Abstract
Background:
Chromosome 8q24 has emerged as an important genetic susceptibility region for several cancers, including prostate cancer; however, little is known about the contribution of DNA methylation in this region to risk.
Methods:
We prospectively evaluated DNA methylation at 8q24 in relation to prostate cancer using pre-diagnostic blood samples from 694 prostate cancer cases (including 172 aggressive cases) and 703 controls in the Prostate, Lung, Colorectal and Ovarian Cancer Screening Trial. We used logistic regression to estimate odds ratios and 95% confidence intervals.
Results:
Although none remained significant after adjustment for multiple testing (q>0.05), of the 50 CpG sites meeting quality control, we identified 8 sites that were nominally associated with prostate cancer (Ptrend<0.05), including 6 correlated (Spearman ρ: 0.20–0.52) sites in POU5F1B and 2 intergenic sites (most significant site: Chr8:128428897 in POU5F1B, Ptrend=0.01). We also identified two correlated (ρ=0.39) sites in MYC (Chr8:128753187 and Chr8:128753154) that were associated with aggressive (Ptrend=0.02 and 0.03), but not non-aggressive disease (Ptrend=0.70 and 0.20; Pheterogeneity=0.01 and 4.6 × 10−3). These findings persisted after adjustment for the top 8q24 prostate cancer variants in our study.
Conclusions:
Although requiring replication, our findings provide some evidence that 8q24 DNA methylation levels may be associated with prostate cancer risk.
Similar content being viewed by others
Main
Genetic variants in several regions at chromosome 8q24 have been found to be associated with the risk of a variety of cancers, underscoring the importance and complexity of this region in relation to cancer risk. Chromosome 8q24 is of particular importance for prostate cancer, as genome-wide association studies (GWAS) and related studies have identified associations for multiple independent single nucleotide polymorphisms (SNPs) (Amundadottir et al, 2006; Gudmundsson et al, 2007; Haiman et al, 2007; Yeager et al, 2007; Eeles et al, 2008; Thomas et al, 2008; Al Olama et al, 2009; Schumacher et al, 2011) and at least one rare variant (Gudmundsson et al, 2012) at 8q24 with prostate cancer risk.
With the oncogene MYC located more than 200 kb downstream from the closest prostate cancer susceptibility SNP, the 8q24 region has traditionally been described as a gene desert and the mechanism underlying prostate carcinogenesis remains unclear. However, additional genes (e.g., POU5F1B) and non-coding RNAs (e.g., PVT1 and PRNCR1), as well as transcriptional enhancers responsive to androgen, have also been identified in the region and might contribute to risk (Jia et al, 2009; Kastler et al, 2010; Chung et al, 2011; Meyer et al, 2011). It has been proposed that epigenetic mechanisms might have a role based on the identification of gene regulatory elements at 8q24 and evidence of long-range interactions for this locus with MYC or PVT1 in prostate cancer cell lines (Ahmadiyeh et al, 2010; Sotelo et al, 2010; Meyer et al, 2011). Providing further support for this hypothesis, genetic variation at 8q24 was associated with PVT1 expression in prostate tissue in a previous study (Meyer et al, 2011). However, the evidence for an association between 8q24 genetic variants and MYC expression has been mixed and appears to vary by tissue type, with most studies in prostate tissue demonstrating null associations (Pomerantz et al, 2009; Prokunina-Olsson and Hall, 2009; Wright et al, 2010). Notably, a recent study evaluating physical interactions across the genome for several prostate cancer susceptibility regions at 8q24 identified a number of additional intra- and inter-chromosomal gene targets in prostate cancer cell lines that tended to be enriched for important cancer pathways, suggesting that the 8q24 locus may serve as a regulatory hub for a variety of genes in key pathways (Du et al, 2015).
Alteration in DNA methylation is a key epigenetic mechanism that can affect gene expression and may have a role in maintaining genomic stability, both of which are thought to be important in carcinogenesis (Kulis and Esteller, 2010). Interestingly, studies have indicated that DNA methylation may mediate or augment the risk of established genetic loci for health outcomes such as obesity and rheumatoid arthritis (Almen et al, 2012; Liu et al, 2013). We recently reported associations between DNA methylation levels at several specific CpG sites in peripheral blood and established cancer susceptibility SNPs (including several prostate cancer susceptibility SNPs) at 8q24 among cancer-free Caucasian men, suggesting that there may be a role for DNA methylation at this important susceptibility locus in prostate cancer risk (Barry et al, 2014).
In the present study, we aimed to follow-up on these findings by prospectively evaluating whether CpG site DNA methylation at 8q24 in peripheral blood DNA is associated with the subsequent risk of prostate cancer in a nested case-control study within the Prostate, Lung, Colorectal and Ovarian (PLCO) Cancer Screening Trial. We also aimed to evaluate whether the CpG site associations with prostate cancer mediated or were independent of the associations for established prostate cancer susceptibility SNPs at 8q24. In addition, as many prostate cancers will likely never progress to cause death to the patient in his lifetime (Zlotta et al, 2013), we aimed to evaluate whether DNA methylation markers at 8q24 can distinguish between the risk of aggressive and non-aggressive prostate cancer.
Materials and methods
Study population
The PLCO Cancer Screening Trial is a randomised trial designed to assess the impact of specific cancer screening regimens on the risk of mortality from prostate, lung, colorectal and ovarian cancers. The participants include more than 150 000 men and women aged 55 to 74 years at the time of enrollment (1993–2001) from 10 centres in the United States (Hayes et al, 2005). Men randomised to the screening arm were offered both the prostate-specific antigen (PSA) test and digital rectal exam (DRE) at baseline, as well as the PSA test annually for 5 years thereafter and the DRE annually for 3 years thereafter (Prorok et al, 2000; Hayes et al, 2005). Participants were referred to their personal physician for follow-up based on a PSA test result >4 ng ml−1 or suspicion of prostate cancer from a DRE exam. Suspected prostate cancer cases based on screening examination results or self-report on annual follow-up questionnaires were confirmed by medical and pathologic records. The protocol for the PLCO trial was approved by the Institutional Review Boards (IRBs) at the 10 centres and the National Cancer Institute (NCI) and all participants provided informed consent. Approval for the present study was granted by the IRB at the NCI.
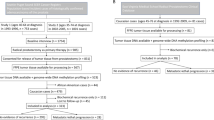
We conducted a nested prostate cancer case-control study within the screening arm of the PLCO Trial using pre-diagnostic peripheral blood samples as described previously (Barry et al, 2015). Briefly, in addition to the availability of a blood specimen, eligibility criteria included completion of the PLCO baseline questionnaire, consent to be included in etiologic studies of cancer and no history of cancer before randomisation (Barry et al, 2015). Cases were non-Hispanic Caucasian men with a pathologically confirmed diagnosis of prostate cancer at least 1 year after blood draw and were preferentially selected from among participants in the Cancer Genetic Markers of Susceptibility initiative (Yeager et al, 2007), where men with aggressive disease were oversampled. Controls were non-Hispanic Caucasian men who had no diagnosis of prostate cancer before the censor date for case diagnosis, 31 December 2007, and were frequency matched to cases on age at randomization (5-year intervals), year of randomisation, year of blood draw and study year of diagnosis/selection (Barry et al, 2015). A total of 707 cases and 707 controls were selected for the study. Following exclusion of participants with insufficient pre-diagnostic DNA, the final sample size was 694 cases and 703 controls (Barry et al, 2015). This included 172 aggressive cases (Stage III/IV or Gleason score ⩾8) and 516 non-aggressive cases (Stage I/II and Gleason score <8); the remaining 6 cases were missing data needed to characterise disease aggressiveness.
DNA methylation assays
We previously designed targeted pyrosequencing assays to quantify and evaluate variability in DNA methylation levels at chromosome 8q24 (Barry et al, 2014). One set of assays was designed to cover regions nearby (within 50 kb) prostate cancer susceptibility SNPs at 8q24 identified in populations of European ancestry and the other set was designed to cover regions in or near the MYC oncogene. For the present study, we carried forward the assays that performed well in our pilot study per data from quality control replicates, including coefficients of variation (CVs) and intraclass correlation coefficients, and that displayed moderate to high between-individual variation, based on the range of DNA methylation values across the pilot samples (Barry et al, 2014). These assays included 63 CpG sites located nearby the cancer susceptibility SNPs at 8q24 or in promoter, exon 2, exon 3 or 3′-regions for MYC (Supplementary Table S1).
DNA was extracted from peripheral blood samples using Qiagen QIAamp DNA Blood Midi/Maxi kits and shipped to EpigenDx, Inc. (Hopkinton, MA, USA), where the DNA was bisulfite converted using a Zymo Research EZ DNA Methylation kit (Zymo Research, Irvine, CA, USA) and PCR amplified within the chromosome 8q24 regions of interest using 45 cycles per PCR (Barry et al, 2015). Each plate included four artificial control samples (one negative control sample (no DNA added), as well as three positive control samples with known global DNA methylation levels: low (0%), partial (50%) and highly methylated (100%)). For quality control, we also included 58 blind replicate samples interspersed within and between plates (Barry et al, 2015). Sequencing was performed using the Pyrosequencing PSQ96 HS System (Pyrosequencing Qiagen). QCpG software (Pyrosequencing Qiagen) was used to determine the methylation status at each CpG site, and the percent of methylation was calculated for each CpG site as methylated cytosine divided by the sum of methylated and unmethylated cytosines (Barry et al, 2015).
Based on the replicate quality control samples, we calculated the CV for each CpG site and we excluded CpG sites that had an overall CV >25. Of the 63 CpG sites evaluated, 50 met our criteria for inclusion in the statistical analysis.
ENCODE/GEO data
We obtained ENCODE and GEO data annotations in a ±2 kb window around each of the CpG sites that were associated with the risk of overall prostate cancer or aggressive prostate cancer (P<0.05) in our study using the hg19 build UCSC ENCODE file browser (http://genome.ucsc.edu/cgi-bin/hgFileSearch) and GEO (http://www.ncbi.nlm.nih.gov/geo/). Using intersectBed (BEDTools package, https://github.com/arq5x/bedtools2; Quinlan and Hall, 2010), we evaluated intersections with transcription factor binding sites (TFBS) based on ChIP-seq data, as well as DNase I hypersensitivity and histone methylation/acetylation, in histologically normal tissue from prostate cancer patients (PrEC) and cancer cell lines of prostate origin (LNCaP, PC3, DU145 and VCaP). Peaks were identified based on previously reported significance thresholds (He et al, 2010; Maurano et al, 2012; Thurman et al, 2012; Chen et al, 2013; Hazelett et al, 2014; Jin et al, 2014; Polak et al, 2014; Puto et al, 2015; Toropainen et al, 2015).
Genotyped SNPs
Cases and controls were genotyped using the Illumina Omni2.5 as part of a larger GWAS of prostate cancer (Berndt et al, 2015) and common SNPs were imputed using 1000 Genomes Project data release version 3 (Abecasis et al, 2012) and IMPUTE2 (Howie et al, 2009). For this study, we evaluated 28 previously reported prostate cancer susceptibility SNPs, which were spread across four different prostate cancer susceptibility regions at 8q24 (Supplementary Table S2). These SNPs were selected on the basis of an association with prostate cancer in a GWAS with a P-value<10−6. We also included one SNP (rs7837328) that was not reported in a GWAS, but that previously was shown to interact with pesticide exposure to increase prostate cancer risk (P-interaction<0.05) in the US Agricultural Health Study (Koutros et al, 2010).
Statistical analysis
For the 50 CpG sites that met our quality control criteria, we categorised DNA methylation levels at each site into quartiles based on the distribution among controls and conducted separate logistic regression models to compute odds ratios (OR) and 95% confidence intervals (CIs) for each CpG site with prostate cancer risk. The lowest quartile was treated as the referent category. We computed P-values for trend by including the DNA methylation level at each CpG site in the model as a continuous variable. We also evaluated the relationship between the CpG sites and prostate cancer risk separately for aggressive and non-aggressive disease, as well as by time from draw to diagnosis (using four categories for the cases: <2 years, 2 to <3 years, 3 to <4 years and ⩾4 years; the comparison group was all controls combined), using polytomous regression and computed P-values for heterogeneity using Wald χ2-tests. In addition, we evaluated potential effect modification of the associations for the CpG sites and prostate cancer by age at diagnosis/selection (<65 or ⩾65 years) using likelihood ratio tests to compare nested models with and without the interaction terms with each CpG site because there is some evidence that DNA methylation levels may vary by age (Florath et al, 2014). All models were adjusted for age at blood draw and year of draw. Additional adjustment for folate intake, BMI, smoking, alcohol intake and family history of prostate cancer did not appreciably alter our findings and thus we present findings from the more parsimonious models here. For each CpG site—prostate cancer association, we also computed q-values reflecting the false discovery rate (FDR) using the Benjamini and Hochberg method (Benjamini and Hochberg, 1995) to account for the number of comparisons. We calculated q-values for all prostate cancer cases combined and then also separately by disease subtype (aggressive and non-aggressive). We used an FDR threshold of 0.05 to define findings that were significant after adjustment for multiple comparisons.
For each of the 28 prostate cancer susceptibility SNPs, we first evaluated the association with overall prostate cancer risk, as well as aggressive prostate cancer risk, using logistic regression, assuming a log-additive genetic model. Although in the full GWAS, the 8q24 locus was significantly associated with risk (Berndt et al, 2015), we wanted to evaluate what the SNPs associations were within this smaller subset of cases and controls. To assess whether the top CpG sites associated with risk (Ptrend<0.05) mediated or were independent of the association with the known prostate cancer susceptibility SNPs at 8q24, we conducted conditional analyses by including both the CpG sites and SNPs in the same model. Specifically, we tested the impact of adding CpG sites to SNP-only models (i.e., impact of CpG sites on SNP–prostate cancer associations) and vice versa, with separate models for each CpG site-SNP combination. For the conditional analyses for overall prostate cancer, we selected the most significant SNP associated with overall prostate cancer risk in our study for each of the four prostate cancer susceptibility regions at 8q24 for individual testing, and we also individually tested all other SNPs that were significantly associated (P<0.05) with overall prostate cancer in our study. We also did the same for SNPs associated with aggressive prostate cancer for the models of aggressive disease (that is, we tested the top SNP in each region and any others that were associated with aggressive prostate cancer with a P<0.05). In addition, we evaluated CpG site–SNP interactions for the top CpG sites and SNPs associated with the risk of overall prostate cancer in our study by using likelihood ratio tests comparing nested models with and without the interaction terms.
Results
Characteristics of the study population are included in Table 1. Compared with controls, cases were more likely to have a family history of prostate cancer and were less likely to have smoked. Cases and controls were similar with respect to age at blood draw and year of blood draw (study matching factors), folate and alcohol intakes and BMI. Of the 28 prostate cancer susceptibility SNPs previously reported for the chromosomal 8q24 region, 13 were associated with overall prostate cancer risk and 2 were associated with aggressive prostate cancer in this nested case-control study, with the most significant SNP for overall prostate cancer being rs4242382 (region 1, P-value=0.01; Supplementary Table S2).
Of the 50 CpG sites evaluated, we identified 8 CpG sites whose DNA methylation levels were associated with the risk of overall prostate cancer (Ptrend<0.05; Table 2), although these findings were no longer significant after correction for multiple testing (q-value>0.05). These included six CpG sites in POU5F1B that were correlated with each other to moderate or moderately high extent (Spearman ρ: 0.20–0.52) and two intergenic CpG sites (Chr8:128012411 and Chr8:128444762) that were weakly to moderately correlated with the other six sites (ρ: 0.05–0.23). The most significant CpG site overall was located at Chr8:128428897 in POU5F1B (OR for the highest compared to the lowest quartile=0.81, 95% CI: 0.60–1.10; Ptrend=0.01; Table 2). There were no significant differences in these associations by time from draw to diagnosis (Pheterogeneity>0.05; data not shown). When we further evaluated CpG site–SNP interactions for the top CpG sites and SNPs in our study, we also observed some evidence of interactions for Chr8:128428897 with rs4242382 and rs6983561 (P interaction=0.004 and 0.02, respectively), such that the association for this CpG site with the risk of overall prostate cancer was restricted to those carrying the variant allele for either of the SNPs (Supplementary Table S3); however, none of the P-values for interaction remained significant after adjustment for multiple comparisons (q-values>0.05; data not shown).
When the cases were stratified by disease aggressiveness, interestingly, we identified two moderately correlated (ρ=0.39) CpG sites in MYC (Chr8:128753187 and Chr8:128753154) that were specifically associated with the risk of aggressive (Ptrend=0.02 and Ptrend=0.03, respectively), but not non-aggressive prostate cancer (Ptrend=0.70 and Ptrend=0.20, respectively; Pheterogeneity=0.01 and Pheterogeneity=4.6 × 10−3, respectively) (Table 3). For the most significant CpG site associated with aggressive prostate cancer (Chr8:128753187), the ORs for aggressive and non-aggressive disease were 1.49 (95% CI: 0.91–2.45; Ptrend=0.02) and 1.05 (95% CI: 0.76–1.44; Ptrend=0.70), respectively; however, these findings did not remain significant after adjustment for multiple comparisons (q-value>0.05; Table 3). There were no significant differences in these associations by time from draw to diagnosis (Pheterogeneity>0.05; data not shown). For the eight CpG sites that were associated with overall prostate cancer risk, none of the associations significantly differed (Pheterogeneity>0.05) between aggressive and non-aggressive disease (Table 3).
To evaluate whether the associations that we observed for the CpG sites with overall prostate cancer mediated or were independent from the previously reported 8q24 SNP associations, we conducted conditional analyses for each CpG site including the top SNPs associated with overall prostate cancer risk in our study for each 8q24 susceptibility region in the model (Region 1: rs4242382, Region 2: rs6983561, Region 3: rs6999921 and Region 4: rs16902094). We also checked the impact of all other 8q24 SNPs that were significantly associated with overall prostate cancer risk in our study (P<0.05). Overall, there was little impact on the CpG site ORs or P-values and most of the top CpG sites remained significant at the 0.05 level after adjustment for the SNPs (Supplementary Table S4). Similarly, there was little impact of additional adjustment for the top SNPs associated with aggressive prostate cancer in our study (Region 1: rs7017300, Region 2: rs10086908, Region 3: rs7000448 and Region 4: rs16902094, which included the two SNPs that were significantly associated with aggressive disease in our study population) for the two MYC CpG sites that we identified as associated with the risk of aggressive prostate cancer, and the findings for both of these sites remained statistically significant (Supplementary Table S5). In addition, when we added the CpG sites to SNP-only models, the ORs and P-values for these prostate cancer susceptibility SNPs did not appreciably change and remained statistically significant after adjustment for the respective individual CpG sites (Supplementary Tables S4 and S5), providing further evidence that the prostate cancer associations observed for the CpG sites and SNPs may be independent of one another.
Given the importance of PSA in prostate cancer screening and detection, we also evaluated whether any of the CpG sites were associated with PSA levels among controls. Of the 10 CpG sites associated with either overall or aggressive prostate cancer, only one site (Chr8:128428869) was significantly associated with PSA at the time of blood draw among controls (Spearman ρ=−0.10, P=0.007).
We also followed up on the 10 CpG sites that were significantly associated with the risk of overall prostate cancer or aggressive prostate cancer in our study by using ENCODE and GEO data to search for TFBS, DNase I hypersensitivity sites (HS) and histone methylation/acetylation marks within 2 kb of the CpG sites in histologically normal tissue from prostate cancer patients and cancer cell lines of prostate origin. We identified a number of significant peaks based on previously reported significance thresholds (He et al, 2010; Maurano et al, 2012; Thurman et al, 2012; Chen et al, 2013; Hazelett et al, 2014; Jin et al, 2014; Polak et al, 2014; Puto et al, 2015; Toropainen et al, 2015) within 2 kb of three of the CpG sites: the intergenic CpG site at Chr8:128012411, and the two MYC CpG sites (Chr8:128753187 and Chr8:128753154) that were associated with the risk of aggressive prostate cancer. For Chr8:128012411, these included DNaseI HS peaks in the PrEC and LNCaP cell lines, a TFBS for DAXX in the PC3 cell line and histone methylation marks (H3K4me2) in the PC3 and VCaP cell lines. For the MYC CpG sites, these findings included DNaseI HS peaks in the PrEC and LNCaP cell lines, a number of histone methylation and acetylation marks (H3K4me1, H3K4me2, H3K4me3 and H3K27ac) in the LNCaP cell line, as well as H3K4me2 in the PC3 line, and TFBS for ETV1 and TCF7L2 (LNCaP line), DAXX (PC3 line) and FoxA1 (DU145 line). We did not observe significant peaks within 2 kb of the other CpG sites evaluated in any of the available prostate cell lines, although this could in part be due to the relatively repetitive nature of the 8q24 locus and the resulting poor alignability that we reported previously (Barry et al, 2014).
Although none of these CpG sites were found to change over time in our previous study (Barry et al, 2014), given the known changes in DNA methylation with age (Florath et al, 2014), we evaluated whether the associations for each of the 50 CpG sites differed by age at diagnosis/selection. Chr8:128428931 in POU5F1B was significantly associated with the risk of prostate cancer diagnosed before age 65 (Ptrend=0.04) and several CpG sites (the intergenic site at Chr8:128444762 and several sites in POU5F1B: Chr8:128428869, Chr8:128428897, Chr8:128428949 and Chr8:128428978) were significantly associated with the risk of prostate cancer diagnosed at age 65 years or older (Ptrend=0.02, 0.05, 0.05, 0.03 and 0.05, respectively). However, none of the P-values for interaction were statistically significant (P interaction>0.05; data not presented).
Discussion
Focusing on the chromosome 8q24 locus, which is known to be important in the genetic susceptibility to a variety of cancers, including prostate cancer, we aimed to evaluate the contribution of variation in DNA methylation at this locus in peripheral blood DNA to the risk of prostate cancer in Caucasian men using a large prospective design. We identified eight specific CpG sites at 8q24, including six correlated sites in the POU5F1B gene, as well as two CpG sites in intergenic regions, whose DNA methylation levels were associated with prostate cancer risk at the 0.05 level. In addition, two CpG sites in the MYC oncogene at 8q24 were associated (P<0.05) with the risk of aggressive, but not non-aggressive prostate cancer. These findings did not appreciably change and tended to remain nominally (P<0.05) statistically significant after adjustment for the top prostate cancer susceptibility SNPs in our study, although none of the 10 CpG site findings were significant after adjustment for multiple comparisons (q-value >0.05). To our knowledge, our study is the first to evaluate whether DNA methylation levels at 8q24 in peripheral blood DNA are associated with the subsequent risk of prostate cancer. In addition, our study is the first to suggest that a blood-based DNA methylation marker may be specifically associated with the risk of aggressive prostate cancer. These findings warrant replication in a large, independent population.
POU5F1B (POU class 5 homeobox 1B gene, also known as OCT4-pg1 and POU5F1P1) was first described as a pseudogene of the gene for octamer-binding transcription factor 4 (OCT4), which has a key role in embryonic development and stem cell pluripotency. However, recent evidence suggests that POU5F1B may encode a functional protein, which is thought to be a transcriptional activator and has been observed to be overexpressed in prostate cancer tissue (Kastler et al, 2010). Notably, some 8q24 cancer susceptibility SNPs have previously been shown to be in linkage disequilibrium with variants in the POU5F1B open reading frame and also correlated with POU5F1B expression in prostate tissue (Breyer et al, 2014). It is possible that the POU5F1B CpG sites that were associated with prostate cancer risk in our study may influence risk by affecting POU5F1B expression, although we did not have gene expression data in our study to directly address this question.
We also identified two intergenic CpG sites at 8q24 that were weakly to moderately correlated with the POU5F1B CpG site hits and were nominally significantly associated with overall prostate cancer (Chr8:128012411 and Chr8:128444762). Chr8:128012411 is part of an AluY SINE and Chr8:128444762 is part of an ERV1 LTR. As these sites are not located in or near a gene, their potential functional roles are less clear. However, interestingly, we observed DNaseI HS peaks, histone methylation marks (specifically H3K4me2, which is often found near promoters) and a TF binding site (specifically DAXX, a transcription repressor) within 2 kb of the site at Chr8:128012411 in various prostate cell lines, characteristics that are common to transcriptional regulatory regions. It is possible that these intergenic CpG sites or strongly correlated CpG sites could influence prostate cancer risk by affecting the expression of various gene targets of the 8q24 locus (Du et al, 2015), but we were unable to directly assess this in our study.
Our finding that DNA methylation levels in the MYC oncogene, specifically for two CpG sites in the exon 3 region, were associated with the risk of aggressive prostate cancer is also intriguing. Although increased DNA methylation in promoter regions is generally thought to correspond to a decrease in gene expression, there is evidence that increased methylation in gene body regions may be associated with an increase in gene expression (Moen et al, 2015), lending some plausibility for the direction of our findings, with increased DNA methylation in MYC associated with an increase in prostate cancer risk. MYC is a transcriptional activator involved in a regulatory network that affects cell growth, apoptosis and differentiation and has been shown to be commonly overexpressed in prostate tumour tissue (Koh et al, 2010). The active expression of MYC in prostate tumour tissue and location of these 2 CpG sites in a coding region of MYC may help explain the overlap of these CpG sites with many active histone markers, TFBS and DNaseI HS regions in prostate cell lines. MYC has long been thought to have a role in prostate cancer progression, although there is evidence for a potential role in tumour initiation as well (Koh et al, 2010). Although whole 8q gain has been observed throughout the spectrum of prostate carcinogenesis, from precursor lesions (i.e., prostatic intraepithelial neoplasia) to metastatic disease (Jenkins et al, 1997), the specific amplification of a narrower region at 8q24 including MYC has been largely reported in metastatic or recurrent prostate cancer (Visakorpi et al, 1995; Jenkins et al, 1997; Sun et al, 2007).
Although our study findings are intriguing, we recognise that chance could have contributed to these results. We did not have available data for a replication set and when we adjusted for multiple comparisons, none of the findings remained statistically significant at the 0.05 level. However, the associations that we observed between established prostate cancer susceptibility SNPs and prostate cancer in our study were also not highly significant and would not have withstood correction for multiple testing, even though they are known to be associated with prostate cancer (Amundadottir et al, 2006; Gudmundsson et al, 2007; Haiman et al, 2007; Yeager et al, 2007; Eeles et al, 2008; Thomas et al, 2008; Al Olama et al, 2009; Schumacher et al, 2011). Thus, we expect that we may have been underpowered to evaluate CpG site associations with prostate cancer, warranting follow-up studies of our noteworthy findings with larger sample sizes.
As our study population was limited to Caucasians, our results may not be generalisable to other populations; however, by restricting our population to Caucasians, we were able to limit the potential for population stratification. We were also limited in our ability to evaluate DNA methylation markers associated with the risk of aggressive prostate cancer as only about 25% of the cases in our study were aggressive; however, our study does suggest that CpG sites in MYC may be worthy of further study for aggressive disease. In addition, although we did not sequence the entire 8q24 region and therefore did not capture all CpG sites, this is the first prospective evaluation of CpG sites in this region in peripheral blood and prostate cancer risk. A further limitation is that we did not have available data in our study for DNA methylation or gene expression in prostate tissue, which is an important question as DNA methylation profiles are thought to be tissue specific (Rakyan et al, 2011). However, there are growing examples in the literature of an association between DNA methylation markers in blood and a variety of solid tumors, including, for example, GSTP1 and prostate cancer (Laird, 2003), suggesting this may be a fruitful area for continued study. If our results are replicated by other studies, then future studies will be needed to investigate the relationship between DNA methylation levels at 8q24 in blood and prostate tumor tissue, and to assess the potential functional impact of variation in DNA methylation levels at these sites (e.g., by incorporating gene expression data).
One of the strengths of our study was the integration of genetic and epigenetic data. However, as some of the established prostate cancer susceptibility SNPs from GWAS did not achieve significance in our present study, we were limited in our ability to assess whether 8q24 DNA methylation may mediate the effects of these particular SNPs on prostate cancer. We anticipate that a larger sample size would be needed to more thoroughly evaluate the possible role of DNA methylation as a mediator of SNP-associated effects at the 8q24 locus.
Another strength of our study was the use of pre-diagnostic blood samples, which allowed us to prospectively study the relationship between DNA methylation levels at 8q24 and the risk of prostate cancer and to reduce potential bias from reverse causality. Additionally, our use of pyrosequencing, which is considered a quantitative, reproducible method that is highly sensitive to detect differences in DNA methylation between individuals, was a further strength.
In summary, our findings in a large prospective study provide some evidence that pre-diagnostic peripheral blood DNA methylation levels at specific CpG sites at 8q24 may be associated with the subsequent risk of prostate cancer and, in particular, that DNA methylation levels at CpG sites in MYC may be associated specifically with the risk of aggressive prostate cancer. Further studies are needed to confirm these findings, but if replicated, they may point to future directions in efforts to identify biomarkers of aggressive prostate cancer.
Change history
23 May 2017
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Abecasis GR, Auton A, Brooks LD, DePristo MA, Durbin RM, Handsaker RE, Kang HM, Marth GT, McVean GA (2012) An integrated map of genetic variation from 1092 human genomes. Nature 491: 56–65.
Ahmadiyeh N, Pomerantz MM, Grisanzio C, Herman P, Jia L, Almendro V, He HH, Brown M, Liu XS, Davis M, Caswell JL, Beckwith CA, Hills A, Macconaill L, Coetzee GA, Regan MM, Freedman M (2010) 8q24 prostate, breast, and colon cancer risk loci show tissue-specific long-range interaction with MYC. Proc Natl Acad Sci USA 107: 9742–9746.
Al Olama AA, Kote-Jarai Z, Giles GG, Guy M, Morrison J, Severi G, Leongamornlert DA, Tymrakiewicz M, Jhavar S, Saunders E, Hopper JL, Southey MC, Muir KR, English DR, Dearnaley DP, Ardern-Jones AT, Hall AL, O'Brien LT, Wilkinson RA, Sawyer E, Lophatananon A, Horwich A, Huddart RA, Khoo VS, Parker CC, Woodhouse CJ, Thompson A, Christmas T, Ogden C, Cooper C, Donovan JL, Hamdy FC, Neal DE, Eeles RA, Easton DF (2009) Multiple loci on 8q24 associated with prostate cancer susceptibility. Nat Genet 41: 1058–1060.
Almen MS, Jacobsson JA, Moschonis G, Benedict C, Chrousos GP, Fredriksson R, Schioth HB (2012) Genome wide analysis reveals association of a FTO gene variant with epigenetic changes. Genomics 99: 132–137.
Amundadottir LT, Sulem P, Gudmundsson J, Helgason A, Baker A, Agnarsson BA, Sigurdsson A, Benediktsdottir KR, Cazier JB, Sainz J, Jakobsdottir M, Kostic J, Magnusdottir DN, Ghosh S, Agnarsson K, Birgisdottir B, Le Roux L, Olafsdottir A, Blondal T, Andresdottir M, Gretarsdottir OS, Bergthorsson JT, Gudbjartsson D, Gylfason A, Thorleifsson G, Manolescu A, Kristjansson K, Geirsson G, Isaksson H, Douglas J, Johansson JE, Balter K, Wiklund F, Montie JE, Yu X, Suarez BK, Ober C, Cooney KA, Gronberg H, Catalona WJ, Einarsson GV, Barkardottir RB, Gulcher JR, Kong A, Thorsteinsdottir U, Stefansson K (2006) A common variant associated with prostate cancer in European and African populations. Nat Genet 38: 652–658.
Barry KH, Moore LE, Liao LM, Huang WY, Andreotti G, Poulin M, Berndt SI (2015) Prospective study of DNA methylation at LINE-1 and Alu in peripheral blood and the risk of prostate cancer. Prostate 75: 1718–1725.
Barry KH, Moore LE, Sampson J, Yan L, Meyer A, Oler AJ, Chung CC, Wang Z, Yeager M, Amundadottir L, Berndt SI (2014) DNA methylation levels at chromosome 8q24 in peripheral blood are associated with 8q24 cancer susceptibility loci. Cancer Prev Res (Phila) 7: 1282–1292.
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Series B Stat Methodol 57: 289–300.
Berndt SI, Wang Z, Yeager M, Alavanja MC, Albanes D, Amundadottir L, Andriole G, Beane Freeman L, Campa D, Cancel-Tassin G, Canzian F, Cornu JN, Cussenot O, Diver WR, Gapstur SM, Gronberg H, Haiman CA, Henderson B, Hutchinson A, Hunter DJ, Key TJ, Kolb S, Koutros S, Kraft P, Le Marchand L, Lindstrom S, Machiela MJ, Ostrander EA, Riboli E, Schumacher F, Siddiq A, Stanford JL, Stevens VL, Travis RC, Tsilidis KK, Virtamo J, Weinstein S, Wilkund F, Xu J, Lilly Zheng S, Yu K, Wheeler W, Zhang H, Sampson J, Black A, Jacobs K, Hoover RN, Tucker M, Chanock SJ (2015) Two susceptibility loci identified for prostate cancer aggressiveness. Nat Commun 6: 6889.
Breyer JP, Dorset DC, Clark TA, Bradley KM, Wahlfors TA, McReynolds KM, Maynard WH, Chang SS, Cookson MS, Smith JA, Schleutker J, Dupont WD, Smith JR (2014) An expressed retrogene of the master embryonic stem cell gene POU5F1 is associated with prostate cancer susceptibility. Am J Hum Genet 94: 395–404.
Chen Y, Chi P, Rockowitz S, Iaquinta PJ, Shamu T, Shukla S, Gao D, Sirota I, Carver BS, Wongvipat J, Scher HI, Zheng D, Sawyers CL (2013) ETS factors reprogram the androgen receptor cistrome and prime prostate tumorigenesis in response to PTEN loss. Nat Med 19: 1023–1029.
Chung S, Nakagawa H, Uemura M, Piao L, Ashikawa K, Hosono N, Takata R, Akamatsu S, Kawaguchi T, Morizono T, Tsunoda T, Daigo Y, Matsuda K, Kamatani N, Nakamura Y, Kubo M (2011) Association of a novel long non-coding RNA in 8q24 with prostate cancer susceptibility. Cancer Sci 102: 245–252.
Du M, Yuan T, Schilter KF, Dittmar RL, Mackinnon A, Huang X, Tschannen M, Worthey E, Jacob H, Xia S, Gao J, Tillmans L, Lu Y, Liu P, Thibodeau SN, Wang L (2015) Prostate cancer risk locus at 8q24 as a regulatory hub by physical interactions with multiple genomic loci across the genome. Hum Mol Genet 24: 154–166.
Eeles RA, Kote-Jarai Z, Giles GG, Olama AA, Guy M, Jugurnauth SK, Mulholland S, Leongamornlert DA, Edwards SM, Morrison J, Field HI, Southey MC, Severi G, Donovan JL, Hamdy FC, Dearnaley DP, Muir KR, Smith C, Bagnato M, Ardern-Jones AT, Hall AL, O'Brien LT, Gehr-Swain BN, Wilkinson RA, Cox A, Lewis S, Brown PM, Jhavar SG, Tymrakiewicz M, Lophatananon A, Bryant SL, Horwich A, Huddart RA, Khoo VS, Parker CC, Woodhouse CJ, Thompson A, Christmas T, Ogden C, Fisher C, Jamieson C, Cooper CS, English DR, Hopper JL, Neal DE, Easton DF (2008) Multiple newly identified loci associated with prostate cancer susceptibility. Nat Genet 40: 316–321.
Florath I, Butterbach K, Muller H, Bewerunge-Hudler M, Brenner H (2014) Cross-sectional and longitudinal changes in DNA methylation with age: an epigenome-wide analysis revealing over 60 novel age-associated CpG sites. Hum Mol Genet 23: 1186–1201.
Gudmundsson J, Sulem P, Gudbjartsson DF, Masson G, Agnarsson BA, Benediktsdottir KR, Sigurdsson A, Magnusson OT, Gudjonsson SA, Magnusdottir DN, Johannsdottir H, Helgadottir HT, Stacey SN, Jonasdottir A, Olafsdottir SB, Thorleifsson G, Jonasson JG, Tryggvadottir L, Navarrete S, Fuertes F, Helfand BT, Hu Q, Csiki IE, Mates IN, Jinga V, Aben KK, van Oort IM, Vermeulen SH, Donovan JL, Hamdy FC, Ng CF, Chiu PK, Lau KM, Ng MC, Gulcher JR, Kong A, Catalona WJ, Mayordomo JI, Einarsson GV, Barkardottir RB, Jonsson E, Mates D, Neal DE, Kiemeney LA, Thorsteinsdottir U, Rafnar T, Stefansson K (2012) A study based on whole-genome sequencing yields a rare variant at 8q24 associated with prostate cancer. Nat Genet 44: 1326–1329.
Gudmundsson J, Sulem P, Manolescu A, Amundadottir LT, Gudbjartsson D, Helgason A, Rafnar T, Bergthorsson JT, Agnarsson BA, Baker A, Sigurdsson A, Benediktsdottir KR, Jakobsdottir M, Xu J, Blondal T, Kostic J, Sun J, Ghosh S, Stacey SN, Mouy M, Saemundsdottir J, Backman VM, Kristjansson K, Tres A, Partin AW, Albers-Akkers MT, Godino-Ivan Marcos J, Walsh PC, Swinkels DW, Navarrete S, Isaacs SD, Aben KK, Graif T, Cashy J, Ruiz-Echarri M, Wiley KE, Suarez BK, Witjes JA, Frigge M, Ober C, Jonsson E, Einarsson GV, Mayordomo JI, Kiemeney LA, Isaacs WB, Catalona WJ, Barkardottir RB, Gulcher JR, Thorsteinsdottir U, Kong A, Stefansson K (2007) Genome-wide association study identifies a second prostate cancer susceptibility variant at 8q24. Nat Genet 39: 631–637.
Haiman CA, Patterson N, Freedman ML, Myers SR, Pike MC, Waliszewska A, Neubauer J, Tandon A, Schirmer C, McDonald GJ, Greenway SC, Stram DO, Le Marchand L, Kolonel LN, Frasco M, Wong D, Pooler LC, Ardlie K, Oakley-Girvan I, Whittemore AS, Cooney KA, John EM, Ingles SA, Altshuler D, Henderson BE, Reich D (2007) Multiple regions within 8q24 independently affect risk for prostate cancer. Nat Genet 39: 638–644.
Hayes RB, Sigurdson A, Moore L, Peters U, Huang WY, Pinsky P, Reding D, Gelmann EP, Rothman N, Pfeiffer RM, Hoover RN, Berg CD (2005) Methods for etiologic and early marker investigations in the PLCO trial. Mutat Res 592: 147–154.
Hazelett DJ, Rhie SK, Gaddis M, Yan C, Lakeland DL, Coetzee SG, Ellipse/GAME-ON Consortium, Practical Consortium, Henderson BE, Noushmehr H, Cozen W, Kote-Jarai Z, Eeles RA, Easton DF, Haiman CA, Lu W, Farnham PJ, Coetzee GA (2014) Comprehensive functional annotation of 77 prostate cancer risk loci. PLoS Genet 10: e1004102.
He HH, Meyer CA, Shin H, Bailey ST, Wei G, Wang Q, Zhang Y, Xu K, Ni M, Lupien M, Mieczkowski P, Lieb JD, Zhao K, Brown M, Liu XS (2010) Nucleosome dynamics define transcriptional enhancers. Nat Genet 42: 343–347.
Howie BN, Donnelly P, Marchini J (2009) A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet 5: e1000529.
Jenkins RB, Qian J, Lieber MM, Bostwick DG (1997) Detection of c-myc oncogene amplification and chromosomal anomalies in metastatic prostatic carcinoma by fluorescence in situ hybridization. Cancer Res 57: 524–531.
Jia L, Landan G, Pomerantz M, Jaschek R, Herman P, Reich D, Yan C, Khalid O, Kantoff P, Oh W, Manak JR, Berman BP, Henderson BE, Frenkel B, Haiman CA, Freedman M, Tanay A, Coetzee GA (2009) Functional enhancers at the gene-poor 8q24 cancer-linked locus. PLoS Genet 5: e1000597.
Jin HJ, Zhao JC, Wu L, Kim J, Yu J (2014) Cooperativity and equilibrium with FOXA1 define the androgen receptor transcriptional program. Nat Commun 5: 3972.
Kastler S, Honold L, Luedeke M, Kuefer R, Moller P, Hoegel J, Vogel W, Maier C, Assum G (2010) POU5F1P1, a putative cancer susceptibility gene, is overexpressed in prostatic carcinoma. Prostate 70: 666–667.
Koh CM, Bieberich CJ, Dang CV, Nelson WG, Yegnasubramanian S, De Marzo AM (2010) MYC and prostate cancer. Genes Cancer 1: 617–628.
Koutros S, Beane Freeman LE, Berndt SI, Andreotti G, Lubin JH, Sandler DP, Hoppin JA, Yu K, Li Q, Burdette LA, Yuenger J, Yeager M, Alavanja MC (2010) Pesticide use modifies the association between genetic variants on chromosome 8q24 and prostate cancer. Cancer Res 70: 9224–9233.
Kulis M, Esteller M (2010) DNA methylation and cancer. Adv Genet 70: 27–56.
Laird PW (2003) The power and the promise of DNA methylation markers. Nat Rev Cancer 3: 253–266.
Liu Y, Aryee MJ, Padyukov L, Fallin MD, Hesselberg E, Runarsson A, Reinius L, Acevedo N, Taub M, Ronninger M, Shchetynsky K, Scheynius A, Kere J, Alfredsson L, Klareskog L, Ekstrom TJ, Feinberg AP (2013) Epigenome-wide association data implicate DNA methylation as an intermediary of genetic risk in rheumatoid arthritis. Nat Biotechnol 31: 142–147.
Maurano MT, Humbert R, Rynes E, Thurman RE, Haugen E, Wang H, Reynolds AP, Sandstrom R, Qu H, Brody J, Shafer A, Neri F, Lee K, Kutyavin T, Stehling-Sun S, Johnson AK, Canfield TK, Giste E, Diegel M, Bates D, Hansen RS, Neph S, Sabo PJ, Heimfeld S, Raubitschek A, Ziegler S, Cotsapas C, Sotoodehnia N, Glass I, Sunyaev SR, Kaul R, Stamatoyannopoulos JA (2012) Systematic localization of common disease-associated variation in regulatory DNA. Science 337: 1190–1195.
Meyer KB, Maia AT, O'Reilly M, Ghoussaini M, Prathalingam R, Porter-Gill P, Ambs S, Prokunina-Olsson L, Carroll J, Ponder BA (2011) A functional variant at a prostate cancer predisposition locus at 8q24 is associated with PVT1 expression. PLoS Genet 7: e1002165.
Moen EL, Mariani CJ, Zullow H, Jeff-Eke M, Litwin E, Nikitas JN, Godley LA (2015) New themes in the biological functions of 5-methylcytosine and 5-hydroxymethylcytosine. Immunol Rev 263: 36–49.
Polak P, Lawrence MS, Haugen E, Stoletzki N, Stojanov P, Thurman RE, Garraway LA, Mirkin S, Getz G, Stamatoyannopoulos JA, Sunyaev SR (2014) Reduced local mutation density in regulatory DNA of cancer genomes is linked to DNA repair. Nat Biotechnol 32: 71–75.
Pomerantz MM, Beckwith CA, Regan MM, Wyman SK, Petrovics G, Chen Y, Hawksworth DJ, Schumacher FR, Mucci L, Penney KL, Stampfer MJ, Chan JA, Ardlie KG, Fritz BR, Parkin RK, Lin DW, Dyke M, Herman P, Lee S, Oh WK, Kantoff PW, Tewari M, McLeod DG, Srivastava S, Freedman ML (2009) Evaluation of the 8q24 prostate cancer risk locus and MYC expression. Cancer Res 69: 5568–5574.
Prokunina-Olsson L, Hall JL (2009) No effect of cancer-associated SNP rs6983267 in the 8q24 region on co-expression of MYC and TCF7L2 in normal colon tissue. Mol Cancer 8: 96.
Prorok PC, Andriole GL, Bresalier RS, Buys SS, Chia D, Crawford ED, Fogel R, Gelmann EP, Gilbert F, Hasson MA, Hayes RB, Johnson CC, Mandel JS, Oberman A, O'Brien B, Oken MM, Rafla S, Reding D, Rutt W, Weissfeld JL, Yokochi L, Gohagan JK (2000) Design of the Prostate, Lung, Colorectal and Ovarian (PLCO) Cancer Screening Trial. Control Clin Trials 21: 273S–309S.
Puto LA, Benner C, Hunter T (2015) The DAXX co-repressor is directly recruited to active regulatory elements genome-wide to regulate autophagy programs in a model of human prostate cancer. Oncoscience 2: 362–372.
Quinlan AR, Hall IM (2010) BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26: 841–842.
Rakyan VK, Down TA, Balding DJ, Beck S (2011) Epigenome-wide association studies for common human diseases. Nat Rev Genet 12: 529–541.
Schumacher FR, Berndt SI, Siddiq A, Jacobs KB, Wang Z, Lindstrom S, Stevens VL, Chen C, Mondul AM, Travis RC, Stram DO, Eeles RA, Easton DF, Giles G, Hopper JL, Neal DE, Hamdy FC, Donovan JL, Muir K, Al Olama AA, Kote-Jarai Z, Guy M, Severi G, Gronberg H, Isaacs WB, Karlsson R, Wiklund F, Xu J, Allen NE, Andriole GL, Barricarte A, Boeing H, Bueno-de-Mesquita HB, Crawford ED, Diver WR, Gonzalez CA, Gaziano JM, Giovannucci EL, Johansson M, Le Marchand L, Ma J, Sieri S, Stattin P, Stampfer MJ, Tjonneland A, Vineis P, Virtamo J, Vogel U, Weinstein SJ, Yeager M, Thun MJ, Kolonel LN, Henderson BE, Albanes D, Hayes RB, Feigelson HS, Riboli E, Hunter DJ, Chanock SJ, Haiman CA, Kraft P (2011) Genome-wide association study identifies new prostate cancer susceptibility loci. Hum Mol Genet 20: 3867–3875.
Sotelo J, Esposito D, Duhagon MA, Banfield K, Mehalko J, Liao H, Stephens RM, Harris TJ, Munroe DJ, Wu X (2010) Long-range enhancers on 8q24 regulate c-Myc. Proc Natl Acad Sci USA 107: 3001–3005.
Sun J, Liu W, Adams TS, Sun J, Li X, Turner AR, Chang B, Kim JW, Zheng SL, Isaacs WB, Xu J (2007) DNA copy number alterations in prostate cancers: a combined analysis of published CGH studies. Prostate 67: 692–700.
Thomas G, Jacobs KB, Yeager M, Kraft P, Wacholder S, Orr N, Yu K, Chatterjee N, Welch R, Hutchinson A, Crenshaw A, Cancel-Tassin G, Staats BJ, Wang Z, Gonzalez-Bosquet J, Fang J, Deng X, Berndt SI, Calle EE, Feigelson HS, Thun MJ, Rodriguez C, Albanes D, Virtamo J, Weinstein S, Schumacher FR, Giovannucci E, Willett WC, Cussenot O, Valeri A, Andriole GL, Crawford ED, Tucker M, Gerhard DS, Fraumeni JF Jr, Hoover R, Hayes RB, Hunter DJ, Chanock SJ (2008) Multiple loci identified in a genome-wide association study of prostate cancer. Nat Genet 40: 310–315.
Thurman RE, Rynes E, Humbert R, Vierstra J, Maurano MT, Haugen E, Sheffield NC, Stergachis AB, Wang H, Vernot B, Garg K, John S, Sandstrom R, Bates D, Boatman L, Canfield TK, Diegel M, Dunn D, Ebersol AK, Frum T, Giste E, Johnson AK, Johnson EM, Kutyavin T, Lajoie B, Lee BK, Lee K, London D, Lotakis D, Neph S, Neri F, Nguyen ED, Qu H, Reynolds AP, Roach V, Safi A, Sanchez ME, Sanyal A, Shafer A, Simon JM, Song L, Vong S, Weaver M, Yan Y, Zhang Z, Zhang Z, Lenhard B, Tewari M, Dorschner MO, Hansen RS, Navas PA, Stamatoyannopoulos G, Iyer VR, Lieb JD, Sunyaev SR, Akey JM, Sabo PJ, Kaul R, Furey TS, Dekker J, Crawford GE, Stamatoyannopoulos JA (2012) The accessible chromatin landscape of the human genome. Nature 489: 75–82.
Toropainen S, Malinen M, Kaikkonen S, Rytinki M, Jaaskelainen T, Sahu B, Janne OA, Palvimo JJ (2015) SUMO ligase PIAS1 functions as a target gene selective androgen receptor coregulator on prostate cancer cell chromatin. Nucleic Acids Res 43: 848–861.
Visakorpi T, Kallioniemi AH, Syvanen AC, Hyytinen ER, Karhu R, Tammela T, Isola JJ, Kallioniemi OP (1995) Genetic changes in primary and recurrent prostate cancer by comparative genomic hybridization. Cancer Res 55: 342–347.
Wright JB, Brown SJ, Cole MD (2010) Upregulation of c-MYC in cis through a large chromatin loop linked to a cancer risk-associated single-nucleotide polymorphism in colorectal cancer cells. Mol Cell Biol 30: 1411–1420.
Yeager M, Orr N, Hayes RB, Jacobs KB, Kraft P, Wacholder S, Minichiello MJ, Fearnhead P, Yu K, Chatterjee N, Wang Z, Welch R, Staats BJ, Calle EE, Feigelson HS, Thun MJ, Rodriguez C, Albanes D, Virtamo J, Weinstein S, Schumacher FR, Giovannucci E, Willett WC, Cancel-Tassin G, Cussenot O, Valeri A, Andriole GL, Gelmann EP, Tucker M, Gerhard DS, Fraumeni JF Jr, Hoover R, Hunter DJ, Chanock SJ, Thomas G (2007) Genome-wide association study of prostate cancer identifies a second risk locus at 8q24. Nat Genet 39: 645–649.
Zlotta AR, Egawa S, Pushkar D, Govorov A, Kimura T, Kido M, Takahashi H, Kuk C, Kovylina M, Aldaoud N, Fleshner N, Finelli A, Klotz L, Sykes J, Lockwood G, van der Kwast TH (2013) Prevalence of prostate cancer on autopsy: cross-sectional study on unscreened Caucasian and Asian men. JNCI 105: 1050–1058.
Acknowledgements
We thank Drs Christine Berg and Philip Prorok, Division of Cancer Prevention at the National Cancer Institute, the screening centre investigators and staff of the PLCO Cancer Screening Trial, Mr Thomas Riley, Mr Craig Williams and Mr Michael Furr at Information Management Services, Inc., and Ms Barbara O’Brien and staff at Westat, Inc. for their contributions to the PLCO Cancer Screening Trial. We also thank the PLCO study participants for their contributions to making this study possible. This study was supported by the Intramural Research Programs of the National Cancer Institute and the National Institute of Allergy and Infectious Diseases, National Institutes of Health, USA.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
L Yan, A Meyer and M Reddy are employed by EpigenDx, Inc., and L Yan is the major stockholder of EpigenDx. All other authors declare no conflicts of interest.
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License.
Supplementary Information accompanies this paper on British Journal of Cancer website
Supplementary information
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Barry, K., Moore, L., Sampson, J. et al. Prospective study of DNA methylation at chromosome 8q24 in peripheral blood and prostate cancer risk. Br J Cancer 116, 1470–1479 (2017). https://doi.org/10.1038/bjc.2017.104
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2017.104
Keywords
This article is cited by
-
Exploring the effects of genetic variation on gene regulation in cancer in the context of 3D genome structure
BMC Genomic Data (2022)
-
BOP1 Silencing Suppresses Gastric Cancer Proliferation through p53 Modulation
Current Medical Science (2021)