Abstract
Background:
Proteomic discovery of cancer biomarkers in body fluids is challenging because of their low abundance in a complex background. Altered gene expression in tumours may not reflect protein levels in body fluids. We have tested combining gene expression profiling of tumours with proteomic analysis of cancer cell line secretomes as a strategy to discover urinary biomarkers for bladder cancer.
Methods:
We used shotgun proteomics to identify proteins secreted by three bladder cancer cell lines. Secreted proteins with high mRNA levels in bladder tumours relative to normal urothelium were assayed by ELISA in urine samples from 642 patients.
Results:
Midkine and HAI-1 were significantly increased in bladder cancer patients, with the highest levels in invasive disease (area under the receiver operating characteristic curve 0.89 vs non-cancer). The urinary concentration of both proteins was too high to be explained by bladder cancer associated haematuria and most likely arises by direct tumour secretion.
Conclusions:
This ‘dual-omic’ strategy identified tumour secreted proteins whose urine concentrations are increased significantly by bladder cancer. Combined secretome-transcriptome analysis may be more useful than direct proteomic analysis of body fluids for biomarker discovery in both bladder cancer and other tumour types.
Similar content being viewed by others
Main
Despite intensive efforts to discover useful biomarkers for the non-invasive detection of cancer using increasingly powerful ‘omic’ technologies there is still a dearth of diagnostic markers available (Anderson, 2010). If such markers do actually exist they have evaded discovery due to their low concentration in very complex backgrounds. The methodological difficulties such samples present, particularly in proteomic work, should not be underestimated (Ray et al, 2011). A recently adopted approach has been to compare proteomic analysis of proteins secreted from cancer-specific cell lines to changes in gene expression in cancer tissue from the same cancer type (Chang et al, 2010, 2011; Wang et al, 2011; Yu et al, 2011). We now apply this method, for the first time, to the discovery of urinary biomarkers for the detection of bladder cancer.
Bladder cancer is the fifth most common cancer in Western populations (van Rhijn et al, 2009) with over 10 000 new cases and 5000 deaths attributed to bladder cancer each year in the UK alone (www.cancerresearchuk.org). Over 90% of these bladder cancers are urothelial carcinomas of the bladder (UCB) (Kaufman et al, 2009). At presentation over 70% of patients with UCB have non-muscle-invasive disease (stages Ta/T1/Tis), which is treated primarily by endoscopic resection (Kaufman et al, 2009). Muscle-invasive UCB (stages T2–4) is a life-threatening disease, which requires more radical treatment (chemotherapy and/or cystectomy/radiotherapy). Patients with non-muscle-invasive UCB require long-term surveillance to detect recurrent tumours with the potential to progress to muscle-invasive disease. The mainstay of surveillance is flexible cystoscopy combined with urine cytology (Babjuk et al, 2011). Flexible cystoscopy is considered the gold standard, although recent optical enhancement modalities have shown that the sensitivity of white light cystoscopy is limited (Cauberg et al, 2009). Cytology is very specific but has low sensitivity for early-stage disease (Lotan and Roehrborn, 2003). Several urine biomarker tests have been commercialised including NMP22 (nuclear matrix protein 22), BTA TRAK/stat (based on complement factor H) and UBC (cytokeratins 8 and 18) (Tilki et al, 2011), but have not proven sufficiently reliable to gain widespread acceptance.
Proteomic analyses of urine have revealed additional UCB biomarkers such as calreticulin (Kageyama et al, 2004), Reg-1 (Orenes-Piñero et al, 2007) and ADAM 28 (Yang et al, 2011); however, many of the reported ‘biomarkers’ are serum proteins (Irmak et al, 2005; Chen et al, 2010; Yang et al, 2011; Lindén et al, 2012) and are therefore unlikely to be UCB specific. Urine is a variable and difficult biospecimen for proteomic analysis, comprising of plasma proteins in addition to proteins arising from the kidneys and urothelium (Nagaraj and Mann, 2011). In addition, haematuria is a complicating factor to a greater or lesser extent in many UCBs (Bryan et al, 2011). An alternative source of candidate biomarkers is proteomic analysis of the conditioned media from cell lines or ‘secretomes’ (Volmer et al, 2005; Grønborg et al, 2006). Selecting relevant candidates from the large number of proteins typically identified is often based on quantitative comparisons between cell lines. In the case of bladder cancer, Kawanishi et al (2008) searched for markers of aggressive bladder cancer by comparing the secretomes of an invasive cell line (T24) with a less invasive cell line (RT112) and Makridakis et al, 2010 compared the secretomes of the T24 cell line with a derivative cell line (T24M) with a more aggressive phenotype. However, although this approach aims to detect aggression specific proteins, it is not appropriate for the detection of biomarkers for early disease. To achieve this, a comparator is required which represents non-cancer tissue, that is, primary cell culture or an immortalised ‘normal’ cell line (usually achieved by hTERT overexpression). Primary cells have limited lifespans and may be less resistant to the serum-free conditions required to harvest secreted proteins, and while immortalised normal cells are easier to culture they have not been widely used in comparative proteomic analyses of secretomes.
Microarray, and more recently next generation sequencing technologies have generated comprehensive, publically available gene expression profiles for many cancer types. Such data may not accurately reflect the concentrations of all proteins in cancers and are even less likely to predict protein secretion rates. Therefore, gene expression data per se may not predict which proteins could be useful as biomarkers in body fluids. Proteins which are both over expressed and detectable in cell line secretomes may be more useful for this purpose. Recently, this approach has been applied successfully to head and neck, lung and pancreatic cancers (Chang et al, 2010, 2011; Wang et al, 2011; Yu et al, 2011).
We hypothesised that a combined secretome-transcriptome strategy could reveal novel biomarkers for the detection of UCB. Ten candidate biomarkers with elevated gene expression in UCB and that were detected reproducibly in the secretome of multiple UCB-derived cell lines were shortlisted. Two of these candidates, midkine and HAI-1, were assayed in the urine of over 600 patients and as a control we also measured a third protein that was detected in the secretomes, but was not found to have an elevated gene expression in UCB.
Materials and Methods
Bladder cell line secretomes
5637 and HB-CLS-2 cells were purchased from CLS Cell Lines Service GmbH (Eppelheim, Germany) and EJ28 cells were a kind gift from Elizabeth Hodgkins, University of Birmingham. The cells were prepared in SILAC media (Life Technologies, Paisley, UK) to enable quantitative analysis although this was not used in this study. Cells were grown in SILAC RPMI 1640 Flex-Media supplemented with glutamine, glucose, 10% dialysed FCS and 12C6lysine and 12C6arginine (5637 and HB-CLS-2) or 13C6lysine and 13C6arginine (EJ28). Each cell line was grown to 80% confluence in 4 × 75 cm2 flasks, washed four times with PBS and four times with serum-free DMEM (phenol red free) and incubated for 18 h at 37 °C (5% CO2) in serum-free media. The conditioned media were filtered using 0.22 μm filters and concentrated using 15 ml 3 kDa spin-filters. Proteins were then reduced (50 mM DTT), alkylated (100 mM iodoacetamide) and digested with trypsin overnight using the FASP method (Wiśniewski et al, 2009). The resulting peptides were separated into 15 fractions by mixed-mode anion exchange/reverse-phase HPLC using a 2.1 × 50 mm2 Acclaim Mixed-mode WAX-1 HPLC column (Dionex, Camberley, UK) and a gradient of 0–40% B over 40 min (A: 20 mm ammonium formate pH 6.5, 3% acetonitrile, B: 2 mm ammonium formate pH 3.0, 80% acetonitrile). The fractions were dried under vacuum, dissolved in 1% formic acid and analysed by LC-MS/MS using a 90 min 0–40% acetonitrile gradient in 0.1% formic acid (75 μm × 25 cm C18 Pepmap column, Dionex) and a micrOTOF-Q II mass spectrometer (Bruker Daltonics, Bremen, Germany). Proteins extracted from the corresponding cell pellets were trypsinised using the FASP method and analysed in triplicate using a 2-h gradient on a maXis impact mass spectrometer (Bruker Daltonics) without pre-fractionation. For both types of experiment peptides were identified using MASCOT 2.3 to search a custom database containing SWISSPROT human and bovine sequences and randomised versions thereof (bovine sequences were included as the secretomes will inevitably contain traces of FCS). Mass tolerances for parent and fragment ions were 20 p.p.m. and 0.05 Da, respectively and the minimum peptide Mowse score was 20. Modifications considered were carbamidomethylation (fixed), methionine oxidation (variable) and 13C6lysine and 13C6arginine (variable, EJ28 cells only). Protein identifications were filtered using both a 1% false discovery threshold and a requirement for two or more peptides using Proteinscape 2.1 software.
Microarray data
We re-analysed, GSE3167 (Dyrskjøt et al, 2004) and GSE7476 (Mengual et al, 2009), two publicly available UCB microarray data sets, to identify mRNAs upregulated in stage Ta UCB. Affymetrix Human Genome U133A and U133 Plus 2.0 arrays were used to generate these data. Both data sets can be obtained from the Gene Expression Omnibus. Data set GSE7474 profiled 12 pooled samples, each containing RNA from bladder tissue from 4 or 5 patients. There were three pools each of normal bladder, low grade Ta UCB, high-grade T1 UCB and T2+ UCB. Data set GSE3167 profiled bladder tissue from 60 patients including 14 normal bladders, 15 Ta UCB, 18 Ta or T1 with CIS and 13 T2+ UCB (the Ta UCBs with surrounding CIS were not included in our analysis).
Patient samples
Urine samples were prospectively collected for biomarker research between 2006 and 2009 as part of the Bladder Cancer Prognosis Programme (BCPP, ethics approval 06/MRE04/65) (Zeegers et al, 2009). Patients were enrolled into the study on the basis of initial findings suggestive of primary UCB (predominantly abnormal cystoscopy). Mid-stream urine was collected before commencing treatment. Urine samples were placed on ice, centrifuged at 2000 r.p.m. for 10 min within 8 h of collection and the supernatant stored in 1 ml aliquots at −80 °C. As patient recruitment occurred before definitive histopathological confirmation of UCB following transurethral resection, a proportion of the patients were ultimately diagnosed with non-UCB conditions and these serve as our non-cancer ‘controls’. Patient information is summarised in Table 1.
Biomarker validation
Candidate biomarkers were measured in urine and serum sample using commercially available ELISA kits (DY258, DY1048 and DY1298, R&D systems, Abingdon, UK) according to the manufacturer’s instructions. Standards and samples were diluted in PBS containing 1% BSA and 20–40 μl of urine or 20 μl of serum was used per assay. Samples were randomised across multiple ELISA plates. Urinary creatinine, albumin, total protein and haematuria were determined previously (Bryan et al, 2011).
Data analysis
The significance of protein concentrations in different patient groups was assessed by Mann–Whitney test. Biomarker potential was assessed by receiver operating characteristic (ROC) analysis. For microarray data, probe level quantile normalisation (Bolstad et al, 2003) and RMA (robust multi-array analysis) (Irizarry et al, 2003) were performed using the affy package of the Bioconductor project (http://www.bioconductor.org). Differentially expressed probe sets were identified using limma (Smyth, 2004) using criteria of fold change >2.5 and P<0.001. Some genes had multiple probe sets and were considered as differentially expressed as long as one of their probe sets met these criteria.
Results
Proteomic analysis of UCB cell line secretomes
In total, 1001 human proteins were identified by two or more peptides in the secretomes of one or more of the three cell lines (EJ28, 5637 and HB-CLS-2). There was considerable overlap between the secretomes with 480 proteins being identified in at least two out of three secretomes and 256 detected in all three (all identifications are listed in Supplementary Information). The secretomes were substantially enriched for extracellular proteins compared with the corresponding cell pellets (Figure 1).
Classification of the proteins identified in the conditioned medium of UCB cell lines. The proteins identified in the conditioned media and in the corresponding cells were divided into categories using the GO Cellular Components function in STRING 9.0 (www.string-db.org). The number of proteins in each analysis was: 5637=622/546, EJ28=649/717, HB-CLS-2=822/462 (cell pellet/secretome).
Analysis of UCB microarray data
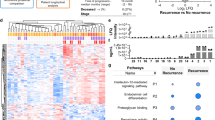
Data sets GSE7476 and GSE3167 were examined to identify substantially elevated mRNAs in early-stage (Ta) UCB. Although the experimental design is slightly different (see Materials and Methods), both data sets compare 12–14 normal bladders with 15 Ta UCBs. We used stringent criteria (2.5-fold change and P<0.001) to select 44 genes upregulated in both data sets. This list of genes was then compared with the list of proteins identified by two or more peptides in at least two of the cell line secretomes (Figure 2). Using this approach, we arrived at a shortlist of 10 biomarker candidates (Table 2).
Biomarker validation
Two of the candidate biomarkers listed in Table 2 (midkine and HAI-1) were assessed for their potential as urinary biomarkers for the detection of UCB. A third protein, ULBP-2, that was identified in all three secretomes but not substantially elevated in the microarray data was assayed as a control. The urine samples were randomly selected from the BCPP study and approximate to the population presenting with primary UCB. The distributions of HAI-1, midkine and ULBP-2 in the control subjects and UCB patients with Ta, T1 and T2+ disease are shown in Figure 3. Urinary levels of midkine and HAI-1 were both significantly elevated in patients with UCB relative to control subjects (P<0.0001). Midkine was significantly elevated in Ta (P=0.0422, 1.3-fold increase), T1 (P<0.0001, 2.4-fold increase) and T2+ UCB (P<0.0001, 5.4-fold increase). HAI-1 was significantly elevated in T1 (P<0.0001, 2.2-fold increase) and T2+ UCB (P<0.0001, 4.0-fold increase) but not Ta UCB (P=0.0890). The urinary concentration of ULBP-2 was not significantly influenced by the presence of Ta, T1 or T2+ UCB (P>0.05). We also analysed the data with regard to grade of disease: HAI-1 and midkine are both most markedly elevated in higher-grade disease (Figure 4). Midkine was significantly elevated in G2 (P=0.0017, 1.5-fold increase) and G3 (P<0.0001, 3.9-fold increase) but not G1 UCB (P=0.5398). Similarly, HAI-1 was significantly elevated in G2 (P=0.0006, 1.4-fold increase) and G3 UCB (P<0.0001, 3.0-fold increase) but not G1 UCB (P=0.5220). Neither HAI-1 nor midkine were significantly affected by age or gender and both have potential as markers for the detection of UCB as shown by the ROC plots in Figure 5. Areas under the ROC curves for discrimination between Ta, T1 and T2+ UCB and non-UCB using midkine were 0.574 (0.522–0.625 95% CI), 0.708 (0.644–0.766) and 0.851 (0.795–0.897), using HAI-1 the areas were 0.566 (0.514–0.618), 0.748 (0.687–0.803) and 0.886 (0.834–0.926). At 90% specificity, the sensitivities for discriminating between Ta, T1 and T2+ UCB and non-UCB were 18.1% (13.8–23.1%), 42.3% (34.2–50.6) and 66.7% (57.7–74.8) using midkine and 16.7% (12.6–21.6), 47.0 (38.8–55.3) and 74.6% (66.1–81.9) using HAI-1. The urinary levels of both proteins increase with stage such that, in this cohort of patients with UCB-like lesions at cystoscopy, midkine and HAI-1 can discriminate between the presence and absence of muscle-invasive UCB with areas under the ROC curve of 0.763 (0.728–0.796) and 0.791 (0.758–822), respectively. These data could be used to stratify patients as to their risk of muscle-invasive UCB: one third of the patients in this study have an urinary HAI-1 concentration greater than 0.9 pg μg−1 creatinine and a 42% risk of having muscle-invasive UCB, whereas the two thirds of patients with HAI-1 below this threshold have an 8% risk of muscle-invasive UCB.
Serum midkine and HAI-1 and effects of haematuria on urinary MDK and HAI-1
A major cause of false-positive results with urinary biomarkers for UCB is haematuria. To address this we have stratified urinary MDK and HAI-1 levels according to the level of haematuria determined by dipstick testing (Table 3). Both proteins show an UCB associated increase in concentration in urine even in the absence of haematuria. Insufficient haematuric controls were available to draw reliable conclusions about the effect of haematuria in the absence of UC (Supplementary Figure 1). Within each stage of UCB there is a significant increase in urinary midkine and HAI-1 concentrations with increasing haematuria. However, urinary midkine and HAI-1 correlate poorly with each other (r2=0.29) and very poorly with urinary albumin (r2=0.17 in both cases) compared with the correlation between urinary total protein and albumin concentrations (r2=0.85), suggesting that haematuria does not directly cause the increased urinary midkine and HAI-1 concentrations observed in UCB. To further investigate whether haematuria per se could be responsible for elevated urinary midkine and HAI-1 we measured both proteins in serum from 30 non-cancer controls and 50 UCB patients with elevated urinary midkine and/or HAI-1 (25 NMIBC and 25 MIBC). The median serum concentrations of midkine were 3.1 and 3.2 ng ml−1 for this subset of control and cancer patients, respectively, compared with median urine concentrations of 3.1 and 54.4 ng ml−1 in the same patients. What is more, in all 50 cancer patients the urine concentration of midkine was higher than the serum concentration. Median serum concentrations of HAI-1 were 2580 and 2126 pg ml−1 for this subset of control and cancer patients compared with median urine concentrations of 666 and 2759 pg ml−1. In 23 of the 50 cancer patients their serum HAI-1 concentration was higher than their urine concentration although a >5-fold ratio was only seen in eight patients and in these cases urinary HAI-1 was low (median 740 pg ml−1). We estimate that even in the most haematuric samples <10% of the urine volume is comprised of plasma (based on albumin concentrations) and that, therefore, it is very unlikely that haematuria directly causes elevated urinary midkine in UCB patients or that haematuria directly accounts for more than a small component of the elevated urinary HAI-1 seen in UCB patients (Supplementary Figure 2). Increased release of HAI-1 and midkine and causation of blood/plasma leakage into the urine may be shared characteristics of some, but not all, bladder tumours.
Discussion
We have used a combination of proteomics (identification of proteins secreted by UCB cell lines) and transcriptomics (publicly available microarray data) to identify candidate biomarkers for UCB. The tissue transcriptome has advantages over the tissue proteome for this purpose because a greater proportion of the genome is typically covered, and upregulated secreted proteins may have elevated mRNA levels but not accumulate as proteins in the tissue. Furthermore, there are many microarray data sets in the public domain that are suitable for the purpose outlined here (www.ncbi.nlm.nih.gov/geo). Two of the proteins identified using this method, HAI-1 and midkine, were selected for further analysis on the basis that their cellular localisation is ‘secreted’ (www.uniprot.org). Both showed a substantial elevation in the urine of UCB patients, although not in the urine of those with early-stage low-grade tumours. Other proteins on the shortlist have already shown potential as urinary biomarkers for UCB. E-cadherin has previously been measured in a large cohort of UCB patients and generated an ROC area of 0.72 (Shariat et al, 2005). Cytokeratin 19 (also known as CYFRA 21-1) is elevated in the urine of UCB patients (Senga et al, 1996). Cytokeratin 8 is measured in conjunction with cytokeratin 18 by the UBC test, which shows elevation in the urine of UCB patients (Sánchez-Carbayo et al, 1999). Thus, at least 5 of the 10 candidate biomarkers on our shortlist are elevated in the urine of UCB patients.
HAI-1 inhibits the proteases hepatocyte growth factor activator and matripase and is a membrane-associated protein that can be shed into the extracellular milieu (Kataoka et al, 2000). HAI-1 may have a role in the progression of many cancers (Parr and Jiang, 2006) and has been suggested as a serum marker for prostate cancer (Nagakawa et al, 2006). Expression of HAI-1 is inversely related to invasion and metastasis (Parr and Jiang, 2001), and low expression is considered to be an indicator of poor prognosis in some cancers (Nakamura et al, 2009). Midkine is an angiogenic growth factor normally expressed in the foetus but also re-expressed in many types of cancer (Muramaki et al, 2003). Midkine mRNA has been reported to be upregulated in UCB tissue and associated with late-stage disease and poor prognosis (O’Brien et al, 1996). The two microarray data sets used in this study, however, both show elevated midkine RNA in Ta UCB, which is maintained through the later stages. Ikematsu et al (2003) showed that urinary levels of midkine are elevated in several different malignant conditions. Ikematsu’s study found that midkine was elevated in 13 out of 15 UCB patients. Although UCB stage was not reported by the group, the data are consistent with our data for muscle-invasive disease. Although Ikematsu’s data support our findings, our data are the first large-scale assessment of midkine as a urinary marker for UCB.
In the two microarray data sets that we have re-analysed both HAI-1 and midkine mRNA expression is elevated in Ta UCB but does not increase further with increasing stage (Supplementary Figure 3). However, the urinary concentrations of both proteins show clear correlation with stage and grade. It is not clear why the urinary protein levels and mRNA do not correlate across disease stages: there is not a strong correlation between the protein levels and tumour size (data not shown) so it may reflect an increase in the rate of protein synthesis or secretion in more aggressive tumours. Regardless of mechanism, the marked increase in HAI-1 and midkine in T2+ UCB indicates potential utility as biomarkers for muscle-invasive disease. Although one of the key objectives of bladder cancer research may be the discovery of accurate and reliable biomarkers for diagnosing UCB and reducing reliance upon cystoscopy for surveillance, no such biomarkers have been discovered. It is likely that a multiple biomarker assay will need to be developed to satisfactorily address these objectives, to which HAI-1 and midkine may contribute. In-depth proteomic analysis of secretomes and cell surface proteomes of more UCB cell lines, comparison with transcriptomic data and validation of more candidates in clinical samples should further contribute to the development of a useful urine test for UCB. However, protein biomarkers for detecting muscle-invasive disease have immediate clinical utility in a number of settings, either alongside cystoscopic surveillance or in the intervals between cystoscopic surveillance. Firstly, for monitoring patients with high-grade T1 tumours who are being treated with intravesical therapy (usually BCG) (Babjuk et al, 2011) to guide clinicians on the need for closer monitoring or more aggressive treatment (for example, radical cystectomy). Similarly, for monitoring patients who have been treated with radiotherapy (or chemoradiotherapy) (James et al, 2012) for muscle-invasive disease. However, we feel that the greatest utility for these biomarkers could lie with the ‘stratified investigation’ of patients suspected of having bladder cancer in order to streamline their diagnosis and staging, and hence speed up their definitive treatment: instead of the usual series of investigations for haematuria (urine cytology, upper tract imaging, flexible cystoscopy) followed by TURBT and then staging CT scanning for patients with muscle-invasive disease, those patients with positive biomarkers at the outset could be ‘fast-tracked’ to immediate CT scanning that would both image their upper urinary tracts and stage the disease before TURBT (which has a disruptive effect on local staging (Salo et al, 1985)). Patients could thus skip one of the processes in the diagnostic pathway and, hence, proceed to definitive treatment more rapidly with the added benefit of more accurate local staging.
ULBP-2 was detected in all three secretomes but was not upregulated in the UCB microarray data. ULBP-2 is a membrane bound ligand for the NKG2D receptor and is widely expressed in tumours but not normal tissues (Onda et al, 2001) and shed by proteases (Waldhauer and Steinle, 2006). A recent study found serum ULBP-2 to perform at least as well as CA19-9 for detecting pancreatic cancer (Chang et al, 2011). In contrast to HAI-1 and midkine, we did not observe a significant increase in urinary ULBP-2 in patients with UCB. This is consistent with the notion that upregulation in transcriptome data is a useful way to select candidate biomarkers from secretome data, and also that the increased levels of midkine and HAI-1 in the urine of UCB patients do not arise from a non-specific leakage of cellular proteins from the tumour. In addition, plasma levels of HAI-1 and midkine are not high enough for tumour associated haematuria to account for the elevated urinary levels of these proteins and the two proteins are increased to different extents in different samples and do not correlate well with haematuria (data not shown).
In conclusion, proteins that are both identified in UCB cell line secretomes and upregulated at the mRNA level in UCB may be useful as urinary biomarkers for UCB as exemplified by the elevation of both midkine and HAI-1 in the urine of UCB patients. This method avoids the difficulties of in-depth proteomic analysis of body fluids and the lack of suitable comparator cell lines to select cancer-specific markers from cancer cell line secretomes. This approach has now proven useful in several types of tumour and may be of widespread applicability in the cancer biomarker field.
Change history
14 May 2013
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Anderson N (2010) The clinical plasma proteome: a survey of clinical assays for proteins in plasma and serum. Clin Chem 56: 177–185.
Babjuk M, Oosterlinck W, Sylvester R, Kaasinen E, Böhle A, Palou-Redorta J, Rouprêt M European Association of Urology (2011) EAU guidelines on non-muscle-invasive urothelial carcinoma of the bladder, the 2011 update. Eur Urol 59: 997–1008.
Bolstad BM, Irizarry R, Astrand M, Speed T (2003) A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19: 185–193.
Bryan R, Wei W, Shimwell N, Collins S, Hussain S, Billingham L, Murray P, Deshmukh N, James N, Wallace D, Johnson P, Zeegers M, Cheng K, Martin A, Ward D (2011) Assessment of high-throughput high-resolution MALDI-TOF-MS of urinary peptides for the detection of muscle-invasive bladder cancer. Proteomics Clin Appl 5: 493–503.
Cauberg E, de Bruin D, Faber D, van Leeuwen T, de la Rosette J, de Reijke T (2009) A new generation of optical diagnostics for bladder cancer: technology, diagnostic accuracy, and future applications. Eur Urol 56: 287–296.
Chang K, Wu C, Chen H, Chen S, Peng P, Tsang N, Lee L, Liu S, Liang Y, Lee Y, Hao S, Chang Y, Yu J (2010) Identification of candidate nasopharyngeal carcinoma serum biomarkers by cancer cell secretome and tissue transcriptome analysis: potential usage of cystatin A for predicting nodal stage and poor prognosis. Proteomics 10: 2644–2660.
Chang Y, Wu C, Shyr Y, Chen T, Hwang T, Yeh T, Chang K, Liu H, Liu Y, Tsai M, Chang Y, Yu J (2011) Secretome-based identification of ULBP2 as a novel serum marker for pancreatic cancer detection. PLoS One 6: e20029.
Chen Y, Chen C, Chen H, Chung T, Wu C, Chen C, Hsu C, Chen M, Tsui K, Chang P, Chang Y, Yu J (2010) Discovery of novel bladder cancer biomarkers by comparative urine proteomics using iTRAQ technology. J Proteome Res 9: 5803–5815.
Dyrskjøt L, Kruhøffer M, Thykjaer T, Marcussen N, Jensen J, Møller K, Ørntoft T (2004) Gene expression in the urinary bladder: a common carcinoma in situ gene expression signature exists disregarding histopathological classification. Cancer Res 64: 4040–4048.
Grønborg M, Kristiansen T, Iwahori A, Chang R, Reddy R, Sato N, Molina H, Jensen O, Hruban R, Goggins M, Maitra A, Pandey A (2006) Biomarker discovery from pancreatic cancer secretome using a differential proteomic approach. Mol Cell Proteomics 5: 157–171.
Ikematsu S, Okamoto K, Yoshida Y, Oda M, Sugano-Nagano H, Ashida K, Kumai H, Kadomatsu K, Muramatsu H, Muramatsu T, Sakuma S (2003) High levels of urinary midkine in various cancer patients. Biochem Biophys Res Commun 306: 329–332.
Irizarry RA, Bolstad BM, Collin F, Cope L, Hobbs B, Speed TP (2003) Summaries of Affymetrix GeneChip probe level data. Nucleic Acids Res 31: e15.
Irmak S, Tilki D, Heukeshoven J, Oliveira-Ferrer L, Friedrich M, Huland H, Ergün S (2005) Stage-dependent increase of orosomucoid and zinc-alpha2-glycoprotein in urinary bladder cancer. Proteomics 5: 4296–4304.
James N, Hussain S, Hall E, Jenkins P, Tremlett J, Rawlings C, Crundwell M, Sizer B, Sreenivasan T, Hendron C, Lewis R, Waters R, Huddart R, Investigators B (2012) Radiotherapy with or without chemotherapy in muscle-invasive bladder cancer. N Engl J Med 366: 1477–1488.
Kageyama S, Isono T, Iwaki H, Wakabayashi Y, Okada Y, Kontani K, Yoshimura K, Terai A, Arai Y, Yoshiki T (2004) Identification by proteomic analysis of calreticulin as a marker for bladder cancer and evaluation of the diagnostic accuracy of its detection in urine. Clin Chem 50: 857–866.
Kataoka H, Shimomura T, Kawaguchi T, Hamasuna R, Itoh H, Kitamura N, Miyazawa K, Koono M (2000) Hepatocyte growth factor activator inhibitor type 1 is a specific cell surface binding protein of hepatocyte growth factor activator (HGFA) and regulates HGFA activity in the pericellular microenvironment. J Biol Chem 275: 40453–40462.
Kaufman D, Shipley W, Feldman A (2009) Bladder cancer. Lancet 374: 239–249.
Kawanishi H, Matsui Y, Ito M, Watanabe J, Takahashi T, Nishizawa K, Nishiyama H, Kamoto T, Mikami Y, Tanaka Y, Jung G, Akiyama H, Nobumasa H, Guilford P, Reeve A, Okuno Y, Tsujimoto G, Nakamura E, Ogawa O (2008) Secreted CXCL1 is a potential mediator and marker of the tumor invasion of bladder cancer. Clin Cancer Res 14: 2579–2587.
Lindén M, Lind S, Mayrhofer C, Segersten U, Wester K, Lyutvinskiy Y, Zubarev R, Malmström P, Pettersson U (2012) Proteomic analysis of urinary biomarker candidates for nonmuscle invasive bladder cancer. Proteomics 12: 135–144.
Lotan Y, Roehrborn C (2003) Sensitivity and specificity of commonly available bladder tumor markers versus cytology: results of a comprehensive literature review and meta-analyses. Urology 61: 109–118.
Makridakis M, Roubelakis M, Bitsika V, Dimuccio V, Samiotaki M, Kossida S, Panayotou G, Coleman J, Candiano G, Anagnou N, Vlahou A (2010) Analysis of secreted proteins for the study of bladder cancer cell aggressiveness. J Proteome Res 9: 3243–3259.
Mengual L, Burset M, Ars E, Lozano J, Villavicencio H, Ribal M, Alcaraz A (2009) DNA microarray expression profiling of bladder cancer allows identification of noninvasive diagnostic markers. J Urol 182: 741–748.
Muramaki M, Miyake H, Hara I, Kamidono S (2003) Introduction of midkine gene into human bladder cancer cells enhances their malignant phenotype but increases their sensitivity to antiangiogenic therapy. Clin Cancer Res 9: 5152–5160.
Nagakawa O, Yamagishi T, Akashi T, Nagaike K, Fuse H (2006) Serum hepatocyte growth factor activator inhibitor type I (HAI-I) and type 2 (HAI-2) in prostate cancer. Prostate 66: 447–452.
Nagaraj N, Mann M (2011) Quantitative analysis of the intra- and inter-individual variability of the normal urinary proteome. J Proteome Res 10: 637–645.
Nakamura K, Abarzua F, Hongo A, Kodama J, Nasu Y, Kumon H, Hiramatsu Y (2009) The role of hepatocyte growth factor activator inhibitor-1 (HAI-1) as a prognostic indicator in cervical cancer. Int J Oncol 35: 239–248.
Onda H, Ohkubo S, Shintani Y, Ogi K, Kikuchi K, Tanaka H, Yamamoto K, Tsuji I, Ishibashi Y, Yamada T, Kitada C, Suzuki N, Sawada H, Nishimura O, Fujino M (2001) A Novel Secreted Tumor Antigen with a Glycosylphosphatidylinositol-Anchored Structure Ubiquitously Expressed in Human Cancers. Biochem Biophys Res Commun 285: 235–243.
Orenes-Piñero E, Cortón M, González-Peramato P, Algaba F, Casal I, Serrano A, Sánchez-Carbayo M (2007) Searching urinary tumor markers for bladder cancer using a two-dimensional differential gel electrophoresis (2D-DIGE) approach. J Proteome Res 6: 4440–4448.
O’Brien T, Cranston D, Fuggle S, Bicknell R, Harris A (1996) The angiogenic factor midkine is expressed in bladder cancer, and overexpression correlates with a poor outcome in patients with invasive cancers. Cancer Res 56: 2515–2518.
Parr C, Jiang W (2001) Expression of hepatocyte growth factor/scatter factor, its activator, inhibitors and the c-Met receptor in human cancer cells. Int J Oncol 19: 857–863.
Parr C, Jiang W (2006) Hepatocyte growth factor activation inhibitors (HAI-1 and HAI-2) regulate HGF-induced invasion of human breast cancer cells. Int J Cancer 119: 1176–1183.
Ray S, Reddy P, Jain R, Gollapalli K, Moiyadi A, Srivastava S (2011) Proteomic technologies for the identification of disease biomarkers in serum: advances and challenges ahead. Proteomics 11: 2139–2162.
Salo J, Kivisaari L, Lehtonen T (1985) CT in determining the depth of infiltration of bladder tumors. Urol Radiol 7: 88–93.
Sánchez-Carbayo M, Herrero E, Megías J, Mira A, Soria F (1999) Initial evaluation of the new urinary bladder cancer rapid test in the detection of transitional cell carcinoma of the bladder. Urology 54: 656–661.
Senga Y, Kimura G, Hattori T, Yoshida K (1996) Clinical evaluation of soluble cytokeratin 19 fragments (CYFRA 21-1) in serum and urine of patients with bladder cancer. Urology 48: 703–710.
Shariat S, Matsumoto K, Casella R, Jian W, Lerner S (2005) Urinary levels of soluble e-cadherin in the detection of transitional cell carcinoma of the urinary bladder. Eur Urol 48: 69–76.
Smyth G (2004) Linear models and empirical Bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol 3, Article 3.
Tilki D, Burger M, Dalbagni G, Grossman H, Hakenberg O, Palou J, Reich O, Rouprêt M, Shariat S, Zlotta A (2011) Urine markers for detection and surveillance of non-muscle-invasive bladder cancer. Eur Urol 60: 484–492.
van Rhijn B, Burger M, Lotan Y, Solsona E, Stief C, Sylvester R, Witjes J, Zlotta A (2009) Recurrence and progression of disease in non-muscle-invasive bladder cancer: from epidemiology to treatment strategy. Eur Urol 56.
Volmer M, Stühler K, Zapatka M, Schöneck A, Klein-Scory S, Schmiegel W, Meyer H, Schwarte-Waldhoff I (2005) Differential proteome analysis of conditioned media to detect Smad4 regulated secreted biomarkers in colon cancer. Proteomics 5: 2587–2601.
Waldhauer I, Steinle A (2006) Proteolytic release of soluble UL16-binding protein 2 from tumor cells. Cancer Res 66: 2520–2526.
Wang C, Wang C, Wang C, Chen C, Wu C, Liang Y, Tsai Y, Chang Y, Yu J, Yu C (2011) Importin subunit alpha-2 is identified as a potential biomarker for non-small cell lung cancer by integration of the cancer cell secretome and tissue transcriptome. Int J Cancer 128: 2364–2372.
Wiśniewski J, Zougman A, Nagaraj N, Mann M (2009) Universal sample preparation method for proteome analysis. Nat Methods 6: 359–362.
Yang M, Chu P, Chen S, Chung T, Chen W, Tan L, Kan W, Wang H, Su S, Tyan Y (2011) Characterization of ADAM28 as a biomarker of bladder transitional cell carcinomas by urinary proteome analysis. Biochem Biophys Res Commun 411: 714–720.
Yang N, Feng S, Shedden K, Xie X, Liu Y, Rosser C, Lubman D, Goodison S (2011) Urinary glycoprotein biomarker discovery for bladder cancer detection using LC/MS-MS and label-free quantification. Clin Cancer Res 17: 3349–3359.
Yu C, Chang K, Chang Y, Hsu C, Liang Y, Yu J, Chi L, Chang Y, Wu C (2011) Identification of guanylate-binding protein 1 as a potential oral cancer marker involved in cell invasion using omics-based analysis. J Proteome Res 10: 3778–3788.
Zeegers MP, Bryan RT, Langford C, Billingham L, Murray P, Deshmukh NS, Hussain S, James N, Wallace DM, Cheng KK (2009) The West Midlands Bladder Cancer Prognosis Programme: rationale and design. BJU Int 105: 784–788.
Acknowledgements
DGW is funded by Birmingham Science City. BCPP is funded by Cancer Research UK and the University of Birmingham, supported by the Birmingham and Black Country and West Midlands North and South Comprehensive Local Research Networks, and sponsored by the University of Birmingham. We thank all the West Midlands’ Consultant Urologists and their units who are involved with BCPP (Mr P Anderson, Mr R Bhatt, Mr G Choudhry, Mr P Cooke, Mr K Desai, Mr M Foster, Mr C Kadow, Mr C Luscombe, Mr A Makar, Mr J Parkin, Mr B Sarmah), as well as our BCPP research nurses (G Heritage, J Maiden, A McGuire, J Robinson, J Allison, L Moore, H Shackleford, K Castro, C Slater, T Martin, J Goodall), co-investigators (LJ Billingham, PG Murray, SA Hussain, DMA Wallace), and R Abid, J Barnwell, L Benson, D Nekeman, and R Reulen for contributing to these studies and for recruiting patients.
Author information
Authors and Affiliations
Corresponding author
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License.
Supplementary Information accompanies this paper on British Journal of Cancer website
Supplementary information
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Shimwell, N., Bryan, R., Wei, W. et al. Combined proteome and transcriptome analyses for the discovery of urinary biomarkers for urothelial carcinoma. Br J Cancer 108, 1854–1861 (2013). https://doi.org/10.1038/bjc.2013.157
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2013.157