sapiens miRBase 13.0, Hummingbird Diagnostics, Heidelberg, Germany).
By comparing probe annotations and sequences between the Nanostring and febit platforms, 28 of the MEL38 miRNAs (74%) were matched. The MEL38 SVM algorithm was applied to the validation data set in two ways; (1) using the exact SVM gene weights obtained from the discovery series; and to partially compensate for the incomplete overlap, (2) re-calculation of SVM gene weights using leave-one-out cross validation. A box plot of the method (2) SVM classification score for each patient/control individual in this series, grouped by disease state and clinical stage, is shown in Figure 1B.
Both SVM application approaches resulted in statistically significant stratification of melanoma and normal profiles. Method 1 gave a sensitivity of 71% and specificity of 86% for prediction of melanoma. ROC analysis was performed and an AUC of 0.79 (P<0.001, Figure 1C – solid line) was observed. Method 2 gave a higher sensitivity of 89% and the same specificity of 86%. The AUC of method (2) was 0.94 (P<0.001, Figure 1C – dotted line). Permutation analysis performed during the LOOCV process method 2 revealed that the probability of observing this level of classification accuracy by chance alone was P<0.01.
As these results are based on (i) an incomplete subset of the total MEL38 gene signature, (ii) from a study using whole blood rather than plasma only, and (iii) were generated using microarray rather than Nanostring profiling, it is likely that the performance of the signature will improve as additional clinical validation series are performed. Despite the caveats of this validation exercise, a robust and statistically significant association was observed between a SVM classifier based on a 28/38 genes in MEL38 and the likelihood of an individual having cutaneous melanoma.
General linear model analysis of MEL38 SVM scores in relation to demographic and clinical variables
Statistical analyses of the MEL38 circulating miRNA classification score for individuals in the training and independent validation series were performed using general linear models (Supplementary Table 2). The discovery series SVM scores showed a highly significant difference between melanoma and normal samples (P<0.001), with no difference associated with age or gender (P>0.05). In the independent validation series, the 28-gene subset SVM score was also significantly different between disease status, with or without partial retraining (P<0.001 and P=0.002, respectively).
In the melanoma patient-only subset of each series, the relationship between the SVM score and tumour thickness, melanoma type (superficial, nodal, or amelanotic), disease stage (I–IV), age, and sex (where available) was also investigated. None of these variables were found to be statistically significant in any of the general linear models (P>0.05).
These results confirm that the MEL38 signature is significantly associated with melanoma state (i.e., the presence or absence of melanoma), over and above any association with patient age, gender or tumour thickness, subtype, and stage of progression.
In vitro assessment of MEL38 expression in exosomes isolated from melanoma and normal skin cell lines.
An additional in vitro validation of the MEL38 gene signature was carried out using Affymetrix miRNA GeneChip profiles of normal melanocytes cell line HEM-LP, melanoma cell line A375, and the isolated exosomes of each (GEO ID: GSE35387). These data were generated by Xiao and colleagues. who developed a method to purify exosomes from cell culture supernatant using multiple rounds of centrifugation and filtration, ensuring removal of whole cells and debris, before verifying the presence of pure exosomes using transmission electron microscopy and performing Affymetrix miRNA analysis of their contents (Xiao et al, 2012).
Affymetrix probes corresponding to 26 of the MEL38 genes (68%) were identified by probe annotation and sequence comparison. Hierarchical clustering using these genes showed similar expression profiles of matched cells and exosomes, with larger differences between normal and melanoma states (Figure 2A). A cross-validated SVM classifier was trained on the 26-gene subset, using LOOCV, the results of which showed clear stratification of melanoma (cells or exosomes) vs normal (cells or exosomes) (Figure 2B). These findings show that MEL38 genes exist at similar relative levels both with, and external to, their cell of origin.
Additional independent validation series. (A) Hierarchical clustering of microRNA expression levels in melanoma cell line A375, normal melanocyte cell line and exosomes experimentally isolated from both cell lines shows separation between disease status phenotypes. (B) MEL38 SVM score calculated from microRNA expression data from normal skin and melanoma cell lines, and their respective exosomes isolated from tissue culture. (C) Hierarchical clustering of MEL38 measured in melanoma and nevus FFPE tissue, profiled using Agilent microRNA microarrays. Clear separation based on disease status can be seen, supporting the hypothesis that genes in the MEL38 signature originate from melanoma or nevi cells. (D) MEL38 SVM scores calculated on microRNA gene expression profiles generated from FFPE nevi and melanoma biopsies. Case IDs are shown on x axis, MEL38 SVM classification score on Y axis. SVM=support vector machine.
MEL38 expression in melanoma compared to other cancer types
To further evaluate the melanoma-specific association of the MEL38 signature circulating miRNA profiles of individuals with melanoma were compared with eight other malignancies (colon, lung, ovarian, prostate, breast, renal, stomach, and wilms tumour; N=393, GEO ID GSE61741). These data were generated by Keller et al (2014), Saarland University (Hamburg, Germany) using a the febit custom miRNA analysis platform on which 28 of the MEL38 genes are present. T-tests of differential expression between melanoma and ‘other’ cancers were calculated. Fifteen miRNAs (54%) exhibited significantly different levels of expression between melanoma vs other cancer types (P<0.05; range P=4.3 × 10−6 to P=0.036).
This analysis shows that a majority of the tested MEL38 genes have patterns of expression that are specific to melanoma vs other common cancer types.
MEL38 expression in melanoma vs melanocytic nevi biopsy tissue
MicroRNA profiles generated from formalin-fixed, paraffin-embedded melanoma (n=16) and melanocytic nevi biopsies (n=3) were downloaded from EBI ArrayExpress (accession E-MTAB-4915). These data were generated by Komina et al (2016) using tissue obtained from individuals ranging from 35 to 81 years old, with tumours from 2–7 mm thick and Clark’s level III–IV. All of the MEL38 genes were found on the Affymetrix microRNA GeneChip 4.1 (mirBase v20) used for this study.
T-tests were performed between melanoma and nevi sample classes and thirteen (34%) of the MEL38 genes were significantly different (P<0.05). Twenty-one MEL38 genes (55%) had a fold-change difference of >1.5. Hierarchical clustering of samples using the MEL38 set resulted in perfect separation of melanoma and benign samples (Figure 2C). A cross-validated MEL38 SVM classifier was trained on these data (without gene re-selection) and the SVM score for each individual is shown in Figure 2D. These results show that MEL38 gene signature also has the ability to discriminate between melanoma and benign biopsy nevi tissue, further supporting the skin-cell origin of these circulating miRNAs and suggesting an additional utility for the signature.
Comparison with previously published circulating microRNA signatures of melanoma
Friedman et al (2012) discovered a set of five serum miRNAs with potential as serum-based biomarkers for recurrence in melanoma using RT–PCR methods. Three of these candidate genes (miR-199a-5p, miR-33a-5p, and miR-424-5p) were also significantly differentially expressed in the Nanostring plasma miRNA discovery series (mirBase v21) generated for this study (P<0.05).
Leidinger et al (2010) identified an optimal set of 16 miRNAs from their whole-blood custom-microarray data set (mirBase v12). Eight of the 16 genes in their signature met, or approached, statistical significance in the Nanostring plasma discovery series (five genes: P<0.05, three genes: P<0.1).
Margue et al (2015) published a set of eight miRNAs profiled on the Affymetrix miRNA GeneChip 1.0 (mirBase v11) which were differentially expressed in late stage melanoma patients when compared to healthy individuals. Of the seven signature genes that are present in the Nanostring Human MicroRNA V.3 assay, four are differentially expressed in the discovery series (57%).
Stark et al measured the expression of 16 miRNAs in melanoma tissue and serum (Stark et al, 2010, 2015) that were previously identified as being differentially expressed between melanoma cell lines when compared to 14 ‘other’ solid malignancies. Thirteen of these genes are technically present in the Nanostring assay and 6/13 (46%) were significantly differentially expressed in the MEL38 discovery series.
Of the total combined set of miRNAs from these previously published melanoma miRNA signatures, a majority were found to be differentially expressed in the Nanostring plasma discovery series. Three of these satisfied the combination of both statistical (P-value) and biological (fold-change) significance we employed to select MEL38; miR-205-5p, miR-424-5p, and miR-301a-3p.
Circulating microRNA and melanoma stage of progression
Finally, we examined whether the expression of any miRNAs in our discovery series were correlated AJCC clinical stage. Quality-filtered plasma profiles from melanoma patients were grouped by stage I, II, III, and IV (Table 1). Voom+limma analysis was performed to identify individual miRNAs with differential expression between any two stages (FDR <0.1 and fold-change >2). Next, the correlation coefficient for each gene vs stage was calculated and those <−0.7 or >0.7 were selected. This resulted in a final set of 18 miRNAs, eight of which overlapped with MEL38 (Figure 3B). The average fold change of each gene relative to its expression in stage I is shown in Figure 3B.
Circulating microRNA and melanoma clinical stage. (A) Venn diagram of MEL38 (diagnostic) and MEL18 (staging) signatures. (B) Line plot of MEL18 gene expression fold-changes relative to melanoma stage I. Each gene in MEL18 exhibits significant difference between one or more stages of melanoma progress and has a correlation coefficient of +0.7 or −0.7 with stage I–IV disease.
This 18-gene-signature may be useful, as an adjunct to the binary diagnostic capability of MEL38, in providing additional information about the extent of melanoma progression. With further validation, this signature (MEL18) may assist in reducing the time between diagnosis and treatment.
Discussion
To address the need for improved melanoma detection and monitoring tools, we have identified a novel set of thirty eight circulating miRNAs by performing Nanostring nCounter analysis of forty eight plasma samples from individuals with or without cutaneous melanoma (MEL38). The ability of the signature to predict disease status independently validated, on multiple previously published data sets, including a series of 57 individuals with or without melanoma. Application of a 28-gene subset classifier to these data showed a high degree of accuracy; 71–97% sensitivity (without and with partial SVM retraining, respectively) and 86% specificity. We anticipate improved performance on future validation studies, where fewer technical differences in experimental design will be present.
These results compare favourably to the average sensitivity and specificity of melanoma detection in general practice, which is reported to be approximately 62% and 84%, respectively, however individual studies report wide ranges (Aitken et al, 2006; Herschorn, 2012). Furthermore, these numbers do not take into account the impact of imprecision in histological examination (akin to technical reproducibility), which results in 17% of cases being misdiagnosed (8% of cases over-interpreted, 9% under-interpreted). As the technical imprecision of genomic profiling has been shown to be negligible (1–3%), MEL38 shows strong potential to improve the accuracy of melanoma diagnosis if incorporated alongside conventional techniques (Ach et al, 2007; van Laar et al, 2014).
To test the cellular origin hypothesis of the circulating miRNAs signature developed, analysis of melanoma cell line, normal melanocyte, and purified exosome miRNA data was performed. Clustering and SVM modelling using MEL38 in these data showed strong separation of normal vs melanoma profiles, with remarkable expression similarity between whole cells and their respective exosomes. The MEL38 signature were also compared between melanoma and benign FFPE tissue biopsies. Hierarchical clustering and SVM classification of these data again showed separation basis of disease state, further supporting the cellular origin hypothesis and biomarker potential of the signature.
A majority of the genes that make up four previously published miRNA signatures of melanoma, also exhibited differential expression in our Nanostring discovery series. This is despite being generated using different technology platforms, patient populations, and specimen processing protocols. The commonality of differential expression between this and previous melanoma miRNA profiling studies addresses a common criticism of genomics, that there is often limited consistency in findings between experiments performed using multiple platforms.
Despite the substantial overlap in gene signatures and the MEL38 discovery series, only three of the miRNAs contained in the previously published signatures were actually part of the final 38 gene selection. This is most likely due to the combination of biological (fold-change) and statistical (FDR) significance criteria used, which represent a unique element of this study. These three miRNAs have well defined roles in melanoma oncogenesis and progression. miR-424-5p has shown promise as a prognostic serum biomarker, being significantly upregulated in patients with poor prognosis. (Rosa et al, 2007; Ghosh et al, 2010). miR-205-5p is differentially expressed between primary and metastatic melanomas and has a role in regulating epithelial to mesenchymal transition, an key step in the metastatic process (Xu et al, 2012). Finally, miR-301a-3p is known to control the expression of the tumour suppressor PTEN. This miRNA significantly up-regulated in melanoma vs benign biopsy tissue and is also correlated poor prognosis (Cui et al, 2016).
The molecular pathways and functional themes represented by MEL38 further support the disease- and tissue-specificity of the signature. Notably, the T-cell activation pathway is the highest ranked for significance of overlap with MEL38 (Supplementary Table 1). This observation aligns with recent advances in the understanding of how melanoma cells modulate the anti-tumour immune response, thus facilitating their growth and progression (Schatton et al, 2015). In addition to genes involved in resisting the body’s defense system, MEL38 miRNAs have involvement in angiogenesis (e.g., hsa-miR-497-3p, which inhibits vascular endothelial growth factor A), metastasis (e.g., miR-3928-3p, which targets ERBB3; required for metastasis formation), and tumour growth control (e.g., miR-548a-5p, a negative regulator of the tumour inhibitor gene Tg737 (Tiwary et al, 2014; Yan et al, 2015; Zhao et al, 2016).
The role of miRNA-rich exosomes as mediators of melanoma development and progression, both at the primary and metastatic site, has only recently been elucidated. Melanoma exosomes have been demonstrated to cause vascular leakiness at pre-metastatic sites and to modify bone marrow progenitor cells with a pro-angiogenic phenotype (Peinado et al, 2012). They also allow melanoma cells to modify the stromal niche, with miRNA stimulation of cancer associated fibroblasts, paving the way for invasion and metastatic spread (Dror et al, 2016). There is a growing body of evidence that cancer-derived exosomes circulate through the body containing pro-metastatic, anti-immune-response, genetic instructions, and thus have a key role in all stages of melanoma progression.
In conclusion, a signature of melanoma-specific circulating miRNAs has been identified in human plasma and independently validated in multiple data sets representing miRNA profiles of whole blood, cell lines, and solid tissue. The signature was developed using a technology platform which due to its high specificity, low specimen requirements and cost efficiency, is readily adaptable to clinical use (Chatterjee et al, 2015). A majority of the genes in the signature have previously been associated melanoma development or progression and are known regulators of relevant oncogenic processes. Extensive additional clinical and technical validation studies will be required to further define the true diagnostic performance of an assay based on MEL38 and its utility in melanoma diagnosis, treatment response monitoring and relapse detection. We believe a non-invasive clinically available biomarker for melanoma such as MEL38 has the potential to improve diagnostic accuracy, reduce healthcare spending and reverse the mortality rate of a highly treatable disease.
Change history
20 March 2018
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Ach RA, Floore A, Curry B, Lazar V, Glas AM, Pover R, Tsalenko A, Ripoche H, Cardoso F, d'Assignies MS, Bruhn L, Van't Veer LJ (2007) Robust interlaboratory reproducibility of a gene expression signature measurement consistent with the needs of a new generation of diagnostic tools. BMC Genomics 8: 148.
Aitken JF, Janda M, Elwood M, Youl PH, Ring IT, Lowe JB (2006) Clinical outcomes from skin screening clinics within a community-based melanoma screening program. J Am Acad Dermatol 54 (1): 105–114.
Australian Institute of Health and Welfare (2017) Australian Cancer Incidence and Mortality (ACIM) books: Australian Government. https://www.aihw.gov.au/reports/cancer/acim-books/contents/acim-books.
Backes C, Khaleeq QT, Meese E, Keller A (2016) miEAA: microRNA enrichment analysis and annotation. Nucleic Acids Res 44 (W1): W110–W116.
Ben-Dor A, Bruhn L, Friedman N, Nachman I, Schummer M, Yakhini Z (2000) Tissue classification with gene expression profiles. J Comput Biol 7 (3-4): 559–583.
Bhattacharya A, Young A, Wong A, Stalling S, Wei M, Hadley D (2017) Precision diagnosis of melanoma and other skin lesions from digital images. AMIA Jt Summits Transl Sci Proc 2017: 220–226.
Carpi S, Polini B, F S, Romanini A (2016) Management of Malignant Melanoma. SMGE Books: SM Group: Dover, DE, USA.
Chatterjee A, Leichter AL, Fan V, Tsai P, Purcell RV, Sullivan MJ, Eccles MR (2015) A cross comparison of technologies for the detection of microRNAs in clinical FFPE samples of hepatoblastoma patients. Sci Rep 5: 10438.
Cui L, Li Y, Lv X, Li J, Wang X, Lei Z, Li X (2016) Expression of microRNA-301a and its functional roles in malignant melanoma. Cell Physiol Biochem 40 (1–2): 230–244.
Dror S, Sander L, Schwartz H, Sheinboim D, Barzilai A, Dishon Y, Apcher S, Golan T, Greenberger S, Barshack I, Malcov H, Zilberberg A, Levin L, Nessling M, Friedmann Y, Igras V, Barzilay O, Vaknine H, Brenner R, Zinger A, Schroeder A, Gonen P, Khaled M, Erez N, Hoheisel JD, Levy C (2016) Melanoma miRNA trafficking controls tumour primary niche formation. Nat Cell Biol 18 (9): 1006–1017.
Elmore JG, Barnhill RL, Elder DE, Longton GM, Pepe MS, Reisch LM, Carney PA, Titus LJ, Nelson HD, Onega T, Tosteson ANA, Weinstock MA, Knezevich SR, Piepkorn MW (2017) Pathologists’ diagnosis of invasive melanoma and melanocytic proliferations: observer accuracy and reproducibility study. BMJ 357: j2813.
Elliott TM, Whiteman DC, Olsen CM, Gordon LG (2017) Estimated Healthcare Costs of Melanoma in Australia Over 3 Years Post-Diagnosis. Appl Health Econ Health Policy 15 (6): 805–816.
Fransen M, Karahalios A, Sharma N, English DR, Giles GG, Sinclair RD (2012) Non-melanoma skin cancer in Australia. Med J Aust 197 (10): 565–568.
Friedman EB, Shang S, de Miera EV, Fog JU, Teilum MW, Ma MW, Berman RS, Shapiro RL, Pavlick AC, Hernando E, Baker A, Shao Y, Osman I (2012) Serum microRNAs as biomarkers for recurrence in melanoma. J Transl Med 10: 155.
Gentleman RC, Carey VJ, Bates DM, Bolstad B, Dettling M, Dudoit S, Ellis B, Gautier L, Ge Y, Gentry J, Hornik K, Hothorn T, Huber W, Iacus S, Irizarry R, Leisch F, Li C, Maechler M, Rossini AJ, Sawitzki G, Smith C, Smyth G, Tierney L, Yang JY, Zhang J (2004) Bioconductor: open software development for computational biology and bioinformatics. Genome Biol 5 (10): R80.
Ghosh G, Subramanian IV, Adhikari N, Zhang X, Joshi HP, Basi D, Chandrashekhar YS, Hall JL, Roy S, Zeng Y, Ramakrishnan S (2010) Hypoxia-induced microRNA-424 expression in human endothelial cells regulates HIF-α isoforms and promotes angiogenesis. J Clin Invest 120 (11): 4141–4154.
Herschorn A (2012) Dermoscopy for melanoma detection in family practice. Can Fam Physician 58: 740–745.
Keller A, Leidinger P, Bauer A, Elsharawy A, Haas J, Backes C, Wendschlag A, Giese N, Tjaden C, Ott K, Werner J, Hackert T, Ruprecht K, Huwer H, Huebers J, Jacobs G, Rosenstiel P, Dommisch H, Schaefer A, Müller-Quernheim J, Wullich B, Keck B, Graf N, Reichrath J, Vogel B, Nebel A, Jager SU, Staehler P, Amarantos I, Boisguerin V, Staehler C, Beier M, Scheffler M, Büchler MW, Wischhusen J, Haeusler SF, Dietl J, Hofmann S, Lenhof HP, Schreiber S, Katus HA, Rottbauer W, Meder B, Hoheisel JD, Franke A, Meese E (2011) Toward the blood-borne miRNome of human diseases. Nat Methods 8 (10): 841–843.
Keller A, Leidinger P, Vogel B, Backes C, ElSharawy A, Galata V, Mueller SC, Marquart S, Schrauder MG, Strick R, Bauer A, Wischhusen J, Beier M, Kohlhaas J, Katus HA, Hoheisel J, Franke A, Meder B, Meese E (2014) miRNAs can be generally associated with human pathologies as exemplified for miR-144. BMC Med 12: 224.
Kolbert CP, Feddersen RM, Rakhshan F, Grill DE, Simon G, Middha S, Jang JS, Simon V, Schultz DA, Zschunke M, Lingle W, Carr JM, Thompson EA, Oberg AL, Eckloff BW, Wieben ED, Li P, Yang P, Jen J (2013) Multi-platform analysis of microRNA expression measurements in RNA from fresh frozen and FFPE tissues. PLoS One 8 (1): e52517.
Komina A, Palkina N, Aksenenko M, Tsyrenzhapova S, Ruksha T (2016) Antiproliferative and pro-apoptotic effects of MiR-4286 inhibition in melanoma cells. PLoS One 11 (12): e0168229.
Kozomara A, Griffiths-Jones S (2014) miRBase: annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res 42: D68–D73.
Law CW, Chen Y, Shi W, Smyth GK (2014) voom: precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol 15 (2): R29.
Leidinger P, Keller A, Borries A, Reichrath J, Rass K, Jager SU, Lenhof HP, Meese E (2010) High-throughput miRNA profiling of human melanoma blood samples. BMC Cancer 10: 262.
Luftos P, Wilslow R (2015) FDA approves Bristol-Myers’s Yervoy, Opdivo for treatment of melanoma. In The Wall Street Journal. Dow Jones & Company: New York.
Margue C, Reinsbach S, Philippidou D, Beaume N, Walters C, Schneider JG, Nashan D, Behrmann I, Kreis S (2015) Comparison of a healthy miRNome with melanoma patient miRNomes: are microRNAs suitable serum biomarkers for cancer? Oncotarget 6(14): 12110–12127.
Peinado H, Alečković M, Lavotshkin S, Matei I, Costa-Silva B, Moreno-Bueno G, Hergueta-Redondo M, Williams C, García-Santos G, Ghajar C, Nitadori-Hoshino A, Hoffman C, Badal K, Garcia BA, Callahan MK, Yuan J, Martins VR, Skog J, Kaplan RN, Brady MS, Wolchok JD, Chapman PB, Kang Y, Bromberg J, Lyden D (2012) Melanoma exosomes educate bone marrow progenitor cells toward a pro-metastatic phenotype through MET. Nat Med 18 (6): 883–891.
R Core Team (2014). R: A language and environment for statistical computing. R Foundation for Statistical Computing: Vienna, Austria. http://www.R-project.org/.
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK (2015) limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 43 (7): e47.
Rosa A, Ballarino M, Sorrentino A, Sthandier O, De Angelis FG, Marchioni M, Masella B, Guarini A, Fatica A, Peschle C, Bozzoni I (2007) The interplay between the master transcription factor PU.1 and miR-424 regulates human monocyte/macrophage differentiation. Proc Natl Acad Sci USA 104 (50): 19849–19854.
Schatton T, Schütte U, Frank MH (2015) Effects of malignant melanoma initiating cells on T-cell activation. In: Methods in Molecular Biology. Humana Press Springer: New York, NY, USA.
Schoonjans F, Zalata A, Depuydt CE, Comhaire FH (1995) MedCalc: a new computer program for medical statistics. Comput Methods Programs Biomed 48 (3): 257–262.
Stark MS, Klein K, Weide B, Haydu LE, Pflugfelder A, Tang YH, Palmer JM, Whiteman DC, Scolyer RA, Mann GJ, Thompson JF, Long GV, Barbour AP, Soyer HP, Garbe C, Herington A, Pollock PM, Hayward NK (2015) The prognostic and predictive value of melanoma-related microRNAs using tissue and serum: a microRNA expression analysis. EBioMedicine 2 (7): 671–680.
Stark MS, Tyagi S, Nancarrow DJ, Boyle GM, Cook AL, Whiteman DC, Parsons PG, Schmidt C, Sturm RA, Hayward NK (2010) Characterization of the melanoma miRNAome by deep sequencing. PLoS One 5 (3): e9685.
Tiwary S, Preziosi M, Rothberg PG, Zeitouni N, Corson N, Xu L (2014) ERBB3 is required for metastasis formation of melanoma cells. Oncogenesis 3: e110.
Tomasetti M, Lee W, Santarelli L, Neuzil J (2017) Exosome-derived microRNAs in cancer metabolism: possible implications in cancer diagnostics and therapy. Exp Mol Med 49 (1): e285.
van Laar R, Flinchum R, Brown N, Ramsey J, Riccitelli S, Heuck C, Barlogie B, Shaughnessy Jr J (2014) Translating a gene expression signature for multiple myeloma prognosis into a robust high-throughput assay for clinical use. BMC Med Genomics 7 (1): 25.
Vapnik V (1995) The Nature of Statistical Learning Theory. Springer: Springer-Verlag: New York, NY, USA.
Williams LH, Shors AR, Barlow WE, Solomon C, White E (2011) Identifying persons at highest risk of melanoma using self-assessed risk factors. J Clin Exp Dermatol Res 2: 6.
Xiao D, Ohlendorf J, Chen Y, Taylor DD, Rai SN, Waigel S, Zacharias W, Hao H, McMasters KM (2012) Identifying mRNA, microRNA and protein profiles of melanoma exosomes. PLoS One 7 (10): e46874.
Xu Y, Brenn T, Brown ER, Doherty V, Melton DW (2012) Differential expression of microRNAs during melanoma progression: miR-200c, miR-205 and miR-211 are downregulated in melanoma and act as tumour suppressors. Br J Cancer 106 (3): 553–561.
Yan JJ, Zhang YN, Liao JZ, Ke KP, Chang Y, Li PY, Wang M, Lin JS, He XX (2015) MiR-497 suppresses angiogenesis and metastasis of hepatocellular carcinoma by inhibiting VEGFA and AEG-1. Oncotarget 6 (30): 29527–29542.
Zhao G, Wang T, Huang QK, Pu M, Sun W, Zhang ZC, Ling R, Tao KS (2016) MicroRNA-548a-5p promotes proliferation and inhibits apoptosis in hepatocellular carcinoma cells by targeting Tg737. World J Gastroenterol 22 (23): 5364–5373.
Acknowledgements
We acknowledge the physicians, patients, and control individuals who contributed to this study and to K Eisenbud and A Zielinska for helpful discussions and manuscript editing assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Geneseq Biosciences is a privately-held start-up company, aiming to develop a novel diagnostic biomarker for melanoma detection and management. A provisional patent application has been filed on the work described herein and we are actively pursuing clinical and technical validation partners locally and nationally.
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License.
Supplementary Information accompanies this paper on British Journal of Cancer website
Supplementary information
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Van Laar, R., Lincoln, M. & Van Laar, B. Development and validation of a plasma-based melanoma biomarker suitable for clinical use. Br J Cancer 118, 857–866 (2018). https://doi.org/10.1038/bjc.2017.477
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2017.477
Keywords
This article is cited by
-
Impact of modeled microgravity stress on innate immunity in a beneficial animal-microbe symbiosis
Scientific Reports (2024)
-
Circulating microRNAs as diagnostic biomarkers for melanoma: a systematic review and meta-analysis
BMC Cancer (2023)
-
Modeled microgravity alters apoptotic gene expression and caspase activity in the squid-vibrio symbiosis
BMC Microbiology (2022)
-
Pre-diagnostic DNA methylation in blood leucocytes in cutaneous melanoma; a nested case–control study within the Norwegian Women and Cancer cohort
Scientific Reports (2022)
-
Promising Blood-Based Biomarkers for Melanoma: Recent Progress of Liquid Biopsy and Its Future Perspectives
Current Treatment Options in Oncology (2022)