Abstract
Background:
Breast cancer is the leading cause of cancer death in women living in the western hemisphere. Despite major advances in first-line endocrine therapy of advanced oestrogen receptor (ER)-positive breast cancer, the frequent recurrence of resistant cancer cells represents a serious obstacle to successful treatment. Understanding the mechanisms leading to acquired resistance, therefore, could pave the way to the development of second-line therapeutics. To this end, we generated an ER-positive breast cancer cell line (MCF-7) with resistance to the therapeutic anti-oestrogen fulvestrant (FUL) and studied the molecular changes involved in resistance.
Methods:
Naive MCF-7 cells were treated with increasing FUL concentrations and the gene expression profile of the resulting FUL-resistant strain (FR.MCF-7) was compared with that of naive cells using GeneChip arrays. After validation by real-time PCR and/or western blotting, selected resistance-associated genes were functionally studied by siRNA-mediated silencing or pharmacological inhibition. Furthermore, general mechanisms causing aberrant gene expression were investigated.
Results:
Fulvestrant resistance was associated with repression of GPER and the overexpression of CDK6, whereas ERBB2, ABCG2, ER and ER-related genes (GREB1, RERG) or genes expressed in resistant breast cancer (BCAR1, BCAR3) did not contribute to resistance. Aberrant GPER and CDK6 expression was most likely caused by modification of DNA methylation and histone acetylation, respectively. Therefore, part of the resistance mechanism was loss of RB1 control. The hSWI/SNF (human SWItch/Sucrose NonFermentable) chromatin remodelling complex, which is tightly linked to nucleosome acetylation and repositioning, was also affected, because as a stress response to FUL treatment-naive cells altered the expression of five subunits within a few hours (BRG1, BAF250A, BAF170, BAF155, BAF47). The aberrant constitutive expression of BAF250A, BAF170 and BAF155 and a deviant stress response of BRG1, BAF170 and BAF47 in FR.MCF-7 cells to FUL treatment accompanied acquired FUL resistance. The regular and aberrant expression profiles of BAF155 correlated directly with that of CDK6 in naive and in FR.MCF-7 cells corroborating the finding that CDK6 overexpression was due to nucleosome alterations.
Conclusion:
The study revealed that FUL resistance is associated with the dysregulation of GPER and CDK6. A mechanism leading to aberrant gene expression was most likely unscheduled chromatin remodelling by hSWI/SNF. Hence, three targets should be conceptually addressed in a second-line adjuvant therapy: the catalytic centre of SWI/SNF (BRG1) to delay the development of FUL resistance, GPER to increase sensitivity to FUL and the reconstitution of the RB1 pathway to overcome resistance.
Similar content being viewed by others
Main
Oestrogen is the main stimulant for the growth of breast cancer cells. As a consequence, oestrogen receptor alpha (ERα) is the most important target in breast cancer treatment (Jensen and Jordan, 2003). Over the past 30 years, antagonists of steroid hormones are clinically important in the management of receptor-positive breast cancer and the ER antagonist tamoxifen (TAM), which is a non-steroidal selective ER modulator (SERM), is widely used as the gold standard for antihormonal therapy. However, the duration of response of advanced breast cancer is limited (progression-free survival <10 month) because of the development of hormone-independent tumours in virtually all cases (Badia et al, 2000; Miyoshi et al, 2010). Therefore, anti-oestrogen resistance is frequently observed in patients after long-term treatment with TAM, stressing the development of resistance to endocrine therapy as a clinically important problem. Reportedly, resistance to TAM correlated significantly with CpG hypomethylation of ERβ (Chang et al, 2005) and its overexpression (Speirs et al, 1999) and resistance to FUL with ERα downregulation (Fan et al, 2006). Selective ER modulator resistance is associated with the acquisition of oestrogen-independent growth (Badia et al, 2000; Schiff et al, 2004; Sabnis et al, 2005; Miyoshi et al, 2010), which is accomplished in particular by the upregulation of ERBB2 (Hu and Mokbel, 2001; Chung et al, 2002; Shou et al, 2004; Gutierrez et al, 2005). Also, increased ABCG2 levels cause TAM resistance (Selever et al, 2011).
It was shown that TAM resistance could be overcome by another SERM, fulvestrant (FUL; synonym: faslodex, ICI 182 780; Shaw et al, 2006), which is a pure anti-oestrogen without agonistic and solely antagonistic features. As FUL follows TAM, we addressed the question regarding the mechanisms that become induced when breast cancer cells develop resistance to FUL.
Materials and methods
Cell culture
The MCF-7 breast cancer cell line was purchased from ATCC (Rockville, MD, USA) and was cultivated in DMEM/F-12 1 : 1 medium supplemented with 10% heat-inactivated FCS, 1% L-glutamine and 1% penicillin/streptomycin. Fulvestrant resistance was achieved by treating MCF-7 cells with increasing concentrations (up to 1 μ M) of FUL over a period of 6 months and the resistant cell line (FR.MCF-7) was maintained in the above-described medium and in the presence of 500 nM FUL. All cells were grown at 37 °C in a humidified atmosphere containing 5% CO2. If not mentioned otherwise, all media and supplements were obtained from Invitrogen (Karlsruhe, Germany).
Reagents and antibodies
Fulvestrant, 5-aza-2′-deoxycytidine (AZA), trichostatin A (TSA), fumitremorgin C (FTC), 17-β oestradiol (E2) and CDK6 inhibitor PD0332991 were purchased from Sigma-Aldrich (Munich, Germany), and trastuzumab (TRA) from Roche (Basel, Switzerland). Amersham ECLPlus Western Blotting Detection System was from GE Healthcare (Buckinghamshire, UK).
Mouse monoclonal anti-β-actin (ascites fluid; clone AC-15, Cat, no. A5441) was from Sigma-Aldrich. Anti-cyclin D1 (M-20; Cat. no. sc-718), p21 (C-19; Cat. no. sc-397), cyclin A (H-432; Cat. no. sc-751), cyclin E (M-20; Cat. no. sc-481), CDK 4 (C-22; Cat. no. sc-260), α-tubulin (TU-02; Cat. no. sc-8035) and β-tubulin (H-235; Cat. no. sc-9104) were from Santa Cruz Biotechnologies Inc. (Santa Cruz, CA, USA). Anti-MYC (AB-02, 9E10; Cat. no. MS-139-D1) was purchased from Neomarkers (Fremont, CA, USA), and p53 antibody (Cat. no. 1767) from Immunotech (Marseille, France). Anti-CDK 6 (Cat. no. 3136), retinoblastoma (RB1; Cat. no. 9309), phospho(Ser780)RB1 (Cat. no. 3590), SmarcC2/BAF170 (Cat. no. 8829) and SmarcC1/BAF 155 (Cat. no. 9053) were from Cell Signalling (Danvers, MA, USA). Anti-ARID1A/BAF250A (Cat. no. ab-50878) and SmarcA4/BRG1 (Cat. no. ab-4081) and anti-SNF5/BAF47 (EPR6966; Cat. no. ab-126734) were from Abcam (Cambridge, UK), anti-SmarcE1/BAF57 (Cat. no. NB100-2591) from Novus Biologicals (Cambridge, UK), anti-ACTL6A/BAF53A (Cat. no. A301-391 A) from Bethyl Antibodies (Montgomery, TX, USA) and anti-mouse and anti-rabbit IgGs were from Dako (Glostrup, Denmark).
SDS gel electrophoresis and western blotting
Cells were grown in petri dishes (6 cm diameter) to 80% confluence and treated with 500 nM FUL or 50 nM PD0332991. Then, cells were washed two times with cold PBS and lysed in a buffer containing 150 mM NaCl, 50 mM Tris (pH 8.0), 1% Triton X-100, 1 mM phenylmethylsulphonyl fluoride and 1 mM Protease Inhibitor Cocktail (Sigma-Aldrich). The lysates were centrifuged at 12 000 r.p.m. for 20 min at 4 °C and the supernatants transferred to 1.5 ml tubes and stored at −20 °C until further analysis. Equal amounts of protein lysate were mixed with (2 × )SDS (sodium dodecyl sulphate) sample buffer, separated by 10% polyacrylamide gel electrophoresis containing SDS (SDS–PAGE) and electrotransferred onto PVDV membranes (Hybond-P; Amersham), 4 °C overnight. Western blotting was performed according to the protocol described by Giessrigl et al (2012). In short, staining membranes with Ponceau S controlled equal sample loading, and after washing with Tris-buffered saline (TBS) (pH 7.6), membranes were blocked in 5% non-fat dry milk in TBS containing 0.1% Tween-20 for 1 h. Membranes were incubated with the first antibody (in blocking solution, dilution 1 : 500–1 : 1000) by gently rocking at 4 °C overnight, washed with TBS containing 0.1% Tween-20 and further incubated with the second antibody (peroxidase-conjugated swine anti-rabbit IgG or rabbit anti-mouse IgG, dilution 1 : 2000–1 : 5000 in blocking solution) for 1 h. Chemoluminescence was developed by the ECL plus detection kit (GE Healthcare, Buckinghamshire, UK) and analysed using a Lumi-Imager F1 Workstation (Roche, Basel, Switzerland).
Proliferation analysis
MCF-7 cells were seeded in 24-well plates at a concentration of 1 × 105 cells per ml allowing logarithmic growth within 96 h. Afterwards, cells were incubated with FUL or PD0332991. The cell number was determined using an electronic cell counter (CASY; Roche Applied Science, Mannheim, Germany). Proliferation rates were calculated as described (Maier et al, 2006; Strasser et al, 2006). Cell duplication was calculated as follows:
doubling time=log 2 × h cultivation time/log N−log N0, where h is the cultivation time, N is the cell number after the time of cultivation and N0 the cell number at the beginning of cultivation.
Quantitative RT-PCR
MCF-7 and FR.MCF-7 cells (1 × 105) were seeded in six wells, cultivated for 24 h, harvested and homogenised using Qia-shredder (Qiagen, Hilden, Germany) and further processed according to the instructions of RNeasy Mini Kit (Qiagen). The total RNA concentration was measured using a NanoDrop Fluorospectrometer (Thermo Fisher Scientific Inc., Waltham, MA, USA). Complementary DNA synthesis from 1 μg RNA was performed using Superscript-first-strand synthesis systems for RT–PCR (Invitrogen, Carlsbad, CA, USA). The transcript levels of GREB1, RERG, GPER, BCAR1, BCAR3, CDK6, ERBB2 and ABCG2 were investigated by real-time PCR using Taqman detection system (Applied Biosystems, Carlsbad, CA, USA). The expression of the housekeeping gene glyceralaldehyde 3-phosphate dehydrogenase (GAPDH) served as an internal control. Assay ID numbers of the Taqman gene expression kits were: GAPDH, HS99999905_m1; GREB1, HS00536409; RERG, HS00922947; GPER, HS01116133; BCAR1, HS01547079; BCAR3, HS00981957; CDK6, HS01026371; and ERBB2, HS01001580. Cycle programme (95 °C for 10 min to activate polymerase followed by 40 cycles of 95 °C for 15 s and 60 °C for 1 min) was started on an Abi Prism 7000 Sequence Detection System (Applied Biosystems). Real-time PCR was performed in duplicate for each gene investigated. Negative controls, containing water instead of cDNA, confirmed the absence of RNA/DNA in all reagents applied in the assay.
siRNA knockdown
MCF-7 and FR.MCF-7 cells (1.0 × 105) were seeded onto 6 cm cell culture plates and cultivated for 24 h. On the day of transefction, 3.6 μg siRNA (corresponding to 360 nM final concentration) was diluted in 500 μl medium. Forty-three microlitres of RNAiFect transfection reagent (Qiagen) was added to the diluted siRNA and the mixture incubated for 15 min at room temperature to allow the formation of transfection complexes. Then, the solution was added dropwise onto the cells. After incubation for 16 h at 37 °C, the medium was changed, and after further 24 h, cells were used for experiments. The siRNAs were from Applied Biosystems (Life Technologies Inc., Carlsbad, CA, USA) and the Silencer Select siRNA IDs were: GREB1 (s18650), RERG (s224986), GPER (s6503), BCAR1 (s18371) and BCAR3 (s228334); negative control cat. no. 4390843.
Gene expression analysis by Affymetrix
Total RNA was extracted from 1 × 105 cells (grown in six-well plates) by using Qiagen RNeasy Kit (Qiagen). All purified RNA samples were quality controlled by measuring the optical density at 230, 260 and 280nm and by analysing an aliquot of the RNA preparation on an Agilent 2100 Bioanalyzer using RNA 6000 Nano chips (Agilent Technologies, Santa Clara, CA, USA). The Bioanalyzer Expert software (Agilent) was used to calculate RNA integrity numbers, which were above 9.0 in all samples. For GeneChip analysis, we followed the standard protocol provided by Affymetrix (Santa Clara, CA, USA) for converting the polyA+ fraction of ∼1.5 μg total RNA into double-stranded cDNA, which was purified with a GeneChip sample cleanup module column and then used as template for in vitro transcription into biotin-labelled complementary RNA (cRNA). We used reagents and materials contained in the GeneChip Expression 3′ Amplification One-Cycle Target Labelling Kit (Affymetrix). Both cRNA preparations and fragmented cRNA samples were quality controlled by analysing on an Agilent 2100 Bioanalyzer. Hybridization was performed to HG-U133 Plus 2.0 GeneChips (Affymetrix) for 16 h at 45 °C with constant rotation at 60 r.p.m. Washing, staining and scanning of the chips was performed using the Fluidics 450 Station and the GeneChip 3000 7G Scanner following the manufacturer’s protocols. Scanned raw data images were processed with GeneChip Operating Software 1.4 (Affymetrix). A quality control report was subsequently made using Bioconductor (open source software for bioinformatics; hosted by Fred Hutchinson Cancer Research Center, Seattle, WA, USA), and normalization and signal extraction was carried out with the RMA (Robust Multichip Average) approach (Bolstadt et al, 2003).
Statistical analyses
For statistical analyses, Prism 5 software package (GraphPad, San Diego, CA, USA) was used. The values were expressed as mean±s.e.m. and the Student’s t-test was used to compare differences between controls and individual samples, whereas analyses of variance (one-way ANOVA together with Dunnett’s post-test) was used to analyse treatment groups. Statistical significance level was set to P<0.05.
Results
Resistance to FUL occurs in the presence of functional ER
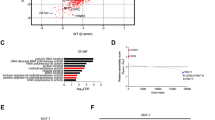
Naive MCF-7 cells were exposed to increasing concentrations of FUL (application schedule described in ‘Materials and Methods’) to generate FUL-resistant cells (FR.MCF-7; Figure 1A). After 1, 2, 4 and 6 months of FUL treatment, the gene expression profiles were analysed by GeneChip (Affymetrix) (Table 1). Gene expression profiling showed that ERα, ERβ and its target progesterone receptor (PGR) were not downregulated after long-term treatment, but expressed at similar levels in FR.MCF-7 and naive MCF-7 cells, which was confirmed by western blotting (data not shown). FR.MCF-7 cells still responded to 100 nM E2 with increased proliferation (Figure 1B) and to TAM with retarded cell growth (Figure 1C). Thus, neither did long-term exposure to FUL downregulate ERs constitutively (Fan et al, 2006) nor did ERα lose its functionality. However, FR.MCF-7 cells were less sensitive to TAM than naive MCF-7 cells, and therefore, FUL resistance partly compromised a common response mechanism.
Sensitivity of MCF-7 and FR.MCF-7 cells to FUL, TAM and E2. (A) MCF-7 and FUL-resistant (FR) MCF-7 cells were exposed to increasing concentrations of FUL or (C) TAM and cell proliferation was measured after 48 h. (B) Fulvestrant-resistant.MCF-7 cells were grown in hormone-deprived (DCC) medium±E2 for 96 h when cells were counted. Experiments were carried out in triplicate and error bars indicate s.e.m. and asterisks denote significance. (A, C) One-way ANOVA and Dunnett’s post-test and (B) t-test.
Aberrant mRNA expression of GPER and CDK6 is reversed by AZA and TSA, respectively
Expression arrays revealed that ERs and PGR levels were unchanged in FR.MCF-7 cells, yet genes associated with oestrogen signalling (RERG, GREB1 and GPER) were repressed. Genes known to contribute to chemoresistance of breast cancer, that is, BCAR1 (poor overall survival when overexpressed), BCAR3 (homologous to BCAR1), ABCG2 and ERBB2 (van Agthoven et al, 1998; van der Flier et al, 2000; Hu and Mokbel, 2001; Selever et al, 2011), were upregulated in FR.MCF-7 cells (Table 1). Notably, CDK6 expression was also increased. However, only the downregulation of GPER and the upregulation of BCAR1, ABCG2 and CDK6 could be confirmed in FR.MCF-7 cells by Q-PCR (Figure 2), but not the aberrant expression levels of GREB1, BCAR3 and ERBB2.
Gene expression upon treatment with AZA and TSA. Fulvestrant-resistant.MCF-7 cells were pretreated for 24 h with 100 ng ml−1 TSA and for 72 h with 2.5 μM AZA. Then, cells were lysed, RNA was extracted, mRNA reverse transcribed to cDNA and Q-PCR performed. The expression levels of GREB1, BCAR1, BCAR3, GPER, CDK6, ABCG2 and ERBB2 were standardised to GAPDH mRNA expression in FR.MCF-7 cells and naive MCF-7 cells. Experiments were carried out in duplicate and error bars indicate s.e.m.
Deprotection of genomic DNA regions can be triggered by modulating the acetylation of histone octamers. In consequence, promoter and enhancer regions, which were previously masked by nucleosomes, become now accessible by methyltransferases. The methylation of CpG islands prevents transcription factor binding and this causes changes in overall protein expression, which may lead to chemoresistance (Chang et al, 2005; Fan et al, 2006; Fiegl et al, 2006). Tamoxifen was shown to epigenetically modulate gene expression that is relevant to acquired resistance (Encarnación et al, 1993; Badia et al, 2000; Stone et al, 2012; Vesuna et al, 2012). Hence, gene expression was examined after treatment of FR.MCF-7 cells with TSA (inhibitor of class I and II mammalian histone deacetylases) and AZA (cytidine analogue that inhibits methyltransferases, thereby hypomethylating the DNA) to modulate nucleosome acetylation and to reverse CpG methylation, respectively. Treatment with AZA restored GPER expression in FR.MCF-7 cells to approximately similar levels as in naive MCF-7 cells, but GPER expression remained virtually unaffected upon TSA treatment in the resistant cell line. Conversely, the increased expression of CDK6 in FR.MCF-7 cells was significantly reversed by TSA but became highly induced by AZA. In contrast, aberrant expression levels of BCAR1 and ABCG2 did not revert to normal levels neither by AZA nor by TSA, but were increased even further. The expression of GREB1 was suppressed by AZA and TSA (Figure 2). Hence, GPER inhibition and CDK6 induction in FR.MCF-7 cells were likely due to methylation and deacetylation events, respectively.
GPER contributes to FUL resistance
The siRNA-mediated downregulation of GPER in naive MCF-7 cells decreased the sensitivity to FUL (Figure 3A), suggesting that repression of GPER contributed to FUL resistance in FR.MCF-7 cells. Notably, siRNA suppressed gene expression only by 50% (data not shown), and therefore, the acquisition of a FUL-resistant phenotype by naive MCF-7 cells was weak, yet significant. For control reasons, the expression of GREB1 and RERG was knocked down by specific siRNAs. As with GPER, the suppression of GREB1 reduced the sensitivity to FUL in naive MCF-7 cells, whereas suppression of RERG did not (Figure 3A). Hence, GREB1 can also contribute to FUL resistance (however, the aberrant GREB1 expression detected by GeneChip could not be confirmed by Q-PCR).
Testing of specific gene products required for the cellular response to FUL. (A) MCF-7 cells were transfected with the indicated small interfering RNAs (siRNAs) or with control RNA in which no complementary cellular RNA exists, were treated with 500 nM FUL or solvent (Co) for 48 h and then cells were counted. (B, C) Fulvestrant-resistant.MCF-7 cells were exposed to 500 nM FUL alone or (B) in combination of 2.5 μ M FTC or (C) in combination with 1 μg ml−1 TRA, and the cell number was measured after 48 h. Experiments were carried out in triplicate, error bars indicate s.e.m. and asterisks denote significance (t-test).
The cause for the aberrant expression of BCAR and ABCG2 remained unclear (Figure 2). Neither the specific inhibition of ABCG2 with FTC (Figure 3B) nor the downregulation of BCAR1 by specific siRNA (data not shown) re-established sensitivity to FUL in FR.MCF-7 cells.
Overexpression and activation of ERBB2 indicates the acquisition of an ER-independent growth mechanism in TAM-insensitive breast cancer (Ghayad et al, 2010), and therefore, it was tested whether ERBB2 (although an increased expression could not be confirmed by Q-PCR) contributed to insensitivity to FUL. For this, FR.MC-7 cells were treated with 1 μg ml−1 TRA, which is a specific inhibitor of ERBB2. Trastuzumab had no effect on the proliferation of FR.MCF-7 cells (Figure 3C), and therefore, basal ERBB2 expression did not desensitise breast cancer cells to FUL.
CDK6 contributes to FUL resistance
The overexpression of CDK6 was confirmed by western blotting (Figure 4A). The expression of CDK4 was unchanged. The study of cell cycle protagonists such as CDK6 by siRNA approaches is difficult, because resulting transfectants cannot be expanded for subsequent analyses. Therefore, we chose a different approach to confirm or disregard whether CDK6 has a role in FUL resistance by analysing the phosphorylation status of serine 780 of retinoblastoma protein (RB1), which is the direct target of CDK6. Indeed, in FR.MCF-7 cells Ser780RB1 was constitutively phosphorylated (thereby inactivating RB1). As a consequence, the indirect downstream effector of inactivated RB1, CCNA1, was also overexpressed (Henglein et al, 1994). Fulvestrant treatment induced CDK6 in naive cells after 24 and 48 h, which remained constitutively high in FR.MCF-7 cells (Figure 4B). Interestingly, the expression of CCNE1 was reduced (Figure 4A) and CCND1 was also slightly downregulated in FR.MCF-7 cells (Table 1, Figure 4B). Thus, FR.MCF7 cells became independent of CCND1 and CCNE1 expression, which was surprising since CCND1 was shown to have a significant role in the development of breast cancer (Dean et al, 2010). Furthermore, FUL treatment induced p21, particularly in FR.MCF-7 cells, and this suggested that the resistant cells managed to bypass effectively the p21 cycle arrest signal (Figure 4B). The loss of CCND1 dependence was supported by the fact that the proliferation of FR.MCF-7 cells was significantly faster than that of naive MCF-7 cells (Figure 4C). Hence, the constitutive downregulation of CCND1 should not be considered as a cause, but rather as a consequence of resistance.
Effects of FUL and PD0332991 on cell cycle regulators. (A) MCF-7 and FR.MCF-7 cells were grown to 70% confluence, or (B) treated with 500 nM FUL or solvent (Co) for the indicated times when they were lysed and similar amounts of protein subjected to electrophoretic separation and western blot analysis with the indicated antibodies. Equal sample loading was confirmed by Ponceau S staining and α-tubulin analysis. (C) MCF-7 and FR.MCF-7 cells were grown for 24 and 48 h when cells were counted and the duplication time calculated. Experiments were carried out in triplicate, error bars indicate s.e.m. and asterisk denotes significance (t-test). (D) MCF-7 and FR.MCF-7 cells were exposed to 25 and 50 nM PD0332991 and cell proliferation was measured after 48 h. Experiments were carried out in triplicate, error bars indicate s.e.m. and asterisks denote significance (t-test). (E) MCF-7 and FR.MCF-7 cells were grown to 70% confluence and treated with 50 nM PD0332911 or solvent (Co) for the indicated times when they were lysed and similar amounts of protein subjected to electrophoretic separation and western blot analysis with the indicated antibodies. Equal sample loading was confirmed by Ponceau S staining and α-tubulin analysis.
The role of high CDK6 expression was tested by treating FR.MCF-7 cells and naive MCF-7 cells with the specific inhibitor PD0332991. The proliferation of FR.MCF-7 cells was significantly more attenuated by 50 nM PD0332991 than that of naive cells (Figure 4D). Rendering FR.MCF-7 cells more susceptible to PD0332991 strongly indicated that the rapid growth of these cells relied on the high expression of CDK6. The functionality of CDK6 in FR.MCF-7 cells was confirmed by the inhibited phosphorylation of Ser780RB1 upon PD0332991 treatment (Figure 4E). Therefore, CDK6 activity contributed to unrestricted cell growth, which was acquired during long-term treatment with FUL. The upregulation of CDK6 during PD0332991 treatment might have been a compensatory feedback mechanism to keep up CDK6 signalling.
Treatment with FUL modulates the expression of subunits of the hSWI/SNF chromatin remodelling complex
The catalytic subunit of hSWI/SNF (human SWItch/Sucrose NonFermentable), the ATPase BRG1, binds directly to RB1 (Strobeck et al, 2000; Zhang et al, 2000), and furthermore, BRG1 is strictly required to maintain ER function (Ichinose et al, 1997; Belandia et al, 2002; Inoue et al, 2002; Reisman et al, 2009). Hence, hSWI/SNF links the RB1 pathway to ER function and may have a role in the acquisition of FUL resistance, which was shown to involve RB1 signalling Thangavel et al, 2011).
Acetylases, deacetylases and the hSWI/SNF chromatin remodelling complex are tightly associated (Zhang et al, 2000; Naidu et al, 2009) with each other, and in dependence of the hSWI/SNF subunit composition, the gene expression pattern changes (Nagl et al, 2007; Jones et al, 2010). Therefore, the expression of constant (BAF47, BAF53A, BAF57, BAF155, BAF170) and of variable (BRG1, BAF250A) subunits of the hSWI/SNF chromatin remodelling complex was analysed. Upon FUL treatment, BAF250A, BAF155 and BAF47 became transiently upregulated and BGR1 and BAF170 downregulated in naive MCF-7 cells (Figure 5). This stress response to FUL was also observed for BRG1, BAF250A and BAF47 in FR.MCF-7 cells, although the response times were attenuated and weaker for BAF47 and accelerated for BRG1. In FR.MCF-7 BAF155 did not respond any longer to FUL and BAF170 became even induced, which was contrary to the response observed in naive MCF-7 cells (Figure 5). The expression of BAF53A and BAF57 remained unchanged in both cell lines. The constitutive upregulation of BAF250A protein in FR.MCF-7 cells was also reflected by the slight increase of the transcript, whereas constitutive upregulation of BAF155 and downregulation of BAF170 was likely due to post-transcriptional/post-translational events, because from the GeneChip data both genes were similarly expressed in naive and resistant cells (Table 1). The constitutive changes of gene expression as well as the aberrant stress response are attributes of the acquired resistance phenotype.
Effects of fulvestrant on SWI/SNF subunits and targets. MCF-7 and FR.MCF-7 cells were incubated with 500 nM FUL and harvested after 2, 8, 24 and 48 h of treatment. Cells were lysed, protein samples subjected to electrophoretic separation and to western blot analysis with the indicated antibodies. Equal sample loading was confirmed by Ponceau S staining and β-tubulin analysis.
The expression of MYC and p53 was shown to be under the control of the hSWI/SNF complex (Albert et al, 2001; Lee et al, 2003, 2005; Chung and Levens, 2005; Chen et al, 2006; Nagl et al, 2006, 2007; Simone, 2006; Sims et al, 2007; Naidu et al, 2009). Fulvestrant treatment downregulated the expression of MYC and p53 in naive MCF-7 cells, which correlated inversely with the expression of BAF47 (BAF47 is bona fide tumour suppressor).
The stress response of MYC and p53 to FUL was suspended in FR.MCF-7 cells and also BAF47 induction was weak and delayed. This indicated that hSWI/SNF regulated MYC and p53 expression in MCF-7 cells when treated with FUL, and that abrogation of MYC and p53 regulation was integral to acquired FUL resistance. Interestingly, the MYC transcript in FR.MCF-7 cells as one-third of that in naive MCF-7cells, yet the protein was almost similarly expressed in both cell lines.
In addition to the direct binding to target genes transient activation of hSWI/SNF was reported to permanently reposition nucleosomes (Schnitzler et al, 2001; Ulyanova and Schnitzler, 2005), leading to long-lasting changes in gene expression patterns. This was corroborated by the fact that FUL induced CDK6 in naive MCF-7 cells and that high CDK6 expression was maintained in FR.MCF-7 cells (Figure 4B). CDK4 expression was not induced by FUL and was similar in naive and FR.MCF-7 cells. Fulvestrant did not further induce CDK6 expression in FR.MCF-7 cells (Figure 4B) as it was already high and CDK6 levels correlated directly with the expression of BAF155 in both cell lines (Figure 5). Thus, affecting nucleosome control by FUL may have been the mechanism responsible for the endurance of CDK6 upregulation, even after withdrawal of FUL, and for acquired resistance.
Discussion
The aim of this investigation was to elucidate the cellular mechanisms that become involved during the acquisition of FUL resistance. Well-known protagonists mediating drug resistance in breast cancer cells, such as activation of ERBB2 (Hu and Mokbel, 2001; Chung et al, 2002; Shou et al, 2004; Gutierrez et al, 2005) or the overexpression of the breast cancer resistance protein BCRP/ABCG2 (Selever et al, 2011), did not seem to contribute to resistance in this model system nor was loss or inactivation of ERs causal for FUL resistance. However, repression of GPER, an endoplasmatic reticulum-bound receptor for oestrogen derivatives (Revankar et al, 2005), caused a significant decrease in the sensitivity to FUL. Hence, GPER partly mediates growth inhibition triggered by FUL. Also GREB1, an oestrogen-regulated gene, exhibited a similar property, although GREB1 did not contribute to resistance in the generated FR.MCF-7 cells.
Recently, it was reported that inactivated RB1 (through phosphorylation of Ser780) caused insensitivity to FUL in LCC9 breast cancer cells and that lack of signature RB1 target regulation is a hallmark of breast cancer cells with spontaneous and acquired resistance to SERM treatment (Lange and Yee, 2011; Thangavel et al, 2011). Disruption of RB1 control was due to maintenance of CCND1 expression, thereby keeping CDK activity high (Thangavel et al, 2011). However, the present study demonstrates that FR.MCF-7 cells became independent of CCND1, because they proliferated significantly faster than naive MCF-7 cells despite reduced CCND1 levels. Instead, CDK6, which is associated with CCND1, was overexpressed, thereby contributing to FUL resistance and CDK6-dependent cell growth made FR.MCF-7 cells more susceptible to the specific inhibitor PD0332991. Thus, resistance of hormone-sensitive breast cancer cells impinges on the RB1 pathway, yet the causal upstream players may vary (Thangavel et al, 2011). Notably, p21 induction by FUL could not arrest FR.MCF-7 cell proliferation underscoring how powerful CDK6-mediated resistance was. This recommends the reconstitution of the RB1 pathway as a subject to target tailored second-line adjuvant therapy. Twenty-one clinical trials testing PD0332991 against different cancer entities are currently recruiting, are active, or have been completed (following trials focus on breast cancer: ‘PD0332991/Paclitaxel in Advanced Breast Cancer’ – ClinicalTrials.gov Identifier: NCT01320592; ‘A Study Of PD-0332991 (Cyclin Dependent Kinase 4/6 Inhibitor) In Japanese Patients With Advanced Solid Tumors’ – ClinicalTrials.gov Identifier: NCT01684215 – both phase I studies, still recruiting; ‘PD 0332991 and Anastrozole for Stage 2 or 3 Estrogen Receptor Positive and HER2 Negative Breast Cancer’ – ClinicalTrials.gov Identifier: NCT01723774; ‘Letrozole and CDK 4/6 Inhibitor for ER Positive, HER2 Negative Breast Cancer in Postmenopausal Women’ – ClinicalTrials.gov Identifier: NCT01709370 – both phase II studies and still recruiting; ‘Study Of Letrozole With Or Without PD 0332991 For The First-Line Treatment Of Hormone-Receptor Positive Advanced Breast Cancer’ – ClinicalTrials.gov Identifier: NCT00721409, a phase I/II study and still active; ‘A Study of PD-0332991+Letrozole vs Letrozole For 1st Line Treatment Of Postmenopausal Women With ER+/HER2− Advanced Breast Cancer’ – ClinicalTrials.gov Identifier: NCT01740427, a phase III study and still recruiting).
The overexpression of CDK6 was reversed by TSA (but not AZA), indicating a functional involvement of nucleosome acetylation in the aberrant expression of CDK6 and in the acquisition of FUL resistance. Acquired TAM resistance was also shown to involve chromatin remodelling through nucleosome acetylation (Badia et al, 2000).
Another mechanism that contributed to FUL resistance was DNA methylation, as the downregulation of GPER was reversed by AZA (but not TSA). Nucleosome (histone) acetylation and DNA (CpG island) methylation requires the accessibility of acetyl transferases and methylases, respectively, to previously protected areas. The positioning of nucleosomes on the DNA (regulated by chromatin remodelling complexes) and the higher order structure of the histone octamer core (controlled by acetyl transferases and deacetylases) facilitate stochastic access for transcription factors to bind promoter regions, or methylases to switch off genes epigenetically. Acetyl transferases, histone deacetylases and chromatin remodelling complexes such as hSWI/SNF were shown to cooperate in this process (Zhang et al, 2000; Naidu et al, 2009). Human SWItch/Sucrose NonFermentable is involved in embryonic development, differentiation and cancer (Reisman et al, 2009) and alters nucleosome positioning (Flaus and Owen-Hughes, 2001; Sims et al, 2007, 2008). Human SWItch/Sucrose NonFermentable consists of ‘core’ and ‘variable’ subunits and changing their composition triggers the movement of nucleosomes from high-affinity (default) positions to different DNA regions (Nagl et al, 2007; Jones et al, 2010). Redistribution of histone octamers within the chromatin by hSWI/SNF causes deprotection or occlusion of DNA regions facilitating or preventing the access to DNA and giving rise to aberrant gene expression. Human SWItch/Sucrose NonFermentable was shown to regulate the expression of p21 and CCNA1 (Murphy et al, 1999) and the subunits BAF250A and BAF47, both described as bona fide tumour suppressors, downregulate MYC (Nagl et al, 2006, 2007) and CCND1 (Rao et al, 2008), as it was observed in our investigation. BAF250A-containing hSWI/SNF complexes repress E2F (Van Rechem et al, 2009) and E2F is required for the expression of MYC (Oswald et al, 1994). Here we show that FUL induced BAF47 and this correlated with MYC downregulation in naive MCF-7 cells, whereas this axis was disturbed in FR.MCF-7 cells.
The catalytic centre of hSWI/SNF, BRG1, is a transcriptional coactivator of p53 and of nuclear hormone receptors (Chen et al, 2006; Simone, 2006). Taken together with BAF170 and BAF155, BRG1 integrates signals of anti-oestrogens directly at the ER promoter (Zhang et al, 2000). This is in agreement with the reduced responsiveness of FR.MCF-7 cells to TAM, because in FR.MCF-7 cells the expression patterns of these subunits was tilted: BRG1 expression was less robust in FR.MCF-7 cells than in naive cells, and BAF170 was repressed and BAF155 overexpressed. Decreased BRG1 levels predispose mice to cancer formation (Klochendler-Yeivin et al, 2002; Simone, 2006), whereas another investigation demonstrates the capacity of BRG1 to induce a tumour-initiating cell phenotype (Okamoto et al, 2011). Therefore, the expression of BRG1 acts in different directions depending on the molecular context.
The composition of the hSWI/SNF subunits is subject to alterations and represents an own level of control to access DNA. Changes in the hSWI/SNF subunit constitution ATP dependently redistributes nucleosomes from default DNA positions to other nucleotide sequences within the chromatin (Schnitzler et al, 2001; Ulyanova and Schnitzler, 2005; Teif and Rippe, 2009) with consequences for chromatin structure and gene expression. Resumption to normal subunit composition may shuttle nucleosomes back to their default positions. This would explain the transient nature of MYC and p53 repression. However, based on the array data, it is more likely that FUL suppressed both genes at a post-transcriptional level. Even transient alterations in hSWI/SNF subunit composition were shown to provoke a steady-state redistribution of nucleosomes (Schnitzler et al, 2001; Ulyanova and Schnitzler, 2005) causing enduring changes in gene expression, that is, when hSWI/SNF subunits become directly affected. Notably, we observed stable overexpression of BAF250A and BAF155 and constitutive suppression of BAF170 in FR.MCF-7 cells and FUL treatment swiftly caused this induction/inhibition of BAF250A/BAF155/BAF170 (respectively) already in naive cells. Hence, the increase of constitutive CDK6 expression (at the transcriptional and translational level) can be explained as an immediate consequence of aberrant hSWI/SNF subunit expression and chromatin remodelling upon FUL treatment, which ultimately resulted in FUL resistance. It is further of note that the expression of BAF155 and CDK6 correlated in both cell strains. BRG1, containing the catalytic centre and being responsible for the general activity of hSWI/SNF, was only transiently downregulated by FUL treatment and remained expressed in FR.MCF-7 cells. Therefore, specifically and temporally inhibiting BRG1 activity throughout adjuvant therapy might prevent the redistribution of nucleosomes through inactivation of hSWI/SNF and possibly the development of a resistance phenotype.
Change history
12 November 2013
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Albert T, Wells J, Funk JO, Pullner A, Raschke EE, Stelzer G, Meisterernst M, Farnham PJ, Eick D (2001) The chromatin structure of the dual c-myc promoter P1/P2 is regulated by separate elements. J Biol Chem 276 (23): 20482–20490.
Badia E, Duchesne MJ, Semlali A, Fuentes M, Giamarchi C, Richard-Foy H, Nicolas JC, Pons M (2000) Long-term hydroxytamoxifen treatment of an MCF-7-derived breast cancer cell line irreversibly inhibits the expression of estrogenic genes through chromatin remodeling. Cancer Res 60 (15): 4130–4138.
Belandia B, Orford RL, Hurst HC, Parker MG (2002) Targeting of SWI/SNF chromatin remodelling complexes to estrogen-responsive genes. EMBO J 21 (15): 4094–4103.
Bolstad BM, Irizarry RA, Astrand M, Speed TP (2003) A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19 (2): 185–193.
Chang HG, Kim SJ, Chung KW, Noh DY, Kwon Y, Lee ES, Kang HS (2005) Tamoxifen-resistant breast cancers show less frequent methylation of the estrogen receptor beta but not the estrogen receptor alpha gene. J Mol Med (Berl) 83 (2): 132–139.
Chen J, Kinyamu HK, Archer TK (2006) Changes in attitude, changes in latitude: nuclear receptors remodeling chromatin to regulate transcription. Mol Endocrinol 20 (1): 1–13.
Chung HJ, Levens D (2005) c-Myc expression: keep the noise down!. Mol Cells 20 (2): 157–166.
Chung YL, Sheu ML, Yang SC, Lin CH, Yen SH (2002) Resistance to tamoxifen-induced apoptosis is associated with direct interaction between Her2/neu and cell membrane estrogen receptor in breast cancer. Int J Cancer 97 (3): 306–312.
Dean JL, Thangavel C, McClendon AK, Reed CA, Knudsen ES (2010) Therapeutic CDK4/6 inhibition in breast cancer: key mechanisms of response and failure. Oncogene 29 (28): 4018–4032.
Encarnación CA, Ciocca DR, McGuire WL, Clark GM, Fuqua SA, Osborne CK (1993) Measurement of steroid hormone receptors in breast cancer patients on tamoxifen. Breast Cancer Res Treat 26 (3): 237–246.
Fan M, Yan PS, Hartman-Frey C, Chen L, Paik H, Oyer SL, Salisbury JD, Cheng AS, Li L, Abbosh PH, Huang TH, Nephew KP (2006) Diverse gene expression and DNA methylation profiles correlate with differential adaptation of breast cancer cells to the antiestrogens tamoxifen and fulvestrant. Cancer Res 66 (24): 11954–11966.
Fiegl H, Millinger S, Goebel G, Muller-Holzner E, Marth C, Laird PW, Widschwendter M (2006) Breast cancer DNA methylation profiles in cancer cells and tumor stroma: association with HER-2/neu status in primary breast cancer. Cancer Res 66 (1): 29–33.
Flaus A, Owen-Hughes T (2001) Mechanisms for ATP-dependent chromatin remodelling. Curr Opin Genet Dev 11 (2): 148–154.
Ghayad SE, Vendrell JA, Ben Larbi S, Dumontet C, Bieche I, Cohen PA (2010) Endocrine resistance associated with activated ErbB system in breast cancer cells is reversed by inhibiting MAPK or PI3K/Akt signaling pathways. Int J Cancer 126 (2): 545–562.
Giessrigl B, Krieger S, Rosner M, Huttary N, Saiko P, Alami M, Messaoudi S, Peyrat JF, Maciuk A, Gollinger M, Kopf S, Kazlauskas E, Mazal P, Szekeres T, Hengstschläger M, Matulis D, Jäger W, Krupitza G (2012) Hsp90 stabilizes Cdc25A and counteracts heat shock-mediated Cdc25A degradation and cell-cycle attenuation in pancreatic carcinoma cells. Hum Mol Genet 21 (21): 4615–4627.
Gutierrez MC, Detre S, Johnston S, Mohsin SK, Shou J, Allred DC, Schiff R, Osborne CK, Dowsett M (2005) Molecular changes in tamoxifen-resistant breast cancer: relationship between estrogen receptor, HER-2, and p38 mitogen-activated protein kinase. J Clin Oncol 23 (11): 2469–2476.
Henglein B, Chenivesse X, Wang J, Eick D, Bréchot C (1994) Structure and cell cycle-regulated transcription of the human cyclin A gene. Proc Natl Acad Sci USA 91 (12): 5490–5494.
Hu JC, Mokbel K (2001) Does c-erbB2/HER2 overexpression predict adjuvant tamoxifen failure in patients with early breast cancer? Eur J Surg Oncol 27 (4): 335–337.
Ichinose H, Garnier JM, Chambon P, Losson R (1997) Ligand-dependent interaction between the estrogen receptor and the human homologues of SWI2/SNF2. Gene 188 (1): 95–100.
Inoue H, Furukawa T, Giannakopoulos S, Zhou S, King DS, Tanese N (2002) Largest subunits of the human SWI/SNF chromatin-remodeling complex promote transcriptional activation by steroid hormone receptors. J Biol Chem 277 (44): 41674–41685.
Jensen EV, Jordan VC (2003) The estrogen receptor: a model for molecular medicine. Clin Cancer Res 9 (6): 1980–1989.
Jones S, Wang TL, IeM Shih, Mao TL, Nakayama K, Roden R, Glas R, Slamon D, Diaz LA Jr, Vogelstein B, Kinzler KW, Velculescu VE, Papadopoulos N (2010) Frequent mutations of chromatin remodeling gene ARID1A in ovarian clear cell carcinoma. Science 330 (6001): 228–231.
Klochendler-Yeivin A, Muchardt C, Yaniv M (2002) SWI/SNF chromatin remodeling and cancer. Curr Opin Genet Dev 12 (1): 73–79.
Lange CA, Yee D (2011) Killing the second messenger: targeting loss of cell cycle control in endocrine-resistant breast cancer. Endocr Relat Cancer 18 (4): C19–C24.
Lee JH, Lee JY, Chang SH, Kang MJ, Kwon H (2005) Effects of Ser2 and Tyr6 mutants of BAF53 on cell growth and p53-dependent transcription. Mol Cells 19 (2): 289–293.
Lee JH, Chang SH, Shim JH, Lee JY, Yoshida M, Kwon H (2003) Cytoplasmic localization and nucleo-cytoplasmic shuttling of BAF53, a component of chromatin-modifying complexes. Mol Cells 16 (1): 78–83.
Maier S, Strasser S, Saiko P, Leisser C, Sasgary S, Grusch M, Madlener S, Bader Y, Hartmann J, Schott H, Mader RM, Szekeres T, Fritzer-Szekeres M, Krupitza G (2006) Analysis of mechanisms contributing to AraC-mediated chemoresistance and re-establishment of drug sensitivity by the novel heterodinucleoside phosphate 5-FdUrd-araC. Apoptosis 11 (3): 427–440.
Miyoshi Y, Murase K, Saito M, Oh K (2010) Prediction of hormone sensitivity for breast cancers. Breast Cancer 17 (2): 86–91.
Murphy DJ, Hardy S, Engel DA (1999) Human SWI-SNF component BRG1 represses transcription of the c-fos gene. Mol Cell Biol 19 (4): 2724–2733.
Nagl NG Jr, Zweitzig DR, Thimmapaya B, Beck GR Jr, Moran E (2006) The c-myc gene is a direct target of mammalian SWI/SNF-related complexes during differentiation-associated cell cycle arrest. Cancer Res 66 (3): 1289–1293.
Nagl NG Jr, Wang X, Patsialou A, Van Scoy M, Moran E (2007) Distinct mammalian SWI/SNF chromatin remodeling complexes with opposing roles in cell-cycle control. EMBO J 26 (3): 752–763.
Naidu SR, Love IM, Imbalzano AN, Grossman SR, Androphy EJ (2009) The SWI/SNF chromatin remodeling subunit BRG1 is a critical regulator of p53 necessary for proliferation of malignant cells. Oncogene 28 (27): 2492–2501.
Okamoto N, Yasukawa M, Nguyen C, Kasim V, Maida Y, Possemato R, Shibata T, Ligon KL, Fukami K, Hahn WC, Masutomi K (2011) Maintenance of tumor initiating cells of defined genetic composition by nucleostemin. Proc Natl Acad Sci USA 108 (51): 20388–20393.
Oswald F, Lovec H, Möröy T, Lipp M (1994) E2F-dependent regulation of human MYC: trans-activation by cyclins D1 and A overrides tumour suppressor protein functions. Oncogene 9 (7): 2029–2036.
Rao M, Casimiro MC, Lisanti MP, D'Amico M, Wang C, Shirley LA, Leader JE, Liu M, Stallcup M, Engel DA, Murphy DJ, Pestell RG (2008) Inhibition of cyclin D1 gene transcription by Brg-1. Cell Cycle 7 (5): 647–655.
Reisman D, Glaros S, Thompson EA (2009) The SWI/SNF complex and cancer. Oncogene 28 (14): 1653–1668.
Revankar CM, Cimino DF, Sklar LA, Arterburn JB, Prossnitz ER (2005) A transmembrane intracellular estrogen receptor mediates rapid cell signaling. Science 307 (5715): 1625–1630.
Sabnis GJ, Jelovac D, Long B, Brodie A (2005) The role of growth factor receptor pathways in human breast cancer cells adapted to long-term estrogen deprivation. Cancer Res 65 (9): 3903–3910.
Schiff R, Massarweh SA, Shou J, Bharwani L, Mohsin SK, Osborne CK (2004) Cross-talk between estrogen receptor and growth factor pathways as a molecular target for overcoming endocrine resistance. Clin Cancer Res 10 (1, Part 2): 331S–336S.
Schnitzler GR, Cheung CL, Hafner JH, Saurin AJ, Kingston RE, Lieber CM (2001) Direct imaging of human SWI/SNF-remodeled mono- and polynucleosomes by atomic force microscopy employing carbon nanotube tips. Mol Cell Biol 21 (24): 8504–8511.
Selever J, Gu G, Lewis MT, Beyer A, Herynk MH, Covington KR, Tsimelzon A, Dontu G, Provost P, Di Pietro A, Boumendjel A, Albain K, Miele L, Weiss H, Barone I, Ando S, Fuqua SA (2011) Dicer-mediated upregulation of BCRP confers tamoxifen resistance in human breast cancer cells. Clin Cancer Res 17 (20): 6510–6521.
Shaw LE, Sadler AJ, Pugazhendhi D, Darbre PD (2006) Changes in oestrogen receptor-alpha and -beta during progression to acquired resistance to tamoxifen and fulvestrant (Faslodex, ICI 182,780) in MCF7 human breast cancer cells. J Steroid Biochem Mol Biol 99 (1): 19–32.
Shou J, Massarweh S, Osborne CK, Wakeling AE, Ali S, Weiss H, Schiff R (2004) Mechanisms of tamoxifen resistance: increased estrogen receptor-HER2/neu cross-talk in ER/HER2-positive breast cancer. J Natl Cancer Inst 96 (12): 926–935.
Simone C (2006) SWI/SNF: the crossroads where extracellular signaling pathways meet chromatin. J Cell Physiol 207 (2): 309–314.
Sims HI, Baughman CB, Schnitzler GR (2008) Human SWI/SNF directs sequence-specific chromatin changes on promoter polynucleosomes. Nucleic Acids Res 36 (19): 6118–6131.
Sims HI, Lane JM, Ulyanova NP, Schnitzler GR (2007) Human SWI/SNF drives sequence-directed repositioning of nucleosomes on C-myc promoter DNA minicircles. Biochemistry 46 (40): 11377–11388.
Speirs V, Malone C, Walton DS, Kerin MJ, Atkin SL (1999) Increased expression of estrogen receptor beta mRNA in tamoxifen-resistant breast cancer patients. Cancer Res 59 (21): 5421–5424.
Stone A, Valdés-Mora F, Gee JM, Farrow L, McClelland RA, Fiegl H, Dutkowski C, McCloy RA, Sutherland RL, Musgrove EA, Nicholson RI (2012) Tamoxifen-induced epigenetic silencing of oestrogen-regulated genes in anti-hormone resistant breast cancer. PLoS One 7 (7): e40466.
Strasser S, Maier S, Leisser C, Saiko P, Madlener S, Bader Y, Bernhaus A, Gueorguieva M, Richter S, Mader RM, Wesierska-Gadek J, Schott H, Szekeres T, Fritzer-Szekeres M, Krupitza G (2006) 5-FdUrd-araC heterodinucleoside re-establishes sensitivity in 5-FdUrd- and AraC-resistant MCF-7 breast cancer cells overexpressing ErbB2. Differentiation 74 (9–10): 488–498.
Strobeck MW, Knudsen KE, Fribourg AF, DeCristofaro MF, Weissman BE, Imbalzano AN, Knudsen ES (2000) BRG-1 is required for RB-mediated cell cycle arrest. Proc Natl Acad Sci USA 97 (14): 7748–7753.
Teif VB, Rippe K (2009) Predicting nucleosome positions on the DNA: combining intrinsic sequence preferences and remodeler activities. Nucleic Acids Res 37 (17): 5641–5655.
Thangavel C, Dean JL, Ertel A, Knudsen KE, Aldaz CM, Witkiewicz AK, Clarke R, Knudsen ES (2011) Therapeutically activating RB: reestablishing cell cycle control in endocrine therapy-resistant breast cancer. Endocr Relat Cancer 18 (3): 333–345.
Ulyanova NP, Schnitzler GR (2005) Human SWI/SNF generates abundant, structurally altered dinucleosomes on polynucleosomal templates. Mol Cell Biol 25 (24): 11156–11170.
van Agthoven T, van Agthoven TLA, Dekker A, van der Spek PJ, Vreede L, Dorssers LCJ (1998) Identification of BCAR3 by a random search for genes involved in antiestrogen resistance of human breast cancer cells. EMBO J 17 (10): 2799–2808.
van der Flier S, Brinkman A, Look MP, Kok EM, Meijer-van Gelder ME, Klijn JGM, Dorssers LCJ, Foekens JA (2000) Bcar1/p130Cas protein and primary breast cancer: prognosis and response to tamoxifen treatment. J Nat Cancer Inst 92 (2): 120–127.
Van Rechem C, Boulay G, Leprince D (2009) HIC1 interacts with a specific subunit of SWI/SNF complexes, ARID1A/BAF250A. Biochem Biophys Res Commun 385 (4): 586–590.
Vesuna F, Lisok A, Kimble B, Domek J, Kato Y, van der Groep P, Artemov D, Kowalski J, Carraway H, van Diest P, Raman V (2012) Twist contributes to hormone resistance in breast cancer by downregulating estrogen receptor-α. Oncogene 31 (27): 3223–3234.
Zhang HS, Gavin M, Dahiya A, Postigo AA, Ma D, Luo RX, Harbour JW, Dean DC (2000) Exit from G1 and S phase of the cell cycle is regulated by repressor complexes containing HDAC-Rb-hSWI/SNF and Rb-hSWI/SNF. Cell 101 (1): 79–89.
Acknowledgements
We thank Toni Jäger for preparing the figures. The work was supported by the Herzfelder Family Foundation to GK.
Author information
Authors and Affiliations
Corresponding author
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License.
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Giessrigl, B., Schmidt, W., Kalipciyan, M. et al. Fulvestrant induces resistance by modulating GPER and CDK6 expression: implication of methyltransferases, deacetylases and the hSWI/SNF chromatin remodelling complex. Br J Cancer 109, 2751–2762 (2013). https://doi.org/10.1038/bjc.2013.583
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2013.583
Keywords
This article is cited by
-
ERα-LBD, an isoform of estrogen receptor alpha, promotes breast cancer proliferation and endocrine resistance
npj Breast Cancer (2022)
-
Distinct mechanisms of resistance to fulvestrant treatment dictate level of ER independence and selective response to CDK inhibitors in metastatic breast cancer
Breast Cancer Research (2021)