Abstract
Background:
In this study, we evaluated the ability of gene expression profiles to predict chemotherapy response and survival in triple-negative breast cancer (TNBC).
Methods:
Gene expression and clinical–pathological data were evaluated in five independent cohorts, including three randomised clinical trials for a total of 1055 patients with TNBC, basal-like disease (BLBC) or both. Previously defined intrinsic molecular subtype and a proliferation signature were determined and tested. Each signature was tested using multivariable logistic regression models (for pCR (pathological complete response)) and Cox models (for survival). Within TNBC, interactions between each signature and the basal-like subtype (vs other subtypes) for predicting either pCR or survival were investigated.
Results:
Within TNBC, all intrinsic subtypes were identified but BLBC predominated (55–81%). Significant associations between genomic signatures and response and survival after chemotherapy were only identified within BLBC and not within TNBC as a whole. In particular, high expression of a previously identified proliferation signature, or low expression of the luminal A signature, was found independently associated with pCR and improved survival following chemotherapy across different cohorts. Significant interaction tests were only obtained between each signature and the BLBC subtype for prediction of chemotherapy response or survival.
Conclusions:
The proliferation signature predicts response and improved survival after chemotherapy, but only within BLBC. This highlights the clinical implications of TNBC heterogeneity, and suggests that future clinical trials focused on this phenotypic subtype should consider stratifying patients as having BLBC or not.
Similar content being viewed by others
Main
Triple-negative breast cancer (TNBC) is characterised by the absence of the therapeutically targetable hormone receptors and HER2 protein overexpression. For this reason, both adjuvant treatment and palliative therapy for metastatic TNBC is limited to chemotherapy. Although TNBC typically has higher rates of chemosensitivity compared with hormone receptor-positive breast cancer, it has a poor overall prognosis (Carey et al, 2007; Liedtke et al, 2008; Silver et al, 2010a) and there is no predictive biomarker of response or survival to allow tailored therapy for these patients.
Over the years, studies based on global gene expression analyses have identified five main intrinsic molecular subtypes of breast cancer known as luminal A, luminal B, HER2 enriched, basal like and claudin low (Perou et al, 2000; Sorlie et al, 2001; Prat et al, 2010, 2013a, 2013b; Prat and Perou, 2011). Among them, the basal-like subtype (BLBC) comprises the majority of TNBC; however, the other 20–30% of TNBCs fall into other subtypes (Prat and Perou, 2011). Thus, significant molecular heterogeneity exists within TNBC and it is likely that improving clinical outcome and tailoring therapy will require further stratification by biologic subtype (Prat et al, 2013a).
In this study, we used gene expression data to classify multiple independent cohorts of patients from cooperative group trials and large multi-institution data sets into the main intrinsic molecular subtypes of breast cancer and then we evaluated the ability of various published gene expression profiles to predict response and/or survival following chemotherapy in TNBC and/or BLBC.
Materials and Methods
Patients, samples and clinical data
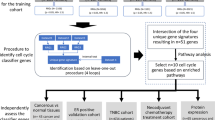
Multiple data sets of TNBC or BLBC were evaluated (Table 1 and Supplementary Figure S1). For response prediction, we evaluated samples at diagnosis from two independent cohorts of patients treated with anthracycline/taxane-based chemotherapy in the neoadjuvant setting: the GEICAM/2006-03 Core-Basal phase II clinical trial (Alba et al, 2012) and a combined cohort of microarray studies previously published by the MDACC group (GSE25066 (Hatzis et al, 2011), GSE16716 (Popovici et al, 2010), GSE20271 (Tabchy et al, 2010), GSE23988 (Iwamoto et al, 2011) and MDACC133 (Hess et al, 2006)). For survival prediction, we evaluated samples from three independent cohorts of patients with primary breast cancer: the GEICAM/9906 and CALGB/9741 phase III clinical trials (Citron et al, 2003; Martín et al, 2008), and the METABRIC data set, which is a UK/Canadian cohort of nearly 2000 primary breast cancers with transcriptomic and outcome data (Curtis et al, 2012).
Characteristics of the patient populations evaluated have previously been described (Citron et al, 2003; Martín et al, 2008; Cheang et al, 2009; Nielsen et al, 2010; Alba et al, 2012) and are summarised in Table 1. The neoadjuvant cohorts had pathological complete response (pCR) in the breast (GEICAM/2006-03) and breast and axilla (MDACC based) as the primary end points. Patients in the GEICAM/2006-03 trial (NCT00432172) were randomised to neoadjuvant epirubicin/cyclophosphamide followed by docetaxel+/−carboplatin (Alba et al, 2012). In the MDACC-based cohort (Hess et al, 2006; Popovici et al, 2010; Tabchy et al, 2010; Hatzis et al, 2011; Iwamoto et al, 2011), all patients received neoadjuvant anthracycline/taxane-based chemotherapy. The adjuvant cohorts had disease-free survival (DFS), disease-specific survival (DSS) or relapse-free survival (RFS) as end points, and included GEICAM/9906 (Martín et al, 2008), in which patients with node-positive disease were randomly assigned to adjuvant 5-fluororacil, epirubicin and cyclophosphamide (FEC) for six cycles vs FEC for four cycles followed by weekly paclitaxel for eight cycles, and CALGB/9741 (Citron et al, 2003), in which patients with node-positive disease were randomly assigned to receive dose dense (every 2 weeks) vs conventional dosing (every 3 weeks) doxorubicin and (or followed by) cyclophosphamide followed by paclitaxel. The METABRIC cohort (Curtis et al, 2012) included patients who received either no adjuvant systemic therapy (AST) or adjuvant chemotherapy, although the exact regimens, doses and schedules are not available.
TNBC definition
The TNBC definition and cut points for oestrogen receptor (ER), progesterone receptor (PR) and HER2 were according to the 2007 and 2010 ASCO/CAP guidelines for HER2 (Wolff et al, 2006) and ER/PR (Hammond et al, 2010), respectively, in GEICAM/2006-03, MDACC-based and GEICAM/9906 data sets. In GEICAM/2006-03 and GEICAM/9906, the TNBC definition as well as Ki-67 immunohistochemical determination (clone MIB-1, DAKO, Carpinteria, CA, USA) were performed at a central laboratory and reviewed by two expert pathologists. In METABRIC and CALGB/9741, the pathological data were not centrally reviewed; thus, we decided to focus on those samples identified as BLBC by gene expression data.
Gene expression data
From GEICAM/2006-03, the PAM50 and claudin-low signatures were derived from a 543-gene set measured using the Nanostring nCounter platform (Nanostring Technologies, Seattle, WA, USA) from formalin-fixed paraffin-embedded (FFPE) tumour samples. For each sample, two 1 mm cores enriched with tumour tissue were obtained from the original tumour block, RNA was purified and ∼100 ng of total was used to measure gene expression. Data were log base 2 transformed and normalised using five house-keeping genes (ACTB, MRPL19, PSMC4, RPLP0 and SF3A1). Raw gene expression data have been deposited in Gene Expression Omnibus (GSE58479).
From the MDACC Affymetrix (Affymetrix Inc., Santa Clara, CA, USA) U133A-based microarray cohort, publicly available gene expression data were obtained and normalised using MAS5 as previously reported (Usary et al, 2013; Prat et al, 2013a). From the METABRIC cohort, normalised microarray data were obtained from the European Genome-Phenome Archive (accession number: EGAS00000000083; Curtis et al, 2012). From GEICAM/9906, expression of the 50 PAM50 genes was measured using the qRT–PCR-based version as described previously (Bastien et al, 2012). Finally, expression of the 50 PAM50 genes, and the same five house-keeping genes used in GEICAM2006-03, was measured from CALGB/9741 using the nCounter platform from FFPE primary tumours (Liu et al, 2012).
Subtypes and gene signatures
All tumours were assigned to an intrinsic molecular subtype of breast cancer (luminal A, luminal B, HER2 enriched, BLBC and claudin low) and the normal-like group using the PAM50 subtype predictor and the claudin-low predictor (Parker et al, 2009; Nielsen et al, 2010; Prat et al, 2010), except for the GEICAM/9906 and CALGB/9741 data sets, in which only PAM50 50-gene data were available (Martín et al, 2008; Bastien et al, 2012). Of note, the same PAM50 and claudin-low training data sets (Parker et al, 2009; Prat et al, 2010) were used for subtype prediction in each test set.
Before subtyping, each individual data set was normalised accordingly (Perou et al, 2010). For GEICAM2006-03 and CALGB9741 data sets, both nCounter based, we had groups of tumour samples representative of each intrinsic subtype, which allowed us to estimate the platform-to-platform bias. For MDACC-based, METABRIC and GEICAM/9906 data sets, all of which have a large number of samples representative of all the intrinsic subtypes, we assumed that differences in the median expression of each gene were due to technical factors. Our subtype calls in MDACC-based and METABRIC data sets were highly concordant (kappa score 0.83 and 0.78) with the ones reported by other investigators in their original publications (Hatzis et al, 2011; Curtis et al, 2012). PAM50 output data for these two data sets are provided in Supplementary Data. To further support that our normalisation approach was appropriate, we have provided a PC1 vs PC2 loading plot of the PAM50 genes and the different data sets evaluated (Supplementary Figure S2), including the PAM50 training data set used in all predictions.
Gene signatures evaluated were obtained from the PAM50 (n=8 signatures) and claudin-low (n=1) subtype predictors (Parker et al, 2009; Nielsen et al, 2010; Prat et al, 2010); from the PAM50 predictor, we evaluated the correlation coefficient of each sample to each subtype centroid (luminal A, luminal B, HER2 enriched, BLBC and normal). In addition, we also evaluated the previously reported proliferation score, which is the mean expression of 11 proliferation-related genes (Nielsen et al, 2010), and the risk of relapse (ROR) score, a continuous prognostic score based on subtype only (ROR-S), and based on subtype and proliferation (ROR-P; Nielsen et al, 2010). Finally, from the claudin-low predictor, we evaluated the Euclidian distance of each sample to the claudin-low 9-cell line centroid.
Statistical analysis
Biologic analysis of gene lists was performed with DAVID annotation tool (http://david.abcc.ncifcrf.gov/; Dennis et al, 2003). Association between the expression of each signature and pCR was assessed by logistic regression analyses adjusted for standard clinical–pathological variables. Survival functions to DSS, DFS and RFS were from the Kaplan–Meier product-limit estimator, with tests of differences by the log-rank test. Cox proportional hazard models adjusted for standard clinical–pathological variables were used to test the independent associations with survival of each gene or signature. All genes and gene signatures were evaluated as continuous variables. Finally, interaction tests between the expression of each signature and the BLBC (vs others) for pCR or survival outcome prediction were also evaluated in multivariable models.
Results
Biologic heterogeneity within TNBC
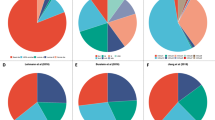
To identify biological entities within TNBC, intrinsic molecular subtyping was performed on three different and independent cohorts (GEICAM/2006-03, MDACC based and GEICAM/9906) of 357 patients with primary TNBC (Table 1). All the intrinsic molecular subtypes were identified within TNBC, with BLBCs predominating (55–81.2%). Of note, in GEICAM/2006-03, eligibility included the ‘core basal’ definition (Nielsen et al, 2004), a more restrictive TNBC definition (that is, TNBC status plus immunostain positive for either cytokeratin 5/6 or EGFR). This cohort showed a significant enrichment for BLBCs compared with the triple-negative-only definition (81.2 vs 56.9%, P<0.001, χ2-test).
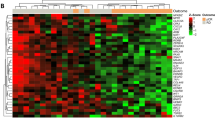
To further explore the heterogeneity of BLBC (Supplementary Figure S3), we focused on the 56 tumours that were both TNBC and of the BLBC in the GEICAM/2006-03 cohort. Within this group, we identified 159 (29.7%) genes whose expression was found to have high variability (that is, >two-fold s.d.; Supplementary Table S1). Of these genes, 87 (54.7%) were also found to have high variability in TNBCs of the BLBC in the MDACC cohort. Among these, we identified a significant enrichment for genes involved in regulation of cell death (for example, CRYAB and INHBA), response to hormone stimulus (for example, FOXA1 and KRT19), cell migration (for example, FOXC1, IL8 and KIT), ectoderm development (for example, KRT14 and KRT5) and cell cycle (for example, CCND1, TP53 and BRCA1). In the GEICAM/2006-03 cohort, we also observed a wide range of Ki-67-positive tumour cells (35–95%; average 74%) in TNBC of the BLBC. Counter to prevailing dogma that all BLBCs are highly proliferative, there was a wide range of proliferation values within TNBC of the BLBC.
Predicting pCR after chemotherapy within TNBC
To identify predictors of chemotherapy response, we first evaluated gene expression-based signatures in the diagnostic (that is, pre-treatment) samples of the GEICAM/2006-03 trial (Alba et al, 2012). Among all patients, none of the signatures or clinical–pathological variables evaluated was found significantly associated with pCR (Figure 1A). Conversely, among patients with BLBC, high expression of the proliferation score, low expression of the luminal A signature and high Ki-67 by IHC were found to be significantly associated with pCR (Figure 1B). Interaction tests between each of these variables and the BLBC (vs others) for pCR showed a trend towards statistical significance for the luminal A signature (inverse relationship, P=0.066), the proliferation score (P=0.062) and no evidence of interaction with Ki-67 (P=0.372).
Adjusted odds ratios (ORs) for pathologic complete response (pCR) of various clinical–pathological variables and gene signatures (for unit increase) in (A) TNBC patients in GEICAM/2006-03, (B) TNBC and BLBC in GEICAM/2006-03, (C) TNBC patients in MDACC and (D) TNBC and BLBC in MDACC. The PAM50-based signatures represent either a correlation coefficient to a gene expression centroid (for basal like, HER2 enriched, luminal A, luminal B and normal) or a score (for RORS, proliferation score and RORP), and they are evaluated as continuous variables. The claudin-high signature represents an Euclidean distance to the claudin-low centroid, and it is evaluated as a continuous variable. Each signature has been standardised to have a mean of 0 and a s.d. of 1. The size of the square is inversely proportional to the s.e.; horizontal bars represent the 95% CIs of ORs. Statistically significant variables are shown in blue. Each gene signature has been evaluated individually after adjustment for standard clinical–pathological variables. The variables used for adjustment were treatment arm, age at diagnosis, nodal status and tumour size (GEICAM/2006-03); and tumour size, age at diagnosis and histological grade (MDACC).
To confirm the findings obtained from GEICAM/2006-03, we interrogated 188 TNBC patients from the combined MDACC data sets treated neoadjuvantly with anthracycline/taxane-based chemotherapy. Similar to the data obtained in GEICAM/2006-03, none of the signatures was significantly associated with pCR within TNBC (Figure 1C); however, the proliferation score and luminal A signature were again significantly associated with pCR within patients with TNBC of the BLBC (Figure 1D). Interaction tests within TNBC between each signature and the BLBC (vs others) for pCR were statistically significant (luminal A P=0.023; proliferation score P=0.005). Similar results were obtained in terms of DFS (Supplementary Figure S4). Ki-67 IHC data were not available from this data set.
Biological processes associated with pCR within TNBC of the BLBC subtype
To identify single genes whose expression might be associated with pCR, we evaluated the 56 TNBC of the BLBC subtype from the GEICAM/2006-03 trial. Among 535 genes assayed, the expression of 82 (15.3%) genes was significantly associated with pCR (Supplementary Table S2–4). Concordant with the previous findings using gene signatures, high expression of cell cycle-related genes (for example, CCNE1 and FANCA) and low expression of endocrine response-related genes (for example, PGR and FOXA1) were found significantly associated with pCR. In contrast, high expression of genes involved in epithelial-to-mesenchymal transition (for example, TWIST1 and ZEB1) was found to be significantly associated with lack of response (that is, residual disease).
To further explore these findings, we combined gene expression and clinical–pathological data of TNBC of the BLBC subtype from the GEICAM/2006-03 and MDACC-based cohorts (Supplementary Figure S5) and performed a multivariable logistic regression analysis for pCR prediction. Among 479 genes in common between these two platforms, 94 genes (19.6%) were found independently associated with pCR (Supplementary Data). Of these, 35 (37.2%) genes were identical to those identified in the GEICAM/2006-03 trial. Finally, the remainder (n=59 genes, 62.8%) tracked similar biology with endocrine response-related genes (for example, CCND1 and IL6) and cell cycle-related genes (for example, CHEK1 and CCNB1) being associated with chemoresistance and chemosensitivity, respectively.
Predicting DFS within TNBC
To evaluate whether the gene signatures predict survival, we evaluated clinical–pathological data and PAM50 50-gene qRT–PCR data from the GEICAM/9906 phase III trial (Martín et al, 2008), which included 100 patients with centrally reviewed TNBCs that received adjuvant chemotherapy. Within TNBC, no significant predictor of DFS was identified (Figure 2A). Conversely, within TNBCs of the BLBC subtype, the two previously identified signatures (that is, high expression of the proliferation score and low expression of luminal A signature) were found significantly associated with DFS, whereas Ki-67 by IHC was not (Figure 2B). Interaction tests within TNBC between each signature and the BLBC subtype (vs others) for DFS were statistically significant (luminal A P=0.035; proliferation score P=0.017).
Adjusted survival HRs of various clinical–pathological variables and gene signatures (for unit increase) in (A) TNBC and (B) TNBC and BLBC treated with adjuvant chemotherapy in GEICAM/9906, (C) BLBC not treated with adjuvant chemotherapy in METABRIC and (D) BLBC treated with adjuvant chemotherapy in METABRIC. The PAM50-based signatures represent either a correlation coefficient to a gene expression centroid (for basal like, HER2 enriched, luminal A, luminal B and normal) or a score (for RORS, proliferation score and RORP), and they are evaluated as continuous variables. The claudin-high signature represents an Euclidean distance to the claudin-low centroid, and it is evaluated as a continuous variable. Each signature has been standardised to have a mean of 0 and a s.d. of 1. The size of the square is inversely proportional to the s.e.; horizontal bars represent the 95% CIs of HRs. Statistically significant variables are shown in blue. Each gene signature has been evaluated individually after adjustment for standard clinical–pathological variables. The variables used for adjustment were treatment arm, age at diagnosis, nodal status and tumour size (GEICAM/9906); and tumour size, age at diagnosis and nodal status (METABRIC).
To illustrate the results obtained in the GEICAM/9906 data set (Martín et al, 2008), we split the TNBC cohort, as well as the cohort of TNBCs of the BLBC subtype, into tertile groups based on the expression of the proliferation score. As expected, within the TNBC cohort, the ordinal proliferation score groups were not significantly associated with DFS (Figure 3A). Conversely, within those patients with TNBCs of the BLBC subtype, the proliferation tertile groups were found associated with DFS (Figure 3B) with an adjusted hazard ratio (HR) between the high vs low proliferation score tertile groups of 0.194 (0.056–0.669 95% confidence interval (CI)).
Kaplan–Meier survival analysis in GEICAM/9906 and METABRIC data sets based on the PAM50 proliferation score. Patients with (A) TNBC and (B) TNBC and BLBC treated with adjuvant chemotherapy in GEICAM/9906, (C) BLBC not treated with adjuvant systemic chemotherapy (no AST) in METABRIC and (D) BLBC treated with adjuvant chemotherapy (CT) in METABRIC.
Predicting DSS within BLBC
We next evaluated gene expression and clinical–pathological data in the recently reported METABRIC cohort, which includes 185 and 199 patients with primary BLBC disease that received adjuvant chemotherapy and no AST, respectively (Curtis et al, 2012). Interestingly, in the no AST subgroup of patients, no genomic variable was found significantly associated with DSS (Figure 2C). Conversely, within the chemotherapy-treated subset of patients, proliferation score and the luminal A signature were found significantly associated with outcome (Figure 2D). Interaction tests within BLBC disease between each signature and the type of treatment (chemotherapy vs no AST) for DSS were found to be statistically significant (luminal A P=0.014; proliferation score P=0.047). In this data set, exclusion of the claudin-low samples did not alter the survival associations.
To further illustrate these findings, we split the BLBC/no AST subset of patients, and the BLBC/chemotherapy-treated subset of patients, into groups based on the expression of the proliferation score. As expected, within the no AST subset of patients, the groups were not found associated with DSS (Figure 3C) with an adjusted HR of the high vs low tertile group of 1.456 (0.680–3.209 95% CI). Conversely, within the chemotherapy-treated subset of patients, the groups were found associated with DSS (Figure 3D) with an adjusted HR between the high vs low proliferation score groups of 0.410 (0.208–0.808 95% CI).
Independent evaluation in the CALGB/9741 clinical trial
Finally, we evaluated the association of gene expression and clinical–pathological data with RFS in 314 patients with primary BLBC from the CALGB/9741 study (Citron et al, 2003), in which patients were treated with adjuvant anthracycline/taxane-based chemotherapy. As in the previous cohorts, similar associations were observed with low expression of the luminal A signature (P=0.038) and high expression of the proliferation score (P=0.067) and ROR-P (P=0.0493) signatures being associated with better outcome (Figure 4A). To illustrate these results, we split patients into tertiles based on the expression of the proliferation score and plotted their RFS (Figure 4B). The adjusted HR between the high proliferation score group vs the intermediate and low groups was 0.509 (0.321–0.808 95% CI) and 0.611 (0.380–0.984 95% CI), respectively.
Associations of clinical–pathological variables and gene signatures with RFS in 314 patients with BLBC of the CALGB/9741 cohort. (A) Adjusted survival HRs of various clinical–pathological variables and gene signatures (for unit increase). The PAM50-based signatures represent either a correlation coefficient to a gene expression centroid (for basal like, HER2 enriched, luminal A, luminal B and normal) or a score (for RORS, proliferation score and RORP), and they are evaluated as continuous variables. The claudin-high signature represents an Euclidean distance to the claudin-low centroid, and it is evaluated as a continuous variable. Each signature has been standardised to have a mean of 0 and a s.d. of 1. The size of the square is inversely proportional to the s.e.; horizontal bars represent the 95% CIs of HRs. Statistically significant variables are shown in blue. The variables used for adjustment were treatment arm, age at diagnosis, nodal status and tumour size. (B) Kaplan–Meier survival analysis in the subset of patients with BLBC in the CALGB/9741 data set based on the PAM50 proliferation score (using tertiles).
Discussion
Over the years, BLBC has become more commonly known as TNBC; however, not all TNBCs are identified as BLBC by gene expression (Prat and Perou, 2011). In addition, the proportion of BLBCs within TNBC varies substantially from cohort to cohort (Prat and Perou, 2011; Table 1). Although this discrepancy could be explained by false-negative immunohistochemical results, the identification of all the intrinsic subtypes within centrally reviewed TNBC argues otherwise (Cheang et al, 2012). A potential explanation is that determination of three biomarkers for identifying the known intrinsic subtypes captures an inferior amount of biological diversity compared with assays that measure tens or hundreds of genes (Prat et al, 2012). It is now clear that significant biological heterogeneity exists within patients diagnosed with TNBC and additional efforts are needed to elucidate the clinical heterogeneity of this aggressive disease subtype as defined by these three biomarkers.
To address this need, we evaluated a selection of known prognostic signatures in patients with TNBC who received multiagent chemotherapy. In addition, we did so using multiple independent data sets and multiple genomic technologies. We found that all data sets and all technology platforms yielded the same result, namely, that only if we stratified TNBC into BLBC could we then identify patients with significantly better response (that is, pCR) and better survival using the evaluated biomarkers (that is, proliferation and correlation to LumA centroid); however, when using all TNBC patients, these predictive biomarkers were not useful, thus showing the value of first stratifying by biological group, and then second, by biomarker.
The association between high proliferation and chemotherapy benefit was not unexpected. Cytotoxic-based therapies are known to be more effective in highly proliferative tumours than in quiescent, low proliferative tumours (Skipper, 1971; Whitfield et al, 2006). Indeed, tumours with a higher percentage of Ki-67-positive tumour cells in the GEICAM/2006-03 trial responded more to chemotherapy than tumours with lower levels, although this association was not significant in interaction testing while the genomic-based proliferation signature was. This is concordant with two previous reports where patients with TNBC and higher percentage of Ki-67-positive tumour cells showed higher pCR rates than those with lower Ki-67 levels (Adamo and Anders, 2011; Fasching et al, 2011; Keam et al, 2011). Supporting these results, Silver et al (2010b) identified a proliferation-related gene signature of the E2F3-related transcription factor that was associated with response to neoadjuvant cisplatin in TNBC, all of which had a BLBC gene expression profile. In addition, in a pooled cohort study (Ignatiadis et al, 2012), the investigators found that this E2F3-related signature and other signatures of chromosomal instability and PTEN loss were associated with increased pCR in ER−/HER2− tumours; however, these signatures were not found significantly associated with pCR in BLBC (Ignatiadis et al, 2012).
Recently, Lehmann et al (2011) identified seven different subtypes of TNBC by global gene expression analysis (basal 1 (BL1), basal 2 (BL2), immunomodulatory (IM), luminal androgen receptor (LAR), mesenchymal (M), mesenchymal stem cell (MSL) and unstable (UNS)). When compared with the intrinsic subtypes of breast cancer, BL1, BL2, IM and M largely fall within the BLBC subtype, whereas LAR largely falls within the non-BLBC subtypes (Lehmann et al, 2011; Prat et al, 2013a; Mayer et al, 2014). The intrinsic subtype distribution within the MSL group is less clear (Lehmann et al, 2011; Mayer et al, 2014). In any case, the seven-subtype classification of TNBC, as a group, has been found associated with pCR in an independent cohort of 143 patients with TNBC treated with anthracycline/taxane-based chemotherapy (Masuda et al, 2013). Among the different subtypes, BL1, which is characterised by high expression of cell cycle-related genes (Lehmann et al, 2011), showed the highest pCR rates (52%), whereas BL2 and LAR subtypes showed the lowest pCR rates (0 and 10%; Masuda et al, 2013). However, no significant association with survival outcomes was observed among the seven subtypes in this chemotherapy-treated data set (Masuda et al, 2013). We were not able to assay these seven subtypes across all our studies presented here, as some of our studies only had data for the 50 PAM50 genes; however, further studies are needed to determine whether the BLBC tumours with high or low expression of the luminal A or proliferation signature belong to one of these potential molecular entities.
The results presented here may have potential clinical implications for the management of patients with TNBC. First, we provide additional evidence that TNBCs are a heterogeneous disease entity and that subclassification of TNBC into phenotypic subsets can help predict chemotherapy response and survival. This argues that clinical trials evaluating TNBC should consider using gene expression data to stratify patients by the different molecular entities given the impact of this heterogeneity on the primary end points of these trials. Second, high-risk tumours of the BLBC subtype with high proliferation scores are very sensitive to chemotherapy and it may be possible that the typical standard of care treatment for this group is effective (anthracycline/taxane neo/adjuvant regimens). If so, future studies might aim to limit toxicity while maintaining or enhancing efficacy. Conversely, those predicted to be less responsive, or to have a worse prognosis, may be appropriate for studies of novel agents or approaches. Some patients might be particularly sensitive to specific additional drugs like platinum agents, which increase pCR rates in TNBC (Sikov et al, 2013; Von Minckwitz et al, 2013), or other cytotoxics, as will be tested in the upcoming CIBOMA/2004-01/GEICAM/2003-11 phase III clinical trial that focuses on adjuvant capecitabine maintenance therapy after conventional induction chemotherapy in 876 patients with TNBC. Novel therapies that target lower-proliferating cells (that is, mesenchymal/claudin-low-like and/or luminal-like cells) might be warranted in patients with TN tumours of the BLBC subtype that show low expression of proliferation features, and also those TNBCs that are of the luminal lineage.
There are several caveats to our study. First, this is a retrospective study involving heterogeneous patient populations and the data need to be confirmed in prospective clinical trial(s). Second, none of the survival data sets came from a randomised clinical trial of adjuvant chemotherapy vs no adjuvant chemotherapy. Thus, the true predictive value of these signatures was only evaluated in the neoadjuvant setting (GEICAM/2006-03 and MDACC), where pCR has been found to be an intermediate biomarker for outcome but one robust enough, particularly in TNBC, to be considered for registrational trials by the FDA. In support of the predictive nature of these signatures, none predicted outcome in patients with BLBCs who did not receive systemic adjuvant therapy in the METABRIC cohort. Third, in the GEICAM/9906 and CALGB/9741 cohorts, we only had PAM50 50-gene data, and thus we were not able to test whether further identification of the claudin-low subtype (and other TNBC signatures) might have improved the survival predictions. Fourth, the common markers of the applied platforms/cohorts are mainly restricted to genes expressed by the carcinoma cells. Thus, the analyses could miss signals from the microenvironment, which have been shown to be predictive both for pCR (Rody et al, 2009; Ignatiadis et al, 2012) and survival (Rody et al, 2011). Finally, patients from each of the data sets received different anthracycline- or anthracycline/taxane-based chemotherapy regimens, schedules and doses, and thus the ability of the signatures to predict response to particular treatment strategies could not be tested.
Another important consideration of our study is that we did not attempt to identify an optimal cutoff for the various biomarkers (that is, signatures) evaluated but rather focused on the association of the continuous expression of each biomarker with each end point. The main reason is that different gene expression-based platforms and protocols were used in each cohort and thus, standardisation of a biomarker cut point would have been difficult to achieve and most likely unreliable. In any case, the fact that all five data sets gave very similar associations, and were found independently of the platform/protocol used, argues in favour of a true biologically based robust finding.
Genomic signatures of proliferation and luminal subtype are associated with response to chemotherapy in the neoadjuvant setting and with outcome following chemotherapy in TNBC, but only within those belonging to the BLBC subtype. TNBC is biologically heterogeneous, and improving therapy may require tumour stratification based on the BLBC vs non-BLBC gene expression profile.
Accession codes
References
Adamo B, Anders C (2011) Stratifying triple-negative breast cancer: which definition(s) to use? Breast Cancer Res 13: 105.
Alba E, Chacon J, Lluch A, Anton A, Estevez L, Cirauqui B, Carrasco E, Calvo L, Segui M, Ribelles N, Alvarez R, Sanchez-Muñoz A, Sanchez R, Garcia-Asenjo J, Rodriguez-Martin C, Escudero M, Albanell J (2012) A randomized phase II trial of platinum salts in basal-like breast cancer patients in the neoadjuvant setting. Results from the GEICAM/2006-03, multicenter study. Breast Cancer Res Treat 136: 487–493.
Bastien RR, Rodriguez-Lescure A, Ebbert MT, Prat A, Munarriz B, Rowe L, Miller P, Ruiz-Borrego M, Anderson D, Lyons B, Alvarez I, Dowell T, Wall D, Segui M, Barley L, Boucher K, Alba E, Pappas L, Davis C, Aranda I, Fauron C, Stijleman I, Palacios J, Anton A, Carrasco E, Caballero R, Ellis M, Nielsen T, Perou C, Astill M, Bernard P, Martin M (2012) PAM50 breast cancer subtyping by RT-qPCR and concordance with standard clinical molecular markers. BMC Med Genomics 5: 44.
Carey LA, Dees EC, Sawyer L, Gatti L, Moore DT, Collichio F, Ollila DW, Sartor CI, Graham ML, Perou CM (2007) The triple negative paradox: primary tumor chemosensitivity of breast cancer subtypes. Clin Cancer Res 13: 2329–2334.
Cheang M, Martin M, Nielsen T, Prat A, Rodriguez-Lescure A, Ruiz A, Chia S, Shepherd L, Voduc D, Bernard P, Ellis M, Perou C, Di Leo A, Carey L (2012) Quantitative hormone receptors, triple-negative breast cancer (TNBC), and molecular subtypes: a collaborative effort of the BIG-NCI NABCG. Proceedings of the American Society of Clinical Oncology 1–5 June 2012; Chicago, IL, USA, a1008.
Cheang MCU, Chia SK, Voduc D, Gao D, Leung S, Snider J, Watson M, Davies S, Bernard PS, Parker JS, Perou CM, Ellis MJ, Nielsen TO (2009) Ki67 index, HER2 status, and prognosis of patients with luminal B breast cancer. J Natl Cancer Inst 101: 736–750.
Citron ML, Berry DA, Cirrincione C, Hudis C, Winer EP, Gradishar WJ, Davidson NE, Martino S, Livingston R, Ingle JN, Perez EA, Carpenter J, Hurd D, Holland JF, Smith BL, Sartor CI, Leung EH, Abrams J, Schilsky RL, Muss HB, Norton L (2003) Randomized trial of dose-dense versus conventionally scheduled and sequential versus concurrent combination chemotherapy as postoperative adjuvant treatment of node-positive primary breast cancer: first report of Intergroup Trial C9741/Cancer and Leukemia Group B Trial 9741. J Clin Oncol 21: 1431–1439.
Curtis C, Shah SP, Chin S-F, Turashvili G, Rueda OM, Dunning MJ, Speed D, Lynch AG, Samarajiwa S, Yuan Y, Graf S, Ha G, Haffari G, Bashashati A, Russell R, McKinney S, Langerod A, Green A, Provenzano E, Wishart G, Pinder S, Watson P, Markowetz F, Murphy L, Ellis I, Purushotham A, Borresen-Dale A-L, Brenton JD, Tavare S, Caldas C, Aparicio S (2012) The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 486: 346–352.
Dennis G, Sherman BT, Hosack DA, Yang J, Gao W, Lane HC, Lempicki RA (2003) DAVID: database for annotation, visualization, and integrated discovery. Genome Biol 4: R60.
Fasching P, Heusinger K, Haeberle L, Niklos M, Hein A, Bayer C, Rauh C, Schulz-Wendtland R, Bani M, Schrauder M, Kahmann L, Lux M, Strehl J, Hartmann A, Dimmler A, Beckmann M, Wachter D (2011) Ki67, chemotherapy response, and prognosis in breast cancer patients receiving neoadjuvant treatment. BMC Cancer 11: 486.
Hammond ME, Hayes DF, Dowsett M, Allred DC, Hagerty KL, Badve S, Fitzgibbons PL, Francis G, Goldstein NS, Hayes M, Hicks DG, Lester S, Love R, Mangu PB, McShane L, Miller K, Osborne CK, Paik S, Perlmutter J, Rhodes A, Sasano H, Schwartz JN, Sweep FCG, Taube S, Torlakovic EE, Valenstein P, Viale G, Visscher D, Wheeler T, Williams RB, Wittliff JL, Wolff AC (2010) American Society of Clinical Oncology/College of American Pathologists guideline recommendations for immunohistochemical testing of estrogen and progesterone receptors in breast cancer. J Clin Oncol 28: 2784–2795.
Hatzis C, Pusztai L, Valero V, Booser DJ, Esserman L, Lluch A, Vidaurre T, Holmes F, Souchon E, Wang H, Martin M, Cotrina J, Gomez H, Hubbard R, Chacón JI, Ferrer-Lozano J, Dyer R, Buxton M, Gong Y, Wu Y, Ibrahim N, Andreopoulou E, Ueno NT, Hunt K, Yang W, Nazario A, DeMichele A, OShaughnessy J, Hortobagyi GN, Symmans WF (2011) A genomic predictor of response and survival following taxane-anthracycline chemotherapy for invasive breast cancer. JAMA 305: 1873–1881.
Hess KR, Anderson K, Symmans WF, Valero V, Ibrahim N, Mejia JA, Booser D, Theriault RL, Buzdar AU, Dempsey PJ, Rouzier R, Sneige N, Ross JS, Vidaurre T, Gomez HL, Hortobagyi GN, Pusztai L (2006) Pharmacogenomic predictor of sensitivity to preoperative chemotherapy with paclitaxel and fluorouracil, doxorubicin, and cyclophosphamide in breast cancer. J Clin Oncol 24: 4236–4244.
Ignatiadis M, Singhal SK, Desmedt C, Haibe-Kains B, Criscitiello C, Andre F, Loi S, Piccart M, Michiels S, Sotiriou C (2012) Gene modules and response to neoadjuvant chemotherapy in breast cancer subtypes: a pooled analysis. J Clin Oncol 30: 1996–2004.
Iwamoto T, Bianchini G, Booser D, Qi Y, Coutant C, Ya-Hui Shiang C, Santarpia L, Matsuoka J, Hortobagyi GN, Symmans WF, Holmes FA, O'Shaughnessy J, Hellerstedt B, Pippen J, Andre F, Simon R, Pusztai L (2011) Gene pathways associated with prognosis and chemotherapy sensitivity in molecular subtypes of breast cancer. J Natl Cancer Inst 103 (3): 264–272.
Keam B, Im S, Lee K, Han S (2011) Ki-67 can be used for further classification of triple-negative breast cancer into two subtypes with different response and prognosis. Breast Cancer Res 13: R22.
Lehmann BD, Bauer JA, Chen X, Sanders ME, Chakravarthy AB, Shyr Y, Pietenpol JA (2011) Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J Clin Invest 121: 2750–2767.
Liedtke C, Mazouni C, Hess KR, Andre F, Tordai A, Mejia JA, Symmans WF, Gonzalez-Angulo AM, Hennessy B, Green M, Cristofanilli M, Hortobagyi GN, Pusztai L (2008) Response to neoadjuvant therapy and long-term survival in patients with triple-negative breast cancer. J Clin Oncol 26: 1275–1281.
Liu M, Pitcher B, Mardis E, Davies S, Snider J, Vickery T, Reed J, DeSchryver K, Singh B, Friedman P, Gradishar W, Perez E, Martino S, Citron M, Norton L, Winer E, Hudis C, Perou C, Ellis M, Barry W (2012) PAM50 gene signature is prognostic for breast cancer patients treated with adjuvant anthracycline and taxane based chemotherapy. San Antonio Breast Cancer Symposium 4–8 December 2012 San Antonio, TX, USA, P2-10-01.
Martín M, Rodríguez-Lescure Á, Ruiz A, Alba E, Calvo L, Ruiz-Borrego M, Munárriz B, Rodríguez CA, Crespo C, de Alava E, López García-Asenjo JA, Guitián MD, Almenar S, González-Palacios JF, Vera F, Palacios J, Ramos M, Gracia Marco JM, Lluch A, Alvarez I, Seguí MA, Mayordomo JI, Antón A, Baena JM, Plazaola A, Modolell A, Pelegrí A, Mel JR, Aranda E, Adrover E, Ãlvarez JV, García Puche JL, Sánchez-Rovira P, Gonzalez S, López-Vega JM (2008) Randomized phase 3 trial of fluorouracil, epirubicin, and cyclophosphamide alone or followed by paclitaxel for early breast cancer. J Natl Cancer Inst 100: 805–814.
Masuda H, Baggerly KA, Wang Y, Zhang Y, Gonzalez-Angulo AM, Meric-Bernstam F, Valero V, Lehmann BD, Pietenpol JA, Hortobagyi GN, Symmans WF, Ueno NT (2013) Differential response to neoadjuvant chemotherapy among 7 triple-negative breast cancer molecular subtypes. Clin Cancer Res 19: 5533–5540.
Mayer IA, Abramson VG, Lehmann BD, Pietenpol JA (2014) New strategies for triple-negative breast cancer—deciphering the heterogeneity. Clin Cancer Res 20: 782–790.
Nielsen TO, Hsu FD, Jensen K, Cheang M, Karaca G, Hu Z, Hernandez-Boussard T, Livasy C, Cowan D, Dressler L, Akslen LA, Ragaz J, Gown AM, Gilks CB, van de Rijn M, Perou CM (2004) Immunohistochemical and clinical characterization of the basal-like subtype of invasive breast carcinoma. Clin Cancer Res 10: 5367–5374.
Nielsen TO, Parker JS, Leung S, Voduc D, Ebbert M, Vickery T, Davies SR, Snider J, Stijleman IJ, Reed J, Cheang MCU, Mardis ER, Perou CM, Bernard PS, Ellis MJ (2010) A comparison of PAM50 intrinsic subtyping with immunohistochemistry and clinical prognostic factors in tamoxifen-treated estrogen receptor-positive breast cancer. Clin Cancer Res 16: 5222–5232.
Parker JS, Mullins M, Cheang MCU, Leung S, Voduc D, Vickery T, Davies S, Fauron C, He X, Hu Z, Quackenbush JF, Stijleman IJ, Palazzo J, Marron JS, Nobel AB, Mardis E, Nielsen TO, Ellis MJ, Perou CM, Bernard PS (2009) Supervised risk predictor of breast cancer based on intrinsic subtypes. J Clin Oncol 27: 1160–1167.
Perou CM, Parker JS, Prat A, Ellis MJ, Bernard PS (2010) Clinical implementation of the intrinsic subtypes of breast cancer. Lancet Oncol 11: 718–719.
Perou CM, Sorlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA, Pollack JR, Ross DT, Johnsen H, Akslen LA, Fluge O, Pergamenschikov A, Williams C, Zhu SX, Lonning PE, Borresen-Dale A-L, Brown PO, Botstein D (2000) Molecular portraits of human breast tumours. Nature 406: 747–752.
Popovici V, Chen W, Gallas B, Hatzis C, Shi W, Samuelson F, Nikolsky Y, Tsyganova M, Ishkin A, Nikolskaya T (2010) Effect of training-sample size and classification difficulty on the accuracy of genomic predictors. Breast Cancer Res 12: R5.
Prat A, Adamo B, Cheang MCU, Anders CK, Carey LA, Perou CM (2013a) Molecular characterization of basal-like and non-basal-like triple-negative breast cancer. Oncologist 18: 123–133.
Prat A, Cheang MCU, Martín M, Parker JS, Carrasco E, Caballero R, Tyldesley S, Gelmon K, Bernard PS, Nielsen TO, Perou CM (2013b) Prognostic significance of progesterone receptor–positive tumor cells within immunohistochemically defined luminal a breast cancer. J Clin Oncol 31: 203–209.
Prat A, Parker J, Fan C, Perou C (2012) PAM50 assay and the three-gene model for identifying the major and clinically relevant molecular subtypes of breast cancer. Breast Cancer Res Treat 135: 301–306.
Prat A, Parker J, Karginova O, Fan C, Livasy C, Herschkowitz J, He X, Perou C (2010) Phenotypic and molecular characterization of the claudin-low intrinsic subtype of breast cancer. Breast Cancer Res 12: R68.
Prat A, Perou CM (2011) Deconstructing the molecular portraits of breast cancer. Mol Oncol 5: 5–23.
Rody A, Holtrich U, Pusztai L, Liedtke C, Gaetje R, Ruckhaeberle E, Solbach C, Hanker L, Ahr A, Metzler D, Engels K, Karn T, Kaufmann M (2009) T-cell metagene predicts a favorable prognosis in estrogen receptor-negative and HER2-positive breast cancers. Breast Cancer Res 11: R15.
Rody A, Karn T, Liedtke C, Pusztai L, Ruckhaeberle E, Hanker L, Gaetje R, Solbach C, Ahr A, Metzler D, Schmidt M, Muller V, Holtrich U, Kaufmann M (2011) A clinically relevant gene signature in triple negative and basal-like breast cancer. Breast Cancer Res 13: R97.
Sikov W, Berry D, Perou C, Singh B, Cirrincione C, Tolaney S, Kuzma C, Pluard T, Somlo G, Port E, Golshan M, Bellon J, Collyar D, Hahn O, Carey L, Hudis C, Winer E (2013) Impact of the addition of carboplatin (Cb) and/or bevacizumab (B) to neoadjuvant weekly paclitaxel (P) followed by dose-dense AC on pathologic complete response (pCR) rates in triple-negative breast cancer (TNBC): CALGB 40603 (Alliance). San Antonio Breast Cancer Symposium 10–14 December 2013 San Antonio, TX, USA, S5-01.
Silver DP, Richardson AL, Eklund AC, Wang ZC, Szallasi Z, Li Q, Juul N, Leong C-O, Calogrias D, Buraimoh A, Fatima A, Gelman RS, Ryan PD, Tung NM, De Nicolo A, Ganesan S, Miron A, Colin C, Sgroi DC, Ellisen LW, Winer EP, Garber JE (2010a) Efficacy of neoadjuvant cisplatin in triple-negative breast cancer. J Clin Oncol 28: 1145–1153.
Silver DP, Richardson AL, Eklund AC, Wang ZC, Szallasi Z, Li Q, Juul N, Leong C-O, Calogrias D, Buraimoh A, Fatima A, Gelman RS, Ryan PD, Tung NM, De Nicolo A, Ganesan S, Miron A, Colin C, Sgroi DC, Ellisen LW, Winer EP, Garber JE (2010b) Efficacy of neoadjuvant cisplatin in triple-negative breast cancer. J Clin Oncol 28: 1145–1153.
Skipper HE (1971) Kinetic behavior versus response to chemotherapy. Natl Cancer Inst Monogr 34: 2–14.
Sorlie T, Perou C, Tibshirani R, Aas T, Geisler S, Johnsen H, Hastie T, Eisen M, van de Rijn M, Jeffrey S, Thorsen T, Quist H, Matese J, Brown P, Botsein D, Eystein Lonning P, Borresen-Dale A (2001) Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci USA 98: 10869–10874.
Tabchy A, Valero V, Vidaurre T, Lluch A, Gomez H, Martin M, Qi Y, Barajas-Figueroa LJ, Souchon E, Coutant C, Doimi FD, Ibrahim NK, Gong Y, Hortobagyi GN, Hess KR, Symmans WF, Pusztai L (2010) Evaluation of a 30-gene paclitaxel, fluorouracil, doxorubicin, and cyclophosphamide chemotherapy response predictor in a multicenter randomized trial in breast cancer. Clin Cancer Res 16: 5351–5361.
Usary J, Zhao W, Darr D, Roberts PJ, Liu M, Balletta L, Karginova O, Jordan J, Combest A, Bridges A, Prat A, Cheang MCU, Herschkowitz JI, Rosen JM, Zamboni W, Sharpless NE, Perou CM (2013) Predicting drug responsiveness in human cancers using genetically engineered mice. Clin Cancer Res 19: 4889–4899.
Von Minckwitz G, Schneeweiss A, Loibl S, Salat C, Denkert C, Rezai M, Blohmer JU, Jackisch C, Paepke S, Gerber B, Zahm DM, Kümmel S, Eidtmann H, Klare P, Huober J, Costa S, Tesch H, Hanusch C, Hilfrich J, Khandan F, Fasching PA, Sinn BV, Engels K, Mehta K, Nekljudova V, Untch M (2014) Neoadjuvant carboplatin in patients with triple-negative and HER2-positive early breast cancer (GeparSixto; GBG 66): a randomised phase 2 trial. Lancet Oncol 15: 747–756.
Whitfield M, George L, Grant G, Perou C (2006) Common markers of proliferation. Nat Rev Cancer 6: 99.
Wolff AC, Hammond MEH, Schwartz JN, Hagerty KL, Allred DC, Cote RJ, Dowsett M, Fitzgibbons PL, Hanna WM, Langer A, McShane LM, Paik S, Pegram MD, Perez EA, Press MF, Rhodes A, Sturgeon C, Taube SE, Tubbs R, Vance GH, van de Vijver M, Wheeler TM, Hayes DF (2006) American Society of Clinical Oncology/College of American Pathologists guideline recommendations for human epidermal growth factor receptor 2 testing in breast cancer. J Clin Oncol 25: 118–145.
Acknowledgements
This work was supported by funds from the NCI Breast SPORE program Grant No. P50-CA58223-09A1 (CMP), by RO1-CA138255 (CMP), by the Breast Cancer Research Foundation (CMP and MJE), National Cancer Institute (NCI) Strategic Partnering to Evaluate Cancer Signatures Grant No. U01 CA114722-01 (MJE), by the Sociedad Española de Oncología Médica (AP), by FEDER (RETICC-RD12/0036/0051, RD12/0036/0042, RD12/0036/0076, RD12/0036/0070), by Instituto de Salud Carlos III—PI13/01718 (AP), by Banco Bilbao Vizcaya Argentaria (BBVA) Foundation (AP) and by the Alliance Statistics and Data Center (U10-CA33601).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
CMP and MJE are equity stock holders of BioClassifier LLC and University Genomics. CMP, MJE, MCUC and JSP have filed a patent on the PAM50 assay. MLG serves as a consultant for Roche Molecular Systems. Uncompensated advisory role of AP for Nanostring Technologies. The remaining authors declare no conflict of interest.
Additional information
This work was presented, in part, as an oral communication at the ASCO 2012 annual meeting (Abstract #10500).
Supplementary Information accompanies this paper on British Journal of Cancer website
Supplementary information
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Prat, A., Lluch, A., Albanell, J. et al. Predicting response and survival in chemotherapy-treated triple-negative breast cancer. Br J Cancer 111, 1532–1541 (2014). https://doi.org/10.1038/bjc.2014.444
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2014.444
Keywords
This article is cited by
-
Transcriptionally regulated miR-26a-5p may act as BRCAness in Triple-Negative Breast Cancer
Breast Cancer Research (2023)
-
Replenishment of myeloid-derived suppressor cells (MDSCs) overrides CR-mediated protection against tumor growth in a murine model of triple-negative breast cancer
GeroScience (2022)
-
Local treatment for triple-negative breast cancer patients undergoing chemotherapy: breast-conserving surgery or total mastectomy?
BMC Cancer (2021)
-
Immunotherapy in endometrial cancer: rationale, practice and perspectives
Biomarker Research (2021)
-
Progress in systemic therapy for triple-negative breast cancer
Frontiers of Medicine (2021)