Abstract
Background:
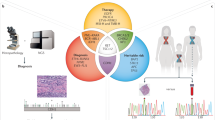
Screening of patients for cancer-driving mutations is now used for cancer prognosis, remission scoring and treatment selection. Although recently emerged targeted next-generation sequencing-based approaches offer promising diagnostic capabilities, there are still limitations. There is a pressing clinical need for a well-validated, rapid, cost-effective mutation profiling system in patient specimens. Given their speed and cost-effectiveness, quantitative PCR mutation detection techniques are well suited for the clinical environment. The qBiomarker mutation PCR array has high sensitivity and shorter turnaround times compared with other methods. However, a direct comparison with existing viable alternatives are required to assess its true potential and limitations.
Methods:
In this study, we evaluated a panel of 117 patient-derived tumour xenografts by the qBiomarker array and compared with other methods for mutation detection, including Ion AmpliSeq sequencing, whole-exome sequencing and droplet digital PCR.
Results:
Our broad analysis demonstrates that the qBiomarker’s performance is on par with that of other labour-intensive and expensive methods of cancer mutation detection of frequently altered cancer-associated genes, and provides a foundation for supporting its consideration as an option for molecular diagnostics.
Conclusions:
This large-scale direct comparison and validation of currently available mutation detection approaches is extremely relevant for the current scenario of precision medicine and will lead to informed choice of screening methodologies, especially in lower budget conditions or time frame limitations.
Similar content being viewed by others
Main
Tumorigenesis is known to progress through a series of genetic alterations. Despite the molecular heterogeneity of the disease, a subset of well-established cancer driver genes are frequently mutated across various solid tumour types at mutation hot spot regions. These genetic alterations are responsible for the initiation and maintenance of the malignancy (Tamborero et al, 2013), and therefore, detection of cancer-driving mutations in clinical samples offers opportunity for improved risk assessment, early cancer detection, therapeutic intervention and tumour surveillance.
Remarkable advances in next-generation sequencing (NGS) technologies have allowed rapid generation of high-quality genetic, transcriptomic and epigenetic data, and have opened up numerous opportunities for translation into the clinic. Due to decreases in sequencing cost and concomitant development of the infrastructure needed to effectively apply individual genomic data in clinical care, whole-genome sequencing (WGS) is likely to become more routinely available over the next few years and may ultimately displace other mutation detection tests. However, until that moment arrives, and for clinical settings that still will not have access to high-end complex technologies, there is an unmet clinical need for a rapid, accurate and affordable assay for cancer-driving mutation profiling.
Studies of cancer-driving genes have flourished over the last decade and a variety of platforms have emerged for surveying the hot spot regions of numerous tumour-promoting and tumour-suppressor genes. Targeted NGS assays, such as CancerSelect-R by Personal Genome Diagnostics (PGDx, Baltimore, MD, USA), seem to be the best approach for profiling tumour-specific mutations with a high sensitivity and specificity. Nevertheless, its clinical application is limited due to the high cost and long turnaround time (3 weeks) required for the data analysis. Moreover, germline variation, alignment artefacts and inherent massively parallel sequencing errors may considerably affect the assay’ sensitivity to accurately detect rare and low-prevalence bona fide somatic events. Ultra-sensitive droplet digital PCR (ddPCR) overcomes these complications, and enables rapid and highly sensitive quantitative genotyping of mutated genes. However, this new technology is not readily available in every research or diagnostic centres, where high cost of establishing ddPCR facilities is still an impediment for the widespread use of this technique. Furthermore, the limited throughput approach further delays its clinical adoption for large-scale mutations screening as a routine personal genomics practice. At present, quantitative PCR (qPCR) instruments are more common in laboratory and diagnostic settings and researchers are more comfortable with handling this platform. qBiomarker, a real-time PCR method for detection of cancer-associated mutations and copy number variations, has been successfully used in both basic research (Sangodkar et al, 2012; Luo et al, 2013) and with diagnostic potential (Jimenez et al, 2013). By combining allele-specific amplification, achieved by Taq polymerase’s discrimination between a match and a mismatch at the 3′ end of the PCR primer (amplification refractory mutation system technology) and hydrolysis probe detection, qBiomarker can detect somatic mutations at 1% minimum mutant allele fraction, making it one of the most sensitive qPCR methods for the detection of DNA mutations. Optimised for robust performance, qBiomarker array is presented as a plate with up to 382 targets analysed at the same reaction. Although this method is an attractive candidate for clinical applications due to its high sensitivity, rapid protocol (<30 min hands-on time), and relatively low cost, a rigorous evaluation and comparison with viable alternatives are required to assess its true potential and limitations.
Patient-derived xenograft (PDX) models are established by directly implanting tumour tissue samples into an immunodeficient mouse and are increasingly recognised as representative in vivo clinical models, as they maintain global gene expression patterns, DNA copy number alterations, mutational status, metastatic potential, clinical predictability and tumour architecture of the parental primary tumours (Rubio-Viqueira et al, 2006; Garber, 2007; Hidalgo et al, 2011; Morelli et al, 2012; Stebbing et al, 2014). Therefore, personalised PDXs can be successfully used as model platforms for cancer-driving mutation screening and improving decision-making in tumour treatment (Gao et al, 2015).
In the current study, qBiomarker somatic mutation PCR array (Qiagen, Germantown, MD, USA) was evaluated in a panel of 117 PDX models of various tumour origins, and its efficacy was compared with that of other methods for mutation detection, such as targeted Ion AmpliSeq sequencing (Thermo Fisher Scientific, Waltham, MA, USA), Whole Exome Sequencing (WES, Illumina, San Diego, CA, USA) and ultra-sensitive ddPCR (Bio-Rad, Hercules, CA, USA). Our comprehensive comparative analysis demonstrates that qBiomarker array accurately detects cancer-associated somatic aberrations and has high concordance with other routinely used, but more complex mutation detecting tests. Moreover, the mutational landscape discovered by the qBiomarker platform in PDX models closely reflects the spectrum of genetic alterations reported in primary tumours, further supporting the credibility of the qBiomarker platform. Taken together, our work demonstrates that qPCR-based qBiomarker, WES, AmpliSeq and ddPCR are reliable methods for cancer-driving mutations assessment and validates qBiomarker array as a simple, rapid and cost-effective mutation detection method that enables potential translational and clinical implementations.
Materials and methods
Generations of PDX models
All patients that agreed to have PDX models generated from their tumours signed informed consent document following federal regulatory requirements. All animals were maintained according to approved Institutional Animal Care and Use Committee protocols. Patient-derived xenograft has been generated as previously described (Bertotti et al, 2011; DeRose et al, 2011; Hidalgo et al, 2011; Villarroel et al, 2011; Morelli et al, 2012; Stebbing et al, 2014). A detailed approach for PDX model generation from a tumour biopsy is available in the Supplementary Materials and Methods.
DNA extraction
Genomic DNA was isolated from fresh-frozen samples by the QIAamp DNA Kit (Qiagen) and quantified with the Nanodrop spectrophotometer (Thermo Fisher Scientific).
qBiomarker somatic mutation PCR array
A Qiagen customised mutation array containing 353 mutations and 29 copy number assays was designed based on comprehensive somatic mutation databases (e.g., COSMIC, cancer.sanger.ac.uk/cosmic) and peer-reviewed scientific literature data on significant known mutations in cancer (Figure 1B). The selected mutations are biologically or therapeutically relevant in different cancer types and for this specific array we have prioritised relevant mutations reported in lung and colorectal tumours. The exact qBiomarker array method and analysis are detailed in the Supplementary Materials and Methods.
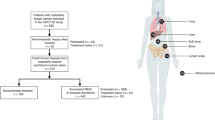
Mutation screening by qBiomarker array and validation with AmpliSeq next-generation sequencing platform in patient-derived tumour xenografts (PDX).(A) Tumour site distribution of 117 PDX models selected for mutation screening. (B) Number of somatic mutations (non-synonymous and indels) covered by the customised qBiomarker array. (C) Number of mutations identified per gene using the qBiomarker array (cyan bars). Mutation rates comparison for the most frequently mutated genes detected by qBiomarker (green inserts) in colorectal cancers, pancreatic carcinomas, non-small cell lung cancers and melanoma with mutation frequencies reported for these genes (including all reported mutations, not only the ones we analysed) in primary tumours by COSMIC (red inserts). (D) List of genes and number of somatic mutations concurrently covered by qBiomarker and AmpliSeq mutation detection platforms. (E) Schematic representation of all mutations detected by qBiomarker and AmpliSeq approaches in 59 PDX models. Green and blue dots represent mutations detected by either qBiomarker or AmpliSeq assay, respectively, whereas red dots indicate mutations concurrently detected by both methods. Cases with no mutation detected are not shown. (F) Venn diagram summarises mutations concurrently detected by both methods and each one of the tested techniques alone.
Ion torrent ampliseq sequencing
From the list of hot spot regions of 50 oncogenes and tumour-suppressor genes included in the Ion Torrent NGS AmpliSeq Cancer Hot spot Panel v2 (Thermo Fisher Scientific), we focused on analysis of 297 non-synonymous genetic aberrations (253 missense/nonsense mutations and 44 indels) across 23 genes. All selected targets overlapped with the qBiomarker panel. Contemporary normal NA12878 cell line was used as a 'normal' tissue. The Ion Torrent AmpliSeq sequencing (Thermo Fisher Scientific) and data analysis are detailed in the Supplementary Materials and Methods.
WES
Genomic DNA from tumour and normal samples were fragmented and whole exome was captured in solution using the Sure Select human all exon kit (Agilent, Santa Clara, CA, USA) according to the manufacturer’s instructions. Paired-end sequencing, resulting in 125 bases from each end of the fragments, was performed using a HiSeq System (Illumina, San Diego, CA, USA). Candidate somatic mutations were identified by applying Genome Analysis Toolkit recommended preprocessing and alignment. Sequence reads were aligned against the human reference genome (GRC37/hg19) and mouse reference genome (GRCm38/mm10) using the Burrows-Wheeler Alignment tool (Li and Durbin, 2009) and reads that do not align to human reference have been removed. Somatic variants were identified using multiple callers: (muTect, Virmid, Strelka, LoFreq, SomaticIndelDet; Saunders et al, 2012; Wilm et al, 2012; Cibulskis et al, 2013; Kim et al, 2013) using a standard normal cell line (NA12878) as matched normal, and all calls were aggregated into a single vcf. Detailed sequencing and data analysis approaches are available in the Supplementary Materials and Methods.
Digital droplet PCR
All ddPCR assays used in this study were designed and optimised to work in the ddPCR QX200 system by Bio-Rad (Hercules, CA, USA). The detailed ddPCR protocol used in this study is available in the Supplementary Materials and Methods.
Statistical analysis
Sensitivity, specificity, negative predictive value and positive predictive value were calculated as the percentage on the basis of KRAS/BRAF mutations correctly detected by ddPCR in 104 patients.
Results
Mutational landscape detected by the qBiomarker platform in human PDX tumour models
We first used a customised qBiomarker array to profile somatic mutation status in 117 early-passaged human PDX models of various tumour origin (Supplementary Figure 1; 35 colorectal cancers, 31 pancreatic cancers, 29 lung cancer, 25 non-small cell lung cancers and 4 small cell lung cancers, 10 melanomas, 7 ovarian cancers and 5 breast cancers; Figure 1A; Supplementary Table 1). The panel of genetic changes selected for the customised assay consists of 353 (260 single-nucleotide substitutions and 63 indels) non-synonymous alterations in 29 well-characterised driver genes common to many solid tumour types (see Materials and Methods for selection criteria; Figure 1B; Supplementary Table 2A).
Our analysis identified 149 genetic alterations across 16 of 29 targeted genes in 86 (73%) of the tumours tested. Over 80% of all identified alterations were attributed to KRAS, TP53, PIK3CA, APC and BRAF (Figure 1C; Supplementary Table 3). These genes are frequently mutated in various solid malignancies and were therefore highly represented in our customised mutation detection assay (Supplementary Table 2A). Notably, mutation frequencies of mostly mutated genes (mutated in >5% of cases) observed in PDX models of different tumour types (Figure 1C, green inserts) highly resembled mutation frequencies reported for these genes in primary human tumours by COSMIC (Figure 1C, red inserts). For example, KRAS and TP53 mutations, the most frequent genetic alterations in colorectal cancer patients (Neumann et al, 2009; Figure 1C, red inserts), were also identified as such by qBiomarker assay in colorectal PDX models (Figure 1C, green inserts). Likewise, mutations in the APC gene, which frequently occur in colorectal carcinomas (Segditsas and Tomlinson, 2006; Anastas and Moon, 2013), were detected in nearly 20% of colorectal xenografts (Anastas and Moon, 2013). Consistent with previous reports, nearly 80% of the pancreatic models harboured mutationally activated KRAS (Bryant et al, 2014), whereas more than half of the melanoma tumours harboured activating BRAF mutations (Ascierto et al, 2012; Figure 1C; Supplementary Table 3). Of note, 7 (20%) of 35 colorectal tumours and 9 (29%) of 31 pancreatic models harboured concurrent KRAS activating (Smith et al, 2010) and TP53 inactivating (Kato et al, 2003) mutations (Supplementary Table 2A). A similar range of coexistent KRAS/TP53 mutant genotype has been recently reported in human colorectal (De Bruijn et al, 2011) and pancreatic (Yachida et al, 2012) tumours, further suggesting that the frequency of driver genes discovered by qBiomarker cancer mutations panel in PDX models is likely reflective of the true frequencies reported in human tumours.
Validation of qBiomarker-detected mutations with ion torrent AmpliSeq assay and WES
We next used Ion Torrent AmpliSeq technology to confirm mutations detected by qBiomarker in a panel of 59 selective PDX tumour models (Supplementary Figure 1). The customised targeted AmpliSeq cancer panel comprises 297 non-synonymous genetic aberrations (253 missense/nonsense mutations and 44 indels in hot spot regions of 25 oncogenes and tumour-suppressor genes), that were also included in the qBiomarker panel (Figure 1D; Supplementary Table 2B). Altogether, both approaches detected 95 genetic aberrations in 16 genes (Supplementary Table 4). Sixty-nine (73%) of these mutations were detected by both methods (Figure 1E), revealing that AmpliSeq and qBiomarker assays identify genetic alterations with high concordance. Whereas 18 and 9 genetic changes were exclusively identified by either the qBiomarker assay or AmpliSeq sequencing respectively (Figure 1F).
To further benchmark the qBiomarker array, we used WES to assess the presence of genetic aberrations detected by qBiomarker approach in 59 randomly selected samples of 117 models used in this study (Figure 2A; Supplementary Figure 1). WES data was analysed for 260 single-nucleotide substitutions targeted by the Biomarker panel (indels were not included). Strikingly, WES confirmed 65 of 71 (92%) mutations detected by the qBiomarker assay (Figure 2A; Supplementary Table 5A) and only 5 mutations positively identified by qBiomarker have been reported as wild-type by deep-exome sequencing (Figure 2B). Notably, there was greater agreement between WES and qBiomarker mutation calls (Figure 2B) than between AmpliSeq and qBiomarker (Figure 1F). To test whether variants that were false positively detected by qBiomarker assay result from probes’ cross-reactivity with actual somatic mutations located within their immediate vicinity, we have analysed our sequencing data for the presence of any genetic aberrations 15 nt up- or downstream from these false-positive calls. A frameshift deletion mutation was detected nearby the false-positive TP53 variant in one out of five PDX tumour models (CTG-0198). These findings suggest that some of the calls unique to qBiomarker may be false positives, resulting from cross-reactivity of probes with the true adjacent genetic aberration.
Validation of qBiomarker-detected mutations with whole-exome sequencing (WES).(A) Schematic representation of all mutations detected by qBiomarker and WES approaches in 59 PDX models. Red dots represent mutations detected by both methods, while green (only qBiomarker) and blue (only WES). Cases with no mutation detected are not shown. (B) Venn diagram summarising the number of mutations identified by both methods or by each technique. (C) Schematic representation of all mutations detected by qBiomarker, WES and AmpliSeq approaches in 43 PDX models. Blue dots represent mutations detected by one of the methods, whereas red crosses represent wild-type loci (no mutation detection). The mutations detected concurrently by all three methods are represented by the red squares. Cases with no mutation detected by any platform are not shown. (D) Venn diagram summarising the number of mutations detected by qBiomarker, WES and AmpliSeq alone, by any combination of two of the methods and the total number of genetic aberrations detected by all three approaches.
Forty-three of 117 PDX tumour models were concurrently analysed for mutation detection by the qBiomarker assay, targeted AmpliSeq sequencing and WES (Figure 2C; Supplementary Figure 1; Supplementary Table 5B). WES data were analysed for 253 missense/nonsense mutations included in the qBiomarker and AmpliSeq target panels (Supplementary Table 2B). Of the 59 mutations discovered across 43 tumours, 47 (80%) mutations have been concomitantly detected by all three platforms, (Figure 2D; Supplementary Table 5B), demonstrating the high level of concurrence between the sequencing- and qPCR-based screening approaches. The fractional abundance of mutations detected by AmpliSeq or WES was at least 20% and the prevalence of the mutant reads were mostly consistent between WES and AmpliSeq assays (Supplementary Table 5B). Nonetheless, there was slightly greater concordance between WES and qBiomarker than AmpliSeq mutation calls. Notably, the AmpliSeq-targeted screening approach appears to be more prone to false negatives, as five mutations discovered by qBiomarker and WES were reported as wildtype by AmpliSeq (Figures 2C and D).
Validation of selective KRAS and BRAF mutations with ultra-sensitive ddPCR
Although our data display high concordance between genetic aberrations calls from qPCR- and NGS-based screening assays (Figures 2C and D), 28% of tested mutations were not consistently detected across all approaches. As the detection of true point mutations requires a high degree of sensitivity, we next utilised ultra-sensitive ddPCR to further validate the accuracy of the qBiomarker assay by probing for three KRAS and two BRAF cancer-associated substitutions (Figure 3A). We first used serially diluted DNA isolated from a tumour model with known KRAS p.G12D mutation to evaluate assay sensitivity. The presence of mutant DNA was detected even at DNA input as low as 0.4 ng, whereas the fractional abundance remained unchanged across all serially diluted samples (Figure 3B). To ensure specific detection of mutations located within the adjacent nucleotides, we next assessed the cross-reactivity of ddPCR probes targeting KRAS p.G12D and p.G12V mutations (hence these probes differ by only one base). DNA with known KRAS p.G12D mutation was probed with either specific ddPCR assay or with a probe designed for detection of p.G12V substitution (Figure 3C). Similarly, DNA isolated from tumour carrying the p.G12V mutation was probed with either a specific or off-target ddPCR probe (Figure 3D). Although in both cases (Figures 3C and D), non-specific probes caused a minor inwards and upwards cluster shift (labelled with a star) compared with the negative droplets (suggesting that some of the non-specific probes bind to the nonperfect sequences and undergo cleavage), specific assays resulted in much greater cluster shifts (blue clusters), indicating that this low level of cross-reactivity will not affect the assay’s ability to specifically detect tumours positive for KRAS p.G12D or p.G12V mutations.
Validation of selective KRAS and BRAF mutations detected by qBiomarker with ultra-sensitive ddPCR.(A) List of cancer-associated KRAS and BRAF mutation loci assessed by qBiomarker and validated with ddPCR assay. (B) Serial dilution curve using DNA with known KRAS mutation (extracted from the PDX tumour CTG-0288). The blue markers indicate the concentration of mutant DNA (copies per μl) and the orange markers indicate the fractional abundance (%) of the KRAS mutated loci in a wild-type DNA background. All error bars generated by QuantaSoft software (Bio-Rad, Hercules, CA, USA) represent a 95% confidence interval. (C) To assess cross-reactivity of ddPCR probes targeting KRAS p.G12D and p.G12V mutations, DNA with known KRAS p.G12D mutation was probed with either specific ddPCR assay or with a probe designed for detection of p.G12V substitution. Alternatively, DNA isolated from tumour carrying the KRAS p.G12V mutation (D) was probed with either a specific or off-target ddPCR probe. Blue dot clusters indicate KRAS mutation detected by the specific assay. Black dot cluster indicates empty droplets. Mutated KRAS p.G12V cases when probed with the assay for p.G12D, and vice-versa, presented with an extra shifted cluster of black dots (identified by a star), probably resulted due to non-specific probes cross-reactivity. Green clusters indicate droplets containing wild-type KRAS alleles. (E) Schematic representation of the comparative analysis of qBiomarker and ddPCR approaches in a panel of 104 PDX tumours. Green and blue dots represent mutations discovered by either qBiomarker or ddPCR, respectively, although red dots indicate mutations concurrently detected by both platforms. Cases with no mutation detected are not shown. (F) Venn diagram summarising the number of KRAS and BRAF cancer-associated mutations detected by qBiomarker and ddPCR alone or concurrently. (G) Summary of detection accuracy (specificity, sensitivity, positive predictive value (PPV) and negative predictive value (NPV) of the qBiomarker approach when referenced to ddPCR.
We next used ddPCR to analyse 104 of 117 tumour models, previously assessed with qBiomarker for presence of five selective cancer-associated KRAS and/or BRAF mutations (Figure 3E; Supplementary Figure 1; Supplementary Table 6). The fractional abundance of cancer-driving mutations detected by ddPCR was at least 20%. Of 41 KRAS/BRAF mutations detected across all tested tumours by ddPCR, 36 mutations (over 87%) were correspondingly discovered by qBiomarker approach (Figure 3F). Due to its high sensitivity, ddPCR might be considered as a gold-standard method for detecting the tumour-associated mutations in research settings (Huggett et al, 2015). Using the ddPCR data as a reference, our findings have further confirmed that qBiomarker array provides a reliable framework for cancer-associated mutations screening (Figure 3G).
Furthermore, the presence of selective KRAS/BRAF substitutions has been assessed in 51 PDX tumour models in parallel by qBiomarker, AmpliSeq and ddPCR techniques (Figure 4A; Supplementary Figure 1; Supplementary Table 7). Notably, 23 (85%) of 27 mutations have been successfully discovered by all three methods (Figure 4B). Finally, the WES data was also available for 31 of these 51 PDX tumours. As indicated in Figure 4C, most (11 of 12) of the KRAS/BRAF mutations assessed in these patients were simultaneously detected by all four mutation detection approaches (Supplementary Table 8), further supporting a high cross-platform concurrency and providing additional evidence that qPCR-based qBiomarker approach is an accurate and reliable method for cancer mutations screening.
KRAS and BRAF mutation detection comparison across the different platforms.(A) Comparison of qBiomarker, AmpliSeq and ddPCR for the detection of clinically relevant KRAS and BRAF mutations in a panel of 51 PDX tumours. Red and blue dots indicate KRAS or BRAF mutations, respectively. Empty squares represent cases with no identified mutations. (B) Venn diagram summarising the number of mutations (KRAS and BRAF combined) detected by one of the methods alone (qBiomarker, AmpliSeq and ddPCR) and concomitantly. (C) Diagram representing KRAS (red dots) and BRAF (blue dots) mutations detected by qBiomarker, AmpliSeq, ddPCR and WES in 31 PDX models. Empty squares represent cases with no identified mutations.
Discussion
The cell growth and survival advantages acquired by malignant tumour cells are consequences of cumulative genetic and epigenetic aberrations (Stratton et al, 2009; Vogelstein et al, 2013). Despite the mutational heterogeneity of neoplastic diseases, a subset of cancer-driving genes are commonly mutated in a high proportion of various solid tumour types, and correlate with disease prognosis and clinical response to therapeutic agents. Over the past decade, genotyping for driver mutations has gained wide acceptance and it is now used for cancer prognosis, remission scoring and treatment selection options. Although recently emerged WES and WGS approaches with promising diagnostic capabilities will probably displace other mutation detection techniques over the next few years, their use as a modality for large-scale population-based screening has recognised limitations, such as: long processing times, cost and most notably, difficulties in calling functional and actionable genetic variants in cancer (Rehm, 2013; Royer-Bertrand and Rivolta, 2015). To make NGS more amenable for diagnostic needs, sequencing kits targeting known cancer-associated genes have been offered by various manufacturers and are CLIA certified for clinical use. Although targeted tests are quicker and provide accurate results due to better coverage (Singh et al, 2013; Tsongalis et al, 2014), the prices have largely remained high, posing a practical barrier to clinical translation. Likewise, although ultra-sensitive ddPCR technology offers an advantageous approach for targeted mutation detection, its clinical implementation requires large volumes of expensive reagents (Huggett et al, 2015) and delays its adoption for mutation screening as a routine practice. On the other hand, the detection sensitivity of most commonly used methods for mutation detection, Sanger sequencing and pyrosequencing, is ∼20% and 5%, respectively (Tsiatis et al, 2010). Therefore, mutations occurring below the detection limit would not be identified.
Given their speed, throughput capabilities and cost-effectiveness, qPCR mutation detection techniques are well suited for the clinical environment and provide a method of choice for various diagnostic uses (Martinez-Llordella et al, 2008; Maurin, 2012; Warhurst et al, 2015). Although several qPCR cancer mutation detection methods have been reported in the literature and are commercially available (Didelot et al, 2012; Bolton et al, 2015), the lack of rigorous validation in tumour specimens has limited their application in the clinical diagnostic setting. This poses a pressing clinical need for a well-validated, rapid and cost-effective mutation profiling method with optimal analytical performance in patient specimens.
In this study, we have directly compared the recently developed qBiomarker (Qiagen) array with NGS-based Ion Torrent AmpliSeq (Thermo Fisher Scientific), WES (Illumina) and ultra-sensitive ddPCR (Bio-Rad) to validate these techniques for cancer-associated mutation analysis in a large panel of early passage PDX tumours. As we have previously demonstrated, PDX models are biologically stable and accurately reflect the patient’s tumour histopathology, mutational landscape and therapeutic response, and represent a unique in vivo setting for preclinical evaluation of cancer-driving events (Rubio-Viqueira et al, 2006; Garber, 2007; Hidalgo et al, 2011; Morelli et al, 2012; Stebbing et al, 2014; Gao et al, 2015). Further supporting this concept, mutational landscape detected by qBiomarker array in our PDX cohort was highly compatible with the spectrum of genetic alterations in primary tumours reported in the comprehensive cancer mutational database COSMIC. Although our data is likely biased, as qBiomarker targets highly selected cancer-associated mutations, these aberrations have greater frequency and represent a large share of all genetic events reported for these genes in human databases. Consequently, these observations support the credibility of the qBiomarker mutation detection approach.
By comparing detection of 297 non-synonymous aberrations, we found an overall 72% agreement between qBiomarker and AmpliSeq in 59 PDX samples. Despite the high concordance, 18.8% and 9.4% of mutations were identified only by qBiomarker or AmpliSeq, respectively. As most of the techniques were performed using the same DNA extraction aliquot to minimise effects of intratumoral heterogeneity, these differences may be attributed to vulnerability of both methods to false-positive and false-negative calls. Results of the qPCR-based approaches may be affected by a number of compounding factors that reduce the specificity of primers and probes leading to a relatively high level of signal background. On the other hand, in general, the amplicon-based approaches, such as AmpliSeq (Thermo Fisher Scientific, Waltham, MA, USA), TruSeq (Illumina, San Diego, CA, USA), Microdroplet PCR (RainDance, Billerica, MA, USA), ddPCR (Bio-Rad, Hercules, CA, USA), real-time PCR and Access Array (Fluidigm, San Francisco, CA, USA), may not be suitable for screening of a large number of targets due to the difficulties associated with primer design for multiplex purposes. As the number of primers in the reaction increases, the level of non-specific amplification caused by the interaction between the primers increases as well, and therefore affects the assay reliability (Mamanova et al, 2010). For the same reason, the addition of targets into an established amplicons panel requires primer redesign to accommodate the new targets and minimise non-specific primer–primer interactions. Accordingly, recent studies have demonstrated that ‘false-positive’ variant calls in clinically relevant genes were frequently detected by an AmpliSeq cancer panel (Zhang et al, 2014) and other amplicon-based approaches (Samorodnitsky et al, 2015). It was reported that hybridisation capture-based methods provide better coverage uniformity and are less likely to nominate false-positive calls than amplicon-based approaches (Samorodnitsky et al, 2015). Notably, results obtained from a direct comparison between qBiomarker, AmpliSeq and capture-based WES for the detection of 253 cancer-associated mutations in 43 PDX models were highly concordant (79.6% detected by all three platforms), and show that qBiomarker assays had greater correspondence with WES than with amplicon-based AmpliSeq platform, further supporting the notion that results from hybridisation-based NGS significantly correlate with those observed by real-time qPCR (Tuononen et al, 2013).
KRAS and BRAF mutations are usually associated with poor prognosis and chemotherapeutic resistance in various solid malignancies (Davies et al, 2002; Pylayeva-Gupta et al, 2011). Although KRAS and BRAF mutations are routinely assessed in tumour tissues, their detection can be challenging due to the high-testing volume frequency and low tumour content. Due to its superior sensitivity, ddPCR has quickly become an approach of choice for KRAS and BRAF genotyping in tumours and bodily fluids (Bettegowda et al, 2014; Diaz and Bardelli, 2014; Oxnard et al, 2014) in research settings. To further evaluate the qBiomarker’s diagnostic accuracy, we analysed 104 of 117 PDX tumour samples for presence of three KRAS and two BRAF clinically relevant mutations with ddPCR assay. The qBiomarker results were confirmed by ddPCR analysis with 88% concordance, demonstrating that qBiomarker assay can detect low-prevalence mutations in a wild-type genomic background. Furthermore, almost 85% of selective KRAS/BRAF mutations were concurrently detected by qBiomarker, AmpliSeq and ddPCR in 51 PDX models and 91.7% of mutations were detected by all four platforms in a panel of 31 tumour samples, demonstrating that the qBiomarker’s performance is on par with that of other routinely used but more complex and expensive methods of BRAF/KRAS mutation detection. Nevertheless, further evaluation is warranted to validate its suitability for noninvasive mutation profiling.
The workflow complexities and turnaround times of the different mutation detecting methods compared in this study differ considerably. Unlike the qBiomarker, the workflow for NGS-based AmpliSeq and WES are lengthier and more labour intensive, owing the different steps needed to construct sequencing libraries and template preparation. An AmpliSeq protocol requires two multiplex PCR amplifications: enrichment with sequence-specific primers followed by barcoding needed for simultaneous sequencing of multiple patient samples. Although the AmpliSeq library construction hands-on time can be considerably shortened (from DNA to results in 2 days) by using an automated library preparation system, Ion Chef, its high list price impedes the widespread access to this technology and it may not be ideal for smaller labs. As qBiomarker PCR instrument-independent workflow involves only one setup step, the status of up to 360 mutations can be screened in <3 h. Although qBiomarker requires a low input of 5–10 ng of fresh-frozen DNA per well, more DNA input is needed for samples extracted from FFPE specimens (depending on the DNA degradation level) to achieve sufficient effective amplification fractions, making this platform less suitable for studies where FFPE-derived input material is limited. As AmpliSeq requires just 10 ng of FFPE DNA, this platform may be preferred over the qPCR for sequencing of hard to obtain FFPE samples. Notwithstanding this limitation, our comprehensive comparative analysis of various cancer mutation detection approaches indicates that the qPCR-based qBiomarker is an accurate, simple, rapid and cost-effective method that compares favourably to more expensive and labour-intensive methodologies that are currently in use. This observation provides a foundation for advancing the use of qBiomarker assay for cancer-driving mutations profiling in clinical diagnostics.
In summary, our comprehensive large-scale direct comparison and validation of currently available mutation detection methods is of extreme importance for the current scenario of precision medicine. Better understanding the sensitivity, turnaround time and cost-effectiveness of mutation detection techniques will lead to informed choice of screening methodologies, especially in lower budget conditions or time frame limitations.
Change history
14 February 2017
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Anastas JN, Moon RT (2013) WNT signalling pathways as therapeutic targets in cancer. Nat Rev Cancer 13 (1): 11–26.
Ascierto PA, Simeone E, Giannarelli D, Grimaldi AM, Romano A, Mozzillo N (2012) Sequencing of BRAF inhibitors and ipilimumab in patients with metastatic melanoma: a possible algorithm for clinical use. J Transl Med 10: 107.
Bertotti A, Migliardi G, Galimi F, Sassi F, Torti D, Isella C, Cora D, Di Nicolantonio F, Buscarino M, Petti C, Ribero D, Russolillo N, Muratore A, Massucco P, Pisacane A, Molinaro L, Valtorta E, Sartore-Bianchi A, Risio M, Capussotti L, Gambacorta M, Siena S, Medico E, Sapino A, Marsoni S, Comoglio PM, Bardelli A, Trusolino L (2011) A molecularly annotated platform of patient-derived xenografts (‘xenopatients’) identifies HER2 as an effective therapeutic target in cetuximab-resistant colorectal cancer. Cancer Discov 1 (6): 508–523.
Bettegowda C, Sausen M, Leary RJ, Kinde I, Wang Y, Agrawal N, Bartlett BR, Wang H, Luber B, Alani RM, Antonarakis ES, Azad NS, Bardelli A, Brem H, Cameron JL, Lee CC, Fecher LA, Gallia GL, Gibbs P, Le D, Giuntoli RL, Goggins M, Hogarty MD, Holdhoff M, Hong SM, Jiao Y, Juhl HH, Kim JJ, Siravegna G, Laheru DA, Lauricella C, Lim M, Lipson EJ, Marie SK, Netto GJ, Oliner KS, Olivi A, Olsson L, Riggins GJ, Sartore-Bianchi A, Schmidt K, Shih LM, Oba-Shinjo SM, Siena S, Theodorescu D, Tie J, Harkins TT, Veronese S, Wang TL, Weingart JD, Wolfgang CL, Wood LD, Xing D, Hruban RH, Wu J, Allen PJ, Schmidt CM, Choti MA, Velculescu VE, Kinzler KW, Vogelstein B, Papadopoulos N, Diaz LA Jr. (2014) Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci Transl Med 6 (224): 224ra24.
Bolton L, Reiman A, Lucas K, Timms J, Cree IA (2015) KRAS mutation analysis by PCR: a comparison of two methods. PLoS One 10 (1): e0115672.
Bryant KL, Mancias JD, Kimmelman AC, Der CJ (2014) KRAS: feeding pancreatic cancer proliferation. Trends Biochem Sci 39 (2): 91–100.
Cibulskis K, Lawrence MS, Carter SL, Sivachenko A, Jaffe D, Sougnez C, Gabriel S, Meyerson M, Lander ES, Getz G (2013) Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat Biotechnol 31 (3): 213–219.
Davies H, Bignell GR, Cox C, Stephens P, Edkins S, Clegg S, Teague J, Woffendin H, Garnett MJ, Bottomley W, Davis N, Dicks E, Ewing R, Floyd Y, Gray K, Hall S, Hawes R, Hughes J, Kosmidou V, Menzies A, Mould C, Parker A, Stevens C, Watt S, Hooper S, Wilson R, Jayatilake H, Gusterson BA, Cooper C, Shipley J, Hargrave D, Pritchard-Jones K, Maitland N, Chenevix-Trench G, Riggins GJ, Bigner DD, Palmieri G, Cossu A, Flanagan A, Nicholson A, Ho JW, Leung SY, Yuen ST, Weber BL, Seigler HF, Darrow TL, Paterson H, Marais R, Marshall CJ, Wooster R, Stratton MR, Futreal PA (2002) Mutations of the BRAF gene in human cancer. Nature 417 (6892): 949–954.
De Bruijn MT, Raats DA, Tol J, Hinrichs J, Teerenstra S, Punt CJ, Rinkes IH, Kranenburg O (2011) Combined KRAS and TP53 mutation status is not predictive in CAPOX-treated metastatic colorectal cancer. Anticancer Res 31 (4): 1379–1385.
DeRose YS, Wang G, Lin YC, Bernard PS, Buys SS, Ebbert MT, Factor R, Matsen C, Milash BA, Nelson E, Neumayer L, Randall RL, Stijleman IJ, Welm BE, Welm AL (2011) Tumor grafts derived from women with breast cancer authentically reflect tumor pathology, growth, metastasis and disease outcomes. Nat Med 17 (11): 1514–1520.
Diaz LA Jr, Bardelli A (2014) Liquid biopsies: genotyping circulating tumor DNA. J Clin Oncol 32 (6): 579–586.
Didelot A, Le Corre D, Luscan A, Cazes A, Pallier K, Emile JF, Laurent-Puig P, Blons H (2012) Competitive allele specific TaqMan PCR for KRAS, BRAF and EGFR mutation detection in clinical formalin fixed paraffin embedded samples. Exp Mol Pathol 92 (3): 275–280.
Gao H, Korn JM, Ferretti S, Monahan JE, Wang Y, Singh M, Zhang C, Schnell C, Yang G, Zhang Y, Balbin OA, Barbe S, Cai H, Casey F, Chatterjee S, Chiang DY, Chuai S, Cogan SM, Collins SD, Dammassa E, Ebel N, Embry M, Green J, Kauffmann A, Kowal C, Leary RJ, Lehar J, Liang Y, Loo A, Lorenzana E, Robert McDonald E 3rd, McLaughlin ME, Merkin J, Meyer R, Naylor TL, Patawaran M, Reddy A, Roelli C, Ruddy DA, Salangsang F, Santacroce F, Singh AP, Tang Y, Tinetto W, Tobler S, Velazquez R, Venkatesan K, Von Arx F, Wang HQ, Wang Z, Wiesmann M, Wyss D, Xu F, Bitter H, Atadja P, Lees E, Hofmann F, Li E, Keen N, Cozens R, Jensen MR, Pryer NK, Williams JA, Sellers WR (2015) High-throughput screening using patient-derived tumor xenografts to predict clinical trial drug response. Nat Med 21 (11): 1318–1325.
Garber K (2007) Personal mouse colonies give hope for pancreatic cancer patients. J Natl Cancer Inst 99 (2): 105–107.
Hidalgo M, Bruckheimer E, Rajeshkumar NV, Garrido-Laguna I, De Oliveira E, Rubio-Viqueira B, Strawn S, Wick MJ, Martell J, Sidransky D (2011) A pilot clinical study of treatment guided by personalized tumorgrafts in patients with advanced cancer. Mol Cancer Ther 10 (8): 1311–1316.
Huggett JF, Cowen S, Foy CA (2015) Considerations for digital PCR as an accurate molecular diagnostic tool. Clin Chem 61 (1): 79–88.
Jimenez C, Sebastian E, Chillon MC, Giraldo P, Mariano Hernandez J, Escalante F, Gonzalez-Lopez TJ, Aguilera C, de Coca AG, Murillo I, Alcoceba M, Balanzategui A, Sarasquete ME, Corral R, Marin LA, Paiva B, Ocio EM, Gutierrez NC, Gonzalez M, San Miguel JF, Garcia-Sanz R (2013) MYD88 L265P is a marker highly characteristic of, but not restricted to, Waldenstrom's macroglobulinemia. Leukemia 27 (8): 1722–1728.
Kato S, Han SY, Liu W, Otsuka K, Shibata H, Kanamaru R, Ishioka C (2003) Understanding the function-structure and function-mutation relationships of p53 tumor suppressor protein by high-resolution missense mutation analysis. Proc Natl Acad Sci USA 100 (14): 8424–8429.
Kim S, Jeong K, Bhutani K, Lee J, Patel A, Scott E, Nam H, Lee H, Gleeson JG, Bafna V (2013) Virmid: accurate detection of somatic mutations with sample impurity inference. Genome Biol 14 (8): R90.
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25 (14): 1754–1760.
Luo Y, Kaz AM, Kanngurn S, Welsch P, Morris SM, Wang J, Lutterbaugh JD, Markowitz SD, Grady WM (2013) NTRK3 is a potential tumor suppressor gene commonly inactivated by epigenetic mechanisms in colorectal cancer. PLoS Genet 9 (7): e1003552.
Mamanova L, Coffey AJ, Scott CE, Kozarewa I, Turner EH, Kumar A, Howard E, Shendure J, Turner DJ (2010) Target-enrichment strategies for next-generation sequencing. Nat Methods 7 (2): 111–118.
Martinez-Llordella M, Lozano JJ, Puig-Pey I, Orlando G, Tisone G, Lerut J, Benitez C, Pons JA, Parrilla P, Ramirez P, Bruguera M, Rimola A, Sanchez-Fueyo A (2008) Using transcriptional profiling to develop a diagnostic test of operational tolerance in liver transplant recipients. J Clin Invest 118 (8): 2845–2857.
Maurin M (2012) Real-time PCR as a diagnostic tool for bacterial diseases. Expert Rev Mol Diagn 12 (7): 731–754.
Morelli MP, Calvo E, Ordonez E, Wick MJ, Viqueira BR, Lopez-Casas PP, Bruckheimer E, Calles-Blanco A, Sidransky D, Hidalgo M (2012) Prioritizing phase I treatment options through preclinical testing on personalized tumorgraft. J Clin Oncol 30 (4): e45–e48.
Neumann J, Zeindl-Eberhart E, Kirchner T, Jung A (2009) Frequency and type of KRAS mutations in routine diagnostic analysis of metastatic colorectal cancer. Pathol Res Pract 205 (12): 858–862.
Oxnard GR, Paweletz CP, Kuang Y, Mach SL, O'Connell A, Messineo MM, Luke JJ, Butaney M, Kirschmeier P, Jackman DM, Janne PA (2014) Noninvasive detection of response and resistance in EGFR-mutant lung cancer using quantitative next-generation genotyping of cell-free plasma DNA. Clin Cancer Res 20 (6): 1698–1705.
Pylayeva-Gupta Y, Grabocka E, Bar-Sagi D (2011) RAS oncogenes: weaving a tumorigenic web. Nat Rev Cancer 11 (11): 761–774.
Rehm HL (2013) Disease-targeted sequencing: a cornerstone in the clinic. Nat Rev Genet 14 (4): 295–300.
Royer-Bertrand B, Rivolta C (2015) Whole genome sequencing as a means to assess pathogenic mutations in medical genetics and cancer. Cell Mol Life Sci 72 (8): 1463–1471.
Rubio-Viqueira B, Jimeno A, Cusatis G, Zhang X, Iacobuzio-Donahue C, Karikari C, Shi C, Danenberg K, Danenberg PV, Kuramochi H, Tanaka K, Singh S, Salimi-Moosavi H, Bouraoud N, Amador ML, Altiok S, Kulesza P, Yeo C, Messersmith W, Eshleman J, Hruban RH, Maitra A, Hidalgo M (2006) An in vivo platform for translational drug development in pancreatic cancer. Clin Cancer Res 12 (15): 4652–4661.
Samorodnitsky E, Jewell BM, Hagopian R, Miya J, Wing MR, Lyon E, Damodaran S, Bhatt D, Reeser JW, Datta J, Roychowdhury S (2015) Evaluation of hybridization capture versus amplicon-based methods for whole-exome sequencing. Hum Mutat 36 (9): 903–914.
Sangodkar J, Dhawan NS, Melville H, Singh VJ, Yuan E, Rana H, Izadmehr S, Farrington C, Mazhar S, Katz S, Albano T, Arnovitz P, Okrent R, Ohlmeyer M, Galsky M, Burstein D, Zhang D, Politi K, Difeo A, Narla G (2012) Targeting the FOXO1/KLF6 axis regulates EGFR signaling and treatment response. J Clin Invest 122 (7): 2637–2651.
Saunders CT, Wong WS, Swamy S, Becq J, Murray LJ, Cheetham RK (2012) Strelka: accurate somatic small-variant calling from sequenced tumor-normal sample pairs. Bioinformatics 28 (14): 1811–1817.
Segditsas S, Tomlinson I (2006) Colorectal cancer and genetic alterations in the Wnt pathway. Oncogene 25 (57): 7531–7537.
Singh RR, Patel KP, Routbort MJ, Reddy NG, Barkoh BA, Handal B, Kanagal-Shamanna R, Greaves WO, Medeiros LJ, Aldape KD, Luthra R (2013) Clinical validation of a next-generation sequencing screen for mutational hotspots in 46 cancer-related genes. J Mol Diagn 15 (5): 607–622.
Smith G, Bounds R, Wolf H, Steele RJ, Carey FA, Wolf CR (2010) Activating K-Ras mutations outwith 'hotspot' codons in sporadic colorectal tumours—implications for personalised cancer medicine. Br J Cancer 102 (4): 693–703.
Stebbing J, Paz K, Schwartz GK, Wexler LH, Maki R, Pollock RE, Morris R, Cohen R, Shankar A, Blackman G, Harding V, Vasquez D, Krell J, Ciznadija D, Katz A, Sidransky D (2014) Patient-derived xenografts for individualized care in advanced sarcoma. Cancer 120 (13): 2006–2015.
Stratton MR, Campbell PJ, Futreal PA (2009) The cancer genome. Nature 458 (7239): 719–724.
Tamborero D, Gonzalez-Perez A, Perez-Llamas C, Deu-Pons J, Kandoth C, Reimand J, Lawrence MS, Getz G, Bader GD, Ding L, Lopez-Bigas N (2013) Comprehensive identification of mutational cancer driver genes across 12 tumor types. Sci Rep 3: 2650.
Tsiatis AC, Norris-Kirby A, Rich RG, Hafez MJ, Gocke CD, Eshleman JR, Murphy KM (2010) Comparison of Sanger sequencing, pyrosequencing, and melting curve analysis for the detection of KRAS mutations: diagnostic and clinical implications. J Mol Diagn 12 (4): 425–432.
Tsongalis GJ, Peterson JD, de Abreu FB, Tunkey CD, Gallagher TL, Strausbaugh LD, Wells WA, Amos CI (2014) Routine use of the Ion Torrent AmpliSeq Cancer Hotspot Panel for identification of clinically actionable somatic mutations. Clin Chem Lab Med 52 (5): 707–714.
Tuononen K, Maki-Nevala S, Sarhadi VK, Wirtanen A, Ronty M, Salmenkivi K, Andrews JM, Telaranta-Keerie AI, Hannula S, Lagstrom S, Ellonen P, Knuuttila A, Knuutila S (2013) Comparison of targeted next-generation sequencing (NGS) and real-time PCR in the detection of EGFR, KRAS, and BRAF mutations on formalin-fixed, paraffin-embedded tumor material of non-small cell lung carcinoma-superiority of NGS. Genes Chromosomes Cancer 52 (5): 503–511.
Villarroel MC, Rajeshkumar NV, Garrido-Laguna I, De Jesus-Acosta A, Jones S, Maitra A, Hruban RH, Eshleman JR, Klein A, Laheru D, Donehower R, Hidalgo M (2011) Personalizing cancer treatment in the age of global genomic analyses: PALB2 gene mutations and the response to DNA damaging agents in pancreatic cancer. Mol Cancer Ther 10 (1): 3–8.
Vogelstein B, Papadopoulos N, Velculescu VE, Zhou S, Diaz LA Jr., Kinzler KW (2013) Cancer genome landscapes. Science 339 (6127): 1546–1558.
Warhurst G, Dunn G, Chadwick P, Blackwood B, McAuley D, Perkins GD, McMullan R, Gates S, Bentley A, Young D, Carlson GL, Dark P (2015) Rapid detection of health-care-associated bloodstream infection in critical care using multipathogen real-time polymerase chain reaction technology: a diagnostic accuracy study and systematic review. Health Technol Assess 19 (35): 1–142.
Wilm A, Aw PP, Bertrand D, Yeo GH, Ong SH, Wong CH, Khor CC, Petric R, Hibberd ML, Nagarajan N (2012) LoFreq: a sequence-quality aware, ultra-sensitive variant caller for uncovering cell-population heterogeneityfrom high-throughput sequencing datasets. Nucleic Acids Res 40 (22): 11189–11201.
Yachida S, White CM, Naito Y, Zhong Y, Brosnan JA, Macgregor-Das AM, Morgan RA, Saunders T, Laheru DA, Herman JM, Hruban RH, Klein AP, Jones S, Velculescu V, Wolfgang CL, Iacobuzio-Donahue CA (2012) Clinical significance of the genetic landscape of pancreatic cancer and implications for identification of potential long-term survivors. Clin Cancer Res 18 (22): 6339–6347.
Zhang L, Chen L, Sah S, Latham GJ, Patel R, Song Q, Koeppen H, Tam R, Schleifman E, Mashhedi H, Chalasani S, Fu L, Sumiyoshi T, Raja R, Forrest W, Hampton GM, Lackner MR, Hegde P, Jia S (2014) Profiling cancer gene mutations in clinical formalin-fixed, paraffin-embedded colorectal tumor specimens using targeted next-generation sequencing. Oncologist 19 (4): 336–343.
Acknowledgements
QIAGEN Sciences Inc has provided part of the reagents used in the current study. QIAGEN Sciences Inc company had no role in the design of the study, data collection or analysis; in the interpretation of the results; in the preparation of the manuscript; or in the decision to submit the manuscript for publication. ‘This project has been funded in part with Federal funds from the National Cancer Institute, National Institutes of Health, (Samuel Long) under Contract No. HHSN261200800001E. The content of this publication does not necessarily reflect the views or policies of the Department of Health and Human Services, nor does mention of trade names, commercial products, or organisations imply endorsement by the US Government’.
Author contributions
MB and DS planned and designed the work. MB, EI and DS wrote the manuscript with input from all authors. MB, EI, LTK, SL, EB and DS contributed to the development of methodology. EB, EF, DC, SL, IS, IB, KP, GB and TK performed analysis and interpretation of data (e.g., statistical and computational analysis, sequence data management). EB, GB, TK, IS, IB and KP contributed to PDX generation, DNA extraction and sequencing. MB, EI, DS, LTK, PW and BF participated in development of methodology and provided administrative, technical or material support (i.e., reporting or organising data, constructing databases). EI produced the figures. All authors participated in review, and/or revision of the manuscript. All authors discussed the results, analysed the data and contributed to the manuscript preparation. MB and DS instigated and supervised the project.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Samuel Long was employed by the QIAGEN Sciences Inc. at the initial time of the experiments preparation. The remaining authors declare no conflict of interest.
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License.
Supplementary Information accompanies this paper on British Journal of Cancer website
Supplementary information
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Brait, M., Izumchenko, E., Kagohara, L. et al. Comparative mutational landscape analysis of patient-derived tumour xenografts. Br J Cancer 116, 515–523 (2017). https://doi.org/10.1038/bjc.2016.450
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2016.450