Abstract
Background:
Mutations in GNAQ/11 genes are considered an early event in the development of uveal melanoma that may derive from a pre-existing nevus. The Hippo pathway, by way of YAP activation, rather than MAP kinase, has a role in the oncogenic capacity of GNAQ/11 mutations.
Methods:
We investigated 16 nevi from 13 human eyes for driver GNAQ/11 mutations using droplet digital PCR and determined whether nevi are clonal by quantifying mutant nevus cell fractions. Immunohistochemistry was performed on 15 nevi to analyse YAP activation.
Results:
For 15 out of 16 nevi, a GNAQ/11 mutation was detected in the nevus cells albeit at a low frequency with a median of 13%. Nuclear YAP, a transcriptional co-activator in the Hippo tumour-suppressor pathway, was detected in 14/15 nevi.
Conclusions:
Our analysis suggests that a mutation in GNAQ/11 occurs in a subset of choroidal nevus cells. We hypothesise that GNAQ/11 mutant-driven extracellular mitogenic signalling involving YAP activation leads to accumulation of wild-type nevus cells.
Similar content being viewed by others
Main
Similar to cutaneous melanoma (CM), uveal melanoma (UM) can develop de novo or from a pre-existing nevus (Figure 1). Uveal nevi most commonly occur in the choroid and are seen in 6.5–30% of the population (Sumich et al, 1998; Baderca et al, 2013). However, transformation from a choroidal nevus to a UM requires complex genetic aberrations (gain of chromosome 8q/6p, loss of chromosome 3, absence of BAP1), and has been reported in only about 1 out of 9000 people (Singh et al, 2005). We hypothesise that a choroidal nevus, as a potential precursor of UM, may already harbour clonal driver mutations, just as many cutaneous nevi are reported to present the typical V600E BRAF mutation associated with CM (Pollock et al, 2003; Shain et al, 2015). Mutations in BRAF are found in about 66% of CM while 83–95% of UM carry a mutation in either the GNAQ or GNA11 gene (Davies et al, 2002; Van Raamsdonk et al, 2008, 2010). Although MAPK (mitogen-activated protein kinase) signalling is activated by the BRAF V600E mutation, the oncogenic effect of GNAQ/11 is still to be defined. A recent publication showed that MAPK activation has a minimal role in the oncogenic potential of GNAQ mutations (Mouti et al, 2016); however, a central role has been attributed to the YAP/Hippo pathway. YAP (Yes-associated protein) has not only a role in homoeostasis and organ size during stress, but also in cancer several oncogenes and tumour suppressor genes converge on YAP (Overholtzer et al, 2006). In a sparse cell environment, YAP is expressed in cell nuclei-promoting gene expression. Conversely, in a dense cell environment, YAP is maintained in the cytoplasm and inhibits cell growth. Overall, tissue growth is controlled through the inhibition of YAP and its co-factors. Yu et al showed that GNAQ/11 mutant activity is controlled by YAP activation (nuclear YAP expression) in several UM cell lines (Yu et al, 2014, 2015). In this study, we analysed the frequency of GNAQ/11 mutations in choroidal nevi and analysed YAP activation as a possible downstream effect.
Clinical and histopathological manifestation of a choroidal nevus. (A) Choroidal nevus (N) (72 year old, female). Retina and RPE removed; choroidal vessels visible (asterisk). Size ∼4 mm diameter. (B) Section through nevus showed thickened area of pigmented nevus cells and large blood vessels at the lesion’s edge (arrows) (periodic acid–Schiff stain).
Materials and methods
From post-mortem human eyes obtained from the Lions NSW Eye Bank (with consent and Human Research Ethics Committee approval), choroidal pigmented lesions were identified, dissected and tissue was paraffin-embedded (Supplementary Materials and Methods) Dermal nevus tissues were obtained from the nevus biobank of the Department of Dermatology, Leiden University Medical Centre.
DNA extraction from nevi
DNA was extracted from formalin-fixed and paraffin-embedded tissue of 16 choroidal nevi using the ReliaPrep FFPE gDNA Miniprep System (Promega Corp, Madison, WI, USA), following the manufacturer’s instructions. For DNA extraction from dermal nevi, 20 tissue sections of 25 μm per nevus were used.
Digital PCR
GNAQ/11 mutations (GNAQ Q209L, GNAQ Q209P, GNAQ R1830, GNA11 Q209L and GNA11 R183C) and the BRAF (V600E) mutation were analysed in choroidal nevi and dermal nevi using digital PCR (Versluis et al, 2015). For copy number analysis of chromosome 3 and 8q, essays for PPARG and PTK2 were used. Proprietary probes and primers (Bio-Rad Laboratories, Inc., Hercules, CA, USA) were used.
Immunohistochemistry
Immunohistochemistry was performed on paraffin sections of 15 choroidal nevi using the Ventana Benchmark Ultra automated staining system (Ventana Medical Systems Inc, Tuscon, AZ, USA), following the protocol described previously (van Essen et al, 2014). Anti-Melan-A antibody (Ventana, 790-2290), YAP antibody (Cell Signalling Technology #4912) and the BAP1 antibody (Santa Cruz, SC-28383, Dallas, TX, USA) were used as primary antibodies. The UltraView Universal Alkaline Phosphatase Red detection kit, (Ventana Ref.: 760-501) was used in combination with target amplification (Ventana ref.: 760-080). The slides were counterstained with haematoxylin II for 8 min, followed by an additional counterstaining with blueing reagent (Ventana medical systems Inc.). Due to red chromogen staining on thin tissue slides, we were able to detect positive staining in pigmented cells.
See also Supplementary Materials and Methods.
Results
In the current study, 16 nevi from 13 post-mortem eyes of 12 donors were analysed for GNAQ/11 hotspot mutations. One pair of donor eyes presented nevi in both the right and the left eye (nevi 11A-C), with the left eye containing two nevi (nevi 11A-B). Two other eyes also contained two nevi (nevi 6 and 12) (Table 1).
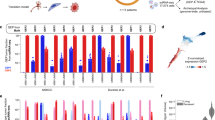
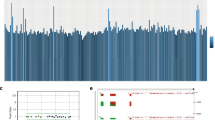
Droplet digital PCR showed that 15 of the 16 choroidal nevi presented one out of the three GNAQ/11 hotspot mutations (Table 1). Mutations in GNAQ and GNA11 were distributed equally in this cohort, similar to our observations in UM (Versluis et al, 2015). None of the nevi showed evidence of multiple mutations. Nevus 12B did not carry either of the GNAQ/11 hotspot mutations. Different nevi from the same eye (nevi 6A-B, 11A-B and 12A-B) never shared the same hotspot mutation. The nevus from the right eye (nevus 11C) of the donor who had nevi in both eyes contained the same mutation as one of the nevi in the left eye (nevus 11B) (Table 1). The fraction of mutated nevus cells varied from 1–55% with a median of 13%. Immunostaining revealed that over 95% of the cells in 12 nevi were Melan-A positive (Table 1), indicating that the tissue examined represented primarily nevus cells with some stromal cells (Supplementary Figure 1B). Four nevi could not be determined for Melan-A due to lack of material.
We examined tissues for the presence of chromosome aberrations as potential progression markers and as the absence of BAP1 is correlated with monosomy 3 in UM, we also performed immunohistochemistry for BAP1 as another progression marker. (van Essen et al, 2014) Neither gain of chromosome 8q/chromosome 6p nor loss of chromosome 3 was observed in the three nevi with the highest mutation fraction (nevi 11A-C) (not shown). The eight nevi stained for BAP1 showed no loss of BAP1 expression (Table 1).
We determined the presence of GNAQ/11 and BRAF mutations in dermal cutaneous nevi (n=5) as a control. None of which contained a GNAQ/11 hotspot mutation; instead, four out of five dermal nevi harboured the BRAF V600E mutation with a median mutant fraction of 45%. The mutant fraction of dermal nevi is in marked contrast with the 13% GNAQ/11 mutant cells in choroidal nevi and emphasises potential differences between BRAF and GNAQ/11-driven nevus development. This suggests besides that either other genetic factors are involved or that GNAQ/11 mutant clones can also drive the proliferation of the wild-type melanocytes in the nevus. The latter mechanism could be explained by YAP activation initiated by GNAQ/11-mutated cells, which could stimulate wild-type cells to proliferate.
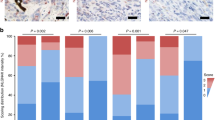
We found heterogeneous YAP nuclear immunostaining in 14 out of 15 choroidal nevi, with no cytoplasmic staining (Table 1 and Figure 1). Although YAP staining tended to be more abundant in nevi with a high mutant fraction, there was no significant correlation between the percentage of positive YAP cells and the mutant cell fraction (Table 1, P=0.334, Spearman correlation coefficient). In the only nevus (12B) without a GNAQ/11 mutation, positive YAP expression in nuclei was also detected. This indicates that YAP activation may occur independently of the presence of a GNAQ/11 mutation. Normal choroidal melanocytes were negative for YAP staining, as was retinal tissue. Sporadically a retinal pigment epithelium cell was found with positive YAP staining (Figure 2).
YAP staining in human choroidal nevi. (A) Nevus 2 with an average of 1.8% YAP-positive cells. A nevus cell with a negative, blue nucleus is seen (white arrow). Magnification of objective × 40. (B) Nevus 12A with an average of 13% YAP-positive cells. Yap+ nuclei in nevus cells are visible (pink, white arrows). A rare RPE cell is found with nuclear YAP staining (blue arrow). Magnification of objective × 40. BM=Bruch’s membrane; cc=choriocapillaris; rpe=retina pigment epithelium.
Discussion
As hypothesised, human choroidal nevi did harbour a GNAQ/11 mutation. No other mutations were found, which suggests that those mutations are clonal. The low GNAQ/11 mutant fraction in choroidal nevi however presents a paradox for GNAQ/11 oncogene function that may be explained by aberrant YAP signalling in choroidal nevi. The oncogenic potential of YAP is broadly recognised and high levels of YAP expression in the nucleus have been associated with several malignancies (Overholtzer et al, 2006). Yu et al (2014) described a positive correlation between mutant GNAQ/11 and aberrant YAP activation (nuclear expression) in UM cell lines. In our study, the nevi with a GNAQ/11 mutation all showed nuclear YAP, confirming this positive correlation between mutant GNAQ/11 and YAP activation. However, this correlation was not significant. Moreover, the one nevus without GNAQ/11 mutations still possessed YAP activity in the nucleus. This is in accordance with the results of Yu et al who also found evidence of nuclear YAP activity in wild-type UM cell lines (Mel285 and Mel290), suggesting that YAP activation is not solely dependent on GNAQ/11 mutations, and that other factors may contribute to YAP activity (Yu et al, 2014).
In summary, we found that GNAQ/11 mutant clones make up a fraction of the cells in choroidal nevi. Nevus cells are furthermore characterised by heterogeneous YAP expression. Combined GNAQ/11 and YAP may constitute a putative precursor tumour pathway with an activated oncogene (GNAQ/11) and downstream effector (YAP).
Change history
05 September 2017
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Baderca F, Solovan C, Boghian L (2013) Epidemiological and morphological data of ocular melanocytic lesions. Rom J Morphol Embryol 54 (1): 77–83.
Davies H, Bignell GR, Cox C, Stephens P, Edkins S, Clegg S, Teague J, Woffendin H, Garnett MJ, Bottomley W, Davis N, Dicks E, Ewing R, Floyd Y, Gray K, Hall S, Hawes R, Hughes J, Kosmidou V, Menzies A, Mould C, Parker A, Stevens C, Watt S, Hooper S, Wilson R, Jayatilake H, Gusterson BA, Cooper C, Shipley J, Hargrave D, Pritchard-Jones K, Maitland N, Chenevix-Trench G, Riggins GJ, Bigner DD, Palmieri G, Cossu A, Flanagan A, Nicholson A, Ho JW, Leung SY, Yuen ST, Weber BL, Seigler HF, Darrow TL, Paterson H, Marais R, Marshall CJ, Wooster R, Stratton MR, Futreal PA (2002) Mutations of the BRAF gene in human cancer. Nature 417 (6892): 949–954.
Mouti MA, Dee C, Coupland SE, Hurlstone AF (2016) Minimal contribution of ERK1/2-MAPK signalling towards the maintenance of oncogenic GNAQQ209P-driven uveal melanomas in zebrafish. Oncotarget 7 (26): 39654–39670.
Overholtzer M, Zhang J, Smolen GA, Muir B, Li W, Sgroi DC, Deng CX, Brugge JS, Haber DA (2006) Transforming properties of YAP, a candidate oncogene on the chromosome 11q22 amplicon. Proc Natl Acad Sci USA 103 (33): 12405–12410.
Pollock PM, Harper UL, Hansen KS, Yudt LM, Stark M, Robbins CM, Moses TY, Hostetter G, Wagner U, Kakareka J, Salem G, Pohida T, Heenan P, Duray P, Kallioniemi O, Hayward NK, Trent JM, Meltzer PS (2003) High frequency of BRAF mutations in nevi. Nat Genet 33 (1): 19–20.
Shain AH, Yeh I, Kovalyshyn I, Sriharan A, Talevich E, Gagnon A, Dummer R, North J, Pincus L, Ruben B, Rickaby W, D’Arrigo C, Robson A, Bastian BC (2015) The genetic evolution of melanoma from precursor lesions. N Engl J Med 373 (20): 1926–1936.
Singh AD, Kalyani P, Topham A (2005) Estimating the risk of malignant transformation of a choroidal nevus. Ophthalmology 112 (10): 1784–1789.
Sumich P, Mitchell P, Wang JJ (1998) Choroidal nevi in a white population: the Blue Mountains Eye Study. Arch Ophthalmol 116 (5): 645–650.
van Essen TH, van Pelt SI, Versluis M, Bronkhorst IH, van Duinen SG, Marinkovic M, Kroes WG, Ruivenkamp CA, Shukla S, de KA, Kilic E, Harbour JW, Luyten GP, van der Velden PA, Verdijk RM, Jager MJ (2014) Prognostic parameters in uveal melanoma and their association with BAP1 expression. Br J Ophthalmol 98 (12): 1738–1743.
Van Raamsdonk CD, Bezrookove V, Green G, Bauer J, Gaugler L, O’Brien JM, Simpson EM, Barsh GS, Bastian BC (2008) Frequent somatic mutations of GNAQ in uveal melanoma and blue naevi. Nature 457 (7229): 599–602.
Van Raamsdonk CD, Griewank KG, Crosby MB, Garrido MC, Vemula S, Wiesner T, Obenauf AC, Wackernagel W, Green G, Bouvier N, Sozen MM, Baimukanova G, Roy R, Heguy A, Dolgalev I, Khanin R, Busam K, Speicher MR, O’Brien J, Bastian BC (2010) Mutations in GNA11 in uveal melanoma. N Engl J Med 363 (23): 2191–2199.
Versluis M, de Lange MJ, van Pelt SI, Ruivenkamp CA, Kroes WG, Cao J, Jager MJ, Luyten GP, van der Velden PA (2015) Digital PCR validates 8q dosage as prognostic tool in uveal melanoma. PLoS One 10 (3): e0116371.
Yu FX, Luo J, Mo JS, Liu G, Kim YC, Meng Z, Zhao L, Peyman G, Ouyang H, Jiang W, Zhao J, Chen X, Zhang L, Wang CY, Bastian BC, Zhang K, Guan KL (2014) Mutant Gq/11 promote uveal melanoma tumorigenesis by activating YAP. Cancer Cell 25 (6): 822–830.
Yu FX, Zhao B, Guan KL (2015) Hippo pathway in organ size control, tissue homoeostasis, and cancer. Cell 163 (4): 811–828.
Acknowledgements
We acknowledge Frank de Gruijl for helpful discussion. This work was supported by grants from the Nelly Reef Fund, the Leiden Ophthalmology Subsidy Fund (03121401.loof.madeleine vader) and the Dutch Cancer Society (KWF: UL2011-4991).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License.
Supplementary Information accompanies this paper on British Journal of Cancer website
Supplementary information
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Vader, M., Madigan, M., Versluis, M. et al. GNAQ and GNA11 mutations and downstream YAP activation in choroidal nevi. Br J Cancer 117, 884–887 (2017). https://doi.org/10.1038/bjc.2017.259
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2017.259
Keywords
This article is cited by
-
The rationale for treating uveal melanoma with adjuvant melatonin: a review of the literature
BMC Cancer (2022)
-
HDAC11 activity contributes to MEK inhibitor escape in uveal melanoma
Cancer Gene Therapy (2022)
-
Palmitoylation of GNAQ/11 is critical for tumor cell proliferation and survival in GNAQ/11-mutant uveal melanoma
Frontiers of Medicine (2022)
-
Scientific and clinical implications of genetic and cellular heterogeneity in uveal melanoma
Molecular Biomedicine (2021)
-
Anticancer effects of mifepristone on human uveal melanoma cells
Cancer Cell International (2021)