Abstract
Background:
Breast cancer is a leading cause of morbidity and mortality worldwide. Although mammography screening is available, there is an ongoing interest in improved early detection and prognosis. Herein, we have analysed a combination of serological biomarkers in a case–control cohort of sera taken before diagnosis.
Methods:
This nested case–control study within the UK Collaborative Trial of Ovarian Cancer Screening (UKCTOCS) used serum samples from 239 women who subsequently developed breast cancer and 239 matched cancer-free controls. Sera were screened by ELISA for 9 candidate markers. Univariate and multivariate analyses were performed to examine associations with clinico-pathological features and between case controls in different time groups before diagnosis.
Results:
Significant associations with clinico-pathological features related to prognosis were found for several candidates (CA15-3, HSP90A and PAI-1). However, there were no consistent differences between cases and controls for any candidate in the lead up to diagnosis. Whilst combination models outperformed single markers, there was no increase in performance towards diagnosis.
Conclusions:
This study using unique pre-diagnosis samples shows that CA15-3, HSP90A and PAI-1 have potential as early prognostic markers and warrant further investigation. However, none of the candidates or combinations would be useful for screening.
Similar content being viewed by others
Main
Breast cancer is the most common neoplasm in women and the second leading cause of cancer-related mortality in females worldwide (Siegel et al, 2014). At present, breast cancer detection relies mostly on mammography, which has been associated with decreased breast cancer mortality (Jatoi, 1999; Gotzsche and Jorgensen, 2013). However, mammography screening has generated controversy due to the risks of false-positive results and over-diagnosis of indolent disease (Baum, 2010; Gotzsche and Jorgensen, 2013; Pace and Keating, 2014; Welch et al, 2016). Mammography also has limited sensitivity for the detection of tumours in dense breast tissue (Boyd et al, 2007). There is thus an urgent need for early biomarkers that could predict disease outcome, providing prognostic information to the clinician for treatment stratification. The addition of a blood-based tumour marker test may also increase patient compliance as blood testing is more acceptable and would also circumvent the problems associated with imaging high-density breast tissue.
Some blood-borne tumour markers have demonstrated ability to detect malignancy before clinical diagnosis and are currently being evaluated in screening trials for certain cancers; for example, CA125 for screening ovarian cancer (Menon et al, 2015). Breast cancer markers in clinical practice are used for predicting response to therapy, monitoring after primary therapy or as prognostic indicators (Harris et al, 2007). However, there are currently no blood-borne biomarkers recommended for breast cancer diagnosis or screening. Although candidate markers such as carcinoembryonic antigen (CEA), the soluble form of MUC1 protein (CA15-3, CA27.29) and circulating cytokeratin fragments (TPA, TPS and CYFRA 21-1) have been suggested as diagnostic markers, they lack sensitivity for early disease detection and/or lack specificity.
Almost all diagnostic biomarker studies to date have used samples collected at or just after diagnosis and therefore may be confounded by late-stage responses to advanced tumours. Ideally, samples taken before clinical diagnosis should be used in the search for biomarkers of pre-symptomatic, early stage disease and for predicting prognosis. Herein, we have sourced such pre-diagnostic samples from the UK Collaborative Trial of Ovarian Cancer Screening (UKCTOCS); a multi-centre RCT (the largest ever undertaken), which aims to assess if screening for ovarian cancer can save lives (Menon et al, 2009, 2015). Serum samples were taken from 239 UKCTOCS women who were diagnosed with invasive ductal carcinoma of the breast, months to years after sample donation. Cases were matched 1 : 1 with samples from non-cancer controls. The serum biomarkers investigated in this study were chosen based on previous evidence of their potential as serum and/or tissue markers of breast cancer: CA15-3 (cancer antigen 15-3) (Hayes et al, 1986; Geraghty et al, 1992; Stieber et al, 2003), RANTES/CCL5 (regulated on activation, normal T cell expressed and secreted/chemokine (C–C motif) ligand 5) (Soria and Ben-Baruch, 2008; Gonzalez et al, 2011), OPN (osteopontin) (Singhal et al, 1997; Fedarko et al, 2001; Bramwell et al, 2014), PAI-1 (plasminogen activator inhibitor-1; Harris et al, 2007; Duffy et al, 2014), SLPI (secretory leukocyte protease inhibitor) (Hu et al, 2004; Bouchard et al, 2006), HSP90A (heat shock protein 90A; Pick et al, 2007; Duran et al, 2008; Cheng et al, 2012), IGFBP3 (insulin-like growth factor-binding protein 3; McCarthy et al, 2009; Worthington et al, 2010), APOC1 (apolipoprotein C-I) (Opstal-van Winden et al, 2011; Devetyarov et al, 2012; Chung et al, 2014; Lee et al, 2015) and PAPPA (pappalysin-1; Loddo et al, 2014; Mansfield et al, 2014). Herein we report on the ability of these serum markers to detect breast cancer cases before their diagnosis and examine associations with clinico-pathological features and prognostic indicators.
Materials and Methods
Ethics, consent and permissions
UKCTOCS participants gave informed written consent at recruitment for the use of their medical notes and serum in ethically approved secondary studies (UK North West Medical Research and Ethics Committee (MREC 00/8/34)). Ethical approval for this nested case–control study was granted by The Joint UCL/UCLH Committees on the Ethics of Human Research (Committee A) REC ref 05/Q0505/57.
Subjects
UKCTOCS participants were post-menopausal women aged 50–74, who had no active malignancy at recruitment (Menon et al, 2008). Notifications of women subsequently diagnosed with invasive ductal carcinoma of the breast were retrieved by querying the Health and Social Care Information Centre (HSCIC) cancer and death registries with the International Classification of Diseases code C50 pertaining to malignant neoplasms of the breast. Cancer notifications were also received via self-reported data completed 3.5 years post-randomisation to the UKCTOCS. Breast cancer notifications were confirmed and characterised by postal questionnaire sent to treating clinicians (consultant, or General Practitioner if details not provided by the volunteer), which was designed to ascertain clinical and histological data on diagnosed cases (date of diagnosis, histology, nodal status, staging, grade, prognosis, ER, PR and HER2 status). Staging for each case was determined using the TNM system, taking into account tumour size (T), spread to the lymph nodes (N), and whether the tumour had metastasised (M). To provide an indication of prognosis, the Nottingham prognostic index (NPI) was calculated for each case using the formula [0.2 × S]+N+G, where S=tumour size in cm, N=node status (0 nodes=1, 1–4 nodes=2, >4 nodes=3) and G=numerical grade. Cases were stratified into good and poor prognosis using an NPI ⩽4 and >4, respectively. Controls were age-matched and were women from the same trial centre who had no history of cancer, and who had donated serum samples on the same date as the matched case. Characteristics of the case–control set are shown in Table 1. There was no significant difference in time to spin (clotting time) or age between cases and controls.
Samples and assays
Blood was collected and serum prepared according to a standardised protocol within the UKCTOCS (Menon et al, 2009) and then shipped frozen to a cryo-repository for long-term storage in liquid nitrogen. For the study, single samples from cases were selected that predated diagnosis by a median time of 1.15 years (13.8 months; range 0.1–53.8 months). Samples were retrieved from storage and shipped to the laboratory on dry ice and thawed at 4 °C for random aliquoting before assay. Serum concentrations of all biomarker candidates were measured using commercial enzyme-linked immunosorbent assays (ELISA) according to the manufacturers’ instructions. Optimum serum dilutions were first determined using a pool of serum from cases and controls and used to determine inter-assay CVs. The ELISA kits used, dilutions and inter-assay CVs were; CA15-3 (Human) ELISA kit (Abnova; 1 : 50; 7.3%), Human CCL5/RANTES Quantikine ELISA (R&D; 1 : 100; 8.1%), Human IGFBP-3 Quantikine ELISA (R&D; 1 : 100; 9.8%), Human PAI-1 ELISA Kit (Invitrogen; 1 : 25; 8.1%), Human Osteopontin (OPN) Quantikine ELISA (R&D; 1 : 12.5; 6.1%), Hsp90α ELISA Kit (Stressgen; 1 : 30; 6.5%), Human SLPI Quantikine ELISA (R&D; 1 : 30; 8.5%), human Pappalysin-1 (PAPPA) ELISA kit (DRG; 1 : 200; 11.3%) and human Apolipoprotein C-I (APOC1) ELISA kit (Abnova; 1 : 100; 11.1%). There were no strong correlations with clotting time or volunteer age for any of the measured analytes.
Statistical analysis
Univariate analysis using GraphPad Prism software (v5.01) was used to test associations between analyte measurements and clinico-pathological features and between cases and matched controls in different time groups before diagnosis. For normally distributed data, the Student t-test was used to assess significance of differences; otherwise the Mann–Whitney U-test was used. P-values of <0.05 were considered significant. Kaplan–Meier analysis was used to examine biomarker levels in relation to survival in cases using time from sample collection to death and various candidate marker cut-off values. The log-rank test was used to test significance between the survival curves. Multivariate logistic regression models were constructed and tested within the R environment (v3.1.1 for Mac OS). Evaluation of performance of each model was based on receiver operating characteristic (ROC) curve analysis, with determination of significance of differences in areas under the curves using the method of DeLong et al (1988). A modelling approach based on the neuro-fuzzy technique was also applied to the data, which combines the mathematical constructions of fuzzy logic (Nauck et al, 1997; Paiva and Dourado, 2004) and neural networks. Briefly, a fuzzy model was generated using half of the data and its parameters estimated to minimise the difference between the model and the experimental data. Model adjustment was based on the sub-gradient modification method for the case of undifferentiated functions, the so-called r-algorithms by Shor (Kiseleva and Shor, 2005), with an accuracy of ɛ=0.001. The model was then validated on the other half of the data. Software implementation of the approach was developed within the Visual C++ environment.
Results
Pre-diagnostic candidate marker levels in discrimination of case–control samples
In this UKCTOCS nested case–control set, volunteers donated serum up to 5 years before clinical diagnosis of breast cancer, with a median time to diagnosis of 1.15 years (13.8 months). We hypothesised that serum levels of the candidate markers may be altered in women with a clinically undetectable tumour. Initially, biomarker data from all case samples were compared with all non-cancer controls and then those taken less than and greater than the median time to diagnosis. No single candidate marker showed a significant difference between cases and controls for these comparisons. Samples were also stratified into time groups that came from cases who developed breast cancer within 0.5 years, 0.5–1 years, 1–2 years, 2–3 years and 3–5 years of sample collection and candidate marker levels were compared against matched-control samples for these groups. Overall, there were no consistent changes in serum concentrations for any marker in cases in the lead up to diagnosis, although some significant differences between cases and controls were apparent for some candidates in different time groups. Thus, cases had elevated levels of IGFBP3 (P=0.044) 0–0.5 years before diagnosis, lower levels of RANTES (P=0.013) 2–3 years before diagnosis and higher levels of APOC1 (P=0.011) and lower levels of PAPPA (P=0.036) at 3–5 years before diagnosis.
Combining markers (including age as a parameter) using logistic regression failed to provide accurate discrimination of cases and controls in any pre-diagnosis time group, although combined models performed better than the single markers for classification (Table 2). There was no trend of improved classification in the lead up to diagnosis and sensitivities were in the range 4–26% at a specificity of >95%. A novel modelling approach based on the neuro-fuzzy technique also failed to provide accurate models, with sensitivities in the range 10–27% (data not shown). These data suggest that these candidates alone or combination are not accurate markers for predicting breast cancer.
Pre-diagnostic candidate marker levels in relation to clinico-pathological features
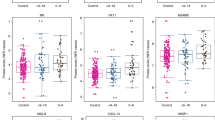
Pre-diagnostic serum levels of CA15-3 were significantly raised in samples from late-stage cases (stages 3/4 combined) vs controls and stage 1 and stage 2 cases (Tables 3A and 4; Figure 1A) and with node positivity within cases (Table 4). The marker was also raised significantly in node positive cancers within cases (Table 4) and vs controls in the subset of cases from which samples were taken <1.15 years before clinical diagnosis (Table 3B). However, CA15.3 levels were lower in grade 1 cancers vs controls or grade 2 cancers. Neither RANTES or OPN discriminated cases from controls for any of the pathological parameters examined (Table 3), and whereas altered levels of both were associated with tumour stage at diagnosis within cases (Table 4), for RANTES, this was not seen in the <1.15 year to diagnosis group and for OPN, the changes were not consistent, with lower levels in stage 2 vs stage 1 or stage 3/4 cases. IGFBP3 also failed to discriminate cases from controls, although was decreased in case samples that went on to be diagnosed with HER2+ breast cancer where there was a strong association for the <1.15 years to diagnosis group (Table 4B; Figure 1B). Serum levels of PAI-1 were significantly lower in samples from women who went on to be diagnosed with grade 3 cancer vs controls and grade 2 cancers and within cases with a higher NPI (Figures 1C and D). Similarly, HSP90A levels were lower in pre-diagnosis serum of grade 3 cases vs controls and in cases with a higher NPI (Figures 1E and F,) with the differences more significant closer to diagnosis (Tables 3 and 4). The association of lowered HSP90A with prognostic index was also significant (P=0.012) when only node-negative cases were considered. SLPI, PAPPA and APOC1 levels did not differ significantly for any of the pathological parameters and there were no differences for any candidate markers in relation to oestrogen receptor, progesterone receptor or triple negative status. The best logistic regression models combining markers for prognosis based on NPI gave AUCs of 0.67 (HSP90A, PAI-1 and RANTES) considering all samples, and 0.77 (HSP90A, PAI-1 and CA15-3), considering samples taken within 1.15 years of diagnosis. Although the AUCs for candidate markers were higher when comparing controls with cases that had subsequently died of breast cancer vs those that had not, a survival analysis indicated that none of the markers were predictive of survival from breast cancer (data not shown). However, this analysis is somewhat limited, as there were only 17 of the 239 cases (7%) where the primary cause of death was attributable to breast cancer (median time to death from sample collection=6.12 years; range 2.1–11.1 years).
Associations of biomarkers with histopathological features. Box and whisker plots showing associations of CA15-3 with tumour stage (A); IGFBP3 with HER2 status in the <1.15 years to diagnosis group (B); PAI-1 with tumour grade (C); PAI-1 with NPI (D); HSP90A with tumour grade (E); and HSP90A with NPI in the <1.15 years to diagnosis group (F).
Discussion
Few studies have tested blood-borne candidate tumour markers for early detection or prognosis in pre-diagnosis samples; the most appropriate sample type for this kind of study. Here, we found that none of the single marker candidates were effective at predicting breast cancer, even close to diagnosis, and there was lack of consistent changes in levels over time for any of the candidates. Although combining the candidate markers improved the detection rat, no combined model performed with a sensitivity that would be acceptable for use in screening. This is in agreement with a previous study using a smaller set of pre-diagnostic samples sourced from the Prospect-EPIC cohort (Opstal-van Winden et al, 2012). The authors reported that no marker combination (of 10 candidates, including CA15-3 and OPN) could accurately discriminate early breast cancer cases from controls. Thus, novel blood-borne biomarkers for the early detection of breast cancer for use in screening still need to be found.
CA15-3 is perhaps the best known, non-invasive marker of breast cancer, although its recommended clinical use is restricted to monitoring of patients with metastatic disease during active therapy (Harris et al, 2007). Herein, CA15-3 levels in serum taken at a median time of 13.8 months before clinical diagnosis showed consistency with respect to stage, grade and nodal status within cases, although it failed to discriminate cases from controls. This suggests that CA15-3 may serve as a pre-diagnostic marker of a more aggressive tumour phenotype, in line with previous reports of the prognostic value of CA15-3 in breast cancer (Ebeling et al, 2002; Gion et al, 2002; Shao et al, 2015). Raised pre-diagnostic serum levels of the chemoattractant RANTES were also associated with stage within cases, supporting previous observations of raised plasma and tissue expression correlating with increasing stage (Niwa et al, 2001). Notably however, this association was not significant in cases where samples were taken closer to diagnosis, suggesting RANTES is not a robust predictor of stage. Similarly, stage-associated changes in serum OPN levels were not consistent, and this bone adhesion molecule could not differentiate cases by grade, node status or prognostic index. This supports previous evidence that OPN is not a useful prognostic marker in early breast cancer (Bramwell et al, 2014).
The fibrinolysis regulator protein PAI-1 has reported roles in invasion, angiogenesis and metastasis with high levels in breast tumour tissue having prognostic value and use in stratification, particularly of node-negative breast cancer (Look et al, 2002; Duffy et al, 2014). Few studies have assessed blood-borne PAI-1 as a breast tumour marker, with one study reporting decreased serum concentrations in cases vs non-cancer controls (Kim et al, 2009). Herein, serum PAI-1 was reduced in women who went on to be diagnosed with grade 3 breast cancer or who had a high prognostic index, though it was unable to discriminate pre-diagnosis cases from controls. Although this data supports serum PAI-1 as an early prognostic factor, its direction of change is at odds with the raised levels seen in tumour tissues. HSP90A is a chaperone protein for several oncogenes (including HER2) and a pro-survival factor of breast cancer cells. We showed lower levels of serum HSP90A in women who went on to be diagnosed with high-grade cancers and with higher prognostic index. High tissue expression of HSP90 has been previously associated with increased HER2 and ER expression, large and high-grade tumours, node positivity and decreased survival (Pick et al, 2007), although another study reported no significant association between serum HSP90A levels and lesion severity (Zagouri et al, 2011). These combined evidences suggest that serum HSP90A may not make a good early prognostic marker, although based on our findings, further investigation of its potential is warranted. Finally, the serum level of the IGF1-binding protein IGFBP3 was significantly lower in samples from HER2+ cases taken within 1.15 years of diagnosis. This observation is in accordance with our previous finding that HER2 overexpression enhances IGF1 signalling via the down-regulation of IGFBP3 (Worthington et al, 2010). However, although increased serum IGF1 levels have been associated with increased breast cancer risk, no association with serum IGFBP3 levels was found (Key et al, 2010). Thus, IGFBP3 is a poor marker for predicting breast cancer, although the relationship between HER2 and IGFBP3 expression warrants further investigation.
Conclusions
The key strength of this study is that a well-characterised, pre-diagnosis set of samples were investigated, allowing an objective assessment of nine candidate serological markers for the early detection and prognostication of breast cancer. One weakness of the study is that more detailed information on the molecular sub-type of the tumours was not available (excepting receptor status), so future studies should address in more detail the possible effect of molecular sub-type on candidate biomarker levels, as recently reported for CA15-3 (Shao et al, 2015). We conclude that CA15-3, PAI-1 and HSP90A have potential as prognostic markers in the pre-diagnosis setting. The next phase of this work would be to validate these candidate biomarkers in a larger independent set of case–control samples sourced from the UKCTOCS with data linked to mammography findings, treatment and survival information for these women. Serial samples from breast cancer cases and matched controls would be used from the multimodal arm of the UKCTOCS, permitting assessment of longitudinal changes in these candidate biomarkers during tumour progression. We envisage, that if validated, these markers could be used as a second line test to mammography to identify tumours of poorer prognosis and thereby allow stratification for treatment. Whether or not such testing would have an impact on survival would need to be assessed in a prospective clinical trial. We also conclude that CA15-3, RANTES, IGFBP3, OPN, PAI-1, SLPI, HSP90A, PAPPA and APOC1 alone, or in combination, cannot be used for accurate prediction of breast cancer and therefore would be of no use in screening. Despite this, our work lays the groundwork for building and assessing longitudinal biomarker algorithms that may give an improvement in performance, such as the Risk of Ovarian Cancer Algorithm used within the UKCTOCS (Menon et al, 2015).
References
Baum M (2010) Should routine screening by mammography be replaced by a more selective service of risk assessment/risk management? Womens Health (Lond) 6: 71–76.
Bouchard D, Morisset D, Bourbonnais Y, Tremblay GM (2006) Proteins with whey-acidic-protein motifs and cancer. Lancet Oncol 7: 167–174.
Boyd NF, Guo H, Martin LJ, Sun L, Stone J, Fishell E, Jong RA, Hislop G, Chiarelli A, Minkin S, Yaffe MJ (2007) Mammographic density and the risk and detection of breast cancer. N Engl J Med 356: 227–236.
Bramwell VH, Tuck AB, Chapman JA, Anborgh PH, Postenka CO, Al-Katib W, Shepherd LE, Han L, Wilson CF, Pritchard KI, Pollak MN, Chambers AF (2014) Assessment of osteopontin in early breast cancer: correlative study in a randomised clinical trial. Breast Cancer Res 16: R8.
Cheng Q, Chang JT, Geradts J, Neckers LM, Haystead T, Spector NL, Lyerly HK (2012) Amplification and high-level expression of heat shock protein 90 marks aggressive phenotypes of human epidermal growth factor receptor 2 negative breast cancer. Breast Cancer Res 14: R62.
Chung L, Moore K, Phillips L, Boyle FM, Marsh DJ, Baxter RC (2014) Novel serum protein biomarker panel revealed by mass spectrometry and its prognostic value in breast cancer. Breast Cancer Res 16: R63.
DeLong ER, DeLong DM, Clarke-Pearson DL (1988) Comparing the areas under two or more correlated receiver operating characteristic curves: a nonparametric approach. Biometrics 44: 837–845.
Devetyarov D, Nouretdinov I, Burford B, Camuzeaux S, Gentry-Maharaj A, Tiss A, Smith C, Luo Z, Chervonenkis A, Hallett R, Vovk V, Waterfield M, Cramer R, Timms JF, Sinclair J, Menon U, Jacobs I, Gammerman A (2012) Conformal predictors in early diagnostics of ovarian and breast cancers. Prog Artif Intell 1: 245–257.
Duffy MJ, McGowan PM, Harbeck N, Thomssen C, Schmitt M (2014) uPA and PAI-1 as biomarkers in breast cancer: validated for clinical use in level-of-evidence-1 studies. Breast Cancer Res 16: 428.
Duran MC, Vega F, Moreno-Bueno G, Artiga MJ, Sanchez L, Palacios J, Ridley A, Timms JF (2008) Characterisation of tumoral markers correlated with ErbB2 (HER2/Neu) overexpression and metastasis in breast cancer. Proteomics Clin Appl 2: 1313–1326.
Ebeling FG, Stieber P, Untch M, Nagel D, Konecny GE, Schmitt UM, Fateh-Moghadam A, Seidel D (2002) Serum CEA and CA 15-3 as prognostic factors in primary breast cancer. Br J Cancer 86: 1217–1222.
Fedarko NS, Jain A, Karadag A, Van Eman MR, Fisher LW (2001) Elevated serum bone sialoprotein and osteopontin in colon, breast, prostate, and lung cancer. Clin Cancer Res 7: 4060–4066.
Geraghty JG, Coveney EC, Sherry F, O’Higgins NJ, Duffy MJ (1992) CA 15-3 in patients with locoregional and metastatic breast carcinoma. Cancer 70: 2831–2834.
Gion M, Boracchi P, Dittadi R, Biganzoli E, Peloso L, Mione R, Gatti C, Paccagnella A, Marubini E (2002) Prognostic role of serum CA15.3 in 362 node-negative breast cancers. An old player for a new game. Eur J Cancer 38: 1181–1188.
Gonzalez RM, Daly DS, Tan R, Marks JR, Zangar RC (2011) Plasma biomarker profiles differ depending on breast cancer subtype but RANTES is consistently increased. Cancer Epidemiol Biomarkers Prev 20: 1543–1551.
Gotzsche PC, Jorgensen KJ (2013) Screening for breast cancer with mammography. Cochrane Database Syst Rev 6: CD001877.
Harris L, Fritsche H, Mennel R, Norton L, Ravdin P, Taube S, Somerfield MR, Hayes DF, Bast RC Jr (2007) American Society of Clinical Oncology 2007 update of recommendations for the use of tumor markers in breast cancer. J Clin Oncol 25: 5287–5312.
Hayes DF, Zurawski VR Jr, Kufe DW (1986) Comparison of circulating CA15-3 and carcinoembryonic antigen levels in patients with breast cancer. J Clin Oncol 4: 1542–1550.
Hu Y, Sun H, Drake J, Kittrell F, Abba MC, Deng L, Gaddis S, Sahin A, Baggerly K, Medina D, Aldaz CM (2004) From mice to humans: identification of commonly deregulated genes in mammary cancer via comparative SAGE studies. Cancer Res 64: 7748–7755.
Jatoi I (1999) Breast cancer screening. Am J Surg 177: 518–524.
Key TJ, Appleby PN, Reeves GK, Roddam AW (2010) Insulin-like growth factor 1 (IGF1), IGF binding protein 3 (IGFBP3), and breast cancer risk: pooled individual data analysis of 17 prospective studies. Lancet Oncol 11: 530–542.
Kim BK, Lee JW, Park PJ, Shin YS, Lee WY, Lee KA, Ye S, Hyun H, Kang KN, Yeo D, Kim Y, Ohn SY, Noh DY, Kim CW (2009) The multiplex bead array approach to identifying serum biomarkers associated with breast cancer. Breast Cancer Res 11: R22.
Kiseleva EM, Shor NZ (2005) Continuous Problems of Optimal Set Partitioning: Theory, Algorithms, Applications. Naukova Dumka: Kyiv.
Lee HB, Kang UB, Moon HG, Lee J, Lee KM, Yi M, Park YS, Lee JW, Yu JH, Choi SH, Cho SH, Lee C, Han W, Noh DY (2015) Development and validation of a novel plasma protein signature for breast cancer diagnosis by using multiple reaction monitoring-based mass spectrometry. Anticancer Res 35: 6271–6279.
Loddo M, Andryszkiewicz J, Rodriguez-Acebes S, Stoeber K, Jones A, Dafou D, Apostolidou S, Wollenschlaeger A, Widschwendter M, Sainsbury R, Tudzarova S, Williams GH (2014) Pregnancy-associated plasma protein A regulates mitosis and is epigenetically silenced in breast cancer. J Pathol 233: 344–356.
Look MP, van Putten WL, Duffy MJ, Harbeck N, Christensen IJ, Thomssen C, Kates R, Spyratos F, Ferno M, Eppenberger-Castori S, Sweep CG, Ulm K, Peyrat JP, Martin PM, Magdelenat H, Brunner N, Duggan C, Lisboa BW, Bendahl PO, Quillien V, Daver A, Ricolleau G, Meijer-van Gelder ME, Manders P, Fiets WE, Blankenstein MA, Broet P, Romain S, Daxenbichler G, Windbichler G, Cufer T, Borstnar S, Kueng W, Beex LV, Klijn JG, O’Higgins N, Eppenberger U, Janicke F, Schmitt M, Foekens JA (2002) Pooled analysis of prognostic impact of urokinase-type plasminogen activator and its inhibitor PAI-1 in 8377 breast cancer patients. J Natl Cancer Inst 94: 116–128.
Mansfield AS, Visscher DW, Hart SN, Wang C, Goetz MP, Oxvig C, Conover CA (2014) Pregnancy-associated plasma protein-A expression in human breast cancer. Growth Horm IGF Res 24: 264–267.
McCarthy K, Laban C, McVittie CJ, Ogunkolade W, Khalaf S, Bustin S, Carpenter R, Jenkins PJ (2009) The expression and function of IGFBP-3 in normal and malignant breast tissue. Anticancer Res 29: 3785–3790.
Menon U, Gentry-Maharaj A, Hallett R, Ryan A, Burnell M, Sharma A, Lewis S, Davies S, Philpott S, Lopes A, Godfrey K, Oram D, Herod J, Williamson K, Seif MW, Scott I, Mould T, Woolas R, Murdoch J, Dobbs S, Amso NN, Leeson S, Cruickshank D, McGuire A, Campbell S, Fallowfield L, Singh N, Dawnay A, Skates SJ, Parmar M, Jacobs I (2009) Sensitivity and specificity of multimodal and ultrasound screening for ovarian cancer, and stage distribution of detected cancers: results of the prevalence screen of the UK Collaborative Trial of Ovarian Cancer Screening (UKCTOCS). Lancet Oncol 10: 327–340.
Menon U, Gentry-Maharaj A, Ryan A, Sharma A, Burnell M, Hallett R, Lewis S, Lopez A, Godfrey K, Oram D, Herod J, Williamson K, Seif M, Scott I, Mould T, Woolas R, Murdoch J, Dobbs S, Amso N, Leeson S, Cruickshank D, McGuire A, Campbell S, Fallowfield L, Skates S, Parmar M, Jacobs I (2008) Recruitment to multicentre trials—lessons from UKCTOCS: descriptive study. BMJ 337: a2079.
Menon U, Ryan A, Kalsi J, Gentry-Maharaj A, Dawnay A, Habib M, Apostolidou S, Singh N, Benjamin E, Burnell M, Davies S, Sharma A, Gunu R, Godfrey K, Lopes A, Oram D, Herod J, Williamson K, Seif MW, Jenkins H, Mould T, Woolas R, Murdoch JB, Dobbs S, Amso NN, Leeson S, Cruickshank D, Scott I, Fallowfield L, Widschwendter M, Reynolds K, McGuire A, Campbell S, Parmar M, Skates SJ, Jacobs I (2015) Risk algorithm using serial biomarker measurements doubles the number of screen-detected cancers compared with a single-threshold rule in the United Kingdom Collaborative Trial of ovarian cancer screening. J Clin Oncol 33 (18): 2062–2071.
Nauck D, Klawonn F, Kruse R (1997) Foundations of Neuro-Fuzzy systems. John Wiley & Sons: New York, NY, USA.
Niwa Y, Akamatsu H, Niwa H, Sumi H, Ozaki Y, Abe A (2001) Correlation of tissue and plasma RANTES levels with disease course in patients with breast or cervical cancer. Clin Cancer Res 7: 285–289.
Opstal-van Winden AW, Krop EJ, Karedal MH, Gast MC, Lindh CH, Jeppsson MC, Jonsson BA, Grobbee DE, Peeters PH, Beijnen JH, van Gils CH, Vermeulen RC (2011) Searching for early breast cancer biomarkers by serum protein profiling of pre-diagnostic serum; a nested case–control study. BMC Cancer 11: 381.
Opstal-van Winden AW, Rodenburg W, Pennings JL, van Oostrom CT, Beijnen JH, Peeters PH, van Gils CH, de Vries A (2012) A bead-based multiplexed immunoassay to evaluate breast cancer biomarkers for early detection in pre-diagnostic serum. Int J Mol Sci 13: 13587–13604.
Pace LE, Keating NL (2014) A systematic assessment of benefits and risks to guide breast cancer screening decisions. JAMA 311: 1327–1335.
Paiva RP, Dourado A (2004) Interpretability and learning in neuro-fuzzy systems. Fuzzy Sets Syst 147: 17–38.
Pick E, Kluger Y, Giltnane JM, Moeder C, Camp RL, Rimm DL, Kluger HM (2007) High HSP90 expression is associated with decreased survival in breast cancer. Cancer Res 67: 2932–2937.
Shao Y, Sun X, He Y, Liu C, Liu H (2015) Elevated levels of serum tumor markers CEA and CA15-3 are prognostic parameters for different molecular subtypes of breast cancer. PLoS One 10: e0133830.
Siegel R, Ma J, Zou Z, Jemal A (2014) Cancer statistics, 2014. CA Cancer J Clin 64: 9–29.
Singhal H, Bautista DS, Tonkin KS, O’Malley FP, Tuck AB, Chambers AF, Harris JF (1997) Elevated plasma osteopontin in metastatic breast cancer associated with increased tumor burden and decreased survival. Clin Cancer Res 3: 605–611.
Soria G, Ben-Baruch A (2008) The inflammatory chemokines CCL2 and CCL5 in breast cancer. Cancer Lett 267: 271–285.
Stieber P, Molina R, Chan DW, Fritsche HA, Beyrau R, Bonfrer JM, Filella X, Gornet TG, Hoff T, Jager W, van Kamp GJ, Nagel D, Peisker K, Sokoll LJ, Troalen F, Untch M, Domke I (2003) Clinical evaluation of the Elecsys CA 15-3 test in breast cancer patients. Clin Lab 49: 15–24.
Welch HG, Prorok PC, O’Malley AJ, Kramer BS (2016) Breast-cancer tumor size, overdiagnosis, and mammography screening effectiveness. N Engl J Med 375: 1438–1447.
Worthington J, Bertani M, Chan HL, Gerrits B, Timms JF (2010) Transcriptional profiling of ErbB signalling in mammary luminal epithelial cells—interplay of ErbB and IGF1 signalling through IGFBP3 regulation. BMC Cancer 10: 490.
Zagouri F, Sergentanis TN, Provatopoulou X, Kalogera E, Chrysikos D, Lymperi M, Papadimitriou CA, Zografos E, Bletsa G, Kalles VS, Zografos GC, Gounaris A (2011) Serum levels of HSP90 in the continuum of breast ductal and lobular lesions. In Vivo 25: 669–672.
Acknowledgements
This research was funded by the National Institute for Health Research (NIHR) University College London Hospitals (UCLH) Biomedical Research Centre. UKCTOCS was core funded by the Medical Research Council, Cancer Research UK, and the Department of Health with additional support from the Eve Appeal, Special Trustees of Bart’s and the London, and Special Trustees of UCLH. OB and JFT also received support from the Eve Appeal Gynaecological Cancer Research Trust and Cancer Research UK PRC Programme Grant A12677.
Author contributions
AK and GM carried out ELISA assays; OB, EK, OMP and JFT carried out statistical analyses and multi-marker model construction; AG-M and AR identified cases from UKCTOCS and retrieved patient data; IJJ, UM and JFT conceived and designed the study; AK, MW and JFT contributed to the drafting of the manuscript. All authors read and approved the final manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
UM and IJJ have financial interest through UCL Business and Abcodia in the third party exploitation of clinical trials biobanks which have been developed through research at UCL. None of the other authors have competing interests.
Rights and permissions
This work is licensed under the Creative Commons Attribution 4.0 International License. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Kazarian, A., Blyuss, O., Metodieva, G. et al. Testing breast cancer serum biomarkers for early detection and prognosis in pre-diagnosis samples. Br J Cancer 116, 501–508 (2017). https://doi.org/10.1038/bjc.2016.433
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2016.433
Keywords
This article is cited by
-
One-pot wet-chemical fabrication of 3D urchin-like core-shell Au@PdCu nanocrystals for electrochemical breast cancer immunoassay
Microchimica Acta (2023)
-
Circulating proteins as predictive and prognostic biomarkers in breast cancer
Clinical Proteomics (2022)
-
Identification and validation of plasma biomarkers for diagnosis of breast cancer in South Asian women
Scientific Reports (2022)
-
Non-invasive optical characterization and detection of CA 15–3 breast cancer biomarker in blood serum using monoclonal antibody-conjugated gold nanourchin and surface-enhanced Raman scattering
Lasers in Medical Science (2022)
-
Design and Fabrication of a DNA-copper Nanocluster-based Biosensor for Multiple Detections of Circulating miRNAs in Early Screening of Breast Cancer
Journal of Fluorescence (2022)