Abstract
Background:
The multi-exon CSPP1 gene, encoding for centrosome and microtubule-associated proteins involved in ciliogenesis and cell division, is a candidate oncogene in luminal breast cancer but expression of CSPP1 proteins remained unexplored.
Methods:
CSPP1 gene and protein expression was examined in normal mammary tissue, human breast cancer cell lines, and primary breast cancer biopsies from two patient cohorts. Cell type and epitope-dependent subcellular-specific CSPP1 staining pattern in normal mammary gland epithelium and cancer biopsies were correlated to molecular and clinical parameters.
Results:
A novel, nuclear localised CSPP1 isoform was exclusively detected in luminal epithelial cells, whereas cytoplasmic CSPP-L was generally expressed in normal mammary epithelium. Luminal cell-related nuclear CSPP1 expression was preserved in type-matched cell lines and carcinomas, and correlated to gene copy number and mRNA expression. In contrast, basal-like carcinomas displayed generally lower CSPP1 mRNA expression. Yet, a subgroup of basal-like breast carcinomas depicted nuclear CSPP1 expression, displayed luminal traits, and differed from nuclear CSPP1 devoid counterparts in expression of eight genes. Eight-gene signature defined groups of basal-like tumours from an independent cohort showed significant differences in survival.
Conclusions:
Differential expression of a nuclear CSPP1 isoform identified biologically and clinically distinct subgroups of basal-like breast carcinoma.
Similar content being viewed by others
Main
Breast cancer is the most frequent cancer in women today, accounting for ∼23% of all new diagnosed female cancers and for ∼14% cancer-related deaths among women (2008, IARC). Breast cancer is a heterogeneous disease reflected by the existence of different subtypes and clinical behaviour. Common treatment modalities consist mainly of surgery, radiation therapy, chemotherapy, hormone therapy, and HER2-targeted treatment. To date, choice of treatment modalities mainly depend on histological evaluation, lymph node assessment, and a few molecular markers (oestrogen receptor, progesterone receptor, KI67, and HER2 (ERBB2)). A decade ago, large-scale gene expression-based molecular classification of breast cancer identified subgroups where the most prominent types (luminal and basal-like subtypes) had similarities with normal mammary epithelia cell types (reviewed in Cheang et al, 2008). This ‘intrinsic’ classification has later been modified by using a set of 50 genes (PAM50) (Parker et al, 2009), and a set of markers for immunohistochemistry (IHC) is considered to be sufficient as surrogate diagnostic biomarkers to distinguish the five defined subgroups: basal-like, HER2-enriched, luminal A, luminal B, and normal-like (Callagy et al, 2003; Blows et al, 2010). Patients with luminal A and luminal B breast cancer generally have better prognosis than patients diagnosed with basal-like carcinoma, although a subset of luminal tumours exhibit a possibility for dormant disease and late recurrences with metastases. Overexpression or amplification of HER2 identifies a patient group with particularly poor disease-specific survival (Chin et al, 2006), but this has improved after the introduction of HER2-targeted therapy. Despite these advances, the need for better prognostic and predictive markers persists, as heterogeneity in patient outcome exists within subgroups. Several large-scale genomic approaches have been employed to gain further insight into the aetiology of breast cancer and to identify clinical outcome predicting factors within subtypes (Reis-Filho and Pusztai, 2011).
Numerical and structural aberrations of centrosomes are frequent in breast cancer and are correlated to tumour aggressiveness and metastatic lesions (D'Assoro et al, 2002; Salisbury et al, 2004; Pujana et al, 2007; Guo et al, 2007). BRCA1 mutations and constitutive oestrogen receptor signalling are promoting factors of centrosome aberrations (Li et al, 2004; Kais and Parvin, 2008). The centrosome functions as an organisation center of the microtubule cytoskeleton and hub for several signalling pathways (Nigg and Raff, 2009). These include in particular proliferation and planar-cell-polarity affecting pathways that are integrated through the centrosome-templated primary cilium (PDGFRα, Wnt-, and Hedgehog (Hh)-signalling). Primary cilia are important for (murine) mammary gland morphogenesis, which is characterised by extensive expansion, branching, and differentiation of the mammary epithelia cells (Visvader, 2009; McDermott et al, 2010). Primary cilia are displayed on myo- and luminal epithelia cells while undergoing branching morphogenesis, but are lost from luminal epithelia cells when branching is completed. Importantly, cilia dysfunction conferred decreased Hh signalling impaired duct extension and branching in a murine model (McDermott et al, 2010). Centrosome dysfunction may, thus, sustain cancer development and progression by aberrant spindle assembly and mis-segregation of chromosomes as well as deregulation of important signalling pathways. Indeed, cilia-dependent Wnt and Hh signalling have been associated with human basal-like carcinomas (Kasper et al, 2009), and specific chromosome segregation/cell division defects may be related to ploidy differences in specific breast cancer subtypes (Van Loo et al, 2010).
In the present study, we investigated the expression and localisation of centrosome/spindle pole-associated protein 1 isoforms (CSPP1). CSPP1 was originally identified as a proto-oncogene in human B-cell lymphoma (Patzke et al, 2005). In a cohort of breast adenocarcinomas, CSPP1 was identified as a candidate oncogene in luminal type breast cancer on the basis of gene dosage correlated overexpression (Adelaide et al, 2007). The CSPP1 locus is a large, multi-exon locus encompassing 13 420 kb on chromosome 8q13.2 (Supplementary Figure S1). Multiple splice isoforms are predicted to be expressed, of which to date, two splice isoforms of CSPP1 (CSPP and CSPP-L) have been characterised. These function in cell cycle control, cilia formation, cytoskeleton organisation, and cell division (Patzke et al, 2005; Patzke et al, 2006; Asiedu et al, 2009; Patzke et al, 2010). Overexpression of CSPP1 proteins in different epithelia cell lines conferred aneuploidy by aberrant spindle formation, whereas CSPP1 depletion promoted cytokinesis failure and loss of primary cilia formation. To investigate the potential role of CSPP1 in mammary gland malignancies, we studied CSPP1 gene and protein expression in the human mammary gland, breast cancer cell lines, and patient cohorts with primary operable breast cancer. We report that an epithelial cell type-dependent CSPP1 protein expression pattern found in the normal mammary gland resulted in identification of subgroups of basal-like breast carcinomas with different outcomes and different molecular properties that may be exploited for pharmaceutical intervention.
Materials and methods
Cell lines, cell culture, and transfection
Breast cancer cell lines used in this study (MCF7, ZR-75-1, BT-474, UACC-812, HCC1937, HCC38, MDA-MB-231, MCF10A), are of the authenticated ATCC ICBP-43 Breast Cancer Panel and were cultivated according to ATCC’s subculturing procedures (#30-4500 K, ATCC, Manassas, VA, USA). Human embryonic kidney 293T (Hek293T) cells were maintained in DMEM including 10% fetal calf serum and antibiotics (Life Technologies, Carlsbad, CA, USA). For transfection, cells were grown in 10-cm plates (Becton and Dickinson, San Jose, CA, USA) and transfected in Optimem (Life Technologies) with 5 μg plasmid DNA using Lipofectamine2000 (Life Technologies) following the manufacturer’s instructions. Knockdown of CSPP1 mRNA expression was performed using shRNA and GFP co-expressing plasmids. For analysis, 72 h post transfection cells were trypsinised and sorted for GFP expression on a FACS Diva instrument (Becton and Dickinson).
Immunofluorescence staining and Imaging
Immunofluorescence microscopy of cells grown on coverslips and general staining procedures were as described earlier (Patzke et al, 2010). Paraffin-embedded tissue sections were treated with 3 × 100% xylene, followed by a double 100% ethanol wash, a 70% ethanol wash, a 50% ethanol wash, and a double water wash. Slides were incubated 30 min in pre-heated 95 °C target retrieval solution (10 mM Tris, 1 mM EDTA; pH9), followed by incubations for 20 min at r.t. and 5 min in running water. Fluorescence images were acquired using appropriate optical filters on an AxioImager Z1 ApoTome microscope system (Carl Zeiss, Jena, Germany) equipped with a × 100 or a × 63 lens (both PlanApo N.A.1.4) and an AxioCam MRm camera. To display the entire cell volume, images are presented as maximal projections of z-stacks using Axiovision 4.8.2 (Carl Zeiss).
Antibodies and plasmids
Antibodies used in this study were as follows: anti-CSPP-L (Proteintech Europe, Manchester, United Kingdom; used for IF at 1 : 1000), anti-CSPP/CSPP-L (described in (Patzke et al, 2010); IF 1 : 200), anti-cytokeratin-8 (AbCam, Cambridge, UK; IF 1 : 500), anti-smooth muscle actin (Abcam; IF 1 : 100), anti-Cyclin A (Santa Cruz Biotechnology, Santa Crux, CA, USA; IF 1 : 1000), anti-γ-Tubulin (Sigma, St Louis, MO, USA; IF 1 : 500), and anti-Pericentrin (Abcam; IF 1 : 400). Secondary fluorochrome conjugated antibodies (Donkey anti-rabbit DyLight488; Donkey anti-mouse DyLight549; IF 1 : 1000) were purchased from Jackson Immuno Research (West Grove, PA, USA). shRNA and GFP co-expression plasmids used for CSPP1 knockdown were obtained from SA Biosciences (part of Qiagen N.V., Venlo, Netherlands): set KH18087G with the following target sequences: shRNA_01: 5′-gcacgaattcagcaggagtat-3′, shRNA_02: 5′-tccttcagttgacagcatcat-3′, shRNA_03: 5′-ggtgccaaagttgacttagat-3′, shRNA_04: 5′-ggaggtgaagatcgagaactt-3′, and shRNA_control: 5′-ggaatctcattcgatgcatac-3′. Tagged CSPP and CSPP-L full-length and truncation protein expression plasmids pCSPPmyc (derived from AJ583433) and pCSPP-Lmyc (derived from AM156947), pCSPP(498-876-eGFP) were described earlier (Patzke et al, 2006).
Immunoblotting
Preparation of cell lysates and immunoblotting was performed as previously described (Patzke et al, 2010).
Patients and tumour specimens
Tissue microarrays suitable for IHC analysis were composed from a series of early stage breast cancer patients (‘Oslo0/ULL’). The clinical-patholigical details of this series is previously published (Langerod et al, 2007). Formalin-fixed and paraffin-embedded tissue were available from 170 out of 212 patients. Three representative 0.6-mm cores were selected from each tumour and composed together in recipient paraffin blocks. Both CSPP1-specific antibodies detect CSPP1 proteins in formalin-fixed, paraffin-embedded tissue sections (Patzke et al, 2010). Epitopes were retrieved in Tris/EDTA high pH (10 mM Tris Base, 1 mM EDTA) and antibody staining (anti-CSPP/CSPP-L (1 : 2000; 0.5 μg ml−1 f.c.) and anti-CSPP-L (1 : 1000; 1.5 μg ml−1 f.c.)) visualised using Envision+peroxidase system (Dako, Denmark A/S, Glostrup, DK). Slides were scanned and archived using TMA-ImageAnalyzer (beta-release; to be available through Room4 Group Ltd. (Crowborough, UK). Localisation and intensity of CSPP1 staining were scored in a semi quantitative manner by two observers (HGR, BR). Cores with discrepant scores were collectively studied by microscopy to obtain consensus. For patients with discrepant results between the three cores, the core with highest score was selected as representative. Cores with <50 tumour cells were treated as non determinable, leaving a total of 135 cases with informative staining.
Gene expression data and DNA copy number data was available for 80 patients in the Oslo0/ULL series and for 115 patients from a second, early stage breast cancer cohort (Oslo1/MicMa) (Langerod et al, 2007; Naume et al, 2007; Russnes et al, 2010; Enerly et al, 2011). Gene expression data was available for further 1974 clinically annotated patients with PAM50-based subtype definition and integrative cluster group assignment (METABRIC, (Curtis et al, 2012)).
Gene annotation mapping
Expression data were annotated using Entrez gene identities. For Oslo0/ULL samples, annotations for Stanford 43k cDNA array were retrieved from SMD SOURCE (http://smd.stanford.edu/cgi-bin/source/sourceSearch; UniGene Build.222). For Oslo1/MicMa and METABRIC samples, Agilent and Illumina-HT12_v3 probes were mapped to Entrez or Ensembl gene IDs, respectively, using BioMart through R library biomaRt (Ensembl release 54/NCBI36 (hg18) human assembly). CSPP1 probes were mapped to each of the individual copy number datasets and expression sets through genomic region and Entrez gene ID, respectively. If multiple probes were mapped to CSPP1, expression values of these were averaged for each sample.
PAM50 classification
Molecular subtype classification was carried out using PAM50 for each of the Oslo cohorts (Parker et al, 2009). Gene median centring was performed on the expression set, where the median of the expression values for a specific gene across all samples was subtracted from that gene. Subtype assignment was based on the nearest of the five centroids (distances calculated using Spearman rank correlation to the centroids) where the assigned subtype on a sample corresponded to the centroid with the nearest correlation. No threshold was set on the correlation when performing subtyping.
Correlation analysis
The Cis-correlation between CSPP1 expression and DNA copy number variation was quantified by Pearson correlation and was also explored by a univariate linear regression model with expression values as variable and copy number variation as response. Two-tail t-test was carried out to compare the CSPP1 mRNA expression between any pair of protein levels. statistical analysis of microarray (SAM) analysis was based on response type of ‘two class unpaired’ and 10 000 permutations.
Survival analysis
Endpoint for the survival analysis was breast cancer-specific death measured from the date of surgery to death of the disease or otherwise censored at the time of the last follow-up visit or noncancer-related death. Kaplan–Meier survival curves for time to breast cancer-specific death were constructed in SPSS (PASW statistics 18), and P-values determined with the log-rank test.
Hierarchical clustering
Hierarchical clustering was performed using Cluster 3.0 software (Eisen et al, 1998). Data were centred to median expression of each gene across the normalised expression dataset, clustered using correlation (uncentered) similarity matrix and the average linkage algorithm, and visualised using TreeView (Eisen lab, University of California at Berkeley, CA, USA). Robustness of cluster group formation was tested by alternative clustering using centroid and complete linkage algorithms (data not shown).
Results
Distinct epithelial expression pattern of CSPP1 proteins in the human mammary gland
Expression of different CSPP1 proteins in the breast epithelium was initially studied using antibodies directed to the common C-terminal domain of CSPP and CSPP-L (a-CSPP/CSPP-L; monoclonal) or to the CSPP-L-specific N-terminal domain (a-CSPP-L; polyclonal; see Supplementary Figure S1). Both antibodies prominently stained epithelial cells in human mastectomy tissue sections (Figure 1 and Supplementary Figure S2). The a-CSPP-L antibody showed exclusively cytoplasmic staining of mammary epithelial cells and endothelial cells of blood vessels, whereas an additional, nuclear expression of CSPP1 proteins was detected solely by the a-CSPP/CSPP-L antibody (Figure 1A). This nuclear expression was apparent in luminal oriented epithelial cells of well-formed ducts, but absent in the myoepithelial cell layer. In less organised ductal structures of terminal duct-lobular units, nuclear CSPP1 expression was found variable, frequently increasing in intensity in centrally positioned cells. The most intense nuclear CSPP1 staining was seen in the luminal cells in lactating glands, suggesting a cell type-dependent nuclear expression of CSPP1 in the epithelium of the mammary gland. We, therefore, co-investigated the nuclear expression of CSPP1 and cytokeratin-8 (Taylor-Papadimitriou et al, 1989), or smooth muscle actin (Deugnier et al, 1995) respectively, by immunofluorescence staining of normal breast epithelium. Cytokeratin-8-positive, luminal epithelial cells showed nuclear and cytoplasmic CSPP1 expression (Figure 1B). Smooth muscle actin-positive, myoepithelial cells showed only cytoplasmic CSPP1 expression supporting the notion of CSPP1 being differentially expressed in lineage-specific epithelial cells (Figure 1C).
Expression of CSPP1 proteins in normal mammary epithelia. (A) Immunohistochemical staining of CSPP1 proteins with a monoclonal antibody against the common C-terminal domain of CSPP and CSPP-L (a-CSPP/CSPP-L) in normal mammary gland tissue showing general CSPP1 expression in epithelia cells with prominent selective nuclear staining in luminal epithelial cells. (B,C) Immunofluorescence co-staining of CSPP1 proteins (red) and Cytokeratin-8 (green, B) or smooth muscle actin (green, C), respectively, showing co-staining of nuclear CSPP1-positive cells with the luminal lineage Cytokeratin-8.
Differential expression and localisation of CSPP1 proteins in human breast cancer cell lines
We next investigated CSPP1 expression in breast cancer cell lines of different subtypes to validate the previously unnoticed nuclear expression. Both antibodies stained CSPP1 proteins at centrosomes (including supernumary centrioles), the kinetochore fibres and the cytokinetic bridge in all tested cell lines (Figure 2A and C). In concordance with staining pattern observed in other epithelial cell lines (hTERT-RPE1; HeLa), both antibodies differed in staining intensity and affinity at the centrosome (Patzke et al, 2010). In agreement with the staining pattern in mammary tissue, nuclear expression of CSPP1 proteins was detected exclusively with the a-CSPP/CSPP-L antibody in luminal type cell lines (MCF7, ZR-75-1, BT-474, UACC-812). Nuclear staining with this antibody was largely diminished or absent in basal-like breast cancer cell lines (HCC1937, MDA-MB-231)(Figure 2A). In contrast, a more prominent CSPP-L staining was found in the two basal-like cell lines than in the four luminal cell lines. In MDA-MB-231 cells and in the non-tumourigenic breast epithelial cell line, MCF10A primary cilia were observed, differing in frequency and length (8.3±1.4%, 4.0±0.5 μm MCF10A; 1.7±0.3%, 2.0±0.5 μm MDA-MB-231). CSPP-L decorated the basal body and the cilia axoneme in both cell lines (Figure 2D). No cilia were observed in luminal cell lines. These data are in agreement with the study of human breast cancer cell lines by Yuan et al, 2010. In correlation with immunofluorescence staining, basal-like cell lines showed higher CSPP-L protein expression than luminal cell lines in immunoblots of total cell lysates and displayed higher frequency of centrosome amplification (Supplementary Figure S3).
Differential expression and localisation of CSPP1 proteins in human breast cancer cell lines. (A) Low-magnification images of immunofluorescence stained luminal type (ZR-75-1, MCF7, UACC-812, BT-474) and basal-like (HCC1937, MDA-MB-231) breast cancer cell lines with the common CSPP and CSPP-L C-terminus directed antibody (a-CSPP/CSPP-L) and a CSPP-L-specific antibody (a-CSPP-L) shows nuclear a-CSPP/CSPP-L staining in luminal type cancer cell lines. Individual channels (a-CSPP-L, green; a-CSPP/CSPP-L, red; DNA, blue) and a colour merged image are shown for each cell line. Images were acquired at identical imaging settings and are presented at identical contrast scales. (B) High-magnification imaging shows centrosomal co-localisation of a-CSPP/CSPP-L staining (red) and a-Pericentrin (centrosome, green) and (C) co-staining of a-CSPP/CSPP-L (red) and a-CSPP-L (green) at the mitotic spindle (ZR-75-1), centrosomes and cytokinetic bridge microtubules (HCC1937). (D) a-CSPP-L (green) staining is also detected at primary cilia (a-glutamylated tubulin, red) in MCF10A and MDA-MB-231, which differs in ciliation frequency and cilia morphology.
Validation and characterisation of nuclear CSPP1 protein expression
The a-CSPP/CSPP-L antibody was raised against the common C-terminal 291 amino acids of CSPP and CSPP-L and centrosome staining by this antibody is sensitive to CSPP1 targeting siRNAs (Patzke et al, 2010). Nuclear staining could have been caused by cross-specificity to a lineage-specific, non-CSPP1 protein. Therefore, ZR-75-1 cells were transfected individually with plasmids co-expressing turbo-EGFP and either a scrambled control or different CSPP1-specific shRNAs (Figure 3A). CSPP1 targeting shRNAs 2 and 4 but not the scrambled control shRNA significantly decreased a-CSPP/CSPP-L nuclear staining in ZR-75-1 transfectants. Knockdown efficacy and specificity of these shRNAs was assessed further by immunoblotting for CSPP-L on GFP-positive Hek293T cell transfectants 72 h post transfection, as only a-CSPP-L detects endogenous levels of CSPP1 protein in total cell lysates and Hek293T cells show high endogenous CSPP-L expression (Patzke et al, 2010). Endogenous expression of CSPP-L was strongly reduced by shRNAs 2 and 4 relative to the expression of γ-tubulin compared with scrambled control shRNA (Figure 3B).
Characterisation of nuclear CSPP1 protein expression. (A) Transient transfection of endogenous nuclear CSPP1 expressing ZR-75-1 cells with selected shRNA and eGFP co-expressing plasmids proof nuclear CSPP1 staining specificity of the a-CSPP/CSPP-L antibody (red). Bar diagram shows quantification of nuclear CSPP1 staining intensity in control and CSPP1 targeting shRNA transfectants co-expressing eGFP (error bars depict s.e.m from three independent experiments (300 eGFP-positive cells/experiment). (B) a-CSPP-L immunoblotting of total cell lysates from Hek293T cells transiently transfected with plasmids expressing distinct CSPP1 mRNA targeting shRNAs or a control shRNA. Co-expression of eGFP allowed flowcytometric sorting of transfectants and identifies CSPP1 targeting shRNAs with highest knockdown efficacy compared with control protein γ-tubulin. (C) Ectopically expressed Myc-tag fusion proteins of CSPP or CSPP-L do not localise to the nucleus of ZR-75-1 cells, but show centrosomal and cytoskeletal staining (a-myc, red; a-Pericentrin, green). (D) The ectopically expressed eGFP-taged common C-terminal domain of CSPP/CSPP-L (CSPP498–876-eGFP, green) shows nuclear localisation in ZR-75-1 and HCC1937 cells and comprises the a-CSPP/CSPP-L target moiety. (E) Nuclear CSPP1 protein expression (a-CSPP/CSPP-L, red) is detected throughout interphase in ZR-75-1 cells, as evaluated by co-staining of cyclin A (green; devoid in G1 phase, increasing nuclear expression in S-phase and high nuclear and cytoplasmic expression in G2 phase). DNA is counterstained in all immunofluoresence experiments by Hoechst 33258 and depicted in blue.
Still, nuclear CSPP1 protein detection could reflect a post-translational regulation of earlier described isoforms, CSPP or CSPP-L, or originate from a yet undefined CSPP1 splice isoform comprising their common C-terminal domain. Therefore, luminal type ZR-75-1 cells were transfected with expression plasmids of C-terminally Myc-tagged CSPP or CSPP-L proteins (pCSPPmyc and pCSPP-Lmyc). Nuclear Myc epitope staining would be expected if these isoforms or a proteolytically derived C-terminal fragment underlie cell type-specific nuclear translocation. Neither of them showed nuclear staining. Similar to results obtained in other epithelial cell lines, both isoforms colocalised with Pericentrin at centrosomes and decorated microtubules (Figure 3C). Having excluded CSPP and CSPP-L from being the nuclear antigen, ZR-75-1 and HCC1937 cells were transfected with an expression plasmid for the common C-terminal 379 amino acids of CSPP and CSPP-L C-terminally fused to eGFP (pCSPP498-876-eGFP) (Patzke et al, 2006). This construct lacks the domains that are required for association with microtubules. Translation of this mRNA (e.g. human cDNA AK026143) is driven by an alternative start codon. This ectopically expressed protein showed nuclear localisation in luminal ZR-75-1 and basal-like HCC1937 transfectants (Figure 3D). Finally, to test for a putative cell cycle phase-dependent regulation of the endogenous nuclear CSPP1 protein, we stained for co-expression of cyclin A in ZR-75-1 cells. Nuclear CSPP1 was detected in all interphase stages based on Cyclin A expression pattern (G1- (cyclin A negative), S- (increasing nuclear cyclin A levels) and G2 phase (cytoplasmic cyclin A); Figure 3E).
CSPP1 in primary operable breast carcinomas
CSPP1 protein expression in 135 patients with primary operable breast cancer (Oslo0/ULL) was determined by IHC on tissue microarrays. CSPP1 expression varied both within and between tumour samples (Figure 4 and Supplementary Table 1). Tumours were grouped into four categories based on percentage of nuclear-positive tumour cells (0=0–2%; 1=2–50%, 2=50–75%, and 3=>75%). Cytoplasmic staining of tumour cells was scored only by intensity. Only 10 cases stained negative in the nucleus, whereof four also stained negative in the cytoplasm. Cases with more abundant nuclear stain (>50%) had a large variation from negative to strong staining of the cytoplasm. No correlations between cytoplasmic expression and clinical relevant parameters were identified (data not shown). The distribution of clinical-pathological parameters in the different IHC groups showed very little variation except for histological type where infiltrative lobular carcinomas always had >50% CSPP1 stained nuclei (Figure 4A and G & Supplementary Table 2).
Immunohistochemical staining of Oslo0/Ull primary operable breast cancer cohort tissue microarray with a-CSPP/CSPP-L and correlation of nuclear CSPP1 protein and CSPP1 mRNA expression in breast cancer subtypes. (A-F) Examples of observed staining pattern in tissue array samples were as follows: nuclear and cytoplasm negative (A), nuclear negative and cytoplasm positive (B), 2–50% fraction of nuclear CSPP1-positive breast cancer cells (C), 50–75% fraction of nuclear CSPP1-positive breast cancer cells (D) >75% fraction of nuclear CSPP1-positive breast cancer cells (E, F). Nuclear staining intensity is consistently high in biopsies displaying >75% fraction of nuclear-positive cancer cells. (G) Summary table of nuclear CSPP1 staining in respect to tumour histology. Cancer biopsies of lobular growth histology do correlate with nuclear CSPP1 expression (P-value Fisher’s exact test). (H) Colour coded Whisker hair plot of mean CSPP1 mRNA expression for 52 cases with immunohistochemistry and gene expression data, (I) Whisker hair plot of mean CSPP1 mRNA expression levels in PAM50 breast cancer subtypes (n=80). Statistically significant (P<0.05) mean gene expression differences are indicated with star symbols (see also Supplementary Figure S3).
Gene copy number and expression data were available for 80 patients. For 52 of these, IHC data were scored, allowing further exploration of CSPP1 expression and regulation. The correlation between gene expression of three probes representing CSPP1 (Supplementary Figure S1) and protein expression was PAM50 subtype dependent (Figure 4H). The protein-negative group showed low relative levels of CSPP1 mRNA and almost exclusively comprised basal-like carcinomas. Nuclear CSPP1-positive carcinomas were sub-dividable into two major trajectories: increasing CSPP1 mRNA expression correlated well with increasing nuclear CSPP1 staining in luminal carcinomas, whereas on the contrary nuclear-positive basal-like carcinomas showed no corresponding increase in mRNA expression. In fact, the group of cases with >75% of nuclear CSPP1-positive cancer cells depicted a prominent variation in mRNA levels (Figure 4H). Consequently, two populations of basal-like tumours with similar, low CSPP1 mRNA expression were distinguishable on the basis of differential nuclear CSPP1 staining (4/14 basal-like carcinomas without and 7/14 basal-like carcinomas with >75% nuclear CSPP1 expressing cancer cells), indicating that differentially regulated nuclear CSPP1 protein expression could define subsets of basal-like breast cancer (see next section). To increase the number of cases for further analysis we looked at the relative gene expression levels of CSPP1 with regard to subtypes (80 patients) and found the same trend; basal-like cases had a very low level of CSPP1 mRNA, whereas luminal cases (and luminal B in particular) showed the highest levels (Figure 4I). This was validated in a second, independent cohort of 115 low-stage breast cancer patients (Oslo1/MicMa (Naume et al, 2007), Supplementary Figure S4). In addition, array CGH copy number data for CSPP1-specific probes were available for both cohorts. Amplification/gain of the CSPP1 locus was most frequent in luminal type tumours and correlated well with mRNA expression in this subtype in both cohorts (Supplementary Figure S4), which is in concordance with previously published data (Adelaide et al, 2007).
Gene expression analysis in nuclear CSPP1-positive vs -negative basal-like tumours
Absence or presence of nuclear CSPP1 protein expression identified two subgroups of basal-like tumours (Figure 4H). Global gene expression data were available for 7 nuclear CSPP1-positive and 4 nuclear CSPP1-negative basal-like carcinomas, and additional 11 basal-like carcinomas of undetermined CSPP1 protein status. Consecutive filtration for gene expression variance and SAMexperiments (SAM; q<10−4) showed that nuclear CSPP1-positive basal-like biopsies were distinguished from nuclear CSPP1-negative counterparts by higher expression of two genes (PAMR1, GRP) and lower expression of six genes (PDCD10, UBD, ATPIF1, SLBP, CHD1, ADAM17) (Figure 5A and Supplementary Figure S5A). Hierarchical clustering analysis of all 22 basal-like tumours by expression data of these eight genes (the eight-gene signature) defined two main groups (Figure 5B and Supplementary Figure S5B). As expected, the nuclear CSPP1-negative cases and highly nuclear CSPP1-positive cases used for SAM analysis clustered group wise into separate arms. Expression of PAMR1, GRP, and UBD showed the most distinct difference between these groups. Nuclear CSPP1-positive tumours showed high levels of PAMR1 and GRP, and low levels of UBD. Nuclear CSPP1-negative tumours depicted the inverse pattern. Clustering of 26 basal-like tumours of the independent Oslo1/MicMa cohort by the eight-gene signature resulted in a similar separation (Supplementary Figure S5C). None of the identified eight genes were among the 50 subtype-classifying PAM50 genes, but in both cohorts, the nuclear CSPP1-positive basal-like cluster group consistently showed elevated PAM50 correlation coefficients towards luminal A centroids than the nuclear CSPP1-negative cluster group (P<0.05) (Figure 5C and Supplementary Figure S5B and C).
Differential gene expression in nuclear CSPP1-postive and-negative basal-like breast carcinomas. (A) Workflow and tabular t-statistics results of differentially expressed genes in nuclear CSPP1-positive (n=7) and nuclear CSPP1-negative (n=4) basal-like breast cancer biopsies determined by significance analysis of microarray experiments (SAM). Eight differentially expressed genes are identified at highest statistical significance. (B) Hierarchical cluster analysis of all basal-like breast carcinomas in the Oslo0/Ull cohort on the basis of the identified differentially expressed genes. Scoring results are indicated for cases with immunohistochemical CSPP1 staining (see also Figure 4H). Cases used for SAM are indicated with # symbol. (C) Box plots of PAM50 breast cancer subtype centroid correlation coefficients of ‘nuclear CSPP1 negative’ and ‘nuclear CSPP1 positive’ cluster group biopsies towards. P-values indicate statistically significant differences in median correlation coefficients (t-test or (*) Mann–Whitney).
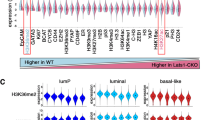
The low number of basal-like carcinomas present in the Oslo0/Ull and Oslo1/MicMa cohorts limited statistical analysis of potential clinical-pathological differences between the identified basal-like subgroups. Recently, integrative copy number and gene expression analysis of samples from almost 2000 breast cancer patients was reported, whereof 329 were of basal-like subtype (METABRIC (Curtis et al, 2012)). Expression of the eight signature genes and CSPP1 revealed no differences between discovery and validation METABRIC cohorts (997 and 995 patients, respectively; Supplementary Figures S6-7, Supplementary Table 3). As in Oslo0/Ull and Oslo1/MicMa, CSPP1 expression was highest in luminal B type breast cancers and lowest in basal-like breast cancers (Figure 6A). Discovery and validation cohorts were combined for further analysis. Notably, PAM50 subtype-wise cluster analysis of mean expression levels of the eight-gene signature genes revealed high expression of GRP and PAMR1 as characteristics of luminal A and normal-like breast cancers, respectively. Conversely, high expression of UBD and ADAM17 characterised basal-like and HER2-enriched breast cancers (Figure 6B).
Expression of CSPP1 and the eight identified signature genes in METABRIC. (A) Whisker hair box plots of mean CSPP1 mRNA expression in PAM50 breast cancer subtypes of the METABRIC discovery (n=996) and validation cohort (n=984), Statistically significant (P<0.05) pair-wise gene expression differences are indicated by ** symbols. (B) Hierarchical clustering of PAM50 breast cancer subtypes and their mean gene expression levels of the eight identified signature genes. Discovery and Validation cohorts showed highly similar gene expression profiles for all investigated genes and were, therefore, combined (Supplementary Figures S6 and S7, Supplementary Table 3).
Also hierarchical clustering analysis of the 329 basal-like cancers revealed the earlier defined PAMR1high, GRPhigh, UBDlow (‘nuclear CSPP1 positive’; designated as Basal_3) and PAMR1low, GRPlow, UBDhigh (‘nuclear CSPP1 negative’, Basal_1) subgroups, but apart from these a third cluster group characterised by PAMR1low, GRPlow, UBDlow expression was evident (designated as Basal_2). Subsequent hierarchical clustering analysis of average gene expression values in these three basal-like subgroups with carcinomas of other subtypes showed close relatedness of the Basal_3 (‘nuclear CSPP1 positive’) group with luminal A and normal-like breast cancers (Figure 7B), whereas Basal_1 and to a lower degree Basal_2 groups related to the HER2-enriched subtype. Importantly, all three basal-like subgroups showed comparable expression of clinically relevant receptor genes (EGFR, ERBB2, PgR and ESR), CSPP1 and the GRP receptor gene (GRPR). Also Integrated cluster groups (IntClust) (Curtis et al, 2012) were similarly distributed, though a slight over-representation of IntClust10 in the Basal_1 and IntClust5 in the Basal_3 subgroup was noted.
Identification of a third basal-like subgroup by hierarchical cluster analysis of basal-like carcinomas in METABRIC. (A) Hierarchical cluster analysis of all PAM50 basal-like breast carcinomas in the Metabric cohort (n=329) on the basis of the identified eight-gene signature subdivides basal-like breast cancers into three main clusters (Basal_1-3), whereof Basal_3 is reminiscent of the ‘nuclear CSPP1 positive’ basal-like subgroup. Integrated cluster annotations are indicated and Basal_1-3 subgroup distribution is shown as table. (B) Hierarchical cluster analysis of identified basal-like subgroups and other PAM50 subgroukps on the basis of mean gene expression values of the identified eight genes. Mean gene expression values for subtype relevant receptor genes and Ki67 are indicated below. Mean gene expression values of individual subgroups are listed as table.
Clinical properties of gene expression defined basal-like subgroups
Breast cancer-specific survival was similar in the biologically defined basal-like subgroups though a statistically significant minor shorter time to breast cancer-specific death was observed for patients of the Basal_2 subgroup when analysis was limited to the genetically defined basal-like carcinomas of the IntClust10 group (P=0.04; Figure 8A). This IntClust group is dominated by basal-like breast cancers and comprises almost two third of all basal-like cancers in METABRIC (Curtis et al, 2012).
Clinical properties of basal-like subgroups in METABRIC. (A) Disease-specific survival probability analysis by Kaplan–Meier plots of identified subgroups across all (n=328) and the genetically defined IntClust10 (n=201) group of basal-like carcinomas in METABRIC. (B) Kaplan–Meier analysis of lymph node affected and unaffected basal-like carcinomas stratified by basal subgroups (C) Kaplan–Meier analysis of all basal-like and individual basal subgroups stratified by lymph node involvement at time of diagnosis.
For patients with lymph node involvement at time of diagnosis, the time to breast cancer-specific death differed significantly between the three basal-like subgroups (Figure 8B). This was in contrast to patients with lymph node-negative disease where no significant difference was observed. The decreased survival probability of the Basal_2 subgroup could, however, not be attributed to skewness in distribution of invasive carcinomas across subgroups. In fact, the importance of lymph node involvement on survival probability varied between basal-like subgroups: highly dependent on the lymph node status in the Basal_3 and Basal_2 subgroups but not the Basal_1 subgroup, which showed best prognosis among the lymph node-positive basal-like carcinomas (P<0.03). Interestingly, the shortest overall time to breast cancer-specific death was observed in lymph node-positive patients of the basal-like subgroup with luminal features, which also showed the highest hazard ratio for lymph node involvement (Figure 8B and C).
Discussion
Previous studies have shown correlated genomic gain and overexpression of CSPP1 in human luminal type breast cancer and indicated oestrogen inducible expression of CSPP1 in human tumour xenografts in mice (Creighton et al, 2006; Harvell et al, 2006; Adelaide et al, 2007). Collectively, these studies suggested a functional importance of CSPP1 proteins in the human mammary gland and a possible involvement in the development and/or progression of human breast cancer. The study presented here is the first to address the expression of CSPP1 proteins (isoforms) in normal mammary tissue and biopsies of primary operable breast cancer at the cellular level.
We identified differential expression of a yet uncharacterised nuclear CSPP1 isoform in myoepithelial and luminal epithelial cells of the mammary gland and this pattern appeared preserved in breast cancer cell lines. Importantly, the unexpected luminal cell-related nuclear CSPP1 expression detected with the monoclonal a-CSPP/CSPP-L antibody was proven CSPP1-specific. Nuclear CSPP1 protein expression is likely to be driven by alternative splicing and/or alternative promoter activity and is not a consequence of post-translational modification or cell type-specific subcellular localisation of the earlier characterised isoforms CSPP and CSPP-L. The mRNA and protein sequence for this isoform remains at present obscure. Multiple mRNA splice isoforms have been reported to be expressed from the CSPP1 locus (17 predicted protein encoding CSPP1 splice isoforms) and are likely to underlie tissue or cell type-specific regulation (Thierry-Mieg and Thierry-Mieg, 2006). Shared mRNA and probe sequences did unfortunately not allow a definite discrimination of CSPP1 isoforms in our tumour gene expression data or public cell line gene expression data (Supplementary Figure S1). Target sequences of shRNAs used in this study, however, indicate that the encoding mRNA encloses common sequence parts with CSPP and CSPP-L transcripts upstream of the C-terminal domain encoding region. GATA3 and FOXA1, two transcription factors regulating luminal epithelial development and highly expressed in luminal type carcinomas (Cancer Genome Atlas Network, 2012; Kouros–Mehr et al, 2008) bind to the CSPP1 promoter region (T47D cells; GSM803514 and GSM803409) and may regulate CSPP1 expression in the mammary gland in addition to oestrogen (Supplementary Figure S8) (Creighton et al, 2006; Harvell et al, 2006). GATA3 and FOXA1 activity may thus account for enhanced CSPP1 mRNA expression in luminal type tumours in addition to gene-dose-dependent effects (Supplementary Figure S4 and (Adelaide et al, 2007)).
Epithelial cell type-dependent nuclear CSPP1 protein expression is, however, unlikely to be regulated by mRNA dosage alone, as some basal tumours showed nuclear staining in spite of low CSPP1 mRNA expression levels. The underlying regulatory mechanism may involve epigenetic mechanisms controlling mammary gland development that remain preserved during malignant transformation (Stingl and Caldas, 2007; Rijnkels et al, 2010). Given the lineage-correlated staining pattern of normal breast epithelia nuclear CSPP1-positive and-negative tumours of the basal-like type may have originated from different progenitors and/or retained a distinct differentiation potential. Heterogeneity in immunohistochemical and histopathological features of basal-like breast cancers are evident. Frequent co-expression of luminal type cytokeratins CK8/18 and CK19 may indicate that some have more luminal features than others (for a recent review: Lavasani and Moinfar, 2012). Further, expression of normal breast myoepithelial markers, such as smooth muscle actin, was only seen at low frequency in a study of basal-like invasive breast cancers (Livasy et al, 2006). Individual carcinomas of the PAM50 basal-like group may thus have originated from transformation events that occurred in epithelial cells of different developmental stages (reviewed in Prat and Perou, 2011). This, as well as the lineage-dependent CSPP1 expression pattern in the mammary gland, supports the idea that nuclear CSPP1-positive basal-like carcinomas have luminal traits. Indeed, nuclear CSPP1-positive basal-like carcinomas were not only found to be reminiscent of luminal epithelial cells and luminal type breast cancer with respect to nuclear CSPP1 expression alone. The ‘nuclear CSPP1 positive’-surrogate eight-gene signature (Basal_3, GRPHigh PAMR1High UBDLow) was delineated from only a limited number of nuclear CSPP1-positive and-negative basal-like cases available for comparative transcriptome analysis, but consistently correlated with increasing PAM50 correlation coefficients towards luminal A centroide in the Oslo1/MicMa validation cohort and closely matched the mean gene expression pattern of the eight-gene signature of luminal A carcinomas in the METABRIC hallmark cohort. Finally, luminal A type breast cancers characteristically display pattern reminiscent of normal (luminal) tissue morphology (Peppercorn et al, 2008; Parker et al, 2009). Similarly, the nuclear CSPP1-positive basal-like breast cancers of the Oslo0/Ull cohort showed more organised morphological features of normal breast epithelia including more differentiated cells, lower histological grade, and lobular histology.
In contrast to this, nuclear CSPP1-negative basal-like carcinomas of the Oslo0/Ull cohort showed lower degree of differentiation and more pleomorphic nuclei. Notably, ‘CSPP1 nuclear negative’-surrogate gene signature basal-like carcinomas had two subclusters, Basal_1 and Basal_2, mainly owing to their difference in UBD expression. This bipartition was possibly unnoticed in Oslo0/Ull and Oslo1/MicMa cohorts owing to low number of cases, but is further supported by the different survival probabilities in lymph node-positive disease. The tumours in the two ‘nuclear CSPP1 negative’ subgroups, Basal_2 and, in particular, Basal_1 clustered more closely with HER2-enriched carcinomas and included higher fractions of IntClust10 carcinomas, the most basal-like-related genomic pattern of carcinomas in METABRIC (90% fraction of IntClust10 are of basal-like type (60% of all basal-like)) (Curtis et al, 2012).
Nuclear CSPP1-positive basal-like (Basal_3) carcinomas had higher mRNA expression of two secreted proteins, PAMR1 and GRP. Mitogenic, migratory and morphogenic roles have been attributed to GRP signalling, including trans-activation of EGFR by Src activation. GRP is frequently overexpressed in human breast cancer (Patel et al, 2006), and GRP receptors are upregulated in the murine mammary gland during lactation phase (Anderson et al, 2007). GRP peptides are currently studied in (breast) cancer diagnostics and experimental therapy (reviewed in Hohla and Schally, 2010). Nuclear CSPP1-negative basal-like breast cancers showed higher expression of PDCD10, ADAM17, ATPIF1, SLBP, CHD1, and UBD. Though ADAM17 did not show major gene expression differences between basal-like subgroups in the METABRIC cohort, we noticed its generally higher expression in basal-like carcinomas. This metalloproteinase is crucial for ductal morphogenesis by release of epithelial amphiregulin to stimulate EGFR signalling on surrounding stroma (Sternlicht et al, 2005) and possibly supports disease exacerbation by sustaining acquired autocrine EGF signalling (Kenny and Bissell, 2007). ADAM17 inhibitors are tested pre-clinically for treatment of triple-negative breast cancers (McGowan et al, 2012) and could, thus, be exploited in combination with GRP-targeting drugs for ‘nuclear CSPP positive’ basal-like cancers. Speculatively, upregulation of GRP induced Src signalling could enhance EGFR signalling in Basal_3 tumours independently of ADAM17, which can be repressed locally by stroma secreted ADAM17 inhibitors. Another interesting identified candidate is the ‘nuclear CSPP1 negative’ group (Basal_1 and Basal_2) distinguishing UBD. Upregulation of this ubiquitin-like modifier is noticed in many epithelial cancers (Lee et al, 2003) and is thought to promote carcinogenesis by increasing mitotic instability (Ren et al, 2006). However, also pro-apoptotic activities have been ascribed (Raasi et al, 2001). The prognostic value of this ubiqitin-like modifier may, thus, be context dependent, which is consistent with the differential hazard in lymph node affected UBDHigh and UBDlow basal-like subgroups. Interestingly, a recent meta-analysis of triple-negative breast cancers (TNBC) determined six subtypes of TNBCs by unsupervised clustering analysis of 587 genes (Lehmann et al, 2011). Though TNBCs not exclusively comprise basal-like carcinomas (for a recent review Foulkes et al, 2010), in congruence with our findings, high UBD expression was correlated with the basal-like TNBC subtype, whereas high GRP expression was correlated with the luminal androgen receptor TNBC subtype. Therefore, further work may address the potential predicative value of nuclear CSPP1 staining and the CSPP1-derived 8-gene signature in TNBC subtypes.
To conclude, our investigation uncovered an unanticipated complex regulation of CSPP1 isoform expression in the human mammary gland and breast carcinomas. The comparison of nuclear CSPP1-positive and-negative basal-like breast cancers provides novel molecular insight into the underlying heterogeneity and encourage further preclinical studies to investigate (1) the possible benefit of inhibition of GRP signalling and ADAM17 activity, and (2) the prognostic value of UBD expression in disseminated disease in respective basal-like subgroups. Further molecular studies are clearly required to deduce the subtype-specific regulation and functional importance of individual CSPP1 isoforms in normal and transformed mammary epithelial cells.
Change history
15 July 2014
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Adelaide J, Finetti P, Bekhouche I, Repellini L, Geneix J, Sircoulomb F, Charafe-Jauffret E, Cervera N, Desplans J, Parzy D, Schoenmakers E, Viens P, Jacquemier J, Birnbaum D, Bertucci F, Chaffanet M (2007) Integrated profiling of basal and luminal breast cancers. Cancer Res 67: 11565–11575.
Anderson SM, Rudolph MC, McManaman JL, Neville MC (2007) Key stages in mammary gland development. Secretory activation in the mammary gland: it's not just about milk protein synthesis!. Breast Cancer Res 9: 204.
Asiedu M, Wu D, Matsumura F, Wei Q (2009) Centrosome/spindle pole-associated protein regulates cytokinesis via promoting the recruitment of MyoGEF to the central spindle. Mol Biol Cell 20: 1428–1440.
Blows FM, Driver KE, Schmidt MK, Broeks A, van Leeuwen FE, Wesseling J, Cheang MC, Gelmon K, Nielsen TO, Blomqvist C, Heikkila P, Heikkinen T, Nevanlinna H, Akslen LA, Begin LR, Foulkes WD, Couch FJ, Wang X, Cafourek V, Olson JE, Baglietto L, Giles GG, Severi G, McLean CA, Southey MC, Rakha E, Green AR, Ellis IO, Sherman ME, Lissowska J, Anderson WF, Cox A, Cross SS, Reed MW, Provenzano E, Dawson SJ, Dunning AM, Humphreys M, Easton DF, Garcia-Closas M, Caldas C, Pharoah PD, Huntsman D (2010) Subtyping of breast cancer by immunohistochemistry to investigate a relationship between subtype and short and long term survival: a collaborative analysis of data for 10,159 cases from 12 studies. PLoS Med 7: e1000279.
Callagy G, Cattaneo E, Daigo Y, Happerfield L, Bobrow LG, Pharoah PD, Caldas C (2003) Molecular classification of breast carcinomas using tissue microarrays. Diagn Mol Pathol 12: 27–34.
Cancer Genome Atlas Network (2012) Comprehensive molecular portraits of human breast tumours. Nature 490: 61–70.
Cheang MC, van de RM, Nielsen TO (2008) Gene expression profiling of breast cancer. Annu Rev Pathol 3: 67–97.
Chin K, DeVries S, Fridlyand J, Spellman PT, Roydasgupta R, Kuo WL, Lapuk A, Neve RM, Qian Z, Ryder T, Chen F, Feiler H, Tokuyasu T, Kingsley C, Dairkee S, Meng Z, Chew K, Pinkel D, Jain A, Ljung BM, Esserman L, Albertson DG, Waldman FM, Gray JW (2006) Genomic and transcriptional aberrations linked to breast cancer pathophysiologies. Cancer Cell 10: 529–541.
Creighton CJ, Cordero KE, Larios JM, Miller RS, Johnson MD, Chinnaiyan AM, Lippman ME, Rae JM (2006) Genes regulated by estrogen in breast tumor cells in vitro are similarly regulated in vivo in tumor xenografts and human breast tumors. Genome Biol 7: R28.
Curtis C, Shah SP, Chin SF, Turashvili G, Rueda OM, Dunning MJ, Speed D, Lynch AG, Samarajiwa S, Yuan Y, Graf S, Ha G, Haffari G, Bashashati A, Russell R, McKinney S, Langerod A, Green A, Provenzano E, Wishart G, Pinder S, Watson P, Markowetz F, Murphy L, Ellis I, Purushotham A, Borresen-Dale AL, Brenton JD, Tavare S, Caldas C, Aparicio S (2012) The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 486: 346–352.
D'Assoro AB, Barrett SL, Folk C, Negron VC, Boeneman K, Busby R, Whitehead C, Stivala F, Lingle WL, Salisbury JL (2002) Amplified centrosomes in breast cancer: a potential indicator of tumor aggressiveness. Breast Cancer Res Treat 75: 25–34.
Deugnier MA, Moiseyeva EP, Thiery JP, Glukhova M (1995) Myoepithelial cell differentiation in the developing mammary gland: progressive acquisition of smooth muscle phenotype. Dev Dyn 204: 107–117.
Eisen MB, Spellman PT, Brown PO, Botstein D (1998) Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci USA 95: 14863–14868.
Enerly E, Steinfeld I, Kleivi K, Leivonen SK, Aure MR, Russnes HG, Ronneberg JA, Johnsen H, Navon R, Rodland E, Makela R, Naume B, Perala M, Kallioniemi O, Kristensen VN, Yakhini Z, Borresen-Dale AL (2011) miRNA-mRNA integrated analysis reveals roles for miRNAs in primary breast tumors. PLoS One 6: e16915.
Foulkes WD, Smith IE, Reis-Filho JS (2010) Triple-negative breast cancer. N Engl J Med 363: 1938–1948.
Guo HQ, Gao M, Ma J, Xiao T, Zhao LL, Gao Y, Pan QJ (2007) Analysis of the cellular centrosome in fine-needle aspirations of the breast. Breast Cancer Res 9: R48.
Harvell DM, Richer JK, Allred DC, Sartorius CA, Horwitz KB (2006) Estradiol regulates different genes in human breast tumor xenografts compared with the identical cells in culture. Endocrinology 147: 700–713.
Hohla F, Schally AV (2010) Targeting gastrin releasing peptide receptors: new options for the therapy and diagnosis of cancer. Cell Cycle 9: 1738–1741.
Kais Z, Parvin JD (2008) Regulation of centrosomes by the BRCA1-dependent ubiquitin ligase. Cancer Biol Ther 7: 1540–1543.
Kasper M, Jaks V, Fiaschi M, Toftgard R (2009) Hedgehog signalling in breast cancer. Carcinogenesis 30: 903–911.
Kenny PA, Bissell MJ (2007) Targeting TACE-dependent EGFR ligand shedding in breast cancer. J Clin Invest 117: 337–345.
Kouros-Mehr H, Kim JW, Bechis SK, Werb Z (2008) GATA-3 and the regulation of the mammary luminal cell fate. Curr Opin Cell Biol 20: 164–170.
Langerod A, Zhao H, Borgan O, Nesland JM, Bukholm IR, Ikdahl T, Karesen R, Borresen-Dale AL, Jeffrey SS (2007) TP53 mutation status and gene expression profiles are powerful prognostic markers of breast cancer. Breast Cancer Res 9: R30.
Lavasani MA, Moinfar F (2012) Molecular classification of breast carcinomas with particular emphasis on "basal-like" carcinoma: a critical review. J Biophotonics 5: 345–366.
Lee CG, Ren J, Cheong IS, Ban KH, Ooi LL, Yong TS, Kan A, Nuchprayoon I, Jin R, Lee KH, Choti M, Lee LA (2003) Expression of the FAT10 gene is highly upregulated in hepatocellular carcinoma and other gastrointestinal and gynecological cancers. Oncogene 22: 2592–2603.
Lehmann BD, Bauer JA, Chen X, Sanders ME, Chakravarthy AB, Shyr Y, Pietenpol JA (2011) Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J Clin Invest 121: 2750–2767.
Li JJ, Weroha SJ, Lingle WL, Papa D, Salisbury JL, Li SA (2004) Estrogen mediates Aurora-A overexpression, centrosome amplification, chromosomal instability, and breast cancer in female ACI rats. Proc Natl Acad Sci U S A 101: 18123–18128.
Livasy CA, Karaca G, Nanda R, Tretiakova MS, Olopade OI, Moore DT, Perou CM (2006) Phenotypic evaluation of the basal-like subtype of invasive breast carcinoma. Mod Pathol 19: 264–271.
McDermott KM, Liu BY, Tlsty TD, Pazour GJ (2010) Primary Cilia regulate branching morphogenesis during mammary gland development. Curr Biol 20 (8): 731–737.
McGowan PM, Mullooly M, Caiazza F, Sukor S, Madden SF, Maguire AA, Pierce A, McDermott EW, Crown J, O'Donovan N, Duffy MJ (2012) ADAM-17: a novel therapeutic target for triple negative breast cancer. Ann Oncol 24 (2): 362–369.
Naume B, Zhao X, Synnestvedt M, Borgen E, Russnes HG, Lingjaerde OC, Stromberg M, Wiedswang G, Kvalheim G, Karesen R, Nesland JM, Borresen-Dale AL, Sorlie T (2007) Presence of bone marrow micrometastasis is associated with different recurrence risk within molecular subtypes of breast cancer. Mol Oncol 1: 160–171.
Nigg EA, Raff JW (2009) Centrioles, centrosomes, and cilia in health and disease. Cell 139: 663–678.
Parker JS, Mullins M, Cheang MC, Leung S, Voduc D, Vickery T, Davies S, Fauron C, He X, Hu Z, Quackenbush JF, Stijleman IJ, Palazzo J, Marron JS, Nobel AB, Mardis E, Nielsen TO, Ellis MJ, Perou CM, Bernard PS (2009) Supervised risk predictor of breast cancer based on intrinsic subtypes. J Clin Oncol 27: 1160–1167.
Patel O, Shulkes A, Baldwin GS (2006) Gastrin-releasing peptide and cancer. Biochim Biophys Acta 1766: 23–41.
Patzke S, Hauge H, Sioud M, Finne EF, Sivertsen EA, Delabie J, Stokke T, Aasheim HC (2005) Identification of a novel centrosome/microtubule-associated coiled-coil protein involved in cell-cycle progression and spindle organization. Oncogene 24: 1159–1173.
Patzke S, Redick S, Warsame A, Murga-Zamalloa CA, Khanna H, Doxsey SJ, Stokke T (2010) CSPP is a ciliary protein interacting with Nephrocystin 8 and required for cilia formation. Mol Biol Cell 21: 2555–2567.
Patzke S, Stokke T, Aasheim HC (2006) CSPP and CSPP-L associate with centrosomes and microtubules and differently affect microtubule organization. J Cell Physiol 209: 199–210.
Peppercorn J, Perou CM, Carey LA (2008) Molecular subtypes in breast cancer evaluation and management: divide and conquer. Cancer Invest 26: 1–10.
Prat A, Perou CM (2011) Deconstructing the molecular portraits of breast cancer. Mol Oncol 5: 5–23.
Pujana MA, Han JD, Starita LM, Stevens KN, Tewari M, Ahn JS, Rennert G, Moreno V, Kirchhoff T, Gold B, Assmann V, Elshamy WM, Rual JF, Levine D, Rozek LS, Gelman RS, Gunsalus KC, Greenberg RA, Sobhian B, Bertin N, Venkatesan K, yivi-Guedehoussou N, Sole X, Hernandez P, Lazaro C, Nathanson KL, Weber BL, Cusick ME, Hill DE, Offit K, Livingston DM, Gruber SB, Parvin JD, Vidal M (2007) Network modeling links breast cancer susceptibility and centrosome dysfunction. Nat Genet 39: 1338–1349.
Raasi S, Schmidtke G, Groettrup M (2001) The ubiquitin-like protein FAT10 forms covalent conjugates and induces apoptosis. J Biol Chem 276: 35334–35343.
Reis-Filho JS, Pusztai L (2011) Gene expression profiling in breast cancer: classification, prognostication, and prediction. Lancet 378: 1812–1823.
Ren J, Kan A, Leong SH, Ooi LL, Jeang KT, Chong SS, Kon OL, Lee CG (2006) FAT10 plays a role in the regulation of chromosomal stability. J Biol Chem 281: 11413–11421.
Rijnkels M, Kabotyanski E, Montazer-Torbati MB, Hue BC, Vassetzky Y, Rosen JM, Devinoy E (2010) The epigenetic landscape of mammary gland development and functional differentiation. J Mammary Gland Biol Neoplasia 15: 85–100.
Russnes HG, Vollan HK, Lingjaerde OC, Krasnitz A, Lundin P, Naume B, Sorlie T, Borgen E, Rye IH, Langerod A, Chin SF, Teschendorff AE, Stephens PJ, Maner S, Schlichting E, Baumbusch LO, Karesen R, Stratton MP, Wigler M, Caldas C, Zetterberg A, Hicks J, Borresen-Dale AL (2010) Genomic architecture characterizes tumor progression paths and fate in breast cancer patients. Sci Transl Med 2: 38ra47.
Salisbury JL, D'Assoro AB, Lingle WL (2004) Centrosome amplification and the origin of chromosomal instability in breast cancer. J Mammary Gland Biol Neoplasia 9: 275–283.
Sternlicht MD, Sunnarborg SW, Kouros-Mehr H, Yu Y, Lee DC, Werb Z (2005) Mammary ductal morphogenesis requires paracrine activation of stromal EGFR via ADAM17-dependent shedding of epithelial amphiregulin. Development 132: 3923–3933.
Stingl J, Caldas C (2007) Molecular heterogeneity of breast carcinomas and the cancer stem cell hypothesis. Nat Rev Cancer 7: 791–799.
Taylor-Papadimitriou J, Stampfer M, Bartek J, Lewis A, Boshell M, Lane EB, Leigh IM (1989) Keratin expression in human mammary epithelial cells cultured from normal and malignant tissue: relation to in vivo phenotypes and influence of medium. J Cell Sci 94 (Pt 3): 403–413.
Thierry-Mieg D, Thierry-Mieg J (2006) AceView: a comprehensive cDNA-supported gene and transcripts annotation. Genome Biol 7 (Suppl 1): S12–S14.
Van Loo P, Nordgard SH, Lingjaerde OC, Russnes HG, Rye IH, Sun W, Weigman VJ, Marynen P, Zetterberg A, Naume B, Perou CM, Borresen-Dale AL, Kristensen VN (2010) Allele-specific copy number analysis of tumors. Proc Natl Acad Sci U S A 107: 16910–16915.
Visvader JE (2009) Keeping abreast of the mammary epithelial hierarchy and breast tumorigenesis. Genes Dev 23: 2563–2577.
Yuan K, Frolova N, Xie Y, Wang D, Cook L, Kwon YJ, Steg AD, Serra R, Frost AR (2010) Primary cilia are decreased in breast cancer: analysis of a collection of human breast cancer cell lines and tissues. J Histochem Cytochem 58: 857–870.
Acknowledgements
We thank Ellen Hellesylt and Inga Hansine Rye for expert advice in immunohistochemical and immunofluorescence staining of formalin-preserved tissue biopsies, and Kari-Anne Myrum Frikstad for excellent technical assistance with experiments for the revised manuscript. SP acknowledges financial support by the Regional Health Authority of South Eastern Norway, the Comprehensive Cancer Cluster Council of OUH-Norwegian Radium Hospital, Radium Hospital Foundation and the Norwegian Cancer Society. HGR is funded by the Norwegian Cancer Society, Radium Hospital Foundation, Raagholdtstiftelsen and Torsteds legat. SN is funded by the Norwegian Cancer Society.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License.
Supplementary Information accompanies this paper on British Journal of Cancer website
Supplementary information
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Sternemalm, J., Russnes, H., Zhao, X. et al. Nuclear CSPP1 expression defined subtypes of basal-like breast cancer. Br J Cancer 111, 326–338 (2014). https://doi.org/10.1038/bjc.2014.297
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2014.297
Keywords
This article is cited by
-
Primary cilia proteins: ciliary and extraciliary sites and functions
Cellular and Molecular Life Sciences (2018)