Abstract
Background:
Accumulating evidence implicates the tumour stroma as an important determinant of cancer progression but the protein constituents relevant for this effect are unknown. Here we utilised a bioinformatics approach to identify an extracellular matrix (ECM) gene signature overexpressed in multiple cancer types and strongly predictive of adverse outcome.
Methods:
Gene expression levels in cancers were determined using Oncomine. Geneset enrichment analysis was performed using the Broad Institute desktop application. Survival analysis was performed using KM plotter. Survival data were generated from publically available genesets.
Results:
We analysed ECM genes significantly upregulated across a large cohort of patients with ovarian, lung, gastric and colon cancers and defined a signature of nine commonly upregulated genes. Each of these nine genes was considerably overexpressed in all the cancers studied, and cumulatively, their expression was associated with poor prognosis across all data sets. Further, the gene signature expression was associated with enrichment of genes governing processes linked to poor prognosis, such as EMT, angiogenesis, hypoxia, and inflammation.
Conclusions:
Here we identify a nine-gene ECM signature, which strongly predicts outcome across multiple cancer types and can be used for prognostication after validation in prospective cancer cohorts.
Similar content being viewed by others
Main
The extracellular matrix (ECM) is a multi-molecular substance that serves functions ranging from cellular adhesion and motility to cell signalling. The extracellular matrix proteins are constructed from a relatively small repertoire of phylogenetically conserved amino-acid domains and genome-wide in silico analysis has led to the categorisation and indexing of the known protein constituents of the ECM. This ECM protein inventory, termed the core matrisome (CM) (Hynes and Naba, 2012), provides a platform for the analysis of physiological and disease-specific patterns of ECM protein expression.
Many solid tumours are characterised by the production of a dense, collagen-rich matrix, the deposition of which is associated with adverse outcome (Erler et al, 2006; Levental et al, 2009; Lu et al, 2012; Naba et al, 2014; Acerbi et al, 2015). The alteration in ECM protein constituents resulting from the development of a cancer causes reciprocal changes in the cancer cell and activates pathways responsible for cell migration, inhibition of apoptosis and proliferation (Pickup et al, 2014). The cancer ECM also promotes the formation of an unstable and chaotic vascular bed, poor oxygen delivery and hypoxia (Gilkes et al, 2014); factors that favour disease progression and metastasis.
Although excessive ECM deposition is a recognised hallmark of cancer, the specific proteins comprising the cancer ECM and their potential contribution to cancer biology and prognosis are less well studied. Here we address these issues using bioinformatics to compare the expression of CM genes in tumours and their normal tissue counterparts. We identify a nine-gene CM signature common to a range of cancers, the expression of which predicts poor prognosis in several solid cancer types. These data provide an impetus to further study proteins of the ECM in order to gain a greater understanding of cancer biology and develop clinical tools for prognostication.
Materials and methods
Ethics
All bioinformatics data were anonymised and required no ethical approval. Commercially available tissue microarrays were produced by US Biomax Inc. under the highest ethical standards with the donor being informed completely and with their consent. No ethical approval was required for this study.
Identification of the CM gene signature
All gene encoding proteins of the CM (http://matrisomeproject.mit.edu/) were included. Gene expression levels were determined in studies comparing lung, breast, ovarian, gastric, oesophageal or colorectal adenocarcinoma with normal tissue in Oncomine (www.oncomine.org/resource/login.html). Here gene expression data are normalised across studies enabling summative gene expression comparisons. Median gene rank (cancer vs normal analysis) was meta-analysed across studies of the same cancer type using Oncomine statistical algorithms. P values of the difference in gene rank were corrected for multiple hypothesis testing using the false discovery rate (FDR) method as described by Storey and Tibshirani (2003). Venn diagrams were generated and analysed using InteractiVenn (Heberle et al, 2015).
Immunohistochemisty
Immunohistochemistry for ECM proteins was performed on colon cancer and matched normal tissue microarrays from 20 patients (US Biomax Inc). Frozen tissues were immediately fixed in ice-cold acetone and blocked in normal serum of the species in which the secondary antibody was raised. Primary antibodies for col11a1 (ab64883), col1a1 (ab34710), col10a1 (ab58632) and spp1 (ab69498) were all obtained from ABCAM. Fluorochrome-conjugated secondary antibodies were obtained from Invitrogen. Tile-scanned images were taken at × 10 magnification using the Nikon Eclipse 90i epifluorescence microscope and were analysed using ImageJ.
Determination of the effect of the nine-gene CM signature on cancer outcome
Survival analysis was performed using KM plotter for ovarian (Gyorffy et al, 2012), gastric (Szász et al, 2016) and lung (Győrffy et al, 2013) cancers or GraphPad Prism for colorectal, renal, bladder or prostate cancer data derived from cBioportal (Cerami et al, 2012; Gao et al, 2013) or data set GSE17538. KM plotter is a manually curated, biannually updated database enabling survival analysis across multiple GEO data sets simultaneously. The GEO data sets used here are shown in Table 1. We used JetSet probes throughout (Li et al, 2011) and patients were divided into two groups on the basis of median expression of the nine-gene signature. For the analysis of colorectal cancer data sets, a z-score threshold of +1 for gene expression was used to define patients. Kaplan–Meier survival curves were constructed and compared using the log-rank method to generate hazard ratios and P values. Survival curves were generated using GraphPad Prism Version 7. Multivariate analysis was performed in lung, gastric and colorectal cancer data sets using SPSS (2017).
Geneset enrichment analysis
Geneset enrichment analysis (GSEA) analysis (Subramanian et al, 2005) was performed using the Broad Institute desktop application (http://software.broadinstitute.org/gsea/downloads.jsp) on RNA-Seq expression data from TCGA colorectal, gastric, ovarian and lung cancer data sets. Phenotypes were defined on the basis of expression of the nine-gene CM signature with samples divided into high or low expression again using a z-score of +1 to define groups. Genesets were identified in the molecular signatures database (http://software.broadinstitute.org/gsea/msigdb/index.jsp), with the exception of the angiogenesis geneset, which was identified in a recent publication describing a meta-analytical approach that identified a transcriptional programme for angiogenesis in human cancers (Masiero et al, 2013) and the EMT geneset (Gröger et al, 2012).
Results
Identification of a CM gene signature expressed by adenocarcinomas
In order to identify ECM proteins important for cancer progression, we compared the expression of genes comprising the CM in adenocarcinomas (ACs) and normal tissues. We focused specifically on ACs because in organs where both squamous cell carcinomas (SCCs) and ACs develop, these tumour types may originate from different cell lineages (Yan et al, 2010; Yuan et al, 2010), are characterised by different genetic landscapes (Contag et al, 2004; Gao et al, 2014) and demonstrate differences in their sensitivity to various treatment modalities (Katanyoo et al, 2012; Chen et al, 2014). These differences may relate in part to differences in the composition of the tumour ECM or regulation of ECM expression, and we did not want this to act as a confounder in the identification of a CM gene signature.
We identified a large number of CM genes expressed at significantly higher levels in ACs compared with their normal tissue counterparts (Figure 1A). Cancers of the oesophagus and lung in particular were highly different from their parent organs, with 110 and 97 of 274 CM genes significantly upregulated in these cancers, respectively. In comparison, ovarian cancers demonstrated less of a difference with only 43 of 274 CM genes significantly upregulated in the tumour (Supplementary Table 1).
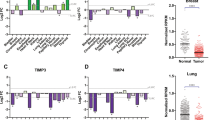
Development of a core matrisome gene signature from multiple cancer data sets. (A) CM gene expression based on gene rank for cancer vs normal tissue in various tumour types. Red squares indicate high rank in the cancer relative to the normal tissue. Grey indicates that the gene was not measured. Genes are listed in order of median rank across the analysis of included studies for that particular cancer type. (B) Venn diagram used to identify common CM genes that are significantly overrepresented throughout all cancer types identified in A. (C) The gene signature derived from the Venn diagram in B displaying the nine common, significantly upregulated genes identified across the analyses of all cancer types from A. (D) The nine-gene CM signature showing median gene rank (red=high expression) in cancer compared with normal tissue for each included study and FDR-corrected P values for the meta-analytical comparison. (E) Fluorescence immunohistochemistry for SPP1, Col10a1, Col1a1 and Col11a1 in colon cancers and matched normal colon with quantification of the area (%) of the microarray core demonstrating positive staining (n=20 per analysis). A full colour version of this figure is available at the British Journal of Cancer journal online.
We next identified genes that were significantly upregulated across all cancer types studied and defined a signature of nine such genes (Figure 1B–D). There was a significant correlation in the expression level of most of these genes in TCGA data sets of colon, gastric, lung and ovarian ACs (Supplementary Figure 1A and B), suggesting that the expression of these genes results from a common regulatory element.
Finally, immunohistochemistry demonstrated a significant increase in the expression of col11a1, col10a1 and spp1 proteins in colon cancer compared with matched normal colon tissues (Figure 1E). The expression of col1a1 was increased in cancer tissues compared with normal colon but this did not reach significance. Importantly, within colon cancers, each protein was identified within the stroma indicating deposition within the ECM. In normal colon, col11a1 was virtually undetectable and col10a1 was identified within the cytoplasm of colonic epithelial cells rather than within the stromal tissue compartment.
The nine-gene CM signature predicts long-term outcome in various cancer types
Given the widespread overexpression of the nine-gene CM signature in ACs compared to normal tissues, we hypothesised that the expression of these genes may be a requirement for cancer. Combined comparison of normalised gene expression data from multiple GSE data sets confirmed this hypothesis, as patients with cancers demonstrating overexpression of the nine-gene signature displayed reduced overall and progression-free survival for gastric, lung and ovarian cancers (Figure 2A). In three large colorectal cancer data sets, overexpression of the nine-gene signature was similarly associated with adverse outcome (Figure 2A) and it was also associated with reduced progression-free survival in a large TCGA breast cancer data set, as it was for cancers not used to generate the nine-gene signature such as those of the prostate and bladder (Supplementary Figure 2). Interestingly, there was no correlation between expression for the CM gene signature and survival in squamous cell carcinoma of the lung, head and neck or oesophagus or for oesophageal AC (Supplementary Figure 2). Multivariate analysis in gastric, lung and colorectal data sets identified the nine-gene CM signature as a factor significantly associated with disease-free survival independent of disease stage or grade (Table 2).
Expression of the core matrisome gene signature predicts survival in various cancer types. (A) Overall survival (top row) and recurrence-free survival (bottom row) for cohorts of patients whose tumours demonstrate overexpression (red) or normal expression (blue) of the nine-gene CM gene signature. Numbers represent hazard ratios (95% confidence intervals). (B) GSEA analysis of colorectal and gastric cancer TCGA data sets analysed for EMT, angiogenesis, hypoxia, inflammation, oxidative phosphorylation, apoptotic regulation and genomic instability geneset enrichment in patients with high or normal expression of the nine-gene CM signature. NES, normalised enrichment score. A full colour version of this figure is available at the British Journal of Cancer journal online.
GSEA analysis identifies biological traits associated with expression of the CM signature
Epithelial–mesenchymal transition (EMT), angiogenesis, hypoxia, inflammation and glycolysis are all features of cancer that have been associated with poor prognosis. Several of these processes have also been linked to ECM deposition (Lu et al, 2012). To gain an insight into the biological mechanisms through which the CM gene signature may define poor prognosis cancers, we performed GSEA analysis to determine whether genes governing these processes are overrepresented in cancers from patients overexpressing the nine-gene CM signature. Strikingly, colorectal, gastric (both Figure 2B), lung and ovarian cancers (both Supplementary Figure 3) expressing the CM gene signature were significantly enriched in EMT, hypoxia, angiogenesis and inflammation genesets, but showed reduced expression of the oxidative phosphorylation geneset. Importantly, several molecular signatures defining other cancer-related processes, including those for apoptosis or genomic instability, were not enriched in cancers expressing the CM signature (Figure 2B and Supplementary Figure 3).
Discussion
Here we present a comprehensive analysis of the difference in expression of CM genes in cancers and normal tissues in order to identify key constituents of the cancer ECM. We have identified commonality in the significant upregulation of nine CM genes across multiple cancer types, suggesting a potential requirement for these CM genes throughout solid tumours. Expression of the nine-gene signature predicted outcome in a broad range of cancers including those not initially used to generate the gene signature. These proteins are therefore associated with cancer progression and their combination may represent a useful biomarker for prognostication. Interestingly, the CM signature failed to predict overall or disease-free survival in squamous cell cancer data sets indicating that the ECM genes in the CM signature are not of relevance for the progression of SCC.
Col11a1 has previously been linked to cancer progression (Fischer et al, 2001; Cheon et al, 2014; Jia et al, 2016; Li et al, 2017) and is expressed by cancer stromal (Galván et al, 2014; Jia et al, 2016) and tumour cells progressed to EMT, where it promotes migration and invasion (Sok et al, 2013; Wu et al, 2014). Expression of secreted phosphoprotein 1 (SPP1, osteopontin) is also reported in cancer (Shevde and Samant, 2014) and is driven by cancer-related signalling pathways including Hedgehog, Wnt/β-catenin and NFκB (Shevde and Samant, 2014). SPP1 is also expressed by tumour-associated macrophages and fibroblasts, where it is linked to angiogenesis (Kale et al, 2014) and the metastatic cascade (Mi et al, 2011), respectively.
The protein products of several genes in our signature have not been thoroughly studied in relation to cancer; however, several are associated with functions of relevance to cancer progression. BGN, for example, interacts with toll-like receptors on the surface of macrophages to promote the synthesis of TNFα and CCL2 (Schaefer et al, 2005; Moreth et al, 2010); both cytokines important for cancer (Balkwill, 2006; Lim et al, 2016). BGN and MXRA5 expression are promoted by the activity of TGFβ1 (Heegaard et al, 2004; Poveda et al, 2017), a key driver of the adverse stromal response in many cancers (Pickup et al, 2013) and COMP binds TGFβ1, enhancing its biological activity (Haudenschild et al, 2011).
In support of these findings, our GSEA analyses link the nine-gene signature to biological processes common to poor prognosis cancers, including EMT, angiogenesis, hypoxia, inflammation and a shift away from oxidative phosphorylation as a means of energy generation. Interestingly, we failed to demonstrate enrichment of gene signatures linked to other cancer-relevant processes including apoptotic regulation and genomic instability. The association with specific cancer-relevant gene signatures may implicate the nine-gene CM signature in their regulation. Nonetheless, it should be noted that the data presented here is only correlative and from its analysis we cannot provide a mechanistic link between the expression of ECM genes and specific biological processes. Moving forward, it will be important to confirm the prognostic relevance of the gene signature in prospective cancer cohorts and look to preclinical models to investigate potential mechanisms through which they might regulate cancer progression.
Change history
06 February 2018
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Acerbi I, Cassereau L, Dean I, Shi Q, Au A, Park C, Chen YY, Liphardt J, Hwang ES, Weaver VM (2015) Human breast cancer invasion and aggression correlates with ECM stiffening and immune cell infiltration. Integr Biol 7: 1120–1134.
Balkwill F (2006) TNF-alpha in promotion and progression of cancer. Cancer Metastasis Rev 25: 409–416.
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, Jacobsen A, Byrne CJ, Heuer ML, Larsson E, Antipin Y, Reva B, Goldberg AP, Sander C, Schultz N (2012) The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov 2: 401–404.
Chen JL-Y, Huang C-Y, Huang Y-S, Chen R-J, Wang C-W, Chen Y-H, JC-H Cheng, Cheng A-L, Kuo S-H (2014) Differential clinical characteristics, treatment response and prognosis of locally advanced adenocarcinoma/adenosquamous carcinoma and squamous cell carcinoma of cervix treated with definitive radiotherapy. Acta Obstet Gynecol Scand 93: 661–668.
Cheon D-J, Tong Y, Sim M-S, Dering J, Berel D, Cui X, Lester J, Beach JA, Tighiouart M, Walts AE, Karlan BY, Orsulic S (2014) A collagen-remodeling gene signature regulated by TGF-β signaling is associated with metastasis and poor survival in serous ovarian cancer. Clin Cancer Res 20: 711–723.
Contag SA, Gostout BS, Clayton AC, Dixon MH, McGovern RM, Calhoun ES (2004) Comparison of gene expression in squamous cell carcinoma and adenocarcinoma of the uterine cervix. Gynecol Oncol 95: 610–617.
Erler JT, Bennewith KL, Nicolau M, Dornhöfer N, Kong C, Le Q-T, Chi J-TA, Jeffrey SS, Giaccia AJ (2006) Lysyl oxidase is essential for hypoxia-induced metastasis. Nature 440: 1222–1226.
Fischer H, Salahshor S, Stenling R, Björk J, Lindmark G, Iselius L, Rubio C, Lindblom A (2001) COL11A1 in FAP polyps and in sporadic colorectal tumors. BMC Cancer 1: 17.
Galván JA, García-Martínez J, Vázquez-Villa F, García-Ocaña M, García-Pravia C, Menéndez-Rodríguez P, González-del Rey C, Barneo-Serra L, de los Toyos JR (2014) Validation of COL11A1/procollagen 11A1 expression in TGF-β1-activated immortalised human mesenchymal cells and in stromal cells of human colon adenocarcinoma. BMC Cancer 14: 867.
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, Sun Y, Jacobsen A, Sinha R, Larsson E, Cerami E, Sander C, Schultz N (2013) Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal 6: pl1.
Gao Y-B, Chen Z-L, Li J-G, Hu X-D, Shi X-J, Sun Z-M, Zhang F, Zhao Z-R, Li Z-T, Liu Z-Y, Zhao Y-D, Sun J, Zhou C-C, Yao R, Wang S-Y, Wang P, Sun N, Zhang B-H, Dong J-S, Yu Y, Luo M, Feng X-L, Shi S-S, Zhou F, Tan F-W, Qiu B, Li N, Shao K, Zhang L-J, Zhang L-J, Xue Q, Gao S-G, He J (2014) Genetic landscape of esophageal squamous cell carcinoma. Nat Genet 46: 3076.
Gilkes DM, Semenza GL, Wirtz D (2014) Hypoxia and the extracellular matrix: drivers of tumour metastasis. Nat Rev Cancer 14: 430–439.
Gröger CJ, Grubinger M, Waldhör T, Vierlinger K, Mikulits W (2012) Meta-analysis of gene expression signatures defining the epithelial to mesenchymal transition during cancer progression. PLoS One 7: e51136.
Gyorffy B, Lánczky A, Szállási Z (2012) Implementing an online tool for genome-wide validation of survival-associated biomarkers in ovarian-cancer using microarray data from 1287 patients. Endocr Relat Cancer 19: 197–208.
Győrffy B, Surowiak P, Budczies J, Lánczky A (2013) Online survival analysis software to assess the prognostic value of biomarkers using transcriptomic data in non-small-cell lung cancer. PLoS One 8: e82241.
Haudenschild DR, Hong E, Yik JHN, Chromy B, Mörgelin M, Snow KD, Acharya C, Takada Y, Di Cesare PE (2011) Enhanced activity of transforming growth factor β1 (TGF-β1) bound to cartilage oligomeric matrix protein. J Biol Chem 286: 43250–43258.
Heberle H, Meirelles GV, da Silva FR, Telles GP, Minghim R (2015) InteractiVenn: a web-based tool for the analysis of sets through Venn diagrams. BMC Bioinformatics 16: 169.
Heegaard A-M, Xie Z, Young MF, Nielsen KL (2004) Transforming growth factor beta stimulation of biglycan gene expression is potentially mediated by sp1 binding factors. J Cell Biochem 93: 463–475.
Hynes RO, Naba A (2012) Overview of the matrisome—an inventory of extracellular matrix constituents and functions. Cold Spring Harb Perspect Biol 4: a004903.
Jia D, Liu Z, Deng N, Tan TZ, RY-J Huang, Taylor-Harding B, Cheon D-J, Lawrenson K, Wiedemeyer WR, Walts AE, Karlan BY, Orsulic S (2016) A COL11A1-correlated pan-cancer gene signature of activated fibroblasts for the prioritization of therapeutic targets. Cancer Lett 382: 203–214.
Kale S, Raja R, Thorat D, Soundararajan G, Patil TV, Kundu GC (2014) Osteopontin signaling upregulates cyclooxygenase-2 expression in tumor-associated macrophages leading to enhanced angiogenesis and melanoma growth via α9β1 integrin. Oncogene 33: 2295–2306.
Katanyoo K, Sanguanrungsirikul S, Manusirivithaya S (2012) Comparison of treatment outcomes between squamous cell carcinoma and adenocarcinoma in locally advanced cervical cancer. Gynecol Oncol 125: 292–296.
Levental KR, Yu H, Kass L, Lakins JN, Egeblad M, Erler JT, Fong SFT, Csiszar K, Giaccia A, Weninger W, Yamauchi M, Gasser DL, Weaver VM (2009) Matrix crosslinking forces tumor progression by enhancing integrin signaling. Cell 139: 891–906.
Li A, Li J, Lin J, Zhuo W, Si J (2017) COL11A1 is overexpressed in gastric cancer tissues and regulates proliferation, migration and invasion of HGC-27 gastric cancer cells in vitro. Oncol Rep 37: 333–340.
Li Q, Birkbak NJ, Gyorffy B, Szallasi Z, Eklund AC (2011) Jetset: selecting the optimal microarray probe set to represent a gene. BMC Bioinformatics 12: 474.
Lim SY, Yuzhalin AE, Gordon-Weeks AN, Muschel RJ (2016) Targeting the CCL2-CCR2 signaling axis in cancer metastasis. Oncotarget 7: 28697–28710.
Lu P, Weaver VM, Werb Z (2012) The extracellular matrix: a dynamic niche in cancer progression. J Cell Biol 196: 395–406.
Masiero M, Simões FC, Han HD, Snell C, Peterkin T, Bridges E, Mangala LS, SY-Y Wu, Pradeep S, Li D, Han C, Dalton H, Lopez-Berestein G, Tuynman JB, Mortensen N, Li J-L, Patient R, Sood AK, Banham AH, Harris AL, Buffa FM (2013) A core human primary tumor angiogenesis signature identifies the endothelial orphan receptor ELTD1 as a key regulator of angiogenesis. Cancer Cell 24: 229–241.
Mi Z, Bhattacharya SD, Kim VM, Guo H, Talbot LJ, Kuo PC (2011) Osteopontin promotes CCL5-mesenchymal stromal cell-mediated breast cancer metastasis. Carcinogenesis 32: 477–487.
Moreth K, Brodbeck R, Babelova A, Gretz N, Spieker T, Zeng-Brouwers J, Pfeilschifter J, Young MF, Schaefer RM, Schaefer L (2010) The proteoglycan biglycan regulates expression of the B cell chemoattractant CXCL13 and aggravates murine lupus nephritis. J Clin Invest 120: 4251–4272.
Naba A, Clauser KR, Lamar JM, Carr SA, Hynes RO (2014) Extracellular matrix signatures of human mammary carcinoma identify novel metastasis promoters. eLife 3: e01308.
Pickup M, Novitskiy S, Moses HL (2013) The roles of TGFβ in the tumour microenvironment. Nat Rev Cancer 13: 788–799.
Pickup MW, Mouw JK, Weaver VM (2014) The extracellular matrix modulates the hallmarks of cancer. EMBO Rep 15: 1243–1253.
Poveda J, Sanz AB, Fernandez-Fernandez B, Carrasco S, Ruiz-Ortega M, Cannata-Ortiz P, Ortiz A, Sanchez-Niño MD (2017) MXRA5 is a TGF-β1-regulated human protein with anti-inflammatory and anti-fibrotic properties. J Cell Mol Med 21: 154–164.
Schaefer L, Babelova A, Kiss E, Hausser H-J, Baliova M, Krzyzankova M, Marsche G, Young MF, Mihalik D, Götte M, Malle E, Schaefer RM, Gröne H-J (2005) The matrix component biglycan is proinflammatory and signals through Toll-like receptors 4 and 2 in macrophages. J Clin Invest 115: 2223–2233.
Shevde LA, Samant RS (2014) Role of osteopontin in the pathophysiology of cancer. Matrix Biol 37: 131–141.
Sok JC, Lee JA, Dasari S, Joyce S, Contrucci SC, Egloff AM, Trevelline BK, Joshi R, Kumari N, Grandis JR, Thomas SM (2013) Collagen type XI α1 facilitates head and neck squamous cell cancer growth and invasion. Br J Cancer 109: 3049–3056.
Storey JD, Tibshirani R (2003) Statistical significance for genomewide studies. Proc Natl Acad Sci USA 100: 9440–9445.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES, Mesirov JP (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102: 15545–15550.
Szász AM, Lánczky A, Nagy Á, Förster S, Hark K, Green JE, Boussioutas A, Busuttil R, Szabó A, Győrffy B, Szász AM, Lánczky A, Nagy Á, Förster S, Hark K, Green JE, Boussioutas A, Busuttil R, Szabó A, Győrffy B (2016) Cross-validation of survival associated biomarkers in gastric cancer using transcriptomic data of 1,065 patients. Oncotarget 7: 49322–49333.
Wu Y-H, Chang T-H, Huang Y-F, Huang H-D, Chou C-Y (2014) COL11A1 promotes tumor progression and predicts poor clinical outcome in ovarian cancer. Oncogene 33: 3432–3440.
Yan W, Wistuba II, Emmert-Buck MR, Erickson HS (2010) Squamous cell carcinoma – similarities and differences among anatomical sites. Am J Cancer Res 1: 275–300.
Yuan P, Kadara H, Behrens C, Tang X, Woods D, Solis LM, Huang J, Spinola M, Dong W, Yin G, Fujimoto J, Kim E, Xie Y, Girard L, Moran C, Hong WK, Minna JD, Wistuba II (2010) Sex determining region Y-Box 2 (SOX2) is a potential cell-lineage gene highly expressed in the pathogenesis of squamous cell carcinomas of the lung. PLoS One 5: e9112.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License.
Supplementary Information accompanies this paper on British Journal of Cancer website
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Yuzhalin, A., Urbonas, T., Silva, M. et al. A core matrisome gene signature predicts cancer outcome. Br J Cancer 118, 435–440 (2018). https://doi.org/10.1038/bjc.2017.458
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2017.458
Keywords
This article is cited by
-
Longevity interventions modulate mechanotransduction and extracellular matrix homeostasis in C. elegans
Nature Communications (2024)
-
The extracellular matrix protein EMILIN-1 impacts on the microenvironment by hampering gastric cancer development and progression
Gastric Cancer (2024)
-
Transcriptome profiling of patient-derived tumor xenografts suggests novel extracellular matrix-related signatures for gastric cancer prognosis prediction
Journal of Translational Medicine (2023)
-
Extracellular matrix educates an immunoregulatory tumor macrophage phenotype found in ovarian cancer metastasis
Nature Communications (2023)
-
CAF-immune cell crosstalk and its impact in immunotherapy
Seminars in Immunopathology (2023)