Abstract
Background:
Eukaryotic translation elongation factor 1A2 (eEF1A2) is a known proto-oncogene. We proposed that stimulation of the eEF1A2 expression in cancer tissues is caused by the loss of miRNA-mediated control.
Methods:
Impact of miRNAs on eEF1A2 at the mRNA and protein levels was examined by qPCR and western blot, respectively. Dual-luciferase assay was applied to examine the influence of miRNAs on 3′-UTR of EEF1A2. To detect miRNA-binding sites, mutations into the 3′-UTR of EEF1A2 mRNA were introduced by the overlap extension PCR.
Results:
miR-663 and miR-744 inhibited the expression of luciferase gene attached to the 3′-UTR of EEF1A2 up to 20% and 50%, respectively. In MCF7 cells, overexpression of miR-663 and miR-744 reduced the EEF1A2 mRNA level by 30% and 50%. Analogous effects were also observed at the eEF1A2 protein level. In resveratrol-treated MCF7 cells the upregulation of mir-663 and mir-744 was accompanied by downregulation of EEF1A2 mRNA. Both miRNAs were able to inhibit the proliferation of MCF7 cells.
Conclusion:
miR-663 and miR-744 mediate inhibition of the proto-oncogene eEF1A2 expression that results in retardation of the MCF7 cancer cells proliferation. Antitumour effect of resveratrol may include stimulation of the miR-663 and miR-744 expression.
Similar content being viewed by others
Main
Eukaryotic translation elongation factor 1A (eEF1A), one of the key players in protein biosynthesis, binds aminoacyl-tRNA and transfers it to the A-site of the ribosome. Human eEF1A exists as two isoforms: eEF1A1, its gene located on 6q13, and eEF1A2, positioned on 20q13 (Knudsen et al, 1993). The two proteins are 92% identical and 98% similar. Both proteins are tissue- and development-specific. Eukaryotic translation elongation factor 1A2 performs nascent polypeptide elongation on the 80S ribosome in neuronal, muscle and cardiac tissues, being the only representative of the elongation factor 1 family (Lee et al, 1992). Other tissues of higher vertebrates employ eEF1A1 for this purpose. Developmentally, the A1 isoform is replaced by A2 in muscles and neurons during the early postnatal period (Pan et al, 2004). Owing to the mutually exclusive tissue-specific character of their expression, the isoforms are rarely found expressed together in the tissues under normal conditions. The mechanisms downregulating A1 or A2 expression are unknown. However, appreciable simultaneous expression of the two isoforms was found in extreme situations, such as muscle injury (Bosutti et al, 2007) or cancerogenesis (Tomlinson et al, 2007), demonstrating a possibility of reversing tissue-specific character of the A1/A2 expression.
The proto-oncogenic nature of eEF1A2 was discovered not long ago in human ovarian cancer (Anand et al, 2002). Since then, the involvement of eEF1A2 in breast, pancreatic and hepatic cancer progression has been detected (Tomlinson et al, 2005, 2007; Pinke et al, 2008; Cao et al, 2009; Lee and Surh, 2009; Li et al, 2010). Importantly, the overexpression of eEF1A2 in tumour tissues was found not to be a consequence of any multiplication of gene loci that might arise from chromosome instability, or from genetic and/or epigenetic changes (Tomlinson et al, 2005, 2007), thus suggesting the possibility of the regulation occurring at the post-transcriptional level. We hypothesised microRNA involvement in the regulation of A1/A2 expression.
MicroRNAs are widely involved in the post-transcriptional regulation of gene expression in both animals and plants (Lagos-Quintana et al, 2001; Reinhart et al, 2002). In mammals, miRNAs are considered to control ∼60% of the protein coding genes (Friedman et al, 2009), participating in the regulation of almost every cellular process, including cancerogenesis (Ambros, 2004). In this paper, we describe the involvement of microRNAs in the regulation of A2, the proto-oncogenic isoform of eEF1A. Our bioinformatic search highlighted several miRNA-binding sites in the 3′-UTR of EEF1A2 mRNA, particularly miR-663 and miR-744.
The existence of miR-744 was initially predicted by Berezikov et al (2006) and later confirmed by Landgraf et al (2007). However, the literature data about the function of this miRNA appeared only in the last few years. It was shown that miR-744 directly targets transforming growth factor beta-1 (TGF-β1) (Martin et al, 2011). MiR-744 is regarded as a potential oncomarker, as it was found to be stably present in mouse serum (Mi et al, 2012) and was shown to be significantly upregulated in the serum of patients with gastric cancer (Song et al, 2012).
Much more is known about miR-663, which is shown to be involved in cancer progression and to have tumour-suppressor properties. Thus, the miR-663 induces mitotic catastrophe growth arrest in human gastric cancer cells (Pan et al, 2010). Also, miR-663 stimulates differentiation of the acute myeloid leukaemia cell line HL-60, and has been proposed to be potentially useable for the anticancer treatment of haematological malignancies (Jian et al, 2011). In patients with lung cancer, miR-663 was found to be dramatically upregulated and to control TGF-β1, P53, Bax and Fas expression (Lewis et al, 2005).
Several articles have been dedicated to the role of miR-663 in the mechanism of resveratrol action. Resveratrol is a natural polyphenol that accumulates in grapes. Recently it was shown that it possess proapoptotic, antiproliferative and anti-inflammatory effects ((Jang et al, 1997; Vang et al, 2011) and was reviewed in Tili and Michaille, 2011). Importantly, Lee et al (2009) reported that resveratrol pretreatment inhibited the induction of eEF1A2 expression in insulin-stimulated PA-1 cells. Also, the treatment with resveratrol inhibited insulin- or serum-induced soft-agar colony formation in the eEF1A2-transfected NIH3T3 cells.
In this article, we show that miR-663 and miR-744 directly target the expression of EEF1A2 mRNA and decrease the amount of the corresponding protein. Moreover, downregulation of eEF1A2 by these miRNAs resulted in the inhibition of MCF7 proliferation. Thus, the unusual appearance of the eEF1A2 isoform observed in the tumour tissues but not in the normal ones may be caused by impairment of the post-transcriptional control of its expression by microRNAs.
We have also found that resveratrol-mediated reduction of the eEF1A2 level in MCF7 cells is caused by the increased expression of miR-663 and miR-744 miRNAs. This suggests that the downregulation of the expression of the putative oncogene eEF1A2 via microRNA-mediated pathways may be one of the mechanisms responsible for the inhibitory effect of resveratrol on the breast cancer progression.
Materials and methods
Cell lines, plasmids, antibodies and resveratrol treatment
T47D, LNCaP and DU145 cells were cultured in RPMI 1640 (Sigma, St Louis, MO, USA), while A549, HEK293 and MCF7 cells were cultured in DMEM (Sigma) growth medium. Both media contained 10% FBS (Sigma) and 1% penicillin/streptomycin (Sigma). Cells were maintained at 37 °C in a humidified atmosphere containing 5% CO2.
The 3′-UTR of the EEF1A2 mRNA was cloned from human leukocyte DNA and inserted into the pSICHECK-2 reporter vector (Promega, Madison, WI, USA). A pCDNA3.1 vector expressing the eEF1A2 ORF was a kind gift of Charlotte R Knudsen (Department of Molecular Biology, University of Aarhus, Gustav Wieds vej 10 C, 8000 Århus C, Denmark).
Antibeta-actin antibodies were from Santa Cruz (Santa Cruz Biotechnologies, Santa Cruz, CA, USA). In-house anti-eEF1A2 polyclonal antibodies were produced as described (Kolesanova et al, 2013).
For resveratrol treatment, 100 μ M resveratrol (in DMSO) (Sigma) was added to the MCF7 cells. After 24 h of incubation, cells were collected, and total RNA was extracted using TRI reagent (Sigma).
Western blot analyses
Total cell lysates were prepared in RIPA lysis buffer supplemented with a protease inhibitor cocktail (Roche, Bazel, Switzerland). Cells were incubated on a rotating wheel at +4 °C for 20 min. Lysates were centrifuged and supernatant was stored at −80 °C. Proteins were separated on a NuPAGE Novex 4–12% Bis-Tris Gel (Invitrogen, Carlsbad, CA, USA) and transferred to nitrocellulose membranes (Millipore, Billerica, MA, USA) in NuPAGE Transfer Buffer (Invitrogen). Membranes were developed using Super Signal West Dura Extended Duration Substrate (Pierce, Rockford, lL, USA).
Mutagenesis
Site-specific mutagenesis of miRNA-binding sites was performed by Overlap Extension PCR (Higuchi et al, 1988). In each case, a restriction site was inserted instead of a miRNA-binding site. Primers used for PCR were: universal forward primer 5′-GCTCTAGAGCCCGCGGCGCGACCCTCCC-3′ universal reverse primer 5′-GCTCTAGAGAGCGTGGCGAGCGCTGGGC-3′; E663 forward 5′-GGGCCCGGGCAAAATAAAATAAAGATATCCGCGCGCCG-3′, E663 reverse 5′-CGGCGCGCGGATATCTTTATTTTATTTTGCCCGGGCCC-3′; H663 forward 5′-CGCCCCGCACAAAGCTTAGGCGCATGT-3′, H663 reverse 5′-ACATGCGCCTAAGCTTTGTGCGGGGCG-3′; K663 forward 5′-GGAAAGGCGCAAAATGGTACCGCTTCCGCGC-3′, K663 reverse 5′-GCGCGGAAGCGGTACCATTTTGCGCCTTTCC-3′; H744 forward 5′-CCGGTCCGGCATAAGCTTCCCCCGCCA-3′, H744 reverse 5′-TGGCGGGGGAAGCTTATGCCGGACCGG-3′.
Reporter gene assay
The day before the experiment, log-phase cells were seeded into 96-well plates at a concentration calculated to reach 80% confluence at the time of experiment. The next day, the cells were transfected with 30 nM miRNA precursors (final concentration in the well) and 25 ng of reporter plasmid using Lipofectamine-2000 (Invitrogen) in accordance to the company protocol. Luciferase was quantified 24 h later, using a Dual-Luciferase reporter assay system (Promega) and a TriStar LB 941 reader (Berthold Technologies, Bad Wildbad, Germany).
Quantitative PCR
Total RNA was isolated using TRI reagent (Sigma). One microgram of RNA was used for cDNA synthesis with a High-Capacity cDNA Reverse Transcription Kit (Applied Biosystems, Foster City, CA, USA). Each reaction was performed in a mix of 20 μl reaction mixture containing 1 μl cDNA, 2 × SYBR Green PCR Master Mix (Applied Biosystems) and 0.3 μ M of each forward and reverse primer. For miRNA quantification, total RNA was transcribed with a TaqMan Reverse transcription kit (Applied Biosystems), and then amplified using TaqMan Universal PCR Master Mix II. The primers for eEF1A2 qPCR were described earlier (Bosutti et al, 2007). Primers for beta-actin were: forward primer 5′-GCGGGAAATCGTGCGTGACATT-3′, reverse primer 5′- GATGGAGTTGAAGGTAGTTTCGTG-3′. MicroRNA primers were purchased from Applied Biosystems. Quantitative PCR was quantified with an ABI PRISM 7500 real-time PCR system (Applied Biosystems). Data were analysed using qPCR Miner 4.0 software (Zhao and Fernald, 2005).
Proliferation assay
On the day before the experiment, 3000 MCF7 cells per well were seeded into 96-well plates. The next day, cells were transfected with 15 nM miRNA precursors or siRNA against eEF1A2 using RNAiMAX (Invitrogen) by standard protocol according to manual. 3000 A549 cells were transfected with 30 nM miRNA precursors or siRNA against eEF1A2 by reverse transfection protocol. After 72 h of incubation, live cells were stained by propidium iodide and quantified. Then, the cells were fixed with paraformaldehyde and stained with DAPI. The number of cells in each well was calculated using the Operetta High Content Imaging System (Perkin Elmer, Waltham, MA, USA).
Results
Identification of miRNAs that potentially target the EEF1A2 mRNA
Several algorithms that take into account conservation of the seed site, distribution through the 3′-UTR, and antisense RNA-binding site accessibility, were applied to identify miRNA that could potentially target the EEF1A2 mRNA. We used TargetScan 5.2, PITA, miRanda and microT v.3.0 programs to find miRNA-seeding sites in the 3′-UTR of EEF1A2 mRNA (Lewis et al, 2005; Kertesz et al, 2007; Betel et al, 2008; Maragkakis et al, 2009). Different algorithms returned different sets of potential miRNA seed sites. However, the most common and the highest scoring microRNAs were: miR-661, miR-663, miR-675 and miR-744 (Figure 1A).
Validation of miRNAs predicted to target the eEF1A2 3′-UTR. (A) Schematic representation of the predicted miRNA seed sites in the eEF1A2 3′-UTR. (B) miRNA-mediated knockdown of the wild-type eEF1A2 3′-UTR reporter in HeLa cells co-transfected with the indicated miRNA precursors. Transfection of scrambled miRNA precursor (Scr.) was used as control. Error bars represent s.d. values. *P<0.05 vs control. n=3. Student’s t-test.
To verify which of predicted miRNAs can regulate eEF1A2 expression, a dual-luciferase assay based on the firefly luciferase reporter gene attached to the EEF1A2 3′-UTR was applied.
Reporter DNA construct was co-transfected with miRNA precursor to the HeLa cells. As shown in Figure 1B, only miR-663 and miR-744 were able to significantly reduce the reporter activity. The effect of co-transfection with miR-663 and miR-744 was similar to the effect of miR-744 alone in luciferase test, suggesting the absence of cooperative action of these miRNAs on EEF1A2 mRNA.
Determination of actual miRNA-binding sites in the 3′-UTR of eEF1A2 mRNA
TargetScan 5.2 predicted multiple sites for both miR-663 and miR-744 in the 3′-UTR of EEF1A2 mRNA (Figure 2A). Consequently, to distinguish between true and false positive sites, we mutated predicted target sites by replacing them with the HindIII, EcoRV or KpnI restriction enzyme sites. As shown in Figure 2B, mutations in the region that contains three miR-663 and two overlapping miR-744 sites (E663) did not influence the reporter gene expression. However, a mutation of each of the two downstream miR-663-binding sites (H663 and K663) resulted in the upregulation of luciferase expression. An increase in the reporter gene expression was also observed when a mutation was introduced in the miR-744-binding site (H744). The results suggest that the EEF1A2 mRNA has two binding sites for miR-663, one of which was predicted as being conserved, and one binding site for miR-744.
Detection of the genuine miRNA-663 and miRNA-744-binding sites in the 3′-UTR of eEF1A2. (A) Scheme representing the mutations of the mir-663 and mir-744 seed sequences in the eEF1A2 3′-UTR miRNA. Mutated regions are shown by rectangles. Names of mutations are in boldface type. (B) Mutations of the mir-663 and mir-744-binding sites increase eEF1A2 3′-UTR reporter activity. MCF7 cells were transfected with wild-type or mutant eEF1A2 reporter constructs, and luciferase activity measured 24 h later. Transfections of MCF7 cells with non-modified pSICheck-2 vector or wild-type eEF1A2 3′-UTR reporter were used as controls. Error bars represent s.d. values. n=3. *P<0.05 vs control. n=3. Student’s t-test.
miR-663 and miR-744 miRNAs decrease eEF1A2 expression at both mRNA and protein levels
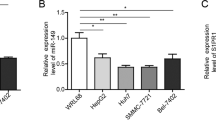
In order to choose a model cell line for our study, we evaluated the expression levels of EEF1A2 mRNA in different cell lines where it had been detected by earlier investigations (Tomlinson et al, 2005). Among those tested, the MCF7 breast cancer epithelial cell line was found to express the greatest amount of eEF1A2 (Figure 3A). Importantly, miR-663 and miR-744 expression was also detected in the MCF7 cell line (Figure 3B).
Transfections of MCF7 cells with miR-663 and miR-744 downregulated EEF1A2 mRNA levels by 30% and 50%, respectively (Figure 4A). This treatment also led to significant decreases in endogenous eEF1A2 protein level (Figure 4B). Analogous effects were observed when the cells were transfected with siRNA against EEF1A2 mRNA (Figures 4A and C).
Repression of eEF1A2 by mir-663, mir-744 or anti-eEF1A2 siRNA. (A) Downregulation of endogenous eEF1A2 transcript by mir-663 and mir-744. MCF7 cells were transfected with miRNA precursors, and the EEF1A2 mRNA level measured 24 h later by qPCR. Error bars represent s.d. values. *P<0.05 vs control. n=3. Student’s t-test. (B) Reduction of the endogenous eEF1A2 protein level by mir-663 and mir-744. (C) Reduction of the endogenous eEF1A2 protein level by siRNA. miRNA or siRNA precursors were transfected into MCF7 cells. Transfection of scrambled miRNA precursor (Scr.) or no-target siRNA (si0) was used as controls. Numbers show the quantification of the eEF1A2 bands intensity (%) with respect to actin.
Effect of miRNA-mediated eEF1A2 downregulation on MCF7 cell proliferation and migration
Previous data showed the possibility of the eEF1A2 input to the cell migration and proliferation processes (Cao et al, 2009; Li et al, 2010). Therefore, it is important to elucidate whether miRNA-dependent downregulation of eEF1A2 may have an impact on these parameters in cancer cells.
Indeed, overexpression of miR-663 and miR-744 downregulated the proliferation of MCF7 cells (Figure 5A). Importantly, treatment of MCF7 cells with the anti-eEF1A2 siRNA resulted in a similar effect on the MCF7 cell proliferation (Figure 5A). To prove that the observed effect of microRNAs on cell proliferation is mediated by eEF1A2, we overexpressed the eEF1A2 ORF before the miRNA transfection. The overexpression of eEF1A2 abolished the antiproliferative effect of the miRNAs (Figure 5B). These data suggest that the EEF1A2 mRNA is one of the targets of miR-663 and miR-744 responsible for the effect on MCF7 cell proliferation. The same trend of proliferation inhibition was observed for lung cancer A549 cells transfected with miRNAs 663, 744 and siRNA against eEF1A2 (Supplementary Figure 1A). Importantly, contribution of cell death into the inhibitory effects observed was rather small (Supplementary Figures 1B and C).
Effect of miRNA-mediated eEF1A2 downregulation on proliferation of MCF7 cells. (A) MCF7 cells were treated with mir-663, mir-744 precursor or siRNA against eEF1A2. (B) Cells were co-transfected with miRNA precursors and a vector expressing the eEF1A2 ORF. After 72 h, the cells were fixed, stained and counted using the OPERETTA High Content Imaging System. Transfections of scrambled miRNA precursor (Scr.) or no-target siRNA (Si0) were used as controls. Error bars represent s.d. values. *P<0.05 vs control. n=3. Student’s t-test.
The effect of miRNA-dependent eEF1A2 downregulation on cell migration was examined in a wound-healing assay. Interestingly, despite the intense inhibition of cell migration by the miR-663 and miR-744 treatment, we did not observe any changes in cell migration when cells were treated with siRNA against eEF1A2 (Supplementary Figure 2). Perhaps, eEF1A2 is not involved in the cell migration process, in which case the effects of miR-663 and miR-744 on cell migration would have to be mediated by other targets, likely it may be TGF-β1 (Gordon and Blobe, 2008; Chow et al, 2011; Imamura et al, 2012).
Resveratrol downregulates the eEF1A2 level through the miRNA pathway
Earlier, it was reported that pretreatment with resveratrol inhibited the induction of eEF1A2 expression in insulin-stimulated PA-1 cells. The treatment with resveratrol also inhibited both insulin- and serum-induced soft-agar colony formations in the eEF1A2-transfected NIH3T3 cells (Lee et al, 2009). On the other hand, Tili et al (2010a) have found upregulation of miR-663 in the resveratrol-treated THP-1 monocytic cells.
We hypothesise that the inhibition of eEF1A2 expression by resveratrol can be mediated by upregulation of miR-663 and miR-744. Indeed, the resveratrol treatment causes 4.5-fold upregulation of miR-663 and two-fold upregulation of miR-744 (Figure 6A). Importantly, the resveratrol treatment also leads to a decrease in the EEF1A2 mRNA level (Figure 6B). To substantiate the specificity of the effect, MCF7 cells were transfected with the miR-744 and miR-663 inhibitors before the resveratrol treatment. Under those conditions, the resveratrol had no effect on the EEF1A2 mRNA level (Figure 6C). As expected, the inhibitors of miR-663 and miR-744 upregulated the EEF1A2 mRNA level, contrary to the irrelative miRNA inhibitors. Thus, resveratrol negatively influences EEF1A2 mRNA post-transcriptionally, via miR-744 and miR-663.
Effect of resveratrol on EEF1A2 mRNA and mir-663/744 expression level. MCF7 cells were treated with 100 μ M of resveratrol. After 24 h of incubation, levels of microRNAs (A) and eEF1A2 (B) were assayed by qPCR. Treatment of MCF7 cells with DMSO was used as control. (C) Before the resveratrol treatment, MCF7 cells were transfected with mir-663 and mir-744 inhibitors. Transfection with irrelevant (irr.) anti-mir, was used as control. Error bars represent s.d. values. *P<0.05 vs control. n=3. Student’s t-test.
Discussion
As cancer can progress in multiple ways, one of the main aims of cancer research nowadays is to identify every possible mechanism involved in cancer induction/progression, in order to develop specific, low toxicity therapy for each specific case (De Palma and Hanahan, 2012). Here, we describe a mechanism of eEF1A2 regulation by two miRNA, miR-663 and miR-744, the loss of which can result in the development of cancer.
MiR-663 and, miR-744 are well-known oncosuppressor miRNAs. MicroRNA-744 amount was found to be significantly upregulated in the serum of patients with gastric cancer (Song et al, 2012). There are only two confirmed targets of this miRNA: the oncogene TGF-β, negative regulation, (Martin et al, 2011) and cyclin B1, positive regulation (Huang et al, 2012). In contrast, miR-663 was found to participate in a large number of cellular processes, including tumour genesis. Peculiarly, miR-663 is involved in mitotic catastrophe growth arrest in human gastric cancer cells (Pan et al, 2010), the same type of cancer in which miR-744 is involved. Recently, it has been proposed that miR-744 could be an appropriate target for anticancer treatments of the haematological malignancies (Jian et al, 2011). Interestingly, in lung cancer, miRNA-663 was discovered to negatively influence the TGF-β1, P53, Bax and Fas expression (Liu et al, 2011) Furthermore, the miR-663 gene was found to be downregulated via methylation in the samples of human hepatocellular carcinoma and breast cancer, as well as in the K-562 leukaemia cell line (Lehmann et al, 2008; Potapova et al, 2011; Yang et al, 2012). There is also one reported example of the opposite effect, where miR-663 stimulated the proliferation and tumorigenesis of nasopharyngeal carcinoma (Yi et al, 2012).
However, the list of direct targets of microRNAs miR-633 and miR-744 is far from complete. The best known target of both miRNAs that could be related to their oncosuppressive properties is TGF-β1 (Liu et al, 2011; Martin et al, 2011). Here, we show that microRNAs miR-663 and miR-744 have a negative impact also on eEF1A2, which is the key component of the translation elongation apparatus. This protein reveals oncogenic properties in some cancer tissues (Anand et al, 2002; Tomlinson et al, 2005). The overexpression of eEF1A2 has been shown to induce the filopodia formation and cell invasion/migration, potentially through cytoskeleton remodelling, in an Akt-and PI3K-dependent manner (Amiri et al, 2007; Jeganathan and Lee, 2007; Li et al, 2010). Eukaryotic translation elongation factor 1A2 possesses antiapoptotic properties, possibly due to the direct interaction with Prdx1 (Chang and Wang, 2007). In addition, the ectopic expression of eEF1A2 altered the three-dimensional morphogenesis of MCF10A cells by influencing the phosphatidylinositol localisation (Pinke and Lee, 2011).
Importantly, the correlation between the low level of eEF1A2 (data not shown) and high level of mir-744 expression (Landgraf et al, 2007) was observed in MCF10 breast, A549 lung and Jurkat leukaemia cells. On the opposite, in the heart tissue where the expression of eEF1A2 is quite high (Lee et al, 1992), the expression of miR-744 was absent (Landgraf et al, 2007). Unfortunately, it is not possible to track the same correlation for miR-663 as no data about its expression are available in MirZ database (Hausser et al, 2009). We postulate that the abnormal occurrence of the eEF1A2 isoform in cancer tissue that is not usually found in normal one is caused, at least partially, by the loss of a microRNA-mediated silencing mechanism. The data on epigenetic inactivation of miR-663 in breast cancer are consistent with this hypothesis (Lehmann et al, 2008).
Our data show that resveratrol may control eEF1A2 expression post-transcriptionally. Resveratrol is a natural phytoalexin, and is found in significant quantities in grapes and, consequently, in red wine. Resveratrol is found to potentially prevent pathologies such as obesity and type 2 diabetes, and possesses cardioprotective and oncosuppressive properties (Jang et al, 1997; Vang et al, 2011). Many authors explain ‘French Paradox’ by regular intake of red wine that is rich in resveratrol and other polyphenols (Ferrieres, 2004; Lippi et al, 2010; Hengst and Yun, 2012). The first evidence of the oncosuppressive action of resveratrol was reported about 15 years ago (Jang et al, 1997). Since then, numerous studies have shown the ability of resveratrol to suppress the development of different human cancers (Vang et al, 2011). However, the mechanism of resveratrol’s antitumour activity is not yet revealed. It is known that miR-663 is upregulated during resveratrol treatment, directly targeting the known proto-oncogenes JunB and JunD, as well as TGF-β1 (Tili et al, 2010a,b); see Tili and Michaille (2011) for review.
Here, we have shown for the first time that resveratrol elevates, by at least two-fold, the expression level of miR-744. Recently, resveratrol has been shown to upregulate also miR-663 (Tili et al, 2010a). Thus, the resveratrol-induced upraise in miR-663 and miR-744 can control the proliferation of cancer cells via the inhibition of proto-oncogene eEF1A2.
In conclusion, it is yet unknown which mechanism determines exclusive tissue-specific expression of the eEF1A2 isoform in mammalian organisms. We believe miR-744 and/or miR-663 may be among the factors contributing to this issue.
Change history
11 June 2013
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Ambros V (2004) The functions of animal microRNAs. Nature 431 (7006): 350–355
Amiri A, Noei F, Jeganathan S, Kulkarni G, Pinke DE, Lee JM (2007) eEF1A2 activates Akt and stimulates Akt-dependent actin remodeling, invasion and migration. Oncogene 26 (21): 3027–3040
Anand N, Murthy S, Amann G, Wernick M, Porter LA, Cukier IH, Collins C, Gray JW, Diebold J, Demetrick DJ, Lee JM (2002) Protein elongation factor EEF1A2 is a putative oncogene in ovarian cancer. Nat Genet 31 (3): 301–305
Berezikov E, van Tetering G, Verheul M, van de Belt J, van Laake L, Vos J, Verloop R, van de Wetering M, Guryev V, Takada S, van Zonneveld AJ, Mano H, Plasterk R, Cuppen E (2006) Many novel mammalian microRNA candidates identified by extensive cloning and RAKE analysis. Genome Res 16 (10): 1289–1298
Betel D, Wilson M, Gabow A, Marks DS, Sander C (2008) The microRNA.org resource: targets and expression. Nucleic Acids Res 36, (database issue): D149–D153
Bosutti A, Scaggiante B, Grassi G, Guarnieri G, Biolo G (2007) Overexpression of the elongation factor 1A1 relates to muscle proteolysis and proapoptotic p66(ShcA) gene transcription in hypercatabolic trauma patients. Metabolism 56 (12): 1629–1634
Cao H, Zhu Q, Huang J, Li B, Zhang S, Yao W, Zhang Y (2009) Regulation and functional role of eEF1A2 in pancreatic carcinoma. Biochem Biophys Res Commun 380 (1): 11–16
Chang R, Wang E (2007) Mouse translation elongation factor eEF1A-2 interacts with Prdx-I to protect cells against apoptotic death induced by oxidative stress. J Cell Biochem 100 (2): 267–278
Chow A, Arteaga CL, Wang SE (2011) When tumor suppressor TGFbeta meets the HER2 (ERBB2) oncogene. J Mammary Gland Biol Neoplasia 16 (2): 81–88
De Palma M, Hanahan D (2012) The biology of personalized cancer medicine: facing individual complexities underlying hallmark capabilities. Mol Oncol 6 (2): 111–127
Ferrieres J (2004) The French paradox: lessons for other countries. Heart 90 (1): 107–111
Friedman RC, Farh KK, Burge CB, Bartel DP (2009) Most mammalian mRNAs are conserved targets of microRNAs. Genome Res 19 (1): 92–105
Gordon KJ, Blobe GC (2008) Role of transforming growth factor-beta superfamily signaling pathways in human disease. Biochimica et Biophysica Acta 1782 (4): 197–228
Hausser J, Berninger P, Rodak C, Jantscher Y, Wirth S, Zavolan M (2009) MirZ: an integrated microRNA expression atlas and target prediction resource. Nucleic Acids Res 37, (web server issue): W266–W272
Hengst JA, Yun JK (2012) Sphingosine Kinase: A key to solving the "French Paradox"? Br J Pharmacol 166 (5): 1603–1604
Higuchi R, Krummel B, Saiki RK (1988) A general method of in vitro preparation and specific mutagenesis of DNA fragments: study of protein and DNA interactions. Nucleic Acids Res 16 (15): 7351–7367
Huang V, Place RF, Portnoy V, Wang J, Qi Z, Jia Z, Yu A, Shuman M, Yu J, Li LC (2012) Upregulation of cyclin B1 by miRNA and its implications in cancer. Nucleic Acids Res 40 (4): 1695–1707
Imamura T, Hikita A, Inoue Y (2012) The roles of TGF-beta signaling in carcinogenesis and breast cancer metastasis. Breast Cancer 19 (2): 118–124
Jang M, Cai L, Udeani GO, Slowing KV, Thomas CF, Beecher CW, Fong HH, Farnsworth NR, Kinghorn AD, Mehta RG, Moon RC, Pezzuto JM (1997) Cancer chemopreventive activity of resveratrol, a natural product derived from grapes. Science 275 (5297): 218–220
Jeganathan S, Lee JM (2007) Binding of elongation factor eEF1A2 to phosphatidylinositol 4-kinase beta stimulates lipid kinase activity and phosphatidylinositol 4-phosphate generation. J Biol Chem 282 (1): 372–380
Jian P, Li ZW, Fang TY, Jian W, Zhuan Z, Mei LX, Yan WS, Jian N (2011) Retinoic acid induces HL-60 cell differentiation via the upregulation of miR-663. J Hematol Oncol 4: 20
Kertesz M, Iovino N, Unnerstall U, Gaul U, Segal E (2007) The role of site accessibility in microRNA target recognition. Nat Genet 39 (10): 1278–1284
Knudsen SM, Frydenberg J, Clark BF, Leffers H (1993) Tissue-dependent variation in the expression of elongation factor-1 alpha isoforms: isolation and characterisation of a cDNA encoding a novel variant of human elongation-factor 1 alpha. Eur J Biochem 215 (3): 549–554
Kolesanova EF, Farafonova TE, Aleshina EY, Pyndyk NV, Veremieva MV, Novosylna OV, Kovalenko MI, Shalak VF, Negrutskii BS (2013) Preparation of monospecific antibodies against isoform 2 of translation elongation factor 1A (eEF1A2). Biochemistry (Moscow) Suppl B Biomedical Chemistry 1: 62–69
Lagos-Quintana M, Rauhut R, Lendeckel W, Tuschl T (2001) Identification of novel genes coding for small expressed RNAs. Science 294 (5543): 853–858
Landgraf P, Rusu M, Sheridan R, Sewer A, Iovino N, Aravin A, Pfeffer S, Rice A, Kamphorst AO, Landthaler M, Lin C, Socci ND, Hermida L, Fulci V, Chiaretti S, Foa R, Schliwka J, Fuchs U, Novosel A, Muller RU, Schermer B, Bissels U, Inman J, Phan Q, Chien M, Weir DB, Choksi R, De Vita G, Frezzetti D, Trompeter HI, Hornung V, Teng G, Hartmann G, Palkovits M, Di Lauro R, Wernet P, Macino G, Rogler CE, Nagle JW, Ju J, Papavasiliou FN, Benzing T, Lichter P, Tam W, Brownstein MJ, Bosio A, Borkhardt A, Russo JJ, Sander C, Zavolan M, Tuschl T (2007) A mammalian microRNA expression atlas based on small RNA library sequencing. Cell 129 (7): 1401–1414
Lee MH, Choi BY, Kundu JK, Shin YK, Na HK, Surh YJ (2009) Resveratrol suppresses growth of human ovarian cancer cells in culture and in a murine xenograft model: eukaryotic elongation factor 1A2 as a potential target. Cancer Res 69 (18): 7449–7458
Lee MH, Surh YJ (2009) eEF1A2 as a putative oncogene. Ann N Y Acad Sci 1171: 87–93
Lee S, Francoeur AM, Liu S, Wang E (1992) Tissue-specific expression in mammalian brain, heart, and muscle of S1, a member of the elongation factor-1 alpha gene family. J Biol Chem 267 (33): 24064–24068
Lehmann U, Hasemeier B, Christgen M, Muller M, Romermann D, Langer F, Kreipe H (2008) Epigenetic inactivation of microRNA gene hsa-mir-9-1 in human breast cancer. J Pathol 214 (1): 17–24
Lewis BP, Burge CB, Bartel DP (2005) Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120 (1): 15–20
Li Z, Qi CF, Shin DM, Zingone A, Newbery HJ, Kovalchuk AL, Abbott CM, Morse HC 3rd (2010) Eef1a2 promotes cell growth, inhibits apoptosis and activates JAK/STAT and AKT signaling in mouse plasmacytomas. PLoS One 5 (5): e10755
Lippi G, Franchini M, Favaloro EJ, Targher G (2010) Moderate red wine consumption and cardiovascular disease risk: beyond the "French paradox". Semin Thromb Hemost 36 (1): 59–70
Liu ZY, Zhang GL, Wang MM, Xiong YN, Cui HQ (2011) MicroRNA-663 targets TGFB1 and regulates lung cancer proliferation. Asian Pac J Cancer Prev 12 (11): 2819–2823
Maragkakis M, Alexiou P, Papadopoulos GL, Reczko M, Dalamagas T, Giannopoulos G, Goumas G, Koukis E, Kourtis K, Simossis VA, Sethupathy P, Vergoulis T, Koziris N, Sellis T, Tsanakas P, Hatzigeorgiou AG (2009) Accurate microRNA target prediction correlates with protein repression levels. BMC bioinformatics 10: 295
Martin J, Jenkins RH, Bennagi R, Krupa A, Phillips AO, Bowen T, Fraser DJ (2011) Post-transcriptional regulation of Transforming Growth Factor Beta-1 by microRNA-744. PLoS One 6 (10): e25044
Mi QS, Weiland M, Qi RQ, Gao XH, Poisson LM, Zhou L (2012) Identification of mouse serum miRNA endogenous references by global gene expression profiles. PLoS One 7 (2): e31278
Pan J, Hu H, Zhou Z, Sun L, Peng L, Yu L, Liu J, Yang Z, Ran Y (2010) Tumor-suppressive mir-663 gene induces mitotic catastrophe growth arrest in human gastric cancer cells. Oncol Rep 24 (1): 105–112
Pan J, Ruest LB, Xu S, Wang E (2004) Immuno-characterization of the switch of peptide elongation factors eEF1A-1/EF-1alpha and eEF1A-2/S1 in the central nervous system during mouse development. Brain Res Develop Brain Res 149 (1): 1–8
Pinke DE, Kalloger SE, Francetic T, Huntsman DG, Lee JM (2008) The prognostic significance of elongation factor eEF1A2 in ovarian cancer. Gynecol Oncol 108 (3): 561–568
Pinke DE, Lee JM (2011) The lipid kinase PI4KIIIbeta and the eEF1A2 oncogene co-operate to disrupt three-dimensional in vitro acinar morphogenesis. Exp Cell Res 317 (17): 2503–2511
Potapova A, Albat C, Hasemeier B, Haeussler K, Lamprecht S, Suerbaum S, Kreipe H, Lehmann U (2011) Systematic cross-validation of 454 sequencing and pyrosequencing for the exact quantification of DNA methylation patterns with single CpG resolution. BMC Biotechnol 11: 6
Reinhart BJ, Weinstein EG, Rhoades MW, Bartel B, Bartel DP (2002) MicroRNAs in plants. Genes & Develop 16 (13): 1616–1626
Song MY, Pan KF, Su HJ, Zhang L, Ma JL, Li JY, Yuasa Y, Kang D, Kim YS, You WC (2012) Identification of serum microRNAs as novel non-invasive biomarkers for early detection of gastric cancer. PLoS One 7 (3): e33608
Tili E, Michaille JJ (2011) Resveratrol, MicroRNAs, Inflammation, and Cancer. J Nucleic Acids 2011: 102431
Tili E, Michaille JJ, Adair B, Alder H, Limagne E, Taccioli C, Ferracin M, Delmas D, Latruffe N, Croce CM (2010a) Resveratrol decreases the levels of miR-155 by upregulating miR-663, a microRNA targeting JunB and JunD. Carcinogenesis 31 (9): 1561–1566
Tili E, Michaille JJ, Alder H, Volinia S, Delmas D, Latruffe N, Croce CM (2010b) Resveratrol modulates the levels of microRNAs targeting genes encoding tumor-suppressors and effectors of TGFbeta signaling pathway in SW480 cells. Biochem Pharmacol 80 (12): 2057–2065
Tomlinson VA, Newbery HJ, Bergmann JH, Boyd J, Scott D, Wray NR, Sellar GC, Gabra H, Graham A, Williams AR, Abbott CM (2007) Expression of eEF1A2 is associated with clear cell histology in ovarian carcinomas: overexpression of the gene is not dependent on modifications at the EEF1A2 locus. British J Cancer 96 (10): 1613–1620
Tomlinson VA, Newbery HJ, Wray NR, Jackson J, Larionov A, Miller WR, Dixon JM, Abbott CM (2005) Translation elongation factor eEF1A2 is a potential oncoprotein that is overexpressed in two-thirds of breast tumours. BMC Cancer 5: 113
Vang O, Ahmad N, Baile CA, Baur JA, Brown K, Csiszar A, Das DK, Delmas D, Gottfried C, Lin HY, Ma QY, Mukhopadhyay P, Nalini N, Pezzuto JM, Richard T, Shukla Y, Surh YJ, Szekeres T, Szkudelski T, Walle T, Wu JM (2011) What is new for an old molecule? systematic review and recommendations on the use of resveratrol. PLoS One 6 (6): e19881
Yang Y, Wang LL, Li YH, Gao XN, Liu Y, Yu L (2012) Effect of CpG island methylation on microRNA expression in the k-562 cell line. Biochem Genet 50 (1–2): 122–134
Yi C, Wang Q, Wang L, Huang Y, Li L, Liu L, Zhou X, Xie G, Kang T, Wang H, Zeng M, Ma J, Zeng Y, Yun JP (2012) MiR-663, a microRNA targeting p21(WAF1/CIP1), promotes the proliferation and tumorigenesis of nasopharyngeal carcinoma. Oncogene 31 (41): 4421–4433
Zhao S, Fernald RD (2005) Comprehensive algorithm for quantitative real-time polymerase chain reaction. J Comput Biol 12 (8): 1047–1064
Acknowledgements
We are grateful to Linda Pritchard for critical reading of the manuscript. AV is thankful to FEBS and French Embassy in the Ukraine for short visiting fellowships. Travel support for BN was provided by the PICS and GDRI programs.
Author information
Authors and Affiliations
Corresponding author
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License.
Supplementary Information accompanies this paper on British Journal of Cancer website
Supplementary information
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Vislovukh, A., Kratassiouk, G., Porto, E. et al. Proto-oncogenic isoform A2 of eukaryotic translation elongation factor eEF1 is a target of miR-663 and miR-744. Br J Cancer 108, 2304–2311 (2013). https://doi.org/10.1038/bjc.2013.243
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2013.243
Keywords
This article is cited by
-
Oncogenic activation of EEF1A2 expression: a journey from a putative to an established oncogene
Cellular & Molecular Biology Letters (2024)
-
Profiling salivary miRNA expression levels in Fanconi anemia patients – a pilot study
Odontology (2024)
-
MiRNA expression deregulation correlates with the Oncotype DX® DCIS score
Breast Cancer Research (2022)
-
Multi-omic brain and behavioral correlates of cell-free fetal DNA methylation in macaque maternal obesity models
Nature Communications (2022)
-
MicroRNA in combination with HER2-targeting drugs reduces breast cancer cell viability in vitro
Scientific Reports (2021)