Abstract
Background:
Recent data suggest a link between blood leukocyte DNA methylation, and cancer risk. However, reports on DNA methylation from a prospective study are unavailable for gastric cancer.
Methods:
We explored the association between methylation in pre-diagnostic blood leukocyte DNA and gastric cancer risk in a case–control study nested in the prospective Shanghai Women's Health Study cohort. Incident gastric cancer cases (n=192) and matched controls (n=384) were included in the study. Methylation of Alu and long interspersed nucleotide elements (LINE)-1 were evaluated using bisulphite pyrosequencing. Odds ratios (ORs) and 95% confidence intervals (CI) were calculated from logistic regression adjusting for potential confounders.
Results:
Alu methylation was inversely associated with gastric cancer risk, mainly among cases diagnosed one or more years after blood collection. After excluding cases diagnosed during the first year of follow-up, the ORs for the third, second, and first quartiles of Alu methylation compared with the highest quartile were 2.43 (1.43–4.13), 1.47(0.85–2.57), and 2.22 (1.28–3.84), respectively. This association appeared to be modified by dietary intake, particularly isoflavone. In contrast, LINE-1 methylation levels were not associated with gastric cancer risk.
Conclusion:
Evidence from this prospective study is consistent with the hypothesis that DNA hypomethylation in blood leukocytes may be related to cancer risk, including risk of gastric cancer.
Similar content being viewed by others
Main
Although its incidence has declined in the past decades, gastric cancer is still the fourth most commonly diagnosed cancer and the second most common cause of cancer death in the world (Jemal et al, 2011). Helicobacter pylori (H. pylori) infection, dietary factors (e.g., low vegetable and fruit intake, and high salt intake), and smoking have been associated with an elevated risk of this disease (Schottenfeld and Fraumeni, 2006). Genetic variations also have been suggested to have a role in gastric cancer development (Gao et al, 2009; Peleteiro et al, 2010). However, the molecular aetiology of gastric cancer remains unclear and we lack biomarkers for risk prediction for this disease. Epigenetic changes, which may represent an archive of past carcinogenic exposure (Jirtle and Skinner, 2007; Skinner, 2011), could be considered such candidates.
DNA methylation, the covalent addition of a methyl group, is an epigenetic event that affects cell function by altering gene expression. The cytosine nucleotide is the major target of methylation reaction and the product is usually 5-methyl-deoxycytidine (5-mdC). Changes in DNA methylation patterns are hypothesised to be involved in carcinogenesis (Ehrlich, 2002). Global DNA methylation, that is, the degree of methyl attachment to cytosine residues of DNA sequences across the genome, is typically reduced in a wide range of tumours (a phenomenon termed as ‘global hypomethylation’), including gastric cancer (Cravo et al, 1996; Hsiung et al, 2007; Moore et al, 2008; Guerrero-Preston et al, 2009). Global DNA methylation was decreased among human gastric cancer cell lines (Kaneda et al, 2004; Jiang et al, 2008), which was usually associated with chromosome instability (Watanabe and Maekawa, 2010). Significantly decreased global methylation has also been shown in gastric cancer tissue (Chalitchagorn et al, 2004; Suzuki et al, 2006) and pre-cancer lesions (Cravo et al, 1996; Yamamoto et al, 2008). Crave et al (1996) observed a graduated decrease in global DNA methylation in gastric specimens from normal, superficial gastritis, chronic atrophic gastritis with intestinal metaplasia, to intestinal type of gastric carcinoma.
Repetitive elements represent a large portion of the human genome and contain much of the CpG methylation found in normal human postnatal somatic tissues (Yang et al, 2004). Long interspersed nucleotide elements (LINE) make up about 15% of human genome with ∼0.5 million copies, and Alu-repetitive elements make up about 10% of human genome with 1.4 million copies. It is estimated that more than a third of DNA methylation occurs in the repetitive sequences (Weisenberger et al, 2005), and methylation levels of LINE-1 and Alu have been used as proxies for global DNA methylation status (Bollati et al, 2007, 2009; Tarantini et al, 2009).
Many studies on DNA methylation and gastric cancer to date have been conducted by comparing tumour tissue DNA with adjacent normal tissue DNA from cancer patients, and found decreased global or repetitive elements DNA methylation in tumour tissue compared with adjacent normal tissue (Chalitchagorn et al, 2004; Suzuki et al, 2006). Relatively fewer studies have been conducted on DNA methylation change in other non-target tissues (Chalitchagorn et al, 2004; Board et al, 2008). Methylation status of some CpG sites could be passed from previous generations as an inherited marker (Kile et al, 2010), or affected by aging or environmental exposures as a whole organism throughout life (Christensen et al, 2009). Changes of methylation in leukocyte DNA have been shown to parallel other somatic tissues (McKay et al, 2011), and have been linked to susceptibility of certain cancers (Marsit et al, 2011). Certainly, we could not rule out the possibility that different tissues have different response to environmental exposure and that methylation status in leukocytes may not fully reflect the changes in the target tissue (McKay et al, 2011). We previously observed suggestive associations between hypomethylation in Alu and LINE-1 in blood leukocyte DNA and gastric cancer risk in a case–control study (Hou et al, 2010). To follow-up this lead, we conducted a nested case–control study within the Shanghai Women's Health Study (SWHS) to prospectively examine the relationship between gastric cancer risk and DNA methylation in LINE-1 and Alu regions from pre-diagnostic blood leukocytes.
Methylation reactions depend on the availability of methyl groups from S-adenosylmethionine, which are mainly derived from foods that contain methionine, folic acid, and choline (Lamprecht and Lipkin, 2003); intake deficiency of these foods has been associated with decreased methylation level in colon and gastric tissues (Kim et al, 2003; Galvan-Portillo et al, 2009). Other lifestyle factors, such as alcohol drinking, cigarette smoking, and vitamins use, have also been linked to DNA methylation change and related with cancer status (Nan et al, 2005; Hillemacher et al, 2008). Furthermore, carriage of H. pylori has been suggested to interact with DNA methylation and lead to gastric carcinogenesis (Chan et al, 2003, 2007; Ushiku et al, 2007; Yamamoto et al, 2008; Hino et al, 2009). We also explored effect modification by these a priori factors on the methylation–cancer association.
Materials and methods
Study population
The SWHS; Zheng et al, 2005) is a prospective cohort comprised of 74 942 women, aged 40–70 years at study entry (1997–2000), from urban Shanghai, China. Participants were interviewed (response rate 93%) to elicit information on demographic characteristics, height and weight history, medical history, family cancer history, menstrual and reproductive history, dietary habits, and other lifestyle factors. Dietary intake was ascertained by a validated food-frequency questionnaire that included frequency of consumption (daily, weekly, monthly, yearly, or not at all) and portion size of each serving (Epplein et al, 2010). Daily consumption of isoflavone and folic acid from food was calculated according to the nutrient content per gram of the food obtained from the Chinese Food Composition Tables (Lee et al, 2009; Epplein et al, 2010). In addition, 56 831 participants provided a 10-ml blood sample collected into an EDTA vacutainer tube. Blood samples were kept at 0–4°C, transported on ice, and saved at −70°C within 6 h after collection. DNA was extracted from blood leukocytes with the standard phenol–chloroform method. Incident cancer patients were identified by biennial in-person follow-up interview and by annual linkage to the population-based Shanghai Cancer Registry.
All incident gastric cancer cases (n=192) diagnosed through December 2009, who provided a blood sample, were included in the current project. For each case, two cancer-free controls were randomly selected from cohort members and matched to the cases by age (±2 years), menopausal status at sample collection, date (±30 days) and time (morning or afternoon) of blood draw, and time since last meal (±2 h).
This study was approved by the institutional review boards of all participating study centers in China and the United States. All participants provided written informed consent.
DNA methylation measurement
For the current study, 500 ng DNA extracted from baseline buffy coat sample for each case and control was treated with bisulphite to convert unmethylated cytosine to uracil. Each assay was compared with an internal control sample to monitor the completion of bisulphite conversion. The bisulphite-treated DNA was then used for PCR amplification of Alu and LINE-1 elements, followed by pyrosequencing, a highly quantitative method as previously described (Tarantini et al, 2009; Baccarelli et al, 2010), to measure the degree of methylation for LINE-1 and Alu. Methylation status was expressed as the percentage of methylated cytosines over the total of methylated and unmethylated cytosines.
All of the 576 samples were tested in eight batches/plates. Matched case and control trio samples were placed within the same batch in random order. One negative control was included in each batch. To assess variability of the assay, duplicates from each of two QC subjects were randomly and blindly dispersed among study samples in each of eight batches (total 32 QC samples). Duplicate assays from bisulphate-converted DNA samples were analysed for each subject and the average was used for statistical analysis. The analytical variability (coefficient of variation calculated from replicates of the QC samples) was 2.2% and 1.0% for Alu methylation and LINE-1 methylation, respectively.
Statistical analysis
DNA methylation levels were classified as quartiles for statistical analysis according to their distributions in controls, and the highest quartile was used as the reference group for risk estimation. Logistic regression was used to estimate odds ratios (ORs) and 95% confidence intervals (95% CI) for the association between DNA methylation and gastric cancer risk. As there was essentially no difference in the risk estimates between conditional and unconditional logistic regressions, we reported results from unconditional logistic regression. Adjustment for age, body mass index (BMI), education, family history of gastric cancer, menopause status, cigarette smoking, alcohol drinking, and use of antibiotics and cold medicine 1 week before blood draw did not modify the risk estimates appreciably (<5%). Among a subset of subjects (67% of total subjects) with H. pylori infection status, further adjustment for this variable did not alter risk estimates either. Therefore, the results were reported from logistic regression adjusted for age only.
Taking advantage of the prospective design of the SWHS, we explored the effect of latency on the association between DNA methylation and gastric cancer risk. For cases, latency was defined as the time difference between blood draw and gastric cancer diagnosis; for controls, latency was defined as the time difference from blood draw to the time their corresponding cases was diagnosed with gastric cancer.
To explore whether the methylation-cancer association was modified by selected exposure factors, we conducted analyses stratified by family history of gastric cancer, menopause status, tea drinking, dietary intake of isoflavone and folic acid, and intake of vegetable and fruit. Dietary intake was dichotomously classified into high vs low intake based on the median in controls. Interactions were evaluated with log-likelihood ratio test comparing models with and without the interaction term. All statistical analyses were conducted with SAS 9.1 statistical software (Rockville, MD, USA) and all tests were two-sided. An analysis with P-value <0.05 was considered statistically significant.
Results
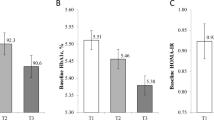
A total of 192 gastric cancer cases and 384 individually matched controls were included in the current analyses. Cases and controls were similar with regard to age, BMI, education, family history of gastric cancer, menopause status, recent non-steroidal anti-inflammatory drug and antibiotics use, vitamin B use, tea drinking, and dietary intake (isoflavones, folic acid, vegetables, and fruits; Table 1). Among subjects with information on H. pylori infection status, over 90% of cases and controls were positive. Smoking and alcohol drinking were not prevalent in Chinese women. The median levels of Alu and LINE-1 were nearly identical in cases and controls.
Compared with the highest quartile, decreased Alu methylation was associated with increased gastric cancer risk overall, with risk estimates of 2.17 (95% CI=1.32–3.56), 1.37 (95% CI=0.82–2.31), and 1.85 (95% CI=1.10–3.11) for the third, second, and the lowest quartiles, respectively. When we stratified the analysis by latency, however, we found that the associations for cases diagnosed within the first year following blood draw were different from those for cases diagnosed at a later time (Table 2). Within the first year of blood draw, hypomethylation of Alu appeared inversely associated with gastric cancer risk, although this observation was based on small numbers of subjects and none of the risk estimates were statistically significant (Table 2). For individuals whose gastric cancer was diagnosed at ⩾1 years after blood collection, risk was elevated with hypomethylation of Alu, although not in a consistent manner with either progression in hypomethylation or increasing years since blood draw. For LINE-1 hypomethylation, there were no consistent associations with gastric cancer risk, regardless of the years since blood collection.
We explored whether selected exposures might modify the association between hypomethylation in Alu and gastric cancer risk. For this analysis, we excluded cases diagnosed within 1 year of blood collection and their controls to minimise the potential impact of preclinical cancer. The inverse associations between the gastric cancer risk and Alu methylation tended to be stronger among women who consumed tea regularly and had high levels of intake of isoflavone, folic acid, or vegetable (Table 3). However, only the interaction with dietary isoflavone intake reached statistical significance (Pinteraction=0.04). In contrast, there were no consistent patterns of effect modification of the association between LINE-1 methylation and gastric cancer risk by selected exposures (Table 4).
Discussion
In this exploratory study, we observed inverse associations between gastric cancer risk and methylation of Alu, but not LINE-1, in pre diagnostic blood leukocytes. The association with Alu methylation appeared to be modified by years since blood collection, with an inverse association observed only for cases diagnosed ⩾1 year after blood collection. Furthermore, this association appeared to be modified by dietary intake, particularly intake of isoflavone.
Our study has several advantages and limitations. First, the study is nested within the population-based SWHS cohort, which is a well-characterised population of women at high risk for gastric cancer. Second, DNA obtained from blood samples at baseline allowed us to assess methylation status before cancer diagnosis, and explore the latency effect on the associations. To our knowledge, this is among the first prospective population-based study to date to examine the association between DNA methylation and cancer risk. In addition, detailed exposure information (i.e., dietary and other life style factors) allowed us to explore potential effect modifications. However, the sample size is relatively small, and we had limited power to detect interactions with potential effect modifiers. Besides, if donated blood samples of the participants of SWHS (75.8% of all participants) were different from those who did not donate blood, it could limit the generalisability of our results to those who did not provide a blood sample.
To date, a few case–control studies have examined the association between global DNA methylation in leukocytes and risk of various cancers, including cancers of the head and neck, stomach, breast, colon and rectum, and bladder (Pufulete et al, 2003; Hsiung et al, 2007; Moore et al, 2008; Choi et al, 2009; Hou et al, 2010; Wilhelm et al, 2010; Cash et al, 2011). Though all of the studies suggested an inverse association between global DNA methylation and cancer risk, not all results were statistically significant (Pufulete et al, 2003; Choi et al, 2009; Cash et al, 2011). Only one study examined gastric cancer risk in relation to both Alu and LINE-1 methylation in a Polish case–control study and found insignificant inverse associations with both (Hou et al, 2010). We did not observe a consistent dose-response effect with Alu methylation, raising the possibility of chance variations. However, the lack of linear association with methylation status has been reported previously, such as the association between bladder cancer and global DNA methylation evaluated directly with 5′-methylcytosine (Moore et al, 2008), which might reflect a threshold effect. Further evidence is needed to clarify the association between peripheral leukocyte DNA methylation and gastric cancer risk.
Though not statistically significant, we found that the association between Alu methylation and gastric cancer risk was stronger among individuals with longer latency. Certainly, this exploratory finding warrants further research. Ideally, it would be most informative if blood samples were available to monitor changes in DNA methylation for the same persons at multiple time points before cancer diagnosis.
We explored effect modification by several a priori factors and found a significant interaction only with dietary isoflavone intake, with a stronger association between gastric cancer risk and Alu hypomethylation among individuals with high isoflavone intake. This observation appears inconsistent with the known inhibitory effect of isoflavones on cancer development (Gilbert and Liu, 2010). It is also possible that this apparent effect modification is a chance occurrence.
Retro transposition (mobile genetic elements inserting into a new location) of LINE-1 and Alu has been reported to increase genomic instability, activate endogenous parasitic sequence, and contribute to the transcriptional silencing of tumour-suppressor genes in tissues, thus, leading to tumour development (Esteller and Herman, 2002; Gilbert et al, 2002; Symer et al, 2002). In addition, Alu and LINE-1 may also participate in regulating immune and inflammatory responses (Li and Schmid, 2001; Hagan and Rudin, 2002). Emerging data from epidemiological studies suggest different functional roles of Alu and LINE-1 in cancer initiation and progression (Bollati et al, 2007; Rusiecki et al, 2008; Jintaridth and Mutirangura, 2010). Though Alu and LINE-1 were correlated to each other in tumour tissue (Choi et al, 2007), a correlation has not always been observed inblood (Bollati et al, 2007; Rusiecki et al, 2008). A study on breast cancer found increased cancer risk associated with lower 5-mdC level in leukocyte DNA but not LINE-1 methylation level; further, LINE-1 methylation was not correlated with 5-mdC (Choi et al, 2009). Direct measurement of global 5-mdC is still needed to examine the role of global DNA methylation in gastric cancer aetiology. In the present study, we observed significantly increased risk of gastric cancer associated with hypomethylation of Alu but not LINE-1, and the correlation between LINE-1 methylation and Alu methylation among controls was modest (correlation coefficient of determination R2=0.12). Our findings seem to support the notion that Alu and LINE-1have different roles in gastric carcinogenesis, but given the small numbers in this study, we cannot rule out chance as an explanation.
In summary, the results from this prospective study suggest that DNA methylation of Alu, but not LINE-1, in blood leukocytes might be inversely associated with risk of gastric cancer. The divergent associations with Alu and LINE-1 raise the possibility that the roles of DNA methylation in these two elements are different in gastric cancer aetiology. However, replication is needed before further inference may be drawn from these observations.
Change history
29 March 2012
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Baccarelli A, Wright R, Bollati V, Litonjua A, Zanobetti A, Tarantini L, Sparrow D, Vokonas P, Schwartz J (2010) Ischemic heart disease and stroke in relation to blood DNA methylation. Epidemiology 21: 819–828
Board RE, Knight L, Greystoke A, Blackhall FH, Hughes A, Dive C, Ranson M (2008) DNA methylation in circulating tumour DNA as a biomarker for cancer. Biomark Insights 2: 307–319
Bollati V, Baccarelli A, Hou L, Bonzini M, Fustinoni S, Cavallo D, Byun HM, Jiang J, Marinelli B, Pesatori AC, Bertazzi PA, Yang AS (2007) Changes in DNA methylation patterns in subjects exposed to low-dose benzene. Cancer Res 67: 876–880
Bollati V, Schwartz J, Wright R, Litonjua A, Tarantini L, Suh H, Sparrow D, Vokonas P, Baccarelli A (2009) Decline in genomic DNA methylation through aging in a cohort of elderly subjects. Mech Ageing Dev 130: 234–239
Cash HL, Tao L, Yuan JM, Marsit CJ, Houseman EA, Xiang YB, Gao YT, Nelson HH, Kelsey KT (2011) LINE-1 hypomethylation is associated with bladder cancer risk among nonsmoking Chinese. Int J Cancer; e-pub ahead of print 25 May 2011, doi:10.1002/ijc.26098
Chalitchagorn K, Shuangshoti S, Hourpai N, Kongruttanachok N, Tangkijvanich P, Thong-ngam D, Voravud N, Sriuranpong V, Mutirangura A (2004) Distinctive pattern of LINE-1 methylation level in normal tissues and the association with carcinogenesis. Oncogene 23: 8841–8846
Chan AO, Chu KM, Huang C, Lam KF, Leung SY, Sun YW, Ko S, Xia HH, Cho CH, Hui WM, Lam SK, Rashid A (2007) Association between Helicobacter pylori infection and interleukin 1beta polymorphism predispose to CpG island methylation in gastric cancer. Gut 56: 595–597
Chan AO, Lam SK, Wong BC, Wong WM, Yuen MF, Yeung YH, Hui WM, Rashid A, Kwong YL (2003) Promoter methylation of E-cadherin gene in gastric mucosa associated with Helicobacter pylori infection and in gastric cancer. Gut 52: 502–506
Choi IS, Estecio MR, Nagano Y, Kim do H, White JA, Yao JC, Issa JP, Rashid A (2007) Hypomethylation of LINE-1 and Alu in well-differentiated neuroendocrine tumors (pancreatic endocrine tumors and carcinoid tumors). Mod Pathol 20: 802–810
Choi JY, James SR, Link PA, McCann SE, Hong CC, Davis W, Nesline MK, Ambrosone CB, Karpf AR (2009) Association between global DNA hypomethylation in leukocytes and risk of breast cancer. Carcinogenesis 30: 1889–1897
Christensen BC, Houseman EA, Marsit CJ, Zheng S, Wrensch MR, Wiemels JL, Nelson HH, Karagas MR, Padbury JF, Bueno R, Sugarbaker DJ, Yeh RF, Wiencke JK, Kelsey KT (2009) Aging and environmental exposures alter tissue-specific DNA methylation dependent upon CpG island context. PLoS Genet 5: e1000602
Cravo M, Pinto R, Fidalgo P, Chaves P, Gloria L, Nobre-Leitao C, Costa Mira F (1996) Global DNA hypomethylation occurs in the early stages of intestinal type gastric carcinoma. Gut 39: 434–438
Ehrlich M (2002) DNA methylation in cancer: too much, but also too little. Oncogene 21: 5400–5413
Epplein M, Shu XO, Xiang YB, Chow WH, Yang G, Li HL, Ji BT, Cai H, Gao YT, Zheng W (2010) Fruit and vegetable consumption and risk of distal gastric cancer in the Shanghai Women's and Men′s Health studies. Am J Epidemiol 172: 397–406
Esteller M, Herman JG (2002) Cancer as an epigenetic disease: DNA methylation and chromatin alterations in human tumours. J Pathol 196: 1–7
Galvan-Portillo MV, Cantoral A, Onate-Ocana LF, Chen J, Herrera-Goepfert R, Torres-Sanchez L, Hernandez-Ramirez RU, Palma-Coca O, Lopez-Carrillo L (2009) Gastric cancer in relation to the intake of nutrients involved in one-carbon metabolism among MTHFR 677 TT carriers. Eur J Nutr 2009; 48: 269–276
Gao L, Nieters A, Brenner H (2009) Cell proliferation-related genetic polymorphisms and gastric cancer risk: systematic review and meta-analysis. Eur J Hum Genet 17: 1658–1667
Gilbert ER, Liu D (2010) Flavonoids influence epigenetic-modifying enzyme activity: structure—function relationships and the therapeutic potential for cancer. Curr Med Chem 17: 1756–1768
Gilbert N, Lutz-Prigge S, Moran JV (2002) Genomic deletions created upon LINE-1 retrotransposition. Cell 110: 315–325
Guerrero-Preston R, Baez A, Blanco A, Berdasco M, Fraga M, Esteller M (2009) Global DNA methylation: a common early event in oral cancer cases with exposure to environmental carcinogens or viral agents. P R Health Sci J 28: 24–29
Hagan CR, Rudin CM (2002) Mobile genetic element activation and genotoxic cancer therapy: potential clinical implications. Am J Pharmacogenomics 2: 25–35
Hillemacher T, Frieling H, Moskau S, Muschler MA, Semmler A, Kornhuber J, Klockgether T, Bleich S, Linnebank M (2008) Global DNA methylation is influenced by smoking behaviour. Eur Neuropsychopharmacol 18: 295–298
Hino R, Uozaki H, Murakami N, Ushiku T, Shinozaki A, Ishikawa S, Morikawa T, Nakaya T, Sakatani T, Takada K, Fukayama M (2009) Activation of DNA methyltransferase 1 by EBV latent membrane protein 2A leads to promoter hypermethylation of PTEN gene in gastric carcinoma. Cancer Res 69: 2766–2774
Hou L, Wang H, Sartori S, Gawron A, Lissowska J, Bollati V, Tarantini L, Zhang FF, Zatonski W, Chow WH, Baccarelli A (2010) Blood leukocyte DNA hypomethylation and gastric cancer risk in a high-risk Polish population. Int J Cancer 127: 1866–1874
Hsiung DT, Marsit CJ, Houseman EA, Eddy K, Furniss CS, McClean MD, Kelsey KT (2007) Global DNA methylation level in whole blood as a biomarker in head and neck squamous cell carcinoma. Cancer Epidemiol Biomarkers Prev 16: 108–114
Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D (2011) Global cancer statistics. CA Cancer J Clin 61: 69–90
Jiang L, Gonda TA, Gamble MV, Salas M, Seshan V, Tu S, Twaddell WS, Hegyi P, Lazar G, Steele I, Varro A, Wang TC, Tycko B (2008) Global hypomethylation of genomic DNA in cancer-associated myofibroblasts. Cancer Res 68: 9900–9908
Jintaridth P, Mutirangura A (2010) Distinctive patterns of age-dependent hypomethylation in interspersed repetitive sequences. Physiol Genomics 41: 194–200
Jirtle RL, Skinner MK (2007) Environmental epigenomics and disease susceptibility. Nat Rev Genet 8: 253–262
Kaneda A, Tsukamoto T, Takamura-Enya T, Watanabe N, Kaminishi M, Sugimura T, Tatematsu M, Ushijima T (2004) Frequent hypomethylation in multiple promoter CpG islands is associated with global hypomethylation, but not with frequent promoter hypermethylation. Cancer Sci 95: 58–64
Kile ML, Baccarelli A, Tarantini L, Hoffman E, Wright RO, Christiani DC (2010) Correlation of global and gene-specific DNA methylation in maternal-infant pairs. PLoS One 5: e13730
Kim DH, Kim SW, Song YJ, Oh TY, Han SU, Kim YB, Joo HJ, Cho YK, Kim DY, Cho SW, Kim MW, Kim JH, Hahm KB (2003) Long-term evaluation of mice model infected with Helicobacter pylori: focus on gastric pathology including gastric cancer. Aliment Pharmacol Ther 18 (Suppl 1): 14–23
Lamprecht SA, Lipkin M (2003) Chemoprevention of colon cancer by calcium, vitamin D and folate: molecular mechanisms. Nat Rev Cancer 3: 601–614
Lee I, Oguma Y (2006) Physical activity. In Cancer Epidemiology and Prevention Schottenfeld D, Fraumeni J (eds) Oxford University Press: New York
Lee SA, Shu XO, Li H, Yang G, Cai H, Wen W, Ji BT, Gao J, Gao YT, Zheng W (2009) Adolescent and adult soy food intake and breast cancer risk: results from the Shanghai Women's Health Study. Am J Clin Nutr 89: 1920–1926
Li TH, Schmid CW (2001) Differential stress induction of individual Alu loci: implications for transcription and retrotransposition. Gene 276: 135–141
Marsit CJ, Koestler DC, Christensen BC, Karagas MR, Houseman EA, Kelsey KT (2011) DNA methylation array analysis identifies profiles of blood-derived DNA methylation associated with bladder cancer. J Clin Oncol 29: 1133–1139
McKay JA, Xie L, Harris S, Wong YK, Ford D, Mathers JC (2011) Blood as a surrogate marker for tissue-specific DNA methylation and changes due to folate depletion in post-partum female mice. Mol Nutr Food Res 55: 1026–1035
Moore LE, Pfeiffer RM, Poscablo C, Real FX, Kogevinas M, Silverman D, Garcia-Closas R, Chanock S, Tardon A, Serra C, Carrato A, Dosemeci M, Garcia-Closas M, Esteller M, Fraga M, Rothman N, Malats N (2008) Genomic DNA hypomethylation as a biomarker for bladder cancer susceptibility in the Spanish Bladder Cancer Study: a case-control study. Lancet Oncol 9: 359–366
Nan HM, Song YJ, Yun HY, Park JS, Kim H (2005) Effects of dietary intake and genetic factors on hypermethylation of the hMLH1 gene promoter in gastric cancer. World J Gastroenterol 11: 3834–3841
Peleteiro B, Lunet N, Carrilho C, Duraes C, Machado JC, La Vecchia C, Barros H (2010) Association between cytokine gene polymorphisms and gastric precancerous lesions: systematic review and meta-analysis. Cancer Epidemiol Biomarkers Prev 19: 762–776
Pufulete M, Al-Ghnaniem R, Leather AJ, Appleby P, Gout S, Terry C, Emery PW, Sanders TA (2003) Folate status, genomic DNA hypomethylation, and risk of colorectal adenoma and cancer: a case control study. Gastroenterology 124: 1240–1248
Rusiecki JA, Baccarelli A, Bollati V, Tarantini L, Moore LE, Bonefeld-Jorgensen EC (2008) Global DNA hypomethylation is associated with high serum-persistent organic pollutants in Greenlandic Inuit. Environ Health Perspect 116: 1547–1552
Skinner MK (2011) Environmental epigenomics and disease susceptibility. EMBO Rep 12: 620–622
Suzuki K, Suzuki I, Leodolter A, Alonso S, Horiuchi S, Yamashita K, Perucho M (2006) Global DNA demethylation in gastrointestinal cancer is age dependent and precedes genomic damage. Cancer Cell 9: 199–207
Symer DE, Connelly C, Szak ST, Caputo EM, Cost GJ, Parmigiani G, Boeke JD (2002) Human l1 retrotransposition is associated with genetic instability in vivo. Cell 110: 327–338
Tarantini L, Bonzini M, Apostoli P, Pegoraro V, Bollati V, Marinelli B, Cantone L, Rizzo G, Hou L, Schwartz J, Bertazzi PA, Baccarelli A (2009) Effects of particulate matter on genomic DNA methylation content and iNOS promoter methylation. Environ Health Perspect 117: 217–222
Ushiku T, Chong JM, Uozaki H, Hino R, Chang MS, Sudo M, Rani BR, Sakuma K, Nagai H, Fukayama M (2007) p73 gene promoter methylation in Epstein-Barr virus-associated gastric carcinoma. Int J Cancer 120: 60–66
Watanabe Y, Maekawa M (2010) Methylation of DNA in cancer. Adv Clin Chem 52: 145–167
Weisenberger DJ, Campan M, Long TI, Kim M, Woods C, Fiala E, Ehrlich M, Laird PW (2005) Analysis of repetitive element DNA methylation by MethyLight. Nucleic Acids Res 33: 6823–6836
Wilhelm CS, Kelsey KT, Butler R, Plaza S, Gagne L, Zens MS, Andrew AS, Morris S, Nelson HH, Schned AR, Karagas MR, Marsit CJ (2010) Implications of LINE1 methylation for bladder cancer risk in women. Clin Cancer Res 16: 1682–1689
Yamamoto E, Toyota M, Suzuki H, Kondo Y, Sanomura T, Murayama Y, Ohe-Toyota M, Maruyama R, Nojima M, Ashida M, Fujii K, Sasaki Y, Hayashi N, Mori M, Imai K, Tokino T, Shinomura Y (2008) LINE-1 hypomethylation is associated with increased CpG island methylation in Helicobacter pylori-related enlarged-fold gastritis. Cancer Epidemiol Biomarkers Prev 17: 2555–2564
Yang AS, Estecio MR, Doshi K, Kondo Y, Tajara EH, Issa JP (2004) A simple method for estimating global DNA methylation using bisulfite PCR of repetitive DNA elements. Nucleic Acids Res 32: e38
Zheng W, Chow WH, Yang G, Jin F, Rothman N, Blair A, Li HL, Wen W, Ji BT, Li Q, Shu XO, Gao YT (2005) The Shanghai Women's Health Study: rationale, study design, and baseline characteristics. Am J Epidemiol 162: 1123–1131
Acknowledgements
This study was supported by the Intramural Research Program of the Division of Cancer Epidemiology and Genetics, National Cancer Institute, National Institutes of Health. Dr Baccarelli is partially supported by New Investigator funding from NIEHS (ES00002). Finally, we acknowledge the study participants for donating their time and making this study possible.
Author contributions
Y Gao and W-H Chow designed the study, analysed data, and wrote the manuscript. A Baccarelli and L Tarantini conducted the DNA methylation assay for this project. All authors read, gave comments, and approved the final version of the manuscript. All authors had full access to all the data in the study and take responsibility for the integrity of the data and the accuracy of the data analysis.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License.
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Gao, Y., Baccarelli, A., Shu, X. et al. Blood leukocyte Alu and LINE-1 methylation and gastric cancer risk in the Shanghai Women's Health Study. Br J Cancer 106, 585–591 (2012). https://doi.org/10.1038/bjc.2011.562
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2011.562
Keywords
This article is cited by
-
DNA methylation and socioeconomic status in a Mexican-American birth cohort
Clinical Epigenetics (2018)
-
LINE-1 Hypermethylation in Serum Cell-Free DNA of Relapsing Remitting Multiple Sclerosis Patients
Molecular Neurobiology (2018)
-
Dietary Ingestion of Calories and Micronutrients Modulates the DNA Methylation Profile of Leukocytes from Older Individuals
The Journal of nutrition, health and aging (2018)
-
LINE-1 hypomethylation is not a common event in preneoplastic stages of gastric carcinogenesis
Scientific Reports (2017)
-
Prospective changes in global DNA methylation and cancer incidence and mortality
British Journal of Cancer (2016)