Abstract
Background:
Over the past years, some members of the family of suppressor of cytokine signalling (SOCS) proteins have emerged as potential tumour suppressors. This study aimed at investigating the clinical significance of SOCS proteins in colorectal carcinoma (CRC).
Methods:
We integrated publicly available microarray expression data on CRC in humans, analysed the expression pattern of SOCSs and assessed the predictive power of SOCS2 and SOCS6 for diagnostic purposes by generating receiver operating characteristic curves. Using laser microdissected patient material we assessed SOCS expression on RNA and protein levels as well as their methylation status in an independent CRC patient cohort. Finally, we investigated the prognostic value of SOCS2 and SOCS6.
Results:
The meta-analysis as well as the independent patient cohort analysis reveal a stage-independent downregulation of SOCS2 and SOCS6 and identify both molecules as diagnostic biomarkers for CRC. We demonstrate a different methylation pattern within the SOCS2 promoter between tumour tissue and normal control tissue in 25% of CRC patients. Furthermore, early CRC stage patients with low expression of SOCS2 display significantly shorter disease-free survival.
Conclusions:
Our data offers evidence that SOCS2 and SOCS6 levels are reduced in CRC and may serve as diagnostic biomarkers for CRC patients.
Similar content being viewed by others
Main
Despite progress made during the past decades, colorectal cancer (CRC) is still one of the most frequent and deadly cancers world-wide in both women and men. The prevention of this disease is, therefore, a significant public health issue. The primary cause of death is the development of distant metastases in organs such as liver and lungs; ∼40–50% of patients who undergo curative surgery relapse and die of metastatic disease (Johnston, 2005). Strikingly, diagnosed in time, CRC can be cured in 9 out of 10 cases. Thus, it is highly important to identify more sensitive and specific CRC markers to strengthen the efficiency of early diagnosis as well as to improve therapeutic strategies.
Aberrant cytokine signalling has been associated with many diseases, including several cancers, disorders in haematopoiesis and autoimmune diseases. Cytokine responses have to be stringently controlled by a number of key regulatory proteins, such as the suppressors of cytokine signalling (SOCSs) family members. SOCS proteins are rapidly induced upon JAK/STAT signalling by activated signal transducer and activator of transcription factors (STAT) to negatively regulate cytokine signalling via a classical feedback loop (Inagaki-Ohara et al, 2013). Taking into consideration that ∼20% of all malignancies are initiated or exacerbated by inflammation, it is not surprising that SOCS proteins are regarded as tumour suppressor-like proteins (Elliott et al, 2008; Culig, 2013). The SOCS family consists of eight proteins, namely SOCS1–SOCS7 and cytokine-inducible SH2-containing protein (CIS). Each of these proteins has a central SH2 domain, an amino-terminal domain of variable length and sequence, and a carboxy-terminal 40 amino-acid region called SOCS box (Inagaki-Ohara et al, 2013). The SOCSs can interact with a series of signalling intermediates through the binding of their SH2 domain to phosphorylated tyrosine residues, particularly those on cytokine receptors and JAKs, leading to the blockade of the signal (Inagaki-Ohara et al, 2013).
Among SOCS family members, SOCS1 and SOCS3 have been most widely studied. SOCS1 and SOCS3 have been shown to suppress cell growth and their expression is frequently downregulated in human cancers (Trengove and Ward, 2013). Low SOCS2 gene expression has been associated with hepatocellular, breast, pulmonary and ovarian cancers (Wikman et al, 2002; Sutherland et al, 2004; Farabegoli et al, 2005; Haffner et al, 2007; Qiu et al, 2013). SOCS4 shows a reduced expression in gastric cancer and is associated with a better clinical outcome in breast cancer patients (Sasi et al, 2010; Kobayashi et al, 2012) whereas loss of SOCS5 expression is correlated with poor prognosis in liver cancer (Yoon et al, 2012). SOCS6 has been reported to be downregulated in carcinomas of prostate, stomach, liver and colon (Lai et al, 2009, 2010; Qiu et al, 2013; Zhu et al, 2013). SOCS7 has a favourable prognostic value in breast cancer (Sasi et al, 2010). Altogether, these findings suggest the involvement of SOCSs in cancer.
In the present study, we examined the expression as well as the methylation status of SOCS proteins in primary colon tumours, its precursor lesions and distant normal tissue. We used laser microdissection (LMD) to select for a highly pure population of epithelial cells in primary CRC tumours as well as in normal colon tissue. Our results point towards the involvement of SOCS2 and SOCS6 in the carcinogenesis of CRC and demonstrate their potential use as biomarkers.
Materials and methods
Patients and samples
Institutional guidelines were followed. All human samples used in the scope of this work were donated freely and informed consent was obtained. Ethical approval was obtained from the Comité National d'Ethique de Recherche, Luxembourg (Reference 201009/09). Primary colon cancer tissue and matched distant non-neoplastic colon tissue (at the farthest longitudinal surgical margin) from 23 CRC patients were collected following the standard preanalytical code for biospecimens by the Integrated Biobank of Luxembourg (IBBL; Betsou et al, 2010) immediately after surgical excision and stored in liquid nitrogen before further processing. This collection contains high dysplasia (n=3), stage I (n=3), stage II (n=10), stage III (n=6), stage IV (n=1) tumour samples according to the TNM Classification of Malignant Tumours (TNM system, American Joint Committee on Cancer; Hari et al, 2013) as well as 23 normal tissue samples matching the corresponding tumour sample. To increase our CRC collection, we also received tumour specimens from stage 1 (n=13) and stage 4 (n=30) from the Ontario Tumour Bank (Ontario Institute for Cancer Research), so that in total our study includes 23 normal tissue samples and 66 CRC samples. The clinical and histopathological data were provided by a pathologist. In total, 66 CRC patients were included, 35 male and 31 female with a median age of 67 years (66.6±12, range 30–89) (Supplementary Table 1). Tumours included in the study were of sporadic origin and restricted to the colon except for one sample located in the rectum. In the bioinformatic analysis the authors of the respective studies used the Dukes staging system, which ranges from A to D. Even if the TNM staging system is preferentially used nowadays we can still correlate both staging methods (Walters et al, 2013). Disease-free survival corresponds to the interval between the date of surgery and the date when recurrence is diagnosed or the date of last follow-up.
Materials
All CRC cell lines were obtained either from the American Type Culture Collection (ATCC, Rockville, MD, USA) or the German Collection of Microorganisms and Cell Cultures (DSMZ, Braunschweig, Germany) and maintained in recommended culture conditions. LS174t and HT-29 were exposed to 5 μ M 5-Aza-2′-deoxycytidine (5-aza-dC/DAC, Sigma-Aldrich, Diegem, Belgium) or vehicle (DMSO) for 4 days followed by RNA extraction.
Tissue processing and laser microdissection
We used LMD to maximise the purity of the epithelial tumour cell population. Briefly, frozen samples were mounted in Tissue-Tek OCT compound, serially sectioned at 10 μm in a cryostat at −20°C (Leica, Wetzlar, Germany), and mounted on prepared membrane slides (Leica mb slides PEN-membrane 2 μm). The parts containing areas of epithelial cells from tumours and normal tissue were identified using haematoxylin and eosin staining (Histogene LCM Frozen section staining kit, Applied Biosystems, Gent, Belgium) and analysed by a histopathologist. A Leica AS LMD microscope (Leica) was then used to select epithelial cells from tumour biopsies and their distant normal counterpart for each patient. Finally, sections were collected in a 0.5 ml microtube with RNA extraction buffer (All Prep Qiagen, Hilden, Germany) containing 1% β-mercaptoethanol.
RNA/DNA extraction and real-time PCR
RNA/DNA extraction was performed using commercially available kits and real-time PCR experiments were performed using TaqMan technology (Applied Biosystems) (Erickson et al, 2009). Details are provided as Supplementary Information to this article.
Methylation analysis
For the monitoring of the methylation pattern of the SOCS2 promoter, pyrosequencing and MassARRAY technology by Sequenom (San Diego, CA, USA) was performed at Varionostics GmbH, Ulm, Germany. The EpiTYPER assay (Sequenom) is a tool for the detection and quantitative analysis of DNA methylation using base-specific cleavage and matrix-assisted laser desorption–ionisation time-of-flight mass spectrometry (MALDI–TOF MS).
Western blot
Frozen tissues from CRC patients were homogenised in RIPA lysis buffer (ThermoFisher Scientific, Erembodegem-Aalst, Belgium) containing protease inhibitors. Beads were added to the homogenate, which was then lysed in a tissue lyser (Qiagen; 2 × 1 min). Proteins were run on a 12% SDS–PAGE gel and transferred to PVDF membranes (Pall Corporation, Port Washington, NY, USA). Membranes were blocked with 10% milk for 1 h at room temperature and were then incubated overnight at 4°C with antibodies directed against SOCS2 (1 : 1000, OriGene Technologies, Uden, Netherlands; TA307336), SOCS6 (1 : 1000, Santa Cruz Biotechnology, Heidelberg, Germany; sc-133058) and GAPDH (1 : 3000, Sigma-Aldrich, G9545). After washing, detection of the protein bands was performed using the Femto kit (SuperSignal West Femto kit, Thermo Scientific, Erembodegem-Aalst, Belgium). Detection was performed using Chemidoc XRS+ imager (Biorad, Nazareth Eke, Belgium).
Immunohistochemical analysis
Tissue sections (4 μm) were cut from FFPE blocks from two patients with stage III to assess immunohistochemical staining for SOCS2 and SOCS6. Briefly, sections were deparaffinised followed by antigen retrieval and incubated with anti-human SOCS2 (Bioss, Woburn, MA, USA; bs-1896R, 1 : 300) and anti-human SOCS6 (Abcam, Cambridge, UK; ab53211, pre-diluted), respectively. The antibody dilution was determined after initial standardisation and specificity of the antibodies was verified by using positive controls. Sections on which the primary antibody was not applied served as negative controls. Secondary detection was performed with a detection kit (ultraView Universal DAB detection kit, Ventana, Basel, Switzerland; ref. 760–500) and counterstain with Hematoxylin II (Ventana, ref. 790–2208) followed by application of bluing reagent (Ventana, ref. 760–2037). Each section was dehydrated by graded concentrations of alcohol before coverslips were applied.
Statistical analysis
GraphPad Prism 5 software (La Jolla, CA, USA) was used for statistical analysis. We used unpaired Student’s t-test to compare expression levels between tumour and normal tissue. Kaplan–Meier curves were performed to assess if SOCS2 and SOCS6 levels correlate to prognosis of patients. For this, datasets GSE39582 and GSE14333 were downloaded from GEO (http://www.ncbi.nlm.nih.gov/geo/) and clinical information extracted (Jorissen et al, 2009; Marisa et al, 2013). For each dataset, gene expression values were grouped in two categories (‘low’ / ‘high’) using the k-mean algorithm. This clustering was performed in each dataset separately to avoid bias in gene expression values between the datasets. When only stages A and B are studied, the clustering was performed after removing patients that were not graded as stage A or B. After determination of the categories, both datasets were merged into a single one, which was used for further analysis. The survival analysis was performed using the survival package in R (http://cran.r-project.org/web/packages/survival/). Statistical significance of this correlation was assessed by log rank test. P-values <0.05 were considered statistically significant.
Bioinformatic analysis
Microarray expression datasets from CRC, colon adenoma and inflammatory bowel disease (IBD) were retrieved from NCBI Gene Expression Omnibus (GEO) Database (http://www.ncbi.nlm.nih.gov/geo/). All the individual CEL files from datasets profiled on HG-U133 plus 2.0 (Affymetrix, Santa Clara, CA, USA) retrieved from GEO (GSE14333; GSE17538; GSE21510; GSE8671; GSE9254; GSE20916; GSE10714; GSE15960; GSE4183; GSE10961) and corresponding to different studies (Sabates-Bellver et al, 2007; Galamb et al, 2008, 2010; LaPointe et al, 2008; Jorissen et al, 2009; Skrzypczak et al, 2010; Smith et al, 2010; Tsukamoto et al, 2011) were integrated into one single global analysis, referred to as meta-analysis (Supplementary Table 2). A preselected set of arrays profiling normal colorectal tissue samples was chosen as control. This large number of arrays was subjected to multiple downstream processing steps. CEL files were preprocessed using the Robust Multiarray Analysis (RMA) algorithm with GC-correction on the commercial software Partek Genome Suite (Version 6.4, St Louis, MO, USA). In order to minimise multiple study expression artifacts, and therefore the technical variability between the different research laboratories, the batch effect was corrected by applying (a) 3′ : 5′ house-keeping gene ratio, which gives an indication on RNA quality (Gentleman et al, 2005); (b) principal component analysis (PCA) allows to determine potential sources of variability in a multidimensional data set; (c) relative log2 expression (RLE) summaries, which are sensitive to technical sources of variability that are large compared to biological variation (Bolstad et al, 2004). Twelve out of 843 arrays had to be withdrawn from the dataset because of insufficient quality. Non-informative gene features were removed by filtering out probe sets with log2 expression level never exceeding 6. An additional summarisation step was performed in order to remove not annotated transcripts and obtain a single expression value for each annotated gene. Statistical significance of the genes of interest was addressed by applying 95% confidence intervals for mean log FC. P-values were adjusted using the Benjamini–Hochberg false discovery rate (FDR) method. Suitability of the potential biomarkers for discrimination of CRC and normal samples was addressed by calculation of the area under receiver operating characteristic (ROC) curves (AUC) using package caTools of R (http://CRAN.R-project.org/package=caTools).
Results
A bioinformatic approach identifies SOCS2 and SOCS6 as potential diagnostic biomarkers
The integration of publicly available data of smaller studies into one meta-analysis increases the power to detect biologically relevant signals specific for the initiation or progression of a disease. Published microarray-based studies profiling human samples on IBD, colon adenoma, CRC, and normal colon tissue were chosen for the present study. Principal component analysis shows that the different disease conditions form well-separated clusters whereas no differential clustering was detected according to age, gender or location of the tumour (data not shown). All FDR values are indicated in Supplementary Table 3. To evaluate the biological significance of the microarray datasets, we analysed the expression profiles of known marker genes (SOCS3, KIAA1199, osteopontin/SPP1, CDX1) specific for IBD, colon adenoma or CRC (Agrawal et al, 2002; Kim et al, 2005; Rigby et al, 2007; Sabates-Bellver et al, 2007). Inflammatory bowel disease is characterised by a chronic recurrent colonic inflammation, which is associated with an increased risk of developing CRC. In our meta-analysis we found SOCS1 and SOCS3 expression to be upregulated in IBD samples compared to normal colorectal mucosa samples (2.94-fold and 3.10-fold, respectively, FDR<0.001, Supplementary Table 3 and Supplementary Figure 1A). Interestingly we find upregulation of all STAT factors except STAT6 in the IBD samples (Figure 1B and Supplementary Table 3). KIAA1199 (colon cancer secreted protein 1) is 7.32-fold upregulated (FDR<0.001) in adenomas and stayed elevated in CRC samples from our meta-analysis (4.48-fold; FDR<0.001; Figure 1A). This is in line with a previous study by Sabates-Bellver et al (2007) who reported KIAA1199 to be upregulated both in colon adenoma and stage independently in CRC patients. Osteopontin/SPP1 a described progression marker for CRC (Agrawal et al, 2002), shows a higher expression in the CRC dataset compared to adenomas and normal mucosal tissue (Figure 1A). CDX1 caudal type homeobox is defined as a differentiation marker in CRC (Kim et al, 2005). Accordingly, its expression is reduced by ∼35% (FDR<0.001) in CRC samples (Figure 1A).
Bioinformatic analysis reveals SOCS2 and SOCS6 downregulation in CRC. (A) Bar plot showing the log2 FC values of the SOCS family genes and marker genes in adenoma and CRC samples compared to normal colorectal mucosa. Error bars correspond to 95% confidence intervals for mean log FC; *FDR<0.05, **FDR<0.01 and ***FDR<0.001. (B) Heatmap representation of SOCS (highlighted in yellow) and STAT (highlighted in grey) family genes as well as marker genes in normal tissues, IBD, adenomas, and CRC stages (A–D).
As the selected control marker genes all showed the expression pattern previously described in the literature, we conclude that our meta-analysis, which contains a high number of microarray datasets, is biologically relevant and can be used in order to address the potential use of SOCS family members as new biomarkers or targets for CRC therapy. When comparing adenoma samples to normal tissue, we found SOCS2 expression to be ∼20% downregulated (linear fold change, FDR<0.001), whereas expression of the other members of the family are either not changed or slightly upregulated (Figure 1A). In CRC, CISH, SOCS2 and SOCS6 show a significant downregulation in CRC by 10%, 45% and 60%, respectively (Figure 1A). Interestingly, the expression of SOCS2 significantly decreases from adenoma to CRC (P<0.001, Figure 1A and B). A detailed analysis of their expression along CRC staging shows that their downregulation is stage independent (Figure 1B, Supplementary Figure 1B). For the remaining SOCS family members, the expression is also not influenced by the CRC stages (Supplementary Table 3). Importantly, expression of the control marker gene SPP1 increases stage dependently from CRC stage A to D by 1.72 (CRC.A); 2.98 (CRC.B), 3.14 (CRC.C) and 3.54 (CRC.D) fold, respectively; FDR<0.001) which is consistent with previous findings (Agrawal et al, 2002). Investigating the expression of the STAT family members, we observe a slight upregulation of STAT1 and a faint upregulation of STAT5A/B, an upstream regulator of SOCS2. Interestingly, we find STAT6 to be downregulated alongside SOCS2 and SOCS6 in CRC samples, albeit to a lesser degree (Figure 1B; Supplementary Table 3). However, a direct link between STAT6 downregulation and the observed reduced expression SOCS2 and SOCS6 is unlikely as STAT6 activation has not yet been associated with the regulation of these two SOCS proteins. Further investigations including STAT protein expression analyses could help clarifying the significance of the observed STAT mRNA changes. We next analysed the predictive power of SOCS2 and SOCS6 expression levels for diagnostic purposes by generating ROC curves and c statistic (AUC; Figure 2). AUC values of 0.910 and 0.958 were observed for SOCS2 and SOCS6, respectively. Furthermore, the distributions of gene expression levels for SOCS2 and SOCS6 clearly separated into two distinct almost non-overlapping peaks. The high AUC values as well as the well-separated distribution profiles support the use for SOCS2 and SOCS6 as biomarkers. Taken together, bioinformatics and biostatistics represent a valuable tool for the analysis of complex human diseases and the identification of novel biomarkers like, in our case, SOCS2 and SOCS6.
SOCS2 and SOCS6 are downregulated at mRNA and protein level in primary human CRC samples
To validate our bioinformatic analysis in a different patient cohort we assessed the expression of SOCS family members in human CRC samples and their respective normal counterparts. 66 CRC tumour samples and 23 normal counterparts were used for the present study (Supplementary Table 1). Using LMD, we selected a highly pure malignant or normal epithelial cell population. This is particularly important for SOCSs as tumour-infiltrating inflammatory cells often express these proteins. Thus, taking bulk material may lead to a wrong estimation of the expression of SOCSs in tumour cells. First we checked the expression of marker genes within our primary human CRC collection. KIAA1199 and SPP1 are both upregulated in CRC samples (P<0.001 and P<0.01, respectively; Figure 3A). CDX1 shows a significantly reduced expression in CRC samples compared to the normal counterparts (P<0.001; Figure 3A). Thus, the marker genes show the same expression pattern as in the bioinformatic analysis. Further, among the SOCS family members, SOCS2 and SOCS6 expression is significantly downregulated (P<0.001) whereas SOCS1 and SOCS3 do not show a statistically relevant difference in expression compared to the normal mucosal tissue (Figure 3B). To analyse whether the SOCS family members show a different expression pattern along the progression of the disease, we separated the samples according to the TNM staging system. As expected SOCS1 and SOCS3 expression does not change between the different CRC stages (Supplementary Figure 2A). Furthermore, we do not observe a change in SOCS2 and SOCS6 expression along CRC staging (Supplementary Figure 2A). Thus, SOCS2 and SOCS6 are significantly downregulated in CRC samples in a stage-independent manner. Western blot analysis and immunohistochemical stainings confirmed that reduced expression of SOCS2 and SOCS6 also result in a decrease in protein levels (Figure 4A and B). These results validate our findings obtained with the bioinformatic approach in a second independent patient cohort and strongly support the use of SOCS2 and SOCS6 as potential diagnostic markers.
Protein levels of SOCS2 and SOCS6 in human CRC. (A) Western blot detection of SOCS2 and SOCS6 in CRC and distant normal counterpart tissues. Caco-2 and Hela cells served as positive control for SOCS2 and SOCS6, respectively. The samples were run on the same gel but the membrane was cut for representative purposes (dotted line). (B) Immunohistochemical staining for SOCS2 and SOCS6 in CRC and normal counterpart for two different patients, magnification × 200.
Mechanisms contributing to reduced SOCS2 expression in colorectal carcinoma
Over the past years, hypermethylation of promoters has been reported for several SOCS family members (Elliott et al, 2008; Culig, 2013). Along this line, we decided to analyse whether promoter methylation could explain the observed downregulation of SOCS2 and SOCS6. First, we investigated the expression of SOCS2 and SOCS6 in CRC cell lines using quantitative real-time PCR. Caco-2 cells show a high expression of SOCS2 whereas HT-29 and LS174t have a low expression level of SOCS2 and SOCS6 (Supplementary Figure 3). To analyse whether methylation could explain the observed results, we treated HT-29 and LS174t with the demethylating agent 5-aza-dC/DAC. DAC treatment barely changed basal expression levels of SOCS6. In contrast, an increase in basal expression was observed for SOCS2 after DAC treatment in HT-29 and LS174t cells (Figure 5A), suggesting that methylation of the SOCS2 promoter is responsible for the reduced expression. However, as DAC demethylates the entire DNA, an indirect effect cannot be excluded. Thus, we decided to analyse the methylation of 16-17 CpG sites within SOCS2 and SOCS6 promoter using pyrosequencing for an initial set of 12 patients. The sequence analysed for SOCS6 did not reveal methylation, which correlates to the unchanged basal expression of SOCS6 after DAC treatment (data not shown). However, SOCS2 pyrosequencing analysis showed methylation in the analysed sequence (−1271 to −1166 from the start codon) for ∼17% (2 out of 12) of the patient samples (data not shown). Of note, methylation was not uniform among all the CpG sites analysed, indicating that the investigation of a small number of CpG sites may in general be problematic. We thus decided to assess SOCS2 promoter methylation in more detail. We used the mass array technology in order to determine the methylation pattern of the entire promoter sequence and analysed the methylation profile of 28 CRC patients as well as 20 normal non-tumour samples. Among these, we had 16 paired tumour/non-tumour samples. Tumour-specific methylation was detected in 3 out of 16 tumour samples from these paired samples in a sequence around the start codon (−158 to +334 from the start codon; Figure 5B and C). Importantly, all of the non-tumour samples showed an absence of methylation (defined by a methylation signal of <5%). Overall, 7 out of 28 (25%) CRC samples were methylated in the SOCS2 promoter region. Furthermore, the same sequence was also highly methylated in HT-29 and LS174t cells expressing low levels whereas no methylation was observed for the Caco-2 cells that express high levels of SOCS2 (data not shown). Strikingly, analysis of the methylated sequence revealed the presence of the consensus motif TTCnnnGAA, which all STAT members (except STAT6) recognise (Kang et al, 2013; Figure 5C). Taken together, methylation of the SOCS2 promoter can partly explain the downregulation of SOCS2 expression in our patient collection.
Methylation analysis of the SOCS2 promoter in primary human CRC samples. (A) Treatment with 5-aza-dC/DAC increases basal SOCS2 expression in LS174t and HT-29 colon cancer cell lines whereas SOCS6 levels are unchanged. Data are representative of four independent experiments and presented as mean±s.d. (B) Representative histogram of the methylation pattern of four primary human CRC samples compared to their respective normal counterpart in a sequence −158 to +334 relative to the start codon. Each CpG site analysed is represented by a dot. (C) Representation of the SOCS2 promoter region that shows differential methylation between tumour and normal control samples. The sequence analysed by mass array is underlined and the CpG sites are highlighted in red. Analysis of the methylated sequence shows the presence of 2 STAT GAS motifs (TTCnnnGAA) highlighted in yellow. The start codon is indicated in green. The primers used in other SOCS2 methylation studies are indicated (Liu et al: pink arrows; Fiegl et al: green arrows; Sutherland et al: blue arrows).
Most interestingly, recent reports have highlighted the crucial role of the glucocorticoid receptor (GR) for the induction of SOCS2 in vivo (Tronche et al, 2004; Martinez et al, 2013). We thus determined the expression levels of GR, which often acts as a cofactor of STAT5 for GH-induced genes, and found that they are also dramatically downregulated in our CRC samples (Supplementary Figure 2B).
SOCS2 has a prognostic value in early colorectal carcinoma
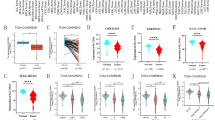
We next wanted to evaluate whether SOCS2 and SOCS6 have prognostic value for CRC. As patients affected by CRC are often already advanced in age at diagnosis and may die from another reason rather than from colon cancer, we decided to analyse the disease-free survival rates in CRC patients. For this we combined two datasets (GSE3958 and GSE14333), which provide follow-up data for the patients. Dividing the data into low and high SOCS2 or SOCS6 expression we observed a difference in disease-free survival for SOCS2, which reached statistical significance (P=0.036). (Figure 6A). We then extended our analysis towards the different CRC stages. Interestingly, while only taking early CRC stages into account, namely stages I and II (lymph node-negative cancer), we confirmed the significantly shorter disease-free survival time associated to patients with low expression compared to patients with high expression of SOCS2 (Figure 6B, P=0.00066). The strong increase in statistical significance for stage I/II patients argues for a prognostic value of SOCS2 expression in early CRC stages.
Prognostic value of SOCS2 and SOCS6 in CRC. Disease-free survival curves for patients with high and low expression of SOCS2 or SOCS6 in primary CRC including (A) all stages or (B) stages I and II only. The number of patients included in each group is mentioned within brackets. Significant P-values are indicated.
Discussion
Selection of the most beneficial treatment regimens in CRC remains challenging due to lack of prognostic markers. In this study, we analysed the expression pattern of the SOCS family members in fresh-frozen primary samples of colon tumours. Laser microdissection was used to dissociate stromal from tumour cells and obtain highly enriched neoplastic cell populations, allowing us to specifically assess the levels of SOCS expression in tumour cells. Here we show for the first time that SOCS2 and SOCS6 display a reduced expression at the mRNA and protein level in CRC patients, identifying them as potential diagnostic markers. Furthermore, SOCS2 downregulation is partly due to methylation and is an independent predictor of shorter disease-free survival.
A bioinformatic study followed by a validation in a second group of fresh-frozen normal and colon cancer tissue specimens identified two potential biomarkers, SOCS2 and SOCS6, for CRC. Importantly, SOCS2 and SOCS6 expression levels allow us to discriminate between healthy and CRC cases, further emphasising their use as diagnostic biomarkers. These data validate and support bioinformatics and biostatistics as an appropriate starting point for the analysis of complex human diseases and discovery of novel biomarkers. SOCS2 has been associated with growth hormone (GH) signalling and is thereby involved in cell growth (Metcalf et al, 2000; Horvat and Medrano, 2001; Greenhalgh et al, 2005). Most importantly, SOCS2 deletion in mice promotes the spontaneous development of intestinal tumours driven by mutations in the APC/β-catenin pathway (Newton et al, 2010). Along this line, it has been shown that forced overexpression of SOCS2 inhibits proliferation of the Caco-2 colon cancer cell line (Miller et al, 2004). Functional studies in mice have further shown that the disruption of one allele of SOCS2 in GH transgene mice leads to an increase in colon and jejunal crypt proliferation, thus favouring the formation of hyperplastic and lymphoid polyps in the colon (Michaylira et al, 2006). These findings provide evidence that SOCS2 normally limits tumour growth and strongly supports its tumour suppressive potential. Accordingly, low SOCS2 gene expression has been associated with diverse cancers (Trengove and Ward, 2013). However, until now there is no report on SOCS2 levels in human CRC. Here we show that SOCS2 levels are downregulated on mRNA as well as on protein levels in CRC patients. Interestingly, SOCS2 protein expression was associated with high differentiation and a low proliferation rate in breast carcinoma (Farabegoli et al, 2005). A low SOCS2 expression in prostate cancer is reported to be associated with an increased incidence of metastasis and SOCS2 mRNA levels decrease during prostate cancer progression (Hendriksen et al, 2006; Iglesias-Gato et al, 2014). However, SOCS2 expression in prostate cancer is controversial among the different studies (Hoefer et al, 2013; Zhu et al, 2013; Iglesias-Gato et al, 2014). In contrast to the studies conducted in breast cancer, there is no progressive decrease in SOCS2 expression along CRC staging. As SOCS2 is already downregulated in the adenoma samples, SOCS2 might be considered as an early diagnostic marker that enables diagnosis and treatment of patients susceptible to CRC at an early stage in cancer development. Furthermore, as we do not detect a progressive decrease in SOCS2 expression along the different cancer stages, our data suggest that reduced expression often occurs during the adenoma to CRC sequence. Altogether, SOCS2 may not be a progression marker gene, but could represent an early diagnostic marker for CRC.
In CRC, loss of SOCS6 gene copy number was found in 54.2% of the patients (Storojeva et al, 2005). SOCS6, by regulating insulin signalling, is thought to participate in CRC progression (Boyd, 2003). In addition, ectopic expression of SOCS6 was shown to suppress cell growth and colony formation (Lai et al, 2010). Accordingly, our study confirms downregulation of SOCS6 in CRC. In contrast to SOCS2 whose expression is downregulated in colon adenoma and more significantly in CRC, expression of SOCS6 is only downregulated in CRC. Thus, SOCS2 could have a higher diagnostic value as it may serve to diagnose CRC at a very early stage. Interestingly, a recent study by Qiu et al (2013) also showed reduced expression of the same two members of the SOCS family, SOCS2 and SOCS6, in hepatocellular carcinoma. This raises the hypothesis that both proteins could be regulated via a common mechanism and that both SOCS proteins may regulate hand in hand growth signalling in cancer.
In contrast to our study, several studies have shown a lower expression of SOCS1 and SOCS3 among other cancer types (Culig, 2013). Thus, their functions seem to be highly dependent on cell and tumour types. Interestingly, SOCS1 and SOCS3 may play an important role in inflammation-induced CRC as both were overexpressed in IBD according to our bioinformatic study. Along this line, SOCS1 and SOCS3 are often associated with inflammatory disorders in humans (Trengove and Ward, 2013).
One possible mechanism that explains downregulation of SOCS proteins in cancer is methylation in their gene promoter region. CpG islands of the SOCS2 gene were shown to be hypermethylated in endometrial cancer (Fiegl et al, 2004), 6.5% of glioblastoma patients, 14% of primary ovarian cancers (Sutherland et al, 2004) and 43–63% of melanoma patients (Marini et al, 2006; Liu et al, 2008). However, methylation of SOCS2 could not be found in human breast cancer patients (Sutherland et al, 2004). In our study, SOCS2 methylation occurred in 25% of CRC patients. Importantly, the sequence throughout the SOCS2 promoter, which showed methylation compared to the normal samples was already previously analysed and showed methylation in three studies performed on endometrial, ovarian and melanoma cancer patients (Fiegl et al, 2004; Sutherland et al, 2004; Liu et al, 2008). Strikingly, a detailed analysis of the methylated sequence revealed the presence of two STAT GAS motifs, which could explain the downregulation of SOCS2 in CRC as STAT factors are the classical inducers of SOCS family members (Kang et al, 2013). It is further suggested that allelic loss and promoter hypermethylation may account for the major mechanisms leading to SOCS6 inactivation (Lai et al, 2009, 2010). However, we could not identify SOCS6 methylation in our CRC patient cohort. We identified a second mechanism that could contribute to the reduced expression of SOCS2 by demonstrating that the expression levels of the GR are reduced in CRC patients. As regulation of SOCS2 is also dependent on a synergistic action of STAT5 and GR (Tronche et al, 2004; Mueller et al, 2012; Martinez et al, 2013), SOCS2 downregulation in CRC patients may additionally be explained by the reduced expression of the coactivator GR. Further studies are needed to shed more light on the diverse mechanisms of SOCS2 downregulation. Growth hormone signalling components such as GH itself or its cell-surface receptor can also impact on SOCS2 expression levels. Initial results indicate that GH receptor is also downregulated in our CRC samples (data not shown) and may thus contribute to the reduced SOCS2 levels. Current investigations focus on the regulation of other GH signalling components in order to clarify the role of this pathway in CRC. In this context the extent of STAT5 activation will also be of interest as increases in STAT5 phosphorylation will most likely impact on SOCS2 expression (Greenhalgh et al, 2002).
A significantly higher disease-free survival time was observed in patients with high compared to low SOCS2 expression in early CRC stages (stages I and II). It can be speculated that patients with especially low expression of SOCS2 at the early CRC stage may display an exaggerated response to growth-promoting signals such as GH. At later stages this signal may lose some of its significance as the growth-promoting mechanism may increasingly drive tumour development. Along this line, patients with high SOCS2 expression had an improved survival rate and high SOCS2 expression proved to be an independent predictor for good prognosis in breast cancer (Haffner et al, 2007). Furthermore, reduced expression of SOCS2 and SOCS6 correlate with poor prognosis in hepatocellular carcinoma patients (Qiu et al, 2013). However, in our study SOCS6 was not associated with disease outcome. This is in line with two studies performed in prostate cancer and CRC, for which no correlation between SOCS6 mRNA expression and patient survival was found (Storojeva et al, 2005; Zhu et al, 2013). Importantly, up to 30% of stage II patients relapse after surgery and many of them will die due to metastatic disease. Therefore, it is highly important to identify clinical and molecular determinants of outcome in order to target treatments to those patients who are destined to relapse. Our data thus suggest that SOCS2 is an early diagnostic marker for CRC and might be useful as a prognostic marker to delineate a high-risk population at an early CRC stage.
References
Agrawal D, Chen T, Irby R, Quackenbush J, Chambers AF, Szabo M, Cantor A, Coppola D, Yeatman TJ (2002) Osteopontin identified as lead marker of colon cancer progression, using pooled sample expression profiling. J Natl Cancer Inst 94: 513–521.
Betsou F, Lehmann S, Ashton G, Barnes M, Benson EE, Coppola D, DeSouza Y, Eliason J, Glazer B, Guadagni F, Harding K, Horsfall DJ, Kleeberger C, Nanni U, Prasad A, Shea K, Skubitz A, Somiari S, Gunter E (2010) Standard preanalytical coding for biospecimens: defining the sample PREanalytical code. Cancer Epidemiol Biomarkers Prev 19: 1004–1011.
Bolstad BM, Collin F, Simpson KM, Irizarry RA, Speed TP (2004) Experimental design and low-level analysis of microarray data. Int Rev Neurobiol 60: 25–58.
Boyd DB (2003) Insulin and cancer. Integr Cancer Ther 2: 315–329.
Culig Z (2013) Suppressors of cytokine signalling-3 and -1 in human carcinogenesis. Front Biosci (Schol Ed) 5: 277–283.
Elliott J, Hookham MB, Johnston JA (2008) The suppressors of cytokine signalling E3 ligases behave as tumour suppressors. Biochem Soc Trans 36: 464–468.
Erickson HS, Albert PS, Gillespie JW, Rodriguez-Canales J, Marston Linehan W, Pinto PA, Chuaqui RF, Emmert-Buck MR (2009) Quantitative RT-PCR gene expression analysis of laser microdissected tissue samples. Nat Protoc 4: 902–922.
Farabegoli F, Ceccarelli C, Santini D, Taffurelli M (2005) Suppressor of cytokine signalling 2 (SOCS-2) expression in breast carcinoma. J Clin Pathol 58: 1046–1050.
Fiegl H, Gattringer C, Widschwendter A, Schneitter A, Ramoni A, Sarlay D, Gaugg I, Goebel G, Müller HM, Mueller-Holzner E, Marth C, Widschwendter M (2004) Methylated DNA collected by tampons—a new tool to detect endometrial cancer. Cancer Epidemiol Biomarkers Prev 13: 882–888.
Galamb O, Györffy B, Sipos F, Spisák S, Németh AM, Miheller P, Tulassay Z, Dinya E, Molnár B (2008) Inflammation, adenoma and cancer: objective classification of colon biopsy specimens with gene expression signature. Dis Markers 25: 1–16.
Galamb O, Spisák S, Sipos F, Tóth K, Solymosi N, Wichmann B, Krenács T, Valcz G, Tulassay Z, Molnár B (2010) Reversal of gene expression changes in the colorectal normal-adenoma pathway by NS398 selective COX2 inhibitor. Br J Cancer 102: 765–773.
Gentleman R, Carey VJ, Huber W, Irizarry RA, Dudoit S, Smyth G (2005) Bioinformatics and Computational Biology Solutions Using R and Bioconductor. Springer-Verlag.
Greenhalgh CJ, Bertolino P, Asa SL, Metcalf D, Corbin JE, Adams TE, Davey HW, Nicola NA, Hilton DJ, Alexander WS (2002) Growth enhancement in suppressor of cytokine signaling 2 (SOCS-2)-deficient mice is dependent on signal transducer and activator of transcription 5b (STAT5b). Mol Endocrinol 16: 1394–1406.
Greenhalgh CJ, Rico-bautista E, Lorentzon M, Thaus AL, Morgan PO, Willson TA, Zervoudakis P, Metcalf D, Street I, Nicola NA, Nash AD, Fabri LJ, Norstedt G, Ohlsson C, Flores-morales A, Alexander WS, Hilton DJ (2005) SOCS2 negatively regulates growth hormone action in vitro and in vivo. J Clin Invest 115: 397–406.
Haffner MC, Petridou B, Peyrat JP, Révillion F, Müller-Holzner E, Daxenbichler G, Marth C, Doppler W (2007) Favorable prognostic value of SOCS2 and IGF-I in breast cancer. BMC Cancer 7: 136.
Hari DM, Leung AM, Lee J-H, Sim M-S, Vuong B, Chiu CG, Bilchik AJ (2013) AJCC Cancer Staging Manual 7th edition criteria for colon cancer: do the complex modifications improve prognostic assessment? J Am Coll Surg 217: 181–190.
Hendriksen PJM, Dits NFJ, Kokame K, Veldhoven A, van Weerden WM, Bangma CH, Trapman J, Jenster G (2006) Evolution of the androgen receptor pathway during progression of prostate cancer. Cancer Res 66: 5012–5020.
Hoefer J, Kern J, Ofer P, Eder IE, Schäfer G, Dietrich D, Kristiansen G, Geley S, Rainer J, Gunsilius E, Klocker H, Culig Z, Puhr M (2013) SOCS2 correlates with malignancy and exerts growth promoting effects in prostate cancer. Endocr Relat Cancer 21: 175–187.
Horvat S, Medrano JF (2001) Lack of Socs2 expression causes the high-growth phenotype in mice. Genomics 72: 209–212.
Iglesias-Gato D, Chuan Y-C, Wikström P, Augsten S, Jiang N, Niu Y, Seipel A, Danneman D, Vermeij M, Fernandez-Perez L, Jenster G, Egevad L, Norstedt G, Flores-Morales A (2014) SOCS2 mediates the cross talk between androgen and growth hormone signaling in prostate cancer. Carcinogenesis 35: 24–33.
Inagaki-Ohara K, Kondo T, Ito M, Yoshimura A (2013) SOCS, inflammation, and cancer. JAKSTAT 2: e24053.
Johnston PG (2005) Stage II colorectal cancer: to treat or not to treat. Oncologist 10: 332–334.
Jorissen RN, Gibbs P, Christie M, Prakash S, Lipton L, Desai J, Kerr D, Aaltonen LA, Arango D, Kruhøffer M, Orntoft TF, Andersen CL, Gruidl M, Kamath VP, Eschrich S, Yeatman TJ, Sieber OM (2009) Metastasis-associated gene expression changes predict poor outcomes in patients with dukes stage B and C colorectal cancer. Clin Cancer Res 15: 7642–7651.
Kang K, Robinson GW, Hennighausen L (2013) Comprehensive meta-analysis of signal transducers and activators of transcription (STAT) genomic binding patterns discerns cell-specific cis-regulatory modules. BMC Genomics 14: 4.
Kim GH, Lee DH, Kim HW, Cheong JY, Seo SB, Heo J, Kang DH, Song GA, Cho M, Yang US, Park DY, Yoo MA (2005) Relationships between the expressions of CDX1 and CDX2 mRNA and clinicopathologic features in colorectal cancers. Korean J Intern Med 20: 317–324.
Kobayashi D, Nomoto S, Kodera Y, Fujiwara M, Koike M, Nakayama G, Ohashi N, Nakao A (2012) Suppressor of cytokine signaling 4 detected as a novel gastric cancer suppressor gene using double combination array analysis. World J Surg 36: 362–372.
Lai R-H, Hsiao Y-W, Wang M-J, Lin H-Y, Wu C-W, Chi C-W, AF-Y Li, Jou Y-S, Chen J-Y (2010) SOCS6, down-regulated in gastric cancer, inhibits cell proliferation and colony formation. Cancer Lett 288: 75–85.
Lai R-H, Wang M-J, Yang S-H, Chen J-Y (2009) Genomic organization and functional characterization of the promoter for the human suppressor of cytokine signaling 6 gene. Gene 448: 64–73.
LaPointe LC, Dunne R, Brown GS, Worthley DL, Molloy PL, Wattchow D, Young GP (2008) Map of differential transcript expression in the normal human large intestine. Physiol Genomics 33: 50–64.
Liu S, Ren S, Howell P, Fodstad O, Riker AI (2008) Identification of novel epigenetically modified genes in human melanoma via promoter methylation gene profiling. Pigment Cell Melanoma Res 21: 545–558.
Marini A, Mirmohammadsadegh A, Nambiar S, Gustrau A, Ruzicka T, Hengge UR (2006) Epigenetic inactivation of tumor suppressor genes in serum of patients with cutaneous melanoma. J Invest Dermatol 126: 422–431.
Marisa L, de Reyniès A, Duval A, Selves J, Gaub MP, Vescovo L, Etienne-Grimaldi M-C, Schiappa R, Guenot D, Ayadi M, Kirzin S, Chazal M, Fléjou J-F, Benchimol D, Berger A, Lagarde A, Pencreach E, Piard F, Elias D, Parc Y, Olschwang S, Milano G, Laurent-Puig P, Boige V (2013) Gene expression classification of colon cancer into molecular subtypes: characterization, validation, and prognostic value. PLoS Med 10: e1001453.
Martinez CS, Piazza VG, Ratner LD, Matos MN, González L, Rulli SB, Miquet JG, Sotelo AI (2013) Growth hormone STAT5-mediated signaling and its modulation in mice liver during the growth period. Growth Horm IGF Res 23: 19–28.
Metcalf D, Greenhalgh CJ, Viney E, Willson TA, Starr R, Nicola NA, Hilton DJ, Alexander WS (2000) Gigantism in mice lacking suppressor of cytokine signalling-2. Nature 405: 1069–1073.
Michaylira CZ, Ramocki NM, Simmons JG, Tanner CK, McNaughton KK, Woosley JT, Greenhalgh CJ, Lund PK (2006) Haplotype insufficiency for suppressor of cytokine signaling-2 enhances intestinal growth and promotes polyp formation in growth hormone-transgenic mice. Endocrinology 147: 1632–1641.
Miller ME, Michaylira CZ, Simmons JG, Ney DM, Dahly EM, Heath JK, Lund PK (2004) Suppressor of cytokine signaling-2: a growth hormone-inducible inhibitor of intestinal epithelial cell proliferation. Gastroenterology 127: 570–581.
Mueller KM, Themanns M, Friedbichler K, Kornfeld J-W, Esterbauer H, Tuckermann JP, Moriggl R (2012) Hepatic growth hormone and glucocorticoid receptor signaling in body growth, steatosis and metabolic liver cancer development. Mol Cell Endocrinol 361: 1–11.
Newton VA, Ramocki NM, Scull BP, Simmons JG, McNaughton K, Lund PK (2010) Suppressor of cytokine signaling-2 gene disruption promotes Apc(Min/+) tumorigenesis and activator protein-1 activation. Am J Pathol 176: 2320–2332.
Qiu X, Zheng J, Guo X, Gao X, Liu H, Tu Y, Zhang Y (2013) Reduced expression of SOCS2 and SOCS6 in hepatocellular carcinoma correlates with aggressive tumor progression and poor prognosis. Mol Cell Biochem 378: 99–106.
Rigby RJ, Simmons JG, Greenhalgh CJ, Alexander WS, Lund PK (2007) Suppressor of cytokine signaling 3 (SOCS3) limits damage-induced crypt hyper-proliferation and inflammation-associated tumorigenesis in the colon. Oncogene 26: 4833–4841.
Sabates-Bellver J, Van der Flier LG, de Palo M, Cattaneo E, Maake C, Rehrauer H, Laczko E, Kurowski MA, Bujnicki JM, Menigatti M, Luz J, Ranalli TV, Gomes V, Pastorelli A, Faggiani R, Anti M, Jiricny J, Clevers H, Marra G (2007) Transcriptome profile of human colorectal adenomas. Mol Cancer Res 5: 1263–1275.
Sasi W, Jiang WG, Sharma A, Mokbel K (2010) Higher expression levels of SOCS 1,3,4,7 are associated with earlier tumour stage and better clinical outcome in human breast cancer. BMC Cancer 10: 178.
Skrzypczak M, Goryca K, Rubel T, Paziewska A, Mikula M, Jarosz D, Pachlewski J, Oledzki J, Ostrowski J, Ostrowsk J (2010) Modeling oncogenic signaling in colon tumors by multidirectional analyses of microarray data directed for maximization of analytical reliability. PLoS One 5: pii: e13091.
Smith JJ, Deane NG, Wu F, Merchant NB, Zhang B, Jiang A, Lu P, Johnson JC, Schmidt C, Bailey CE, Eschrich S, Kis C, Levy S, Washington MK, Heslin MJ, Coffey RJ, Yeatman TJ, Shyr Y, Beauchamp RD (2010) Experimentally derived metastasis gene expression profile predicts recurrence and death in patients with colon cancer. Gastroenterology 138: 958–968.
Storojeva I, Boulay J-L, Ballabeni P, Buess M, Terracciano L, Laffer U, Mild G, Herrmann R, Rochlitz C (2005) Prognostic and predictive relevance of DNAM-1, SOCS6 and CADH-7 genes on chromosome 18q in colorectal cancer. Oncology 68: 246–255.
Sutherland KD, Lindeman GJ, Choong DYH, Wittlin S, Brentzell L, Phillips W, Campbell IG, Visvader JE (2004) Differential hypermethylation of SOCS genes in ovarian and breast carcinomas. Oncogene 23: 7726–7733.
Trengove MC, Ward AC (2013) SOCS proteins in development and disease. Am J Clin Exp Immunol 2: 1–29.
Tronche F, Opherk C, Moriggl R, Kellendonk C, Reimann A, Schwake L, Reichardt HM, Stangl K, Gau D, Hoeflich A, Beug H, Schmid W, Schütz G (2004) Glucocorticoid receptor function in hepatocytes is essential to promote postnatal body growth. Genes Dev 18: 492–497.
Tsukamoto S, Ishikawa T, Iida S, Ishiguro M, Mogushi K, Mizushima H, Uetake H, Tanaka H, Sugihara K (2011) Clinical significance of osteoprotegerin expression in human colorectal cancer. Clin Cancer Res 17: 2444–2450.
Walters S, Maringe C, Butler J, Brierley JD, Rachet B, Coleman MP (2013) Comparability of stage data in cancer registries in six countries: lessons from the International Cancer Benchmarking Partnership. Int J Cancer 132: 676–685.
Wikman H, Kettunen E, Seppänen JK, Karjalainen A, Hollmén J, Anttila S, Knuutila S (2002) Identification of differentially expressed genes in pulmonary adenocarcinoma by using cDNA array. Oncogene 21: 5804–5813.
Yoon S, Yi Y-S, Kim SS, Kim J-H, Park WS, Nam SW (2012) SOCS5 and SOCS6 have similar expression patterns in normal and cancer tissues. Tumour Biol 33: 215–221.
Zhu J, Dai Q, Han Z, He H, Mo R, Chen G, Chen Y, Wu Y, Yang S, Jiang F, Chen W, Sun Z, Zhong W (2013) Expression of SOCSs in human prostate cancer and their association in prognosis. Mol Cell Biochem 381: 51–59.
Acknowledgements
We would like to thank the Fondation Cancer for funding this study and the Fonds National de la Recherche (FNR) for supporting KB under the AFR grant scheme. We are also grateful to the Fondation du Pélican de Mie et Pierre Hippert-Faber, under the aegis of the Fondation de Luxembourg for supporting KB via a Pelican grant. We would also like to thank the contributing surgeons from Centre Hospitalier du Luxembourg and Centre Hospitalier Emile Mayrisch and the nurses of the Clinical and Epidemiological Investigation Center of CRP-Santé for working with the patients, as well as Dr Nikolai Goncharenko and Dr Fay Betsou from the IBBL for the overall set up of the sample collection for the study. We thank Dr Sophie Losciuto for assistance with LMD and Dr Christelle Bahlawane for informative discussions and critical reading of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License.
Supplementary Information accompanies this paper on British Journal of Cancer website
Supplementary information
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Letellier, E., Schmitz, M., Baig, K. et al. Identification of SOCS2 and SOCS6 as biomarkers in human colorectal cancer. Br J Cancer 111, 726–735 (2014). https://doi.org/10.1038/bjc.2014.377
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2014.377
Keywords
This article is cited by
-
Pseudogenes and the associated ceRNA network as potential prognostic biomarkers for colorectal cancer
Scientific Reports (2022)
-
Aspects of Modern Biobank Activity – Comprehensive Review
Pathology & Oncology Research (2018)
-
Loss of Myosin Vb in colorectal cancer is a strong prognostic factor for disease recurrence
British Journal of Cancer (2017)
-
NOD1 modulates IL-10 signalling in human dendritic cells
Scientific Reports (2017)
-
Development and Validation of a Three-gene Prognostic Signature for Patients with Hepatocellular Carcinoma
Scientific Reports (2017)