Abstract
Background:
Meta-analysis shows that women with diabetes have a 20% increased risk of breast cancer and also an increased risk for distant metastasis and mortality. The molecular mechanisms for distant metastasis and mortality in breast cancer patients with diabetes are not very well understood.
Methods:
We compared the effect of physiological (5 mM) and diabetic (10 mM) levels of glucose on malignant breast epithelial cell invasion and stemness capabilities. We performed microRNA array to determine the dysregulated microRNAs in hyperglycaemic conditions and performed functional and molecular analysis of the gene targets.
Results:
Hyperglycaemia leads to hyperactivation of cancer stem cell pool and enhances invasive ability of breast cancer cells. MiR-424 seems to be a key regulator of cancer cell stemness and invasion. Knockdown of miR-424 in cancer cells under euglycaemic conditions leads to enhanced invasion and stem cell activity, whereas ectopic expression of miR-424 in cancer cells under hyperglycaemic conditions results in suppressed invasion and stem cell activity. Cdc42, a target of miR-424, influences cancer stem cell activity by positively regulating prdm14 through activation of pak1 (p-21-activated kinase 1) and stat5.
Conclusions:
Our findings establish miR-424→cdc42→prdm14 axis as a key molecular signalling cascade that might influence breast cancer progression in diabetic patients through hyperactivation of cancer stem cells.
Similar content being viewed by others
Main
Epidemiological findings suggest a strong association between diabetes mellitus (DM) and poor prognosis of breast cancer (Giovannucci et al, 2010). Emerging lines of evidence suggests that pre-existing diabetes increases the probability of distant metastasis in breast cancer patients and reduces the overall survival (Giovannucci et al, 2010). It is believed that altered blood glucose levels can influence the metabolic reprogramming of the cancer cells. In this regard, several factors like the glucose transport channels, and glycolytic and glutamine pathways have been suggested to have a major role (Li et al, 2016).
MicroRNAs are non-coding, single-stranded RNAs that are 20–23 nucleotides in length and are evolutionarily conserved. MicroRNAs are thought to regulate gene expression by reducing the stability of mRNA or by inhibiting translation of a particular target gene through binding of its seed sequence to complementary sequence located at the 3′-untranslated region (UTR) of the target gene (Esquela-Kerscher and Slack, 2006; Cho, 2007; Voorhoeve and Agami, 2007). We here demonstrate a tumour-suppressive role for miR-424, which is significantly downregulated in malignant breast epithelial cells under hyperglycaemic (HG) condition. MiR-424 is an intergenic microRNA belonging to the miR-424/503 cluster, which is an integral component of the cell cycle arrest program in mammary epithelium (Llobet-Navas et al, 2014a, 2014b). MiR-424 has tumour-suppressive properties in breast (Llobet-Navas et al, 2014; Zhang et al, 2015), endometrial (Li et al, 2015; Zhang et al, 2015), renal (Chen et al, 2013), ovarian clear cell (Wu et al, 2017), bladder carcinomas (Wu et al, 2015) and osteosarcoma (Long et al, 2013). Contrasting evidence suggests miR-424 to favour prostate cancer (Dallavalle et al, 2016) and the process of epithelial–mesenchymal transition in breast cancer (Drasin et al, 2015).
In this study, we show that glycaemic levels can negatively influence the expression of miR-424. We also identified complementarity between miR-424 and promoter sequence of cdc42 (cell division control 42). Cdc42 is a small 21 kDa Rho-GTPase that has been associated with enhanced invasive abilities of cancer cells and is often hyperactivated in breast tumours (Fritz et al, 1999, 2002; McHenry and Vargo-Gogola, 2010). We here demonstrate that in hyperglycaemia, impaired inhibitory regulation of miR-424 on cdc42 leads to the activation of prdm14 (PR-domain containing 14). Prdm14, a transcription regulator is a part of the embryonic core regulatory circuit involving Sox2, Nanog and Oct4 (Yamaji et al, 2013). Prdm14 is thought to repress differentiation and support the maintenance of pluripotency.
Our findings in the current study establish a role for reduced miR-424 under hyperglycaemic condition leading to (a) promotion of invasive abilities of breast cancer cells and (b) hyperactivation of the breast cancer stem cell (CSC) pool. It also suggests a molecular mechanism that can be explored further to improve the prognostic outcomes of breast cancer patients with diabetes.
Materials and methods
Cell lines
MDA-MB-231 (ATCC, Manassas, VA, USA; HTB-26), triple-negative metastatic breast cancer cell line, and MCF-10A (ATCC CRL 10317), non-malignant breast epithelial cell line, were purchased from ATCC and maintained in media containing normal physiological levels (NMLs) of glucose (5 mM). The cells were also cultured in 10 mM glucose containing media to mimic hyperglycaemia (HG).
Transfections and stable cell lines
Stable cell lines overexpressing miR-424 (miR-424-MDA231 and miR-424-MCF10A) and miR-424 knockdown (anti-miR-424-MDA231 and anti-miR-424-MCF10A) were generated from parental MDA-MB-231 and MCF-10A cell lines. Both MDA-MB-231 and MCF-10A cell lines were transfected with miR-424-overexpressing vector (pEZX-MR04; cat. no. HmiR-R0396-MR04; Genecopoeia, Rockville, MD, USA) and inhibitor containing vector (pEZX-AM01; cat. no. HmiR-AN0494-AM01; Genecopoeia). Stable clones with miR-424 overexpression and knockdown were drug selected using puromycin (5 μg ml−1; cat. no. A11138-03; Invitrogen, Carlsbad, CA, USA). MDA-MB-231 were also transfected with short interfering RNA (siRNA) for cdc42 (cat. no. SR300714; Origene, Rockville, MD, USA) and plasmid vector harbouring open reading frame for cdc42 (cat. no. RC214076; Origene). For prdm14 promoter reporter assay, pEZX-PG04 plasmid vector containing PRDM14 promoter with Gaussia luciferase (cat. no. HPRM19109-PG04; Genecopoeia) as the reporter gene was used. All plasmid DNA and siRNA transfections were performed using Lipofectamine 2000 transfection reagent (cat. no.11668019; Invitrogen) and Lipofectamine RNAiMAX transfection reagent (cat. no. 13778150), respectively, as per the manufacturer’s instruction.
In situ hybridisation
miR-424 was detected in live cells using SmartFlare RNA Detection Probes (EMD Millipore, Billerica, MA, USA). Briefly, cells in both NML and HG conditions were incubated with miR-424-specific Cy-5-labelled RNA Detection Probe (cat. no. SF-408; EMD Millipore) overnight. The cells were imaged the following day using an inverted fluorescent microscope (Floid Imaging Station; Life Technologies, Carlsbad, CA, USA).
3′-UTR luciferase reporter assay
miR-424-MDA-231 cells were transfected with plasmid vector harbouring the wild-type 3′-UTR (cat. no. HmiT023455-MT06; Genecopoeia) or mutant 3′-UTR of CDC42 (cat. no. CS-HmiT023455-MT06-01; Genecopoeia) using Lipofectamine 2000 transfection reagent (cat. no.11668019; Invitrogen). Luminescence was analysed after 48 h using Dual Luciferase Detection Kit (cat. no. LPFR-P030; Genecopoeia).
Immunoblotting
Proteins from whole-cell lysates for western blotting were extracted with mammalian protein extraction reagent (cat. no. 78501; Thermo Scientific, Rockford, IL, USA), resolved on SDS–PAGE and transferred to polyvinylidene fluoride (PVDF; cat. no. IPVH00010; EMD Millipore) membrane. The membranes were then blocked with 5% bovine serum albumin (BSA; cat. no. A7906; Sigma-Aldrich, St Louis, MO, USA) in Tris-buffered saline with 0.1% Tween-20 (TBST) for 1 h at room temperature. The membranes were then incubated with primary antibody for 1 h at room temperature, followed by rinsing of unbound antibody with TBST and incubating the membranes with appropriate horseradish peroxidase-conjugated secondary antibodies. Primary antibodies for Cdc42 (cat. no. ab64533; Abcam, Cambridge, UK), E-cadherin (cat. no. 3195S, Cell Signaling, Danvers, MA, USA), vimentin (cat. no. 5741S; Cell Signaling), p-PAK1 (p-21-activated kinase 1) (cat. no. ab40852; Abcam), LIMK1 (cat. no. ab95186; Abcam), prdm14 (cat. no. ab91587; Abcam), caspase-3 (cat. no. C8487; Sigma-Aldrich) and β-actin (cat. no. A1978; Sigma-Aldrich) were used. Protein expression was detected using SuperSignal West Femto Maximum Sensitivity Substrate (cat. no. 34095; Thermo Scientific) and the membranes were imaged for chemiluminescence using the ImageQuant LAS 4000 Imager (GE Healthcare Life Sciences, Bio-Sciences, Pittsburgh, PA, USA).
Active cdc42 pull-down assay
Guanosine-5′-triphosphate (GTP)-bound cdc42 was detected using Pull-Down and Detection Kit (cat. no. 16119; Thermo Scientific). Proteins from whole-cell lysates for immunoprecipitation were used. Active cdc42 was pulled down using 20 μg of GST-Pak1-PBD (Pak1 protein-binding domain). The pulled down lysate along with total lysate (input) was then resolved on a SDS–PAGE and transferred to a PVDF membrane. The membrane was processed as per regular immunoblotting protocol and probed with anti-cdc42 antibody provided with the kit. GTPγS or GDP were used as positive and negative controls in the assay.
MicroRNA reverse transcription–PCR array
To analyse the altered microRNAs (associated with breast cancer) in response to hyperglycaemia, a miScript miRNA PCR array was performed (cat. no. MIHS-109Z; Qiagen, Germantown, MD, USA). Good quality RNA (1 μg) was used to generate cDNA using RT2 miRNA First Strand Kit (Qiagen). The cDNA generated was used to perform the PCR. Data were normalised with the six snoRNA/snRNA miScript PCR controls provided in the array. Data analysis was performed using the 2−ΔΔCT relative quantification method.
Real-time quantitative reverse transcription–PCR
Total RNA was extracted using the Trizol reagent (cat. no. 15596-018; Life Technologies) from both cell lines and xenograft tumours. To analyse the expression of miR-424, cDNA was prepared from RNA (500 ng per reaction) using miScript II RT Kit (cat. no. 218161; Qiagen). The quantitative reverse transcription–PCR (qRT–PCR) was performed using miScript SYBR Green PCR Kit (cat. no. 204141; Qiagen) on a StepOnePlus Real-Time PCR System (Applied Biosystems, Carlsbad, CA, USA). Complementary DNA (2 μl per reaction) was used with miR-424-specific primers (SABiosciences, Qiagen). RNU6-2 and SNORD91 were used as internal controls. The data were analysed using the 2−ΔΔCT relative quantification method.
Sphere-forming assay
For sphere-forming assays, the cells were grown in serum-free DMEM media supplemented with B-27 (cat. no. 12587010; Gibco Life Technologies, Grand Island, NY, USA) and MEGM SingleQuots (cat. no. CC-4133; Lonza, Basel, Switzerland). The assay was established using 5000 cells per well on ultra-low attachment 6-well plates (Corning, Corning, NY, USA; cat. no. 3471). Cultures were terminated after 5–7 days and analysed under a phase contrast light microscope for sphere formation (Nikon Eclipse TS100; Nikon, Tokyo, Japan).
Flow cytometric analysis
Flow cytometric analysis was performed to analyse the percentage of CD44+/CD24−/low cells. Post trypsinisation, the cells were probed with anti-CD44-FITC (cat. no. 555478; BD Biosciences, San Jose, CA, USA) and CD24-PE (cat. no. 555428; BD Biosciences) after fixing and blocking with 5% formaldehyde and 0.1% BSA, respectively. After incubating for 30 min at room temperature, the cells were rinsed with 1 × PBS once before acquisition and analysed using flow cytometer (cat. no. 653118; Accuri C6; BD Biosciences). Unstained and single stained cells for CD44 or CD24 were used as compensation controls.
Immunohistochemistry
Formalin-fixed, paraffin-embedded xenograft tumour sections were used for immunohistochemical (IHC). Briefly, the paraffin sections were deparaffinised and the antigens were unmasked using Trilogy (cat. no. 920P-10; Cell Marque, Rocklin, CA, USA). The sections were then blocked for endogenous peroxidase and non-specific antibody binding before being probed with primary antibodies against Ki67, active caspase-3, vimentin, E-cadherin, Cdc42 and p-pak1. The signals from primary antibody were detected using mouse/rabbit polyscan HRP/DAB Detection System (cat. no.956D-40; Cell Marque). The sections were counterstained with haematoxylin, followed by treatment with bluing agent. The sections were imaged using upright microscope (Eclipse 50i; Nikon). The images were analysed and documented using NIS Element Imaging Software (Nikon).
Immunofluorescence
Cells seeded in an 8-well chamber slide were fixed and blocked using 5% formaldehyde and 5% BSA, respectively. They were then stained with antibodies against stat5 (cat. no. sc-836; Santa Cruz, Dallas, TX, USA) and pak1 (cat. no.: ab40852; Abcam). Alexa Fluor 488- and 594-tagged secondary antibodies raised in species appropriate for the primary antibody were used followed by counterstaining with DAPI. All slides were examined using a confocal microscope (Nikon Eclipse Ti; Nikon).
Animals and xenograft assays
All the animal studies conducted were approved by TTUHSC-Institutional Animal Care and Use Committee. Animals used in the study were cared as per the Institutional guidelines. Six- to seven-week-old female athymic nude mice were obtained from Envigo (Indianapolis, IN, USA). All animals used were at the age of 10–12 weeks at the time of transplantation. Experimental group animals (n=6) were subcutaneously transplanted with MDA-MB-231 cells stably over expressing miR-424 (0.5 × 106 cells per flank). Control group (n=6) animals were transplanted with MDA-MB-231 containing the empty vector (EV). The animals were followed till 10 weeks post transplantation. Tumour volumes were calculated according to the formulae 4/3π × r12 × r2. After killing the animal, the tumours were surgically excised, weighed, formalin fixed for IHC and snap frozen for RNA and protein studies.
Metastasis assay
Female athymic nude mice 10–12 weeks of age were used for the assay. Anti-miR-424-MDA231 (n=3) and EV (n=3) containing cells were injected non-surgically through the tail vein route. The animals were weekly monitored for body weight until 8 weeks before terminating them. Necropsy was performed to obtain lung, live, brain, heart and mammary gland for the control and experimental animals.
Invasion assay
Invasive ability of the cells was assessed by performing transwell Matrigel-based invasion assay. Cells in 0.2% FBS were seeded at a density of 8 × 104/well in the upper chamber of 6.5 mm transwell with 8.0 μm pore polycarbonate membrane inserts (Corning) coated with the Matrigel (cat. no. 354234; BD Biosciences). Media containing 10% FBS was added to the lower chamber. After 48 h of incubation in 37 °C/5% CO2, cultures were terminated; the Matrigel from the upper chamber was gently scrapped using Q-tips. The inserts containing invaded cells on the side of the membrane facing the lower chamber was rinsed with 1 × PBS and fixed with 4% paraformaldehyde. They were then stained with 0.1% crystal violet stain for 24 h at 4 °C before washing the inserts well with tap water and imaging the inserts under inverted phase contrast microscope (Eclipse Ti; Nikon).
CSC transcription factor array
CSC Transcription Factor (TF) Profiling Plate Array (cat. no. FA-1004; Signosis Inc., Santa Clara, CA, USA) was used to find TFs affected by miR-424. Briefly, biotin-labelled probes specific for TF DNA binding sites were incubated with cell nuclear extract. The TF–probe complex was separated from the unbound probes using a spin column. Finally, the bound probes were detached from the complex and were allowed to hybridise in the wells precoated with complementary sequence to the probes. Detection of the bound probe was read in a luminometer (Clariostar, BMG Labtech, Cary, NC, USA).
In silico analysis
Predictive miR-424 binding site on CDC42 3′-UTR were identified by using the TargetScan online portal (http://www.targetscan.org/;version6.2). The predictions were also validated using other online platforms like miRbase and microRNA.org.
Statistical analysis
Data are represented as mean±s.d. and analysed by paired Student’s t-test. P<0.05 was considered as statistically significant.
Results
Hyperglycaemia promotes invasion and stemness in breast cancer cells
To investigate the effect of hyperglycaemia on triple-negative breast cancer (TNBC) cells; invasion and sphere-forming assays were performed. Our findings suggest that hyperglycaemia enhances the invasive behaviour of highly metastatic TNBC cells compared with euglycaemic controls (Figure 1A and B). Non-malignant breast epithelial cell line, MCF-10A, under hyperglycaemia also seems to have a similar effect on invasion (Supplementary Figure S1A). Further, it was also noted that hyperglycaemia promoted expansion of stem cell population in TNBC cells. Flow cytometric analysis of CD44+/hi/CD24−/low cells revealed that hyperglycaemia promoted CSCs in TNBC cells (Figure 1C and D). These data were further supported by sphere formation assay (Figure 1E). An increase by two-fold in the sphere-forming ability in hyperglycaemic conditions indicated an elevated CSC activity (Figure 1E and F). Self-renewing ability of TNBC stem cells was also increased significantly under hyperglycaemic condition compared with cells under euglycaemic conditions (Figure 1G). This set of data suggested that hyperglycaemic environment could potentially help cancer cells to attain more aggressive traits by promoting cancer stemness and invasiveness. Similar observations for stem cell activity were made in non-malignant epithelial cells as well (Supplementary Figure S1B). However, no changes were observed in self-renewal ability of non-malignant breast epithelial cells under hyperglycaemic compared with euglycaemic conditions (Supplementary Figure S1C).
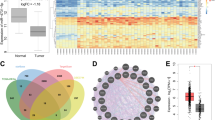
Impact of hyperglycaemia on invasion, migration, stem cell behaviour and on expression of miR-424 in TNBC cells. (A and B) Matrigel-based assay to analyse invasive abilities of TNBC cells in hyperglycaemic (10 mM glucose, HG) compared with euglycaemic (5 mM glucose, NML) culture conditions (n=3). (C and D) Flow cytometry dot plots and cumulative bar graph representation depicting the increase in the percentage of CSCs in hyperglycaemic compared with euglycaemic conditions. (E) Micrograph of mammospheres at NML and HG conditions (n=3). (F) Bar graphs depicting the sphere-forming ability of CSCs in serum-free conditions under hyperglycaemia in contrast to euglycaemic. (G) Self-renewing ability of CSCs in first (G1) and second (G2) generations (n=3). (H) Quantitative reverse transcription–PCR analysis of miR-424 expression levels with RNU-6 as an internal control in TNBC cells in hyperglycaemic conditions (n=3). (I) In situ hybridisation for miR-424 using Cy-5-tagged probes against miR-424 in live breast cancer cells under hyperglycaemic and euglycaemic conditions (n=3). Data are represented as mean ± s.d. *P⩽0.05.
Hyperglycaemia alters microRNA profile in TNBC cells
To analyse the impact of hyperglycaemia on global microRNA expression in TNBC cells, RT–PCR-based microRNA array was performed. MicroRNA profiles of MDA-MB-231 cells under hyperglycaemia were analysed and compared with cells under euglycaemic condition. MiR-424 was found to be one of the most significantly altered (downregulated) microRNA in response to hyperglycaemia in TNBC cells (Supplementary Figure S2). Reverse transcription–PCR and in situ hybridisation analysis also confirmed downregulation of miR-424 in TNBC cells under hyperglycaemic condition (Figure 1H and I).
Hyperglycaemia mediated reduced miR-424 expression leads to promotion of invasion and CSC activity
We now wanted to investigate if enhanced invasion and CSC activity of malignant and non-malignant breast epithelial cells in hyperglycaemia was mediated by miR-424. For this, we established miR-424 stably overexpressing (miR-424-MDA231 and miR-424-MCF10A) and knockdown (anti-miR-424-MDA231 and anti-miR-424-MCF10A) cell lines from parental MDA-MB-231 and MCF-10A cells (Supplementary Figure S3A–D). MiR-424 overexpressing and knockdown cell lines were maintained in hyperglycaemic and euglycaemic culture conditions, respectively. MiR-424-MDA231 cells had marked reduction in their invasive abilities compared with EV controls despite being maintained in hyperglycaemic conditions (Figure 2A and B). In addition, anti-miR-424-MDA231 cells had enhanced invasive abilities compared with EV control in euglycaemic conditions (Figure 2A and B). Similar trends in invasive abilities were observed in non-malignant cells with miR-424 modulation (Supplementary Figure S4A). This set of data points towards the crucial involvement of miR-424 in the regulation of invasive abilities of breast epithelial cells, both malignant and non-malignant in response to glycaemic levels. Findings from sphere-forming assay in miR-424-MDA231 revealed a two-fold reduction in sphere-forming ability compared with EV controls under hyperglycaemic condition (Figure 2C and D). On the contrary, anti-miR-424-MDA231 cells showed an increase in sphere-forming abilities compared with EV control under euglycaemic conditions (Figure 2C and D). Further, in anti-miR-424-MCF10A cells, a statistically significant increase in sphere-forming abilities was observed (three-folds) (Supplementary Figure S4B and C), whereas no such change was observed in miR-424-MCF10A cells. These findings suggested that reduced miR-424 promotes invasion and stem cell activity of both malignant and non-malignant breast epithelial cells.
Role of miR-424 in hyperglycaemia induced effects of enhanced invasion and hyperactivation of CSC pool in TNBC cells. (A and B) Matrigel-based invasion assay showing reintroduction of miR-424 in breast cancer cells under hyperglycaemic conditions leads to significant reduction in cellular invasion, whereas knockdown of miR-424 in breast cancer cells under euglycaemic conditions results in enhanced cellular invasiveness (n=3). Bar diagram represents cumulative data for a number of invaded cells per field for all groups (C and D) Sphere-forming ability of breast cancer cells in serum-free conditions (n=3). Reintroduction of miR-424 in cancer cells under hyperglycaemic condition resulted in reduced number as well as size of spheres. On the contrary, knockdown of miR-424 in cancer cells under euglycaemic conditions led to increased number of spheres formed. (E) In silico analysis demonstrating complementarity between miR-424 and cdc42 at two different sites on 3′-UTR. (F) Bar diagram depicting the luciferase reporter activity in cdc42 wt 3′-UTR with miR-424 overexpression compared with cdc42 mutant 3′-UTR with miR-424 overexpression (n=3). (G) Immunoblotting for cdc42 expression in miR-424-MDA231 and anti-miR-424-MDA231 cells compared with their respective empty vector control (n=3). (H) Pull-down assay using PAK1-binding domain to analyse levels of active GTP-bound cdc42 (n=3). Epidermal growth factor of 50 ng/ml has been used to as an activator of cdc42. Data are represented as mean±s.d. *P⩽0.05.
Cdc42 is a direct target of miR-424
It was important to investigate the gene targets of miR-424 to understand its role in hyperglycaemia. For this, we performed in silico analysis using online platforms like TargetScan, miRBase and microRNA.org. Based on the in silico analysis, we found cdc42 to be a predicted target of miR-424 (Figure 2E). MicroRNAs are believed to regulate gene expression by recognising specific target sites in the 3′-UTRs of target genes. To test the direct interaction between miR-424 and 3′-UTR of cdc42, we performed luciferase-reporter assay. 3′-UTR region of cdc42 including the putative binding site of miR-424 was introduced downstream of the open reading frame of luciferase gene in the reporter plasmid pEZX-PG04 (Genecopoeia). Simultaneously, luciferase reporter constructs containing specific mutation in the putative binding site of miR-424 were used to confirm the interaction between miR-424 and 3′-UTR of cdc42 in MDA231 cells. A significant reduction in luciferase activity was observed in the miR-424-overexpressing MDA231 cells with cdc42 wild-type (wt) 3′-UTR, while no difference was noted with the mutant (mut) 3′-UTR (Figure 2F).
Further, immunoblotting demonstrated a reduced level of cdc42 in miR-424-MDA231 cells under hyperglycaemia (Figure 2G). In contrast, anti-miR-424-MDA231 had a higher level of cdc42 compared with EV controls under euglycaemic condition (Figure 2G). While these results suggested that an increased expression of cdc42 was mediated by reduced miR-424 in cancer cells under hyperglycaemia, it was also important to check the levels of activated cdc42 (GTP-bound). We looked for the levels of active cdc42 through pull-down assays using GST-pak1 protein binding domain. Pak1 is a major downstream effector of cdc42 and its binding domain has high affinity for active cdc42 and negligible affinity for inactive form of cdc42 (Manser et al, 1997). Further, epidermal growth factor (EGF) was used as an exogenous stimulator of cdc42 activity in this experiment. MiR-424-MDA231 under hyperglycaemia had lower levels of active cdc42 than EV controls (Figure 2H). However, EGF stimulation in miR-424-MDA231 cells was able to bring the levels of active cdc42 comparable to EGF-stimulated EV control cells. It was clear that suppression of miR-424 in hyperglycaemic condition leads to increased cdc42 activity. In other words, high expression of miR-424 in euglycaemic condition keeps the level of active cdc42 under control.
MiR-424 influence xenograft tumourigenesis and metastasis
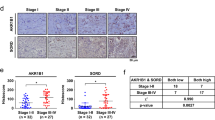
MiR-424 seems to be a key player in invasion and stemness, which are important contributing factors towards tumourigenesis and metastasis. To investigate miR-424-mediated effects on xenograft tumour formation, 0.5 × 106 miR-424-MDA231 or EV cells were injected in both flanks of nude mice (n=6 per group). These animals were followed for 10 weeks post transplantation, and alterations in body weight (Figure 3A) and tumour volume were assessed (Figure 3B and C). It was noted that miR-424-MDA231 group had a significantly reduced tumour growth compared with the control group. All animals in the control group had tumours by week 6, whereas only one animal from the experimental group barely showed signs of tumour formation at week 10. Reverse transcription–PCR of the miR-424-MDA231-derived tumours revealed a stable overexpression of miR-424 (Figure 3D). Haematoxylin and eosin staining indicated an intense cellular phenotype in control compared with miR-424-overexpressing tumours (Figure 3E). Immunohistochemical staining and western blot analysis confirmed the stable suppression of cdc42 and p-pak1 in miR-424-MDA231-derived tumours compared with empty vector control group (Figure 3F and G). MiR-424-MDA231 tumours also demonstrated reduced number of Ki67-positive cells, indicating a lower proliferation index (Figure 3F). On the contrary, higher expression of active caspase-3, an apoptotic marker, was observed in miR-424-overexpressing tumours (Figure 3F and G). This suggested an increased apoptosis in miR-424-overexpressing tumours. Mesenchymal marker vimentin was markedly reduced in miR-424-overexpressing tumours (Figure 3F). In addition, expression of epithelial marker E-cadherin was elevated in miR-424-overexpressing tumours (Figure 3G). This confirmed the role of miR-424 as a suppressor of tumour growth in extremely aggressive TNBC cell lines.
Overexpression of miR-424 in TNBC cells markedly reduced tumour growth. MDA231 cell lines stably overexpressing miR-424 and empty vector (0.5 × 106 cells) were subcutaneously injected in the flanks of experimental and control animals, respectively (n=6 per group). (A) Body weight of control and experimental animals. (B) Tumour volume as measured every 2 weeks for a period of 10 weeks in both control and experimental animals. (C) Representative pictures of mice and xenograft tumours. (D) Reverse transcription–PCR analysis of the miR-424-MDA231-derived xenograft tumours reveals increased levels of miR-424 were maintained throughout the in vivo experimental period. (E) Haematoxylin and eosin (H&E) for miR-424-MDA231-derived xenograft tumour sections showing reduced cell mass in miR-424-MDA231-derived xenograft tumour tissues. (F) Immunohistochemical staining for cdc42 and p-pak1 in xenograft tumour tissues derived from miR-424-MDA231 cells illustrates significantly reduced expression of cdc42 and p-pak1. Immunohistochemical staining for apoptosis marker active caspase-3 demonstrates elevated expression in experimental group, whereas staining for proliferation marker, Ki67, shows reduced expression in miR-424-MDA231-derived xenograft tumours. Expression of vimentin, a mesenchymal marker, is significantly reduced in the experimental group. (G) Immunoblotting analysis for cdc42, active caspase-3 and E-cadherin in tumour sections. Data are represented as mean±s.d. *P⩽0.05.
To investigate the role of miR-424 in metastasis, we used cell lines with stable knockdown of miR-424 (anti-miR-424-MDA231) and EV control cells. A million anti-miR-424-MDA231 cells or EV control cells were non-surgically injected into nude mice (n=3 per group) through the tail vein. Body weight of animals was monitored every week till 4 weeks postinjection (Figure 4A). Metastatic activity was assayed by examination of the organs at necropsy. Histological analysis of lungs isolated at necropsy suggested that animals with miR-424 knockdown had severe metastasis in the lung (Figure 4C) compared with EV control cells (Figure 4B). These findings suggested an antimetastatic role for miR-424.
Reduced expression of miR-424 in TNBC cells strengthens its metastatic ability. Tail vein injection of MDA231 cells with stable knockdown of miR-424 (anti-miR-MDA231) led to generation of more lung metastatic colonisation in the experimental group compared with the control group (injected with empty vector containing cells) (n=3). (A) Body weight for both control and experimental group of animals. (B) Haematoxylin and eosin (H&E)-stained lung section from the control group of animals showing signs of metastatic colony formation. (C) Haematoxylin and eosin-stained lung section from experimental group of animal demonstrating the formation of huge metastatic colonies.
MiR-424→cdc42 signalling axis is a key regulator of invasiveness in hyperglycaemia
Although the role of cdc42 in promotion of cancer cell invasion is well established, it was imperative to understand if hyperglycaemia promotes invasion of cancer cells through miR-424→cdc42 axis. In an attempt to do so, cdc42 was overexpressed in miR-424-MDA231 cells. Cdc42 overexpression was able to rescue the inhibitory effect of miR-424 on cdc42 (Figure 5A). Matrigel-based invasion assay was performed in cdc42-overexpressing miR-424-MDA231 cells (Figure 5B and C). It was observed that cdc42 overexpression in miR-424-MDA231 cells was able to increase the invasive ability of the cells and was comparable to the invasive ability of cdc42-overexpressed EV group. Thus, it was clear that ectopic expression of cdc42 could abrogate the effects of miR-424 overexpression and can elevate the invasion rates in otherwise less invasive miR-424-MDA231 cells. Further, expression levels of p-pak1, cdc42 and limk1 were significantly increased with knockdown of miR-424 (Figure 5D). These data suggested that miR-424 not only suppressed cdc42 activity but also its downstream signalling molecules. To further investigate if hyperglycaemia-mediated hyperactivation of CSCs is through miR-424→cdc42 axis, we silenced cdc42 in anti-miR-424-MDA231 cells and quantified the percentage of CSCs. Silencing of cdc42 was able to oppose the hyperactivation of CSCs as a result of miR-424 knockdown (Figure 5E and F). These findings suggested that hyperglycaemia promotes invasion and CSC activity through miR-424→cdc42 signalling axis.
miR-424–cdc42 is a key signalling axis in hyperglycaemia-induced enhanced invasive abilities of breast cancer cells. Ectopic expression of cdc42 was carried out by transient transfection in EV and in miR-424-MDA231 cells under hyperglycaemic condition. (A) Immunoblotting for cdc42 in control, cdc42, miR-424 and cdc42+miR-424 groups (n=3). (B and C) Matrigel-based invasion assay indicated increased invasion with cdc42 overexpression (n=3). More importantly, cdc42 overexpression in miR-424-MDA231 was able to rescue invasiveness in these cells to a large extent. (D) Immunoblotting for cdc42, p-pak1 and limk1 in anti-miR-424-MDA231 indicating an increased expression of cdc42 downstream proteins (n=3). (E and F) Flow cytometric analysis of CSC (CD44+/hi/CD24−/low) in EV, anti-miR-424 and anti-miR-424+sicdc42 groups (n=3). Data indicate that silencing of cdc42 nullifies the effect of miR-424 knockdown on CSC expansion. *Statistical significance between EV and anti-miR-424 groups. #Statistical significance between anti-miR-424 and anti-miR-424+sicdc42 groups. Data are represented as mean±s.d. */#P⩽0.05.
Hyperglycaemia enhances CSC activity by stimulating prdm14 through cdc42
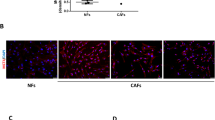
It was imperative to understand the mechanism behind the regulatory influence of miR-424 on breast CSCs. First, we looked into the CSC-associated TFs that are influenced by miR-424 under hyperglycaemia. To test this, we performed a CSC-associated TF activation assay using nuclear extracts from miR-424-MDA231 and compared it to EV control under high glucose culture condition (Supplementary Figure S5). Prdm14 and Nanog were the most significantly suppressed TFs in miR-424-overexpressing cells compared with EV controls under hyperglycaemic conditions (Supplementary Figure S5). Increased nuclear accumulation of prdm14 levels in anti-miR-424-MDA231 cells was noted, which strongly suggested an inhibitory role of miR-424 in prdm14 activation (Figure 6A). To further investigate, if prdm14 activity is mediated by miR-424→cdc42 axis, we overexpressed cdc42 in miR-424-MDA231 cells and looked for the transcript levels of prdm14. Reintroduction of cdc42 in miR-424-MDA231 cells led to a marked increase in the transcript level of prdm14 (Figure 6B). Further, prdm14 promoter reporter assay indicated a definitive role for cdc42 in prdm14 regulation. Knockdown of miR-424 in MDA-MB-231 cells elevated the prdm14 promoter activity by 30% and silencing of cdc42 in anti-miR-424-MDA231 reduced it by almost 50%, suggesting suppression of prdm14 promoter activity in the absence of cdc42 (Figure 6C). These findings strongly suggested miR-424→cdc42 axis to be a key regulator of prdm14. However, the mechanism for regulation of prdm14 by cdc42 was not clear. We noted that prdm14 promoter has a consensus sequence for stat5 binding (binding position: Chr 8: 70 980 676–70 980 685). This led us to hypothesise that cdc42 through phosphorylation of pak1 promotes stat5 activation, which further binds to prdm14 promoter and results in gene expression. To test the involvement of stat5 in cdc42–prdm14 signalling axis, we inhibited stat5 using Stat5 inhibitor (CAS 285986-31-4) in anti-miR-424-MDA231 cells and looked for the transcript levels of prdm14. Anti-miR-424 elevated the prdm14 transcript levels by over eight-folds, whereas with stat5 inhibition prdm14 transcript levels were reduced by three-folds (Figure 6D). This indicates stat5 to be the intermediate player between cdc42 and prdm14. Further, increased nuclear localisation of stat5 was noted in anti-miR-424-MDA231 cells (Figure 6E). Interestingly, p-pak1 (S144) downstream of cdc42 seems to colocalise with stat5 and facilitates its nuclear accumulation. These findings strongly indicate that cdc42 activates stat5, most probably through phosphorylation of pak1, which then leads to prdm14 activation. Based on these findings, we propose a mechanism involving miR-424–cdc42–prdm14 signalling axis, which can significantly contribute towards breast cancer metastasis under hyperglycaemic conditions through enhanced invasion and hyperactivation of CSC pool (Figure 7).
Hyperglycaemia-mediated promotion of TNBC stem cell pool is due to activation of prdm14. Anti-miR-424-MDA231 and miR-424-MDA231 cells were used to investigate the mechanistic link between miR-424 and hyperactivation of TNBC stem cell pool. (A) Immunoblotting for prdm14 in the cytoplasmic and nuclear fractions of anti-miR-424 MDA231 cells (n=3). Data demonstrate increase in nuclear and decrease in cytoplasmic prdm14 with miR-424 knockdown. (B) Reverse transcription–PCR (RT–PCR) analysis for cdc42 and prdm14 in miR-424-MDA231 and miR-424-MDA231+cdc42 groups. The data indicates an enormous increase in the transcript level of prdm14 with the introduction of cdc42 in miR-424-MDA231 cell system (n=3). * and # represents statistical significance for the expression of prdm14 and cdc42 respectively between miR-424-MDA231 and miR-MDA231+cdc42 groups. (C) Prdm14 promoter-reporter (luciferase) assay, indicating activation of prdm14 with miR-424 knock-down and reduced promoter activity with sicdc42 (n=3). * represents statistical significance between control (only prdm14 promoter) and sicdc42 group. # represents statistical significance between sicdc42 and Anti-miR-424 groups. (D) Reverse transcription–PCR analysis in anti-miR-424-MDA231 indicated an increased transcript level for prdm14 compared with EV group (n=3). * represents statistical significance between EV and Anti-miR-424 groups and also between EV and Anti-miR-424+Stat5 Inh groups. # represents statistical significance between Anti-miR-424 and Anti-miR-424+Stat5 Inh groups. More importantly, inhibition of Stat5 in anti-miR-424-MDA231 led to marked decrease in the prdm14 transcript levels, indicating a crucial role of stat5 in signal transduction between miR-424 and prdm14 activation. (E) Immunofluorescence for stat5 and pak1 in anti-miR-424 and EV control cells indicates high levels of nuclear stat5 with miR-424 knockdown (n=3). Further, the data also suggest enhanced nuclear colocalisation between stat5 and pak1 with miR-424 knockdown. Data are represented as mean±s.d. */#P⩽0.05.
Discussion
Diabetes mellitus not only worsens the prognosis (Kaplan et al, 2012; Villarreal-Garza et al, 2012; Yerrabothala et al, 2014) but also the quality of life of breast cancer patients (Tang et al, 2016). Diabetes being a metabolic disorder can affect cancer progression and recurrence through factors such as hyperglycaemia (Villarreal-Garza et al, 2012; Flores-Lopez et al, 2016), hyperinsulinaemia (Flores-Lopez et al, 2016) and hyperlipidaemia (Owiredu et al, 2009; Emaus et al, 2010; Kaye et al, 2002). Antidiabetic drugs such as metformin has shown improvement in cancer-specific survival rates in breast cancer patients with DM (Xu et al, 2015). Earlier we have shown that the effect of hyperglycaemia on proliferation of breast cancer cells is mainly due to activation of JAK-STAT signalling (Lopez et al, 2013). The findings of our current study indicates that hyperglycaemia in addition to proliferation can also significantly affect invasion and CSC behaviour in TNBC through miR-424–cdc42–stat5–prdm14 signalling axis. Invasive ability of non-malignant breast epithelial cells was also significantly influenced by hyperglycaemia.
Several lines of evidence have suggested the role of high glucose levels in enhancing the invasive and migratory capacity of breast cancer (Takatani-Nakase et al, 2014; Flores-Lopez et al, 2016), pancreatic cancer (Han et al, 2013) and lung cancer cells (Kang et al, 2015). Recent literature also suggests crucial roles for microRNAs in various disease pathologies. Through the current study we have directed our efforts to understand the role of miR-424, whose expression is significantly reduced under hyperglycaemic conditions in TNBC cells. Although the role of miR-424 in different cancers is debatable, it is important to note that microRNAs in general act in a context-dependent manner. Further, the fact that miR-424 is regulated by the biphasic signalling cytokine, TGF-β makes it even more complicated to define its role in different cancers (Llobet-Navas et al, 2014). Role of miR-424 in cancer cells under hyperglycaemic conditions has so far not been reported. Our findings present a novel role for reduced miR-424 levels in enhancing the aggressive behaviour of breast cancer by promoting invasiveness and stem cell activity. Further, enhancement in cdc42 activity by reduced miR-424 under hyperglycaemia seems like a crucial signalling axis to promote invasion of cancer cells. Rho-GTPase cdc42 has been previously implicated in invasive behaviour of cancer cells. It is interesting to note that breast cancer cells under euglycaemic conditions with higher levels of miR-424 have reduced levels of cdc42, thereby affecting its overall activation (GTP-bound state). Expression of miR-424 under euglycaemic conditions seems to inhibit invasion and metastatic behaviour of TNBC. This was clearly demonstrated by our in vivo metastasis assay where miR-424 knockdown of TNBC cells had increased metastasis. Apart from playing a crucial role in influencing the invasive abilities of TNBC cells in hyperglycaemia, we have demonstrated miR-424 to be closely associated with the regulation of CSC pool. Reduced miR-424 in hyperglycaemic conditions resulted in hyperactivation of CSC population leading to its undesirable expansion. Drasin et al (2015) has reported the role of miR-424 in the process of breast tumourigenesis and metastasis, wherein they specifically pointed out that miR-424 did not seem to favour tumour-initiating properties. Tumour initiation has been strongly associated with cancer stem cell activation, and in the context of breast cancer under hyperglycaemia, miR-424 seems to play a tumour-suppressive role.
Our observations regarding expansion of CSC population under hyperglycaemic condition can be associated with epidemiological data showing aggravation of breast cancer malignancy and recurrence post therapy in breast cancer patients with diabetes. Mechanistically, we found that knockdown of miR-424 resulted in significant activation of TF, prdm14. Prdm14 is not a very widely studied TF, but its overexpression has been associated with poor prognosis in breast cancer patients (Nishikawa et al, 2007; Moelans et al, 2011, 2014). Interestingly, in one of the studies comparing the copy numbers of oncogenes and tumour suppressors in breast cancer samples suggested prdm14 to be one of the genes that showed significantly higher frequency of gain and copy number amplification in liver and lung mets (Moelans et al, 2014). In yet another study, prdm14 was reported to favour metastasis through MMP/TIMP, leading to increased degradation of extracellular matrix (Lu et al, 2016). Current literature does not provide evidence for direct regulation of prdm14 by miR-424. We proposed and demonstrated that the miR-424 regulation over prdm14 is mediated by cdc42–stat5 axis through pak1 kinase activation under hyperglycaemic condition. In addition to promotion of invasion, activated cdc42-stat5-prdm14 signalling cascade results in hyperactivation of the breast CSC pool.
In conclusion, hyperglycaemia can contribute towards aggravating breast malignancies by promoting invasion and hyperactivation of the CSC pool. Both invasive nature of cancer cells and hyperactivated CSC pool go hand in hand in worsening the prognosis of cancer patients. MiR-424, by its ability to regulate cdc42 expression and activation, can be explored to combat aggressive behaviour of TNBC cells under hyperglycaemic condition. Also, suppressing the activity of cdc42 to target breast cancer invasion and CSC hyperactivity can be a beneficial strategy in this scenario.
Change history
21 November 2017
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Chen B, Duan L, Yin G, Tan J, Jiang X (2013) Simultaneously expressed miR-424 and miR-381 synergistically suppress the proliferation and survival of renal cancer cells – Cdc2 activity is up-regulated by targeting WEE1. Clinics (Sao Paulo) 68: 825–833.
Cho WC (2007) OncomiRs: the discovery and progress of microRNAs in cancers. Mol Cancer 6: 60.
Dallavalle C, Albino D, Civenni G, Merulla J, Ostano P, Mello-Grand M, Rossi S, Losa M, D’Ambrosio G, Sessa F, Thalmann GN, Garcia-Escudero R, Zitella A, Chiorino G, Catapano CV, Carbone GM (2016) MicroRNA-424 impairs ubiquitination to activate STAT3 and promote prostate tumor progression. J Clin Invest 126: 4585–4602.
Drasin DJ, Guarnieri AL, Neelakantan D, Kim J, Cabrera JH, Wang CA, Zaberezhnyy V, Gasparini P, Cascione L, Huebner K, Tan AC, Ford HL (2015) TWIST1-induced miR-424 reversibly drives mesenchymal programming while inhibiting tumor initiation. Cancer Res 75: 1908–1921.
Emaus A, Veierod MB, Tretli S, Finstad SE, Selmer R, Furberg AS, Bernstein L, Schlichting E, Thune I (2010) Metabolic profile, physical activity, and mortality in breast cancer patients. Breast Cancer Res Treat 121: 651–660.
Esquela-Kerscher A, Slack FJ (2006) Oncomirs–microRNAs with a role in cancer. Nat Rev Cancer 6: 259–269.
Flores-Lopez LA, Martinez-Hernandez MG, Viedma-Rodriguez R, Diaz-Flores M, Baiza-Gutman LA (2016) High glucose and insulin enhance uPA expression, ROS formation and invasiveness in breast cancer-derived cells. Cell Oncol (Dordr) 39: 365–378.
Fritz G, Brachetti C, Bahlmann F, Schmidt M, Kaina B (2002) Rho GTPases in human breast tumours: expression and mutation analyses and correlation with clinical parameters. Br J Cancer 87: 635–644.
Fritz G, Just I, Kaina B (1999) Rho GTPases are over-expressed in human tumors. Int J Cancer 81: 682–687.
Giovannucci E, Harlan DM, Archer MC, Bergenstal RM, Gapstur SM, Habel LA, Pollak M, Regensteiner JG, Yee D (2010) Diabetes and cancer: a consensus report. Diabetes Care 33: 1674–1685.
Han L, Peng B, Ma Q, Ma J, Li J, Li W, Duan W, Chen C, Liu J, Xu Q, Laporte K, Li Z, Wu E (2013) Indometacin ameliorates high glucose-induced proliferation and invasion via modulation of e-cadherin in pancreatic cancer cells. Curr Med Chem 20: 4142–4152.
Kang X, Kong F, Wu X, Ren Y, Wu S, Wu K, Jiang Z, Zhang W (2015) High glucose promotes tumor invasion and increases metastasis-associated protein expression in human lung epithelial cells by upregulating heme oxygenase-1 via reactive oxygen species or the TGF-beta1/PI3K/Akt signaling pathway. Cell Physiol Biochem 35: 1008–1022.
Kaplan MA, Pekkolay Z, Kucukoner M, Inal A, Urakci Z, Ertugrul H, Akdogan R, Firat U, Yildiz I, Isikdogan A (2012) Type 2 diabetes mellitus and prognosis in early stage breast cancer women. Med Oncol 29: 1576–1580.
Kaye JA, Meier CR, Walker AM, Jick H (2002) Statin use, hyperlipidaemia, and the risk of breast cancer. Br J Cancer 86: 1436–1439.
Li C, Zhang G, Zhao L, Ma Z, Chen H (2016) Metabolic reprogramming in cancer cells: glycolysis, glutaminolysis, and Bcl-2 proteins as novel therapeutic targets for cancer. World J Surg Oncol 14: 15–016-0769-9.
Li Q, Qiu XM, Li QH, Wang XY, Li L, Xu M, Dong M, Xiao YB (2015) MicroRNA-424 may function as a tumor suppressor in endometrial carcinoma cells by targeting E2F7. Oncol Rep 33: 2354–2360.
Llobet-Navas D, Rodriguez-Barrueco R, Castro V, Ugalde AP, Sumazin P, Jacob-Sendler D, Demircan B, Castillo-Martin M, Putcha P, Marshall N, Villagrasa P, Chan J, Sanchez-Garcia F, Pe’er D, Rabadan R, Iavarone A, Cordon-Cardo C, Califano A, Lopez-Otin C, Ezhkova E, Silva JM (2014a) The miR-424(322)/503 cluster orchestrates remodeling of the epithelium in the involuting mammary gland. Genes Dev 28: 765–782.
Llobet-Navas D, Rodriguez-Barrueco R, de la Iglesia-Vicente J, Olivan M, Castro V, Saucedo-Cuevas L, Marshall N, Putcha P, Castillo-Martin M, Bardot E, Ezhkova E, Iavarone A, Cordon-Cardo C, Silva JM (2014b) The microRNA 424/503 cluster reduces CDC25A expression during cell cycle arrest imposed by transforming growth factor beta in mammary epithelial cells. Mol Cell Biol 34: 4216–4231.
Long XH, Mao JH, Peng AF, Zhou Y, Huang SH, Liu ZL (2013) Tumor suppressive microRNA-424 inhibits osteosarcoma cell migration and invasion via targeting fatty acid synthase. Exp Ther Med 5: 1048–1052.
Lopez R, Arumugam A, Joseph R, Monga K, Boopalan T, Agullo P, Gutierrez C, Nandy S, Subramani R, de la Rosa JM, Lakshmanaswamy R (2013) Hyperglycemia enhances the proliferation of non-tumorigenic and malignant mammary epithelial cells through increased leptin/IGF1R signaling and activation of AKT/mTOR. PLoS One 8: e79708.
Lu Y, Wan Z, Zhang X, Zhong X, Rui L, Li Z (2016) PRDM14 inhibits 293T cell proliferation by influencing the G1/S phase transition. Gene 595: 180–186.
Manser E, Huang HY, Loo TH, Chen XQ, Dong JM, Leung T, Lim L (1997) Expression of constitutively active alpha-PAK reveals effects of the kinase on actin and focal complexes. Mol Cell Biol 17: 1129–1143.
McHenry PR, Vargo-Gogola T (2010) Pleiotropic functions of Rho GTPase signaling: a Trojan horse or Achilles’ heel for breast cancer treatment? Curr Drug Targets 11: 1043–1058.
Moelans CB, de Wegers RA, Monsuurs HN, Maess AH, van Diest PJ (2011) Molecular differences between ductal carcinoma in situ and adjacent invasive breast carcinoma: a multiplex ligation-dependent probe amplification study. Cell Oncol (Dordr) 34: 475–482.
Moelans CB, van der Groep P, Hoefnagel LD, van de Vijver MJ, Wesseling P, Wesseling J, van der Wall E, van Diest PJ (2014) Genomic evolution from primary breast carcinoma to distant metastasis: Few copy number changes of breast cancer related genes. Cancer Lett 344: 138–146.
Nishikawa N, Toyota M, Suzuki H, Honma T, Fujikane T, Ohmura T, Nishidate T, Ohe-Toyota M, Maruyama R, Sonoda T, Sasaki Y, Urano T, Imai K, Hirata K, Tokino T (2007) Gene amplification and overexpression of PRDM14 in breast cancers. Cancer Res 67: 9649–9657.
Owiredu WK, Donkor S, Addai BW, Amidu N (2009) Serum lipid profile of breast cancer patients. Pak J Biol Sci 12: 332–338.
Takatani-Nakase T, Matsui C, Maeda S, Kawahara S, Takahashi K (2014) High glucose level promotes migration behavior of breast cancer cells through zinc and its transporters. PLoS One 9: e90136.
Tang Z, Wang J, Zhang H, Sun L, Tang F, Deng Q, Yu J (2016) Associations between diabetes and quality of life among breast cancer survivors. PLoS One 11: e0157791.
Villarreal-Garza C, Shaw-Dulin R, Lara-Medina F, Bacon L, Rivera D, Urzua L, Aguila C, Ramirez-Morales R, Santamaria J, Bargallo E, Mohar A, Herrera LA (2012) Impact of diabetes and hyperglycemia on survival in advanced breast cancer patients. Exp Diabetes Res 2012: 732027.
Voorhoeve PM, Agami R (2007) Classifying microRNAs in cancer: the good, the bad and the ugly. Biochim Biophys Acta 1775: 274–282.
Wu CT, Lin WY, Chang YH, Lin PY, Chen WC, Chen MF (2015) DNMT1-dependent suppression of microRNA424 regulates tumor progression in human bladder cancer. Oncotarget 6: 24119–24131.
Wu X, Ruan Y, Jiang H, Xu C (2017) MicroRNA-424 inhibits cell migration, invasion, and epithelial mesenchymal transition by downregulating doublecortin-like kinase 1 in ovarian clear cell carcinoma. Int J Biochem Cell Biol 85: 66–74.
Xu H, Chen K, Jia X, Tian Y, Dai Y, Li D, Xie J, Tao M, Mao Y (2015) Metformin use is associated with better survival of breast cancer patients with diabetes: a meta-analysis. Oncologist 20: 1236–1244.
Yamaji M, Ueda J, Hayashi K, Ohta H, Yabuta Y, Kurimoto K, Nakato R, Yamada Y, Shirahige K, Saitou M (2013) PRDM14 ensures naive pluripotency through dual regulation of signaling and epigenetic pathways in mouse embryonic stem cells. Cell Stem Cell 12: 368–382.
Yerrabothala S, Shaaban H, Capo G, Maroules M, Debari VA (2014) The impact of diabetes mellitus on breast cancer outcomes: a single center retrospective study. Pathol Oncol Res 20: 209–214.
Zhang H, Wang X, Chen Z, Wang W (2015) MicroRNA-424 suppresses estradiol-induced cell proliferation via targeting GPER in endometrial cancer cells. Cell Mol Biol (Noisy-le-grand) 61: 96–101.
Zhang L, Xu Y, Jin X, Wang Z, Wu Y, Zhao D, Chen G, Li D, Wang X, Cao H, Xie Y, Liang Z (2015) A circulating miRNA signature as a diagnostic biomarker for non-invasive early detection of breast cancer. Breast Cancer Res Treat 154: 423–434.
Acknowledgements
We thank Texas Tech University Health Sciences Center El Paso for supporting this project.
Author contributions
SBN: Conceived the idea, designed and carried out the experiments and drafted the manuscript; AO: helped with cell culture and WBs; RL-V and RR: helped with microRNA array and analysis; RS and AA: shared knowledge to improve scientific quality of the manuscript; AKD: statistics consultant; VS, GP, SJ and RL: helped with various experimental techniques; RL: conceived the idea, supervised the project and helped in finalising the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License.
Supplementary Information accompanies this paper on British Journal of Cancer website
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Nandy, S., Orozco, A., Lopez-Valdez, R. et al. Glucose insult elicits hyperactivation of cancer stem cells through miR-424–cdc42–prdm14 signalling axis. Br J Cancer 117, 1665–1675 (2017). https://doi.org/10.1038/bjc.2017.335
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2017.335
Keywords
This article is cited by
-
Rho GTPases in cancer radiotherapy and metastasis
Cancer and Metastasis Reviews (2020)
-
PRDM14 promotes malignant phenotype and correlates with poor prognosis in colorectal cancer
Clinical and Translational Oncology (2020)
-
H19X-encoded miR-424(322)/-503 cluster: emerging roles in cell differentiation, proliferation, plasticity and metabolism
Cellular and Molecular Life Sciences (2019)
-
Effect of Different Glucose Concentrations on Small RNA Levels and Adherence of Streptococcus mutans
Current Microbiology (2019)