Abstract
Background:
The revolution in cancer genomics shows that the dominant mutations are CG->TA transitions. The sources of these mutations are probably two host cell cytidine deaminases APOBEC3A and APOBEC3B. The former in particular can access nuclear DNA and monotonously introduce phenomenal numbers of C->T mutations in the signature 5′TpC context. These can be copied as G->A transitions in the 5′GpA context.
Methods:
DNA hypermutated by an APOBEC3 enzyme can be recovered by a technique called 3DPCR, which stands for differential DNA denaturation PCR. This method exploits the fact that APOBEC3-edited DNA is richer in A+T compared with the reference. We explore explicitly 3DPCR error using cloned DNA.
Results:
Here we show that the technique has a higher error rate compared with standard PCR and can generate DNA strands containing both C->T and G->A mutations in a 5′GpCpR context. Sequences with similar traits have been recovered from human tumour DNA using 3DPCR.
Conclusions:
Differential DNA denaturation PCR cannot be used to identify fixed C->T transitions in cancer genomes. Presently, the overall mutation frequency is ∼104–105 base substitutions per cancer genome, or 0.003–0.03 kb−1. By contrast, the 3DPCR error rate is of the order of 4–20 kb−1 owing to constant selection for AT DNA and PCR-mediated recombination. Accordingly, sequences recovered by 3DPCR harbouring mixed C->T and G->A mutations associated with the 5′GpC represent artefacts.
Similar content being viewed by others
Main
The advent of cancer genomics has shown that there are far more somatic mutations than hitherto imagined. Although large karyotypic changes have been known for decades, there are a myriad of small rearrangements, whereas the number of mutations can exceed 105 per genome (Greenman et al, 2007; Stephens et al, 2009, 2012; Pleasance et al, 2010). Apart from cancers involving mutagens that leave genetic signatures, such as aflatoxin or benzo[a]pyrene, overall there is an excess of CG->TA transitions in cancer genomes (Greenman et al, 2007). For some genomes there is a pronounced 5′ base context, notably TpC for C->T transitions (Nik-Zainal et al, 2012; Stephens et al, 2012; Alexandrov et al, 2013). Of course, the G->A transition is merely a C->T transition on the opposite strand, suggesting a single transition.
The human APOBEC3 (A3) family of seven genes encoding six functional cytidine deaminases (APOBEC3A, A3B, A3C, A3F, A3G and A3H) has come to the fore in recent years as host cell mutators of viral DNA (Jarmuz et al, 2002; Sheehy et al, 2002; Harris et al, 2003; Lecossier et al, 2003; Mangeat et al, 2003; Mariani et al, 2003). They are closely related to activation-induced deaminase (AID), which is responsible for immunoglobulin class switch recombination and somatic hypermutation or rearranged VDJ genes (Di Noia and Neuberger, 2007). Indeed, it is thought that the A3 locus emerged by gene duplication of AID (Conticello et al, 2005). Like AID, the substrate specificity of the A3 enzymes is the cytidine base in single-stranded DNA (ssDNA), the product being uridine, which base pairs as thymidine. As such, they effectively hypermutate DNA, the mutational load being so great that the viral genetic information is irretrievably lost. That several A3 genes are upregulated by type I and II interferons lent credence to their being novel restriction factors and part of a broad innate immune response to microbes (Bonvin et al, 2006; Koning et al, 2009; Refsland et al, 2010; Stenglein et al, 2010). These APOBEC3 mutator enzymes must be tightly controlled as at least four (A3A, A3B, A3C and A3H) can access the nucleus (Vartanian et al, 2008; Stenglein et al, 2010; Landry et al, 2011; Suspène et al, 2011a; Aynaud et al, 2012; Shinohara et al, 2012; Burns et al, 2013a, 2013b).
More recently, it has emerged that APOBEC3A (A3A) and probably APOBEC3B (A3B) can mutate nuclear DNA (nuDNA) (Suspène et al, 2011a; Aynaud et al, 2012; Shinohara et al, 2012; Burns et al, 2013a). A3A can edit both cytidine and 5-methylcytidine residues in ssDNA (Carpenter et al, 2012; Wijesinghe and Bhagwat, 2012; Suspène et al, 2013) and can generate DNA double-strand breaks (Landry et al, 2011; Mussil et al, 2013). Low levels of A3 mutation that do not overwhelm DNA mismatch repair are probably the source of the CG->TA mutations in cancer genomes. By contrast, hypermutation is proapoptotic and represents a novel pathway for DNA catabolism (Suspène et al, 2011a; Mussil et al, 2013). Hypermutated nuDNA with hundreds of mutations per kilobase can be found in both pathological and physiological settings (Suspène et al, 2011a). Invariably the mutations are influenced by the base 5′ with a TpC and CpC being the preferred targets (the edited cytidine residue being underlined) (Suspène et al, 2011a; Nik-Zainal et al, 2012; Alexandrov et al, 2013). Only A3G shows a strong penchant for CpC (Beale et al, 2004; Bishop et al, 2004; Suspène et al, 2004; Henry et al, 2009; Vartanian et al, 2010). By contrast, AID prefers a purine residue in the 5′ position, where GpC>ApC (Conticello et al, 2005; Vartanian et al, 2010). At low levels of nuDNA editing, compatible with DNA repair, these enzymes are probably the source of the C->T transitions seen in cancer genomes (Greenman et al, 2007; Stephens et al, 2009, 2012; Pleasance et al, 2010; Nik-Zainal et al, 2012). While the relative contribution of A3A and A3B to the editing of nuDNA needs to be ascertained, the A3B−/− genotype is particularly prevalent in SE Asia (Kidd et al, 2007). Interestingly, these individuals have a higher odds ratio of developing breast and liver cancer compared with controls (Komatsu et al, 2008; Zhang et al, 2012).
DNA that has been hypermutated by an APOBEC3 (A3) enzyme can be recovered by a technique called 3DPCR, which stands for differential DNA denaturation PCR (Suspène et al, 2005a). This method exploits the fact that A3-edited DNA is richer in A+T compared with the reference. Modulation of the PCR denaturation temperature allows selective amplification of AT-rich DNA, sometimes by up to 104-fold (Suspène et al, 2005b).
While 3DPCR was first used to detect hypermutated viral genomes (Suspène et al, 2005b; Vartanian et al, 2008, 2010; Suspène et al, 2011b), it is being increasingly used to recover low levels of A3-edited nuclear DNA. Two recent papers are noteworthy because they used 3DPCR to recover lightly edited DNA sequences manifesting ∼2–13 CG->TA transitions per kilobase of nuDNA (Shinohara et al, 2012; Burns et al, 2013a). They exploit the use of 4–5 °C temperature gradients across the 12-wells of the heating block, which is small in comparison with the ±0.4 °C temperature stability per well leaving considerable room for experimental variation. One study presented the sequences in sufficient detail – the sequences were not monotonously substituted by C->T or G->A (Shinohara et al, 2012); rather they contained both mutations on the same strand. Furthermore, a preferred 5′GpC editing context was frequently noted, which is more typical of AID rather than an A3 enzyme (Beale et al, 2004). Although ectopic upregulation of AID has indeed been reported (Endo et al, 2007; Matsumoto et al, 2007; Morisawa et al, 2008), we were concerned that such low-editing rates per kilobase might be close to the 3DPCR error threshold. The second study implicated A3B in nuDNA editing (Burns et al, 2013a), yet the signal was only 2–3-fold over 3DPCR background, which we found surprisingly low, particularly as A3B can extensively edit HBV DNA, the editing rates being of the order of >100 kb−1 (Suspène et al, 2005b; Vartanian et al, 2010).
Here we explore explicitly 3DPCR error using cloned DNA. It transpires that the 3DPCR error rate is of the order of 4–20 kb−1 and generates sequences encoding both C->T and G->A substitutions occurring preferentially in the GpC dinucleotide. Sequences with similar traits have been recovered from human DNA (Shinohara et al, 2012; Burns et al, 2013a).
Materials and methods
Blood was obtained from anonymous healthy donors and peripheral blood mononuclear cells (PBMCs) were isolated by Ficoll gradient (Eurobio AbCys, Courtaboeuf, France). Isolation of CD4+ T lymphocytes was performed by incubation with antibody-coated magnetic beads (Miltenyi Biotec, Bergish Glabach, Germany). Purity of CD4+ T lymphocytes was above 90% as checked by flow cytometry (FACSCalibur; Becton Dickinson, Franklin Lakes, NJ, USA). CD4+ T lymphocytes were stimulated with 10 μg ml−1 PHA (Sigma, St Louis, MO, USA), 100 U ml−1 IL-2 (Sigma) and 500 U ml−1 IFN-α (PBL Biomedical Laboratories, Piscataway, NJ, USA) for 48 h. For the detection of hypermutation by 3DPCR, CD4+ T cells were transduced with lentivirus rV2.EF1.UGI, which encodes a codon optimised UNG inhibitor (UGI) under the control of the constitutive human EF1 promoter (Vectalys, Toulouse, France).
A molecular clone corresponding to 376 bp spanning exon 8 and intron 9 of the human TP53 gene was recovered from HeLa cells. For amplification of human TP53, input DNA was ∼1 ng. The first-round reaction parameters were 95 °C for 5 min, followed by 40 cycles (95 °C for 30 s, 60 °C for 30 s and 72 °C for 2 min), and finally 10 min at 72 °C with the following primers: P53ext5 (5′-GAGCTGGACCTTAGGCTCCAGAAAGGACAA-3′) and P53ext3 (5′-GCTGGTGTTGTTGGGCAGTGCTAGGAA-3′). Second-round 3DPCR was performed with 1/50 of the first round as input using an Eppendorf Mastercycler ep Gradient S (Eppendorf AG, Hamburg, Germany) programmed to generate a 4 °C gradient in the denaturation temperature. The reaction parameters were 86–90 °C for 5 min, followed by 42 cycles (86–90 °C for 30 s, 58 °C for 30 s and 72 °C for 2 min), and finally 10 min at 72 °C. The buffer conditions were 2.5 mM MgCl2, 16 mM (NH4)2SO4, 67 mM Tris-HCl (pH 8.8), 0.01% Tween-20, 200 μ M each deoxynucleoside triphosphate, 100 μ M each primer (P53int5, 5′-TTCTCTTTTCCTATCCTGAGTAGTGGTAA-3′; P53int3, 5′-AAAGGTGATAAAAGTGAATCTGAGGCATAA-3′) and 1.5 U of Taq (EurobioTaq+; Eurobio AbCys) or Pfu (PfuUltra II Fusion HS DNA Polymerase; Agilent Technologies, Santa Clara, CA, USA) DNA polymerase.
Conditions of amplification of the β-catenin were 95 °C for 5 min, followed by 35 cycles (95 °C for 30 s, 60 °C for 30 s and 72 °C for 10 min) and finally 20 min at 72 °C with the following primers: 5′βcatout (5′-AGCTGATTTGATGGAGTTGGACA-3′) and 3′βcatout (5′-CCAGCTACTTGTTCTTGAGTGAA-3′). Nested PCR was performed with 1/50 of the first round, conditions were 80–87 °C for 5 min, followed by 35 cycles (80–87 °C for 30 s, 60 °C for 30 s and 72 °C for 10 min) and finally 20 min at 72 °C with the following primers: 5′βcatin (5′-ACATGGCCATGGAACCAGACAGA-3′) and 3′βcatin (5′-GTTCTTGAGTGAAGGACTGAGAA-3′).
For long-range PCR, the first-round reaction parameters were 95 °C for 5 min, followed by 40 cycles (95 °C for 30 s, 60 °C for 30 s and 72 °C for 10 min), and finally 20 min at 72 °C. The reaction parameters for the second-round 3DPCR were 86–90 °C for 5 min, followed by 42 cycles (86–90 °C for 30 s, 58 °C for 30 s and 72 °C for 10 min), and finally 20 min at 72 °C. The buffer conditions were 2.5 mM MgCl2, 1 × LA PCR Buffer ll (TaKaRa LA Taq; TaKaRa, Otsu, Japan), 400 μ M each deoxynucleoside triphosphate (TaKaRa LA Taq; TaKaRa), 100 μ M each primer and 1.5 U of Taq (EurobioTaq+; Eurobio AbCys).
Results and discussion
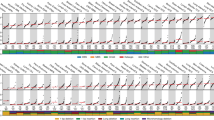
We tested the 3DPCR error rate using the first PCR amplification corresponding to 235 bp. For a 4 °C gradient across the block, the temperature of the last positive amplification varied by 0.9 °C from 89.6 °C for the outer rows A and H to 88.7 °C for the inner rows D and E (Figure 1A), showing that small thermal gradients are not strictly uniform across the block. Two 3DPCR-positive samples identified by asterisks were cloned and sequenced, and the majority (70–80%) of sequences carried both C->T and G->A transitions with up to 5 per clone, with an average of 7.5 kb−1 for sample D8 (88.7 °C) compared with the input sequence (Figure 1B and C), the most divergent being shown in Figure 1D. For A10 (89.6 °C) the overall mutation rate was similar (7.4 kb−1; Figure 1B). While transversions accounted for ∼8% of the total, there was an excess in favour of N->A,T. Such observations are to be expected for a technique that selectively amplifies AT-rich DNA and contrasts strongly with the mutation bias of Taq polymerase, which generates more A->G and T->C transitions than any others (Eckert and Kunkel, 1991).
Variation in 3DPCR Taq polymerase background mutation rate across the PCR block. (A) White spots correspond to PCR-positive samples for a 176 bp fragment of human TP53. Those samples indicated by an asterisk were cloned and sequenced. (B) Mutation matrices for A10- and D8-derived sequences; transitions were invariably of the type N->T,A; the number of bases sequenced is given by n. (C) Distribution of the number of CG->TA transitions per clone. (D) A collection of the most highly mutated sequences. To compact the data, only variable sites are shown, their positions being identified above. Nucleotide positions should be read from top to bottom. To the right are the numbers of mutations per clone (mut) and the minimum number of recombination events (rec) to explain the complexity. Zones of recombination are highlighted in grey shading when possible. (E) 5′ Dinucleotide context for the G->A and C->T transitions, with the expected values shown as horizontal bars.
Most sequences contained both C->T and G->A substitutions, although there were a few with just one type of transition (Figure 1D; the issue of PCR recombination (Meyerhans et al, 1990) is addressed below). Interestingly the 5′ dinucleotide context for these 3DPCR mutations is GpC for both types of transition (Figure 1E). There was a weaker 3′ context that tends to CpR, where R=G or A (Supplementary Figure 1A). As the 5′ and 3′ contexts are the same for both G->A and C->T transitions, this suggests that a single mutation is involved occurring in the motif 5′GpCpR.
Abasic DNA sites can result from slightly acidic conditions and can be copied by inserting an A. This is one of the reasons why long-range PCR buffers showing low pKa variation with temperature are used, unlike the standard Tris-HCl buffer (Eckert and Kunkel, 1993). Accordingly, we repeated the above experiment using long-range buffer, but using the same Taq enzyme as before. For a 4 °C variation there was spatial variation across the heating block, with 3DPCR positivity varying by an apparent 1.7 °C (Figure 2A). Qualitatively the sequences with both G->A and C->T transitions (Figure 2B) were identified while the same preference for 5′GpC was observed (Figure 2C and D) and a weaker 3′ bias for CpR (Supplementary Figure 1B). Interestingly, the overall mutation frequency was approximately two-fold higher (4–20 kb−1) than for the experiment using standard Taq buffer (Figure 2B vs Figure 1B). When Pfu polymerase was used instead of Taq comparable results (mixed sequences, a GpC context bias, mutation frequencies ∼13 kb−1) were obtained including the same GpC bias (Supplementary Figure S2).
Variation in 3DPCR long-range Taq polymerase background mutation rate across the PCR block. (A) White spots correspond to PCR-positive samples for TP53 DNA. Those samples indicated by an asterisk were cloned and sequenced. (B) Mutation matrices for A6-, B3- and C2-derived sequences; transitions were invariably of the type N->T,A; the number of bases sequenced is given by n. (C) 5′ Dinucleotide context for the G->A and C->T transitions, with the expected values shown as horizontal bars. (D) A collection of the most highly mutated sequences. To compact the data, only variable sites are shown, their positions being identified above. Nucleotide positions should be read from top to bottom. To the right are the numbers of mutations per clone (mut) and the minimum number of recombination events (rec) to explain the complexity. Zones of recombination are highlighted in grey shading when possible.
Given these experimental baselines using cloned DNA, we turned to DNA from peritumoral cirrhotic tissue from patients with HBV- or HCV-associated hepatocellular carcinoma (HCC). We had previously shown that numerous A3 genes were upregulated in the liver (Vartanian et al, 2010). As the β-catenin gene, CTNNB1, harbors mutations in residues 32–45 in approximately 10–40% of HCCs (Miyoshi et al, 1998; Datta et al, 2008), we focused on a 155 bp segment that encodes this region. Three cirrhotic samples (nos. 310, 326 and 345) were analyzed along with tissue from one healthy liver (HL2). Sample nos. 310 and 345 harboured CTNNB1 mutations in the accompanying tumour, whereas no. 326 did not. Differential DNA denaturation PCR was performed using a 7 °C denaturation gradient (80–87 °C) and Taq polymerase. DNA from the last positive amplification was cloned and sequenced.
Once again sequences with both C->T and G->A substitutions dominated with up to four such substitutions per sequence, although there were a few sequences encoding just one type of transition (Figure 3A and B). The overall mutation frequency for sample no. 310 was ∼3 × that for sample nos. 326, 345 and HL2 and more than 13 x the plasmid control (Figure 3C). Once again the GpC context predominated (Figure 3D), which argues against the involvement of an A3 enzyme.
Differential DNA denaturation PCR-derived CTNNB1 sequences from cirrhotic and normal liver DNA. (A) A collection of 3DPCR recovered CTNNB1 sequences from viral-associated liver cirrhosis. The amino-acid sequence is shown above where bold case identifies the residues that frequently are substituted in liver cancers. The annotation is as for Figure 1D. (B) Stack-up graph of the number of GC->AT transitions per sequence per liver sample. (C) Mutation frequencies per sample along with the absolute number of mutations and bases sequences (within parentheses). (D) 5′ Dinucleotide context of GC->AT transitions for G->A and C->T transitions respectively.
Is it possible to distinguish lightly A3-edited sequences from the 3DPCR background described above, and what would an unambiguous A3-edited β-catenin look like? As haematopoietic cells generally show high A3 expression levels compared with other tissues (http://www.biogps.org), we analysed DNA from purified CD4+ T cells from PBMCs that had been treated by PHA, IL-2 and interferon-α, which strongly upregulates the A3A gene (Aynaud et al, 2012). To increase detection of A3-edited nuDNA, the cells were transduced with a lentivirus encoding the uracil N-glycosylase inhibitor UGI to block catabolism of dU-containing DNA (Wang and Mosbaugh, 1988). Using a 7 °C gradient 3DPCR amplification of the same β-catenin gene segment revealed a signal differing by 1 °C with respect to the non-stimulated CD4+ T cells as a control (Figure 4A and C). Cloning and sequencing revealed monotonously C->T or G->A substituted sequences (Figure 4A–C) with editing focused on TpC (GpA) dinucleotides, all consistent with editing by A3A and/or A3B (Figure 4E and F).
APOBEC3 hypermutated CTNNB1 sequences from purified human CD4+ T cells. (A) A collection of sequences bearing all the hallmarks of APOBEC3 editing from interferon α treated purified human CD4+ T cells – monotonously substituted with a penchant for GpA. (B) A collection of sequences, this time C->T hypermutants from activated human CD4+ T cells. (C) A collection of sequences recovered from unstimulated CD4+ T cells as control for the above. The first group of 10 shows mixed sequences with both C->T and G->A substitutions, the annotation is as for Figure 1D. The second group of two sequences is monotonously G->A substituted with a penchant for GpA. (D) Three different mutated CTNNB1 molecular clones A, B and C were mixed and amplified by 3DPCR. The lower three sequences are recombinants of the input sequences and denoted as B and C and A and B, the regions of recombination being highlighted in grey. (E and F) 5′ Dinucleotide context for the G->A and C->T transitions, with the expected values shown as horizontal bars, NS, unstimulated.
The non-stimulated control proved interesting. The last positive 3DPCR amplification was at 83.7 °C in the centre of the block to minimise the effects of thermal heterogeneity (Figure 4A and B). A collection of sequences was found that could be broken down into two groups of AT-rich sequences (Figure 4C). The first group of 10 sequences corresponded to what we can call 3DPCR background sequences containing up to five substitutions per locus (32 kb−1) involving both C->T and G->A transitions, there being a bias for transitions in GpC dinucleotides (not shown). The second group of two sequences in Figure 4C were monotonously G->A substituted compared with the plus strand with five and six transitions, most of which were in TpC and CpC. Indeed, the dinucleotide context is very similar to that of the bona fide A3-edited β-catenin sequences recovered from IFN-α-stimulated CD4+ T cells (Figure 4A). Accordingly, it appears that 3DPCR is indeed capable of recovering lightly A3-edited sequences when present.
The constant presence of sequences bearing both C->T and G->A transitions was intriguing, no matter what the source of input DNA was – molecularly cloned, and tissue- or PBMC-derived DNA. We have previously shown that recombination during PCR can contribute to the complexity of amplified DNA (Meyerhans et al, 1990). It usually occurs towards the end of amplification when [DNA]>[Taq], the polymerase simply being unable to complete synthesis of the nascent strands before the next temperature shift to denaturation.
Long elongation times reduce recombination but do not eliminate the problem (Meyerhans et al, 1990). In the above experiments involving TP53 and CTNNB1 loci, 10 min elongation times were used. However, as 3DPCR amplification occurs very close to the denaturation temperature, some templates may not be fully denatured, which would block polymerase elongation, thus rendering amplification less efficient. Indeed, this is reflected by the fact that we regularly use 42 cycles when performing 3DPCR, when normally 30 cycles would be more than sufficient to amplify the TP53 target.
To address the question of PCR recombination, 1 ng of DNA of three different molecularly cloned CTNNB1 sequences were mixed at an equimolar ratio and 3DPCR performed using the same 7 °C gradient. The last positive amplification (84.7 °C) was cloned and sequenced. By comparison of the input and output sequences (Figure 4D), it is clear that PCR-mediated recombination can increase the sequence complexity. The most parsimonious explanation for hypermutated sequences bearing both C->T and G->A transitions is that thermostable polymerases make a spectrum of mutations. For C->T and G->A transitions, one is dominant and occurs either in the context GpCpR (C->T transition) or YpGpC for a G->A transition. Successive cycles of 3DPCR enrich DNA bearing CG->TA transitions, and to a lesser extent N->TA transversions are selectively amplified. The complexity of the sequences is increased by PCR recombination.
Differential DNA denaturation PCR was initially used to recover A3 edited viral genomes and plasmid DNA (Vartanian et al, 2008, 2010; Suspène et al, 2011a, 2011b). Subsequently, it proved crucial to demonstrating that mitochondrial and nuclear DNA could be hyperedited (Suspène et al, 2011a). Recently two studies have reported A3 editing of nuDNA following recovery by 3DPCR (Shinohara et al, 2012; Burns et al, 2013a). The overall mutation rates were within the 3DPCR error range reported here. For one study the difference in the 3DPCR temperature was never >1 °C compared to the control while the mutation frequencies were 6–9 kb−1, only 2 fold greater than the basal error rate (Shinohara et al, 2012). Sequences bearing mixed C->T and G->A substitutions were found while the preferred 5′ dinucleotide context for C->T substitutions was GpC, observations reminiscent of the data shown in Figures 1, 2, 3 and suggests that they are non-physiological 3DPCR generated sequences.
If mixed DNA molecules resulted from A3 editing then the first edited strand would have to be replicated, followed by strand separation and APOBEC3 editing. The crucial difference compared to 3DPCR error would be the 5′ dinucleotide context associated with the C->T transitions. To date we have only seen sequences with large numbers of both C->T and G->A transitions in the context of hepatitis B virus editing by A3G (Suspène et al, 2005b). Here the numbers of C->T and G->A transitions was much greater than the 3–5 per clone reported here. Furthermore, as the dinucleotide context was 5′CpC for the C->T transitions on both strands, such sequences can be attributed to A3G editing and not 3DPCR (Suspène et al, 2005b).
Differential DNA denaturation PCR cannot be used to identify fixed C->T transitions in cancer genomes. Presently, the overall mutation frequency is ∼104–105 base substitutions per cancer genome, or 0.003–0.03 kb−1. These numbers are orders of magnitude lower than those for 3DPCR error, which are of the order of 4–20 mutations per kb; such numbers translate into 12–60 million mutations per diploid genome. As the reported mutation frequencies were of the same order as PCR error, along with mixed sequences associated with the GpC context, suggests that in all probability they result from 3DPCR.
Differential DNA denaturation PCR is very powerful at selectively amplifying A3 hyperedited DNA, but as shown above there is a grey zone where the editing frequency falls below 5–15 substitutions per kb−1. Having worked with it for many years (Suspène et al, 2005a, 2011a; Vartanian et al, 2008, 2010), we would suggest that cloning and sequencing is vital to a correct interpretation of the results. Prima face evidence of A3A and A3B editing of nuclear DNA requires that sequences should be monotonously substituted and coupled to a clear preference for TpC and CpC, the hallmark signature of these enzymes.
Change history
13 May 2014
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Alexandrov LB, Nik-Zainal S, Wedge DC, Campbell PJ, Stratton MR (2013) Deciphering signatures of mutational processes operative in human cancer. Cell Rep 3 (1): 246–259.
Aynaud MM, Suspene R, Vidalain PO, Mussil B, Guetard D, Tangy F, Wain-Hobson S, Vartanian JP (2012) Human Tribbles 3 protects nuclear DNA from cytidine deamination by APOBEC3A. J Biol Chem 287 (46): 39182–39192.
Beale RC, Petersen-Mahrt SK, Watt IN, Harris RS, Rada C, Neuberger MS (2004) Comparison of the differential context-dependence of DNA deamination by APOBEC enzymes: correlation with mutation spectra in vivo. J Mol Biol 337 (3): 585–596.
Bishop KN, Holmes RK, Sheehy AM, Davidson NO, Cho SJ, Malim MH (2004) Cytidine deamination of retroviral DNA by diverse APOBEC proteins. Curr Biol 14 (15): 1392–1396.
Bonvin M, Achermann F, Greeve I, Stroka D, Keogh A, Inderbitzin D, Candinas D, Sommer P, Wain-Hobson S, Vartanian JP, Greeve J (2006) Interferon-inducible expression of APOBEC3 editing enzymes in human hepatocytes and inhibition of hepatitis B virus replication. Hepatology 43 (6): 1364–1374.
Burns MB, Lackey L, Carpenter MA, Rathore A, Land AM, Leonard B, Refsland EW, Kotandeniya D, Tretyakova N, Nikas JB, Yee D, Temiz NA, Donohue DE, McDougle RM, Brown WL, Law EK, Harris RS (2013a) APOBEC3B is an enzymatic source of mutation in breast cancer. Nature 494 (7437): 366–370.
Burns MB, Temiz NA, Harris RS (2013b) Evidence for APOBEC3B mutagenesis in multiple human cancers. Nat Genet 45 (9): 977–983.
Carpenter MA, Li M, Rathore A, Lackey L, Law EK, Land AM, Leonard B, Shandilya SM, Bohn MF, Schiffer CA, Brown WL, Harris RS (2012) Methylcytosine and normal cytosine deamination by the foreign dna restriction enzyme APOBEC3A. J Biol Chem 287 (41): 34801–34808.
Conticello SG, Thomas CJ, Petersen-Mahrt SK, Neuberger MS (2005) Evolution of the AID/APOBEC family of polynucleotide (deoxy)cytidine deaminases. Mol Biol Evol 22 (2): 367–377.
Datta J, Kutay H, Nasser MW, Nuovo GJ, Wang B, Majumder S, Liu CG, Volinia S, Croce CM, Schmittgen TD, Ghoshal K, Jacob ST (2008) Methylation mediated silencing of microRNA-1 gene and its role in hepatocellular carcinogenesis. Cancer Res 68 (13): 5049–5058.
Di Noia JM, Neuberger MS (2007) Molecular mechanisms of antibody somatic hypermutation. Annu Rev Biochem 76: 1–22.
Eckert KA, Kunkel TA (1991) DNA polymerase fidelity and the polymerase chain reaction. PCR Methods Appl 1 (1): 17–24.
Eckert KA, Kunkel TA (1993) Effect of reaction pH on the fidelity and processivity of exonuclease-deficient Klenow polymerase. J Biol Chem 268 (18): 13462–13471.
Endo Y, Marusawa H, Kinoshita K, Morisawa T, Sakurai T, Okazaki IM, Watashi K, Shimotohno K, Honjo T, Chiba T (2007) Expression of activation-induced cytidine deaminase in human hepatocytes via NF-kappaB signaling. Oncogene 26 (38): 5587–5595.
Greenman C, Stephens P, Smith R, Dalgliesh GL, Hunter C, Bignell G, Davies H, Teague J, Butler A, Stevens C, Edkins S, O'Meara S, Vastrik I, Schmidt EE, Avis T, Barthorpe S, Bhamra G, Buck G, Choudhury B, Clements J, Cole J, Dicks E, Forbes S, Gray K, Halliday K, Harrison R, Hills K, Hinton J, Jenkinson A, Jones D, Menzies A, Mironenko T, Perry J, Raine K, Richardson D, Shepherd R, Small A, Tofts C, Varian J, Webb T, West S, Widaa S, Yates A, Cahill DP, Louis DN, Goldstraw P, Nicholson AG, Brasseur F, Looijenga L, Weber BL, Chiew YE, DeFazio A, Greaves MF, Green AR, Campbell P, Birney E, Easton DF, Chenevix-Trench G, Tan MH, Khoo SK, Teh BT, Yuen ST, Leung SY, Wooster R, Futreal PA, Stratton MR (2007) Patterns of somatic mutation in human cancer genomes. Nature 446 (7132): 153–158.
Harris RS, Bishop KN, Sheehy AM, Craig HM, Petersen-Mahrt SK, Watt IN, Neuberger MS, Malim MH (2003) DNA deamination mediates innate immunity to retroviral infection. Cell 113 (6): 803–809.
Henry M, Guetard D, Suspene R, Rusniok C, Wain-Hobson S, Vartanian JP (2009) Genetic editing of HBV DNA by monodomain human APOBEC3 cytidine deaminases and the recombinant nature of APOBEC3G. PLoS One 4 (1): e4277.
Jarmuz A, Chester A, Bayliss J, Gisbourne J, Dunham I, Scott J, Navaratnam N (2002) An anthropoid-specific locus of orphan C to U RNA-editing enzymes on chromosome 22. Genomics 79 (3): 285–296.
Kidd JM, Newman TL, Tuzun E, Kaul R, Eichler EE (2007) Population stratification of a common APOBEC gene deletion polymorphism. PLoS Genet 3 (4): e63.
Komatsu A, Nagasaki K, Fujimori M, Amano J, Miki Y (2008) Identification of novel deletion polymorphisms in breast cancer. Int J Oncol 33 (2): 261–270.
Koning FA, Newman EN, Kim EY, Kunstman KJ, Wolinsky SM, Malim MH (2009) Defining APOBEC3 expression patterns in human tissues and hematopoietic cell subsets. J Virol 83 (18): 9474–9485.
Landry S, Narvaiza I, Linfesty DC, Weitzman MD (2011) APOBEC3A can activate the DNA damage response and cause cell-cycle arrest. EMBO Rep 12 (5): 444–450.
Lecossier D, Bouchonnet F, Clavel F, Hance AJ (2003) Hypermutation of HIV-1 DNA in the absence of the Vif protein. Science 300 (5622): 1112.
Mangeat B, Turelli P, Caron G, Friedli M, Perrin L, Trono D (2003) Broad antiretroviral defence by human APOBEC3G through lethal editing of nascent reverse transcripts. Nature 424 (6944): 99–103.
Mariani R, Chen D, Schrofelbauer B, Navarro F, Konig R, Bollman B, Munk C, Nymark-McMahon H, Landau NR (2003) Species-specific exclusion of APOBEC3G from HIV-1 virions by Vif. Cell 114 (1): 21–31.
Matsumoto Y, Marusawa H, Kinoshita K, Endo Y, Kou T, Morisawa T, Azuma T, Okazaki IM, Honjo T, Chiba T (2007) Helicobacter pylori infection triggers aberrant expression of activation-induced cytidine deaminase in gastric epithelium. Nat Med 13 (4): 470–476.
Meyerhans A, Vartanian JP, Wain-Hobson S (1990) DNA recombination during PCR. Nucleic Acids Res 18 (7): 1687–1691.
Miyoshi Y, Iwao K, Nagasawa Y, Aihara T, Sasaki Y, Imaoka S, Murata M, Shimano T, Nakamura Y (1998) Activation of the beta-catenin gene in primary hepatocellular carcinomas by somatic alterations involving exon 3. Cancer Res 58 (12): 2524–2527.
Morisawa T, Marusawa H, Ueda Y, Iwai A, Okazaki IM, Honjo T, Chiba T (2008) Organ-specific profiles of genetic changes in cancers caused by activation-induced cytidine deaminase expression. Int J Cancer 123 (12): 2735–2740.
Mussil B, Suspene R, Aynaud MM, Gauvrit A, Vartanian JP, Wain-Hobson S (2013) Human APOBEC3A isoforms translocate to the nucleus and induce DNA double strand breaks leading to cell stress and death. PLoS One 8 (8): e73641.
Nik-Zainal S, Alexandrov LB, Wedge DC, Van Loo P, Greenman CD, Raine K, Jones D, Hinton J, Marshall J, Stebbings LA, Menzies A, Martin S, Leung K, Chen L, Leroy C, Ramakrishna M, Rance R, Lau KW, Mudie LJ, Varela I, McBride DJ, Bignell GR, Cooke SL, Shlien A, Gamble J, Whitmore I, Maddison M, Tarpey PS, Davies HR, Papaemmanuil E, Stephens PJ, McLaren S, Butler AP, Teague JW, Jonsson G, Garber JE, Silver D, Miron P, Fatima A, Boyault S, Langerod A, Tutt A, Martens JW, Aparicio SA, Borg A, Salomon AV, Thomas G, Borresen-Dale AL, Richardson AL, Neuberger MS, Futreal PA, Campbell PJ, Stratton MR (2012) Mutational processes molding the genomes of 21 breast cancers. Cell 149 (5): 979–993.
Pleasance ED, Cheetham RK, Stephens PJ, McBride DJ, Humphray SJ, Greenman CD, Varela I, Lin ML, Ordonez GR, Bignell GR, Ye K, Alipaz J, Bauer MJ, Beare D, Butler A, Carter RJ, Chen L, Cox AJ, Edkins S, Kokko-Gonzales PI, Gormley NA, Grocock RJ, Haudenschild CD, Hims MM, James T, Jia M, Kingsbury Z, Leroy C, Marshall J, Menzies A, Mudie LJ, Ning Z, Royce T, Schulz-Trieglaff OB, Spiridou A, Stebbings LA, Szajkowski L, Teague J, Williamson D, Chin L, Ross MT, Campbell PJ, Bentley DR, Futreal PA, Stratton MR (2010) A comprehensive catalogue of somatic mutations from a human cancer genome. Nature 463 (7278): 191–196.
Refsland EW, Stenglein MD, Shindo K, Albin JS, Brown WL, Harris RS (2010) Quantitative profiling of the full APOBEC3 mRNA repertoire in lymphocytes and tissues: implications for HIV-1 restriction. Nucleic Acids Res 38 (13): 4274–4284.
Sheehy AM, Gaddis NC, Choi JD, Malim MH (2002) Isolation of a human gene that inhibits HIV-1 infection and is suppressed by the viral Vif protein. Nature 418 (6898): 646–650.
Shinohara M, Io K, Shindo K, Matsui M, Sakamoto T, Tada K, Kobayashi M, Kadowaki N, Takaori-Kondo A (2012) APOBEC3B can impair genomic stability by inducing base substitutions in genomic DNA in human cells. Sci Rep 2: 806.
Stenglein MD, Burns MB, Li M, Lengyel J, Harris RS (2010) APOBEC3 proteins mediate the clearance of foreign DNA from human cells. Nat Struct Mol Biol 17 (2): 222–229.
Stephens PJ, McBride DJ, Lin ML, Varela I, Pleasance ED, Simpson JT, Stebbings LA, Leroy C, Edkins S, Mudie LJ, Greenman CD, Jia M, Latimer C, Teague JW, Lau KW, Burton J, Quail MA, Swerdlow H, Churcher C, Natrajan R, Sieuwerts AM, Martens JW, Silver DP, Langerod A, Russnes HE, Foekens JA, Reis-Filho JS, van 't Veer L, Richardson AL, Borresen-Dale AL, Campbell PJ, Futreal PA, Stratton MR (2009) Complex landscapes of somatic rearrangement in human breast cancer genomes. Nature 462 (7276): 1005–1010.
Stephens PJ, Tarpey PS, Davies H, Van Loo P, Greenman C, Wedge DC, Nik-Zainal S, Martin S, Varela I, Bignell GR, Yates LR, Papaemmanuil E, Beare D, Butler A, Cheverton A, Gamble J, Hinton J, Jia M, Jayakumar A, Jones D, Latimer C, Lau KW, McLaren S, McBride DJ, Menzies A, Mudie L, Raine K, Rad R, Chapman MS, Teague J, Easton D, Langerod A, Lee MT, Shen CY, Tee BT, Huimin BW, Broeks A, Vargas AC, Turashvili G, Martens J, Fatima A, Miron P, Chin SF, Thomas G, Boyault S, Mariani O, Lakhani SR, van de Vijver M, van 't Veer L, Foekens J, Desmedt C, Sotiriou C, Tutt A, Caldas C, Reis-Filho JS, Aparicio SA, Salomon AV, Borresen-Dale AL, Richardson AL, Campbell PJ, Futreal PA, Stratton MR (2012) The landscape of cancer genes and mutational processes in breast cancer. Nature 486 (7403): 400–404.
Suspène R, Aynaud M, Guétard D, Henry M, Eckhoff G, Marchio A, Pineau P, Dejean A, Vartanian JP, Wain-Hobson S (2011a) Somatic hypermutation of human mitochondrial and nuclear DNA by APOBEC3 cytidine deaminases, a pathway for DNA catabolism. Proc Natl Acad Sci USA 108 (12): 4858–4863.
Suspène R, Aynaud MM, Koch S, Pasdeloup D, Labetoulle M, Gaertner B, Vartanian JP, Meyerhans A, Wain-Hobson S (2011b) Genetic editing of herpes simplex virus 1 and Epstein–Barr herpesvirus genomes by human APOBEC3 cytidine deaminases in culture and in vivo. J Virol 85 (15): 7594–7602.
Suspène R, Aynaud MM, Vartanian JP, Wain-Hobson S (2013) Efficient deamination of 5-methylcytidine and 5-substituted deoxycytidine residues in DNA by human APOBEC3A cytidine deaminase. PLoS One 8 (8): e73641.
Suspène R, Guétard D, Henry M, Sommer P, Wain-Hobson S, Vartanian JP (2005b) Extensive editing of both hepatitis B virus DNA strands by APOBEC3 cytidine deaminases in vitro and in vivo. Proc Natl Acad Sci USA 102 (23): 8321–8326.
Suspène R, Henry M, Guillot S, Wain-Hobson S, Vartanian JP (2005a) Recovery of APOBEC3-edited human immunodeficiency virus G->A hypermutants by differential DNA denaturation PCR. J Gen Virol 86 (Part 1): 125–129.
Suspène R, Sommer P, Henry M, Ferris S, Guétard D, Pochet S, Chester A, Navaratnam N, Wain-Hobson S, Vartanian JP (2004) APOBEC3G is a single-stranded DNA cytidine deaminase and functions independently of HIV reverse transcriptase. Nucleic Acids Res 32 (8): 2421–2429.
Vartanian JP, Guétard D, Henry M, Wain-Hobson S (2008) Evidence for editing of human papillomavirus DNA by APOBEC3 in benign and precancerous lesions. Science 320 (5873): 230–233.
Vartanian JP, Henry M, Marchio A, Suspène R, Aynaud MM, Guétard D, Cervantes-Gonzalez M, Battiston C, Mazzaferro V, Pineau P, Dejean A, Wain-Hobson S (2010) Massive APOBEC3 editing of hepatitis B viral DNA in cirrhosis. PLoS Pathogen 6 (5): e1000928.
Wang Z, Mosbaugh DW (1988) Uracil-DNA glycosylase inhibitor of bacteriophage PBS2: cloning and effects of expression of the inhibitor gene in Escherichia coli. J Bacteriol 170 (3): 1082–1091.
Wijesinghe P, Bhagwat AS (2012) Efficient deamination of 5-methylcytosines in DNA by human APOBEC3A, but not by AID or APOBEC3G. Nucleic Acids Res 40 (18): 9206–9217.
Zhang T, Cai J, Chang J, Yu D, Wu C, Yan T, Zhai K, Bi X, Zhao H, Xu J, Tan W, Qu C, Lin D (2012) Evidence of associations of APOBEC3B gene deletion with susceptibility to persistent HBV infection and hepatocellular carcinoma. Hum Mol Genet 22 (6): 1262–1269.
Acknowledgements
This work was supported by grants from the Institut Pasteur, INCa and the CNRS. VC and MSB were supported by OSEO and the Ligue Nationale Contre le Cancer.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License.
Supplementary Information accompanies this paper on British Journal of Cancer website
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Suspène, R., Caval, V., Henry, M. et al. Erroneous identification of APOBEC3-edited chromosomal DNA in cancer genomics. Br J Cancer 110, 2615–2622 (2014). https://doi.org/10.1038/bjc.2014.176
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2014.176
Keywords
This article is cited by
-
Mouse APOBEC1 cytidine deaminase can induce somatic mutations in chromosomal DNA
BMC Genomics (2019)
-
Genotoxic stress increases cytoplasmic mitochondrial DNA editing by human APOBEC3 mutator enzymes at a single cell level
Scientific Reports (2019)
-
Endogenous APOBEC3B Overexpression Constitutively Generates DNA Substitutions and Deletions in Myeloma Cells
Scientific Reports (2019)
-
APOBEC3A catabolism of electroporated plasmid DNA in mouse muscle
Gene Therapy (2015)
-
A prevalent cancer susceptibility APOBEC3A hybrid allele bearing APOBEC3B 3′UTR enhances chromosomal DNA damage
Nature Communications (2014)