Abstract
Acute myeloid leukemia (AML) is a heterogeneous neoplasm characterized by the accumulation of complex genetic alterations responsible for the initiation and progression of the disease. Translating genomic information into clinical practice remained challenging with conflicting results regarding the impact of certain mutations on disease phenotype and overall survival (OS) especially when clinical variables are controlled for when interpreting the result. We sequenced the coding region for 62 genes in 468 patients with secondary AML (sAML) and primary AML (pAML). Overall, mutations in FLT3, DNMT3A, NPM1 and IDH2 were more specific for pAML whereas UTAF1, STAG2, BCORL1, BCOR, EZH2, JAK2, CBL, PRPF8, SF3B1, ASXL1 and DHX29 were more specific for sAML. However, in multivariate analysis that included clinical variables, only FLT3 and DNMT3A remained specific for pAML and EZH2, BCOR, SF3B1 and ASXL1 for sAML. When the impact of mutations on OS was evaluated in the entire cohort, mutations in DNMT3A, PRPF8, ASXL1, CBL EZH2 and TP53 had a negative impact on OS; no mutation impacted OS favorably; however, in a cox multivariate analysis that included clinical data, mutations in DNMT3A, ASXL1, CBL, EZH2 and TP53 became significant. Thus, controlling for clinical variables is important when interpreting genomic data in AML.
Similar content being viewed by others
Introduction
Acute myeloid leukemia (AML) is a heterogeneous clonal disorder characterized by the acquisition of chromosomal abnormalities and somatic mutations that drive disease phenotype, progression, and resistance to therapies.1, 2, 3 In the last four decades, the discovery of chromosomal abnormalities such as balanced translocations and inversions has illuminated the pathogenesis of AML and confirmed the genetic basis of the disease. Since then, cytogenetic information, along with other clinical variables such as age, disease phenotype (primary AML (pAML) vs secondary AML (sAML), and white blood cell count (WBC) at diagnosis have been used to risk stratify patients.3, 4 Approximately 50% of AML patients have normal karyotype (NK) without any evidence of the structural abnormalities that have subsequently been detected by higher resolution technologies such as high-density comparative genomic hybridization or single-nucleotide polymorphism arrays.5, 6, 7
Advances in genomic technologies have increasingly highlighted the remarkable complexity of genetic and epigenetic alterations in AML.1, 2, 3 Whole genome sequencing studies have identified at least one driver mutation in almost every sample from AML patients, with an average of ~13 mutations per sample.1 Some mutations, such as NPM1, FLT3, CEBPA and DNMT3A, are more common and have been shown to impact overall survival (OS) whereas other mutations, such as IDH1, IDH2 and TET2, occur in a lower frequencies without a clear impact on OS.8 Further, other mutations, such as SF3B1, SRSF2, U2AF1, ZRSR2, ASXL1, EZH2, BOCR and STAG2, occur more specifically in patients with sAML compared with pAML, and can be used to define disease phenotype.9 Whether the specificity of these mutations is retained in all subtypes of AML, such as in patients with complex karyotype or unfavorable risk cytogenetics, has not been established. Further, controversies regarding the impact of somatic mutations on disease phenotype and OS may be related to several factors, including a small sample size in some studies, a small number of genes tested in a given panel, and the lack of careful evaluation of the impact of these mutations on outcome and disease phenotype in the setting of clinical variables such as age, cytogenetics and WBC, that inform prognosis and determination of therapeutic options. Further, some of these studies only included younger patients who received intensive chemotherapy and the application of the results of these studies in older adults who are not eligible to receive such therapy is not established.
In this study, we investigated the interplay between genomic and clinical information in a large cohort of patients with pAML and sAML using a genomic panel of the most recurrent somatic mutations in myeloid malignancies.
Methods
Patients
Clinical and mutational data for patients diagnosed with sAML and pAML according to 2008 World Health Organization criteria and treated at Cleveland Clinic between 1–2003 and 1–2013 were included.10 sAML was defined by histological interpretation of bone marrow biopsy specimens in conjunction with documentation of antecedent myelodysplastic syndromes, aplastic anemia, myelproliferative neoplasm or chronic myelomonocytic leukemia, by experienced hematopathologists not associated with this study. The pAML cohort consisted of 79 patients with complete mutational and clinical data that were randomly selected from our samples database and 168 patients from TCGA atlas (publicly available data).1 Patients with AML and t(15,17), t(8;21)(q22;q22), t(16,16) and inv(16)(p13q22) were excluded since the number of cases was not enough to derive a meaningful conclusions.
Conventional cytogenetic analyses were performed on bone marrow samples obtained at diagnosis by culturing bone marrow cells for 24–48 h using standard techniques. An abnormality was considered clonal when at least 2 metaphases had the same abnormalities, in accordance with the International System of Human Cytogenetic Nomenclature (ISCN 2005).11 Cytogenetic risk grouping was per CALGB/Alliance criteria.4
All samples and clinical data were collected with patient consent and this study was performed under an Institutional Review Board approved protocol in accordance with the Declaration of Helsinki.
Sample processing, DNA sequencing and mutational analysis
DNA was extracted from peripheral blood or bone marrow mononuclear cells in blood samples that were stored at the Stem Cell Tissue Bank at the Cleveland Clinic. Direct sequencing was performed on coding exons of 62 genes (Supplementary Data) using Illumina TrueSeq Custom Amplicon kit. For germ-line confirmation, mutations were analyzed in non-clonal CD3þ cells whenever DNA was available. Bidirectional sequencing was performed by standard techniques using an ABI 3730xl DNA analyzer (Applied Biosystems, Foster City, CA, USA). Putative variants were extracted using GATK3.3 pipeline, following recommended best practices for variant discovery. Variants with at least 10 positive reads and variate allelic frequency of 5% were prioritized for further processing and annotation. VCF files generated were used as an input for Annovar and were annotated with multiple databases (dbSNP138, COSMIC, ExacDb). Variants found in ExacDb with allelic frequency >0.0001 were excluded. Mutations in NPM1 and CEBPA, FLT3 were also tested using standard methods. The sequencing method of the pAML cohort from TCGA database is described previously.1
Treatment
Patients were not treated uniformly since our cohort included elderly patients who were not eligible to receive intensive chemotherapy or allogeneic stem cell transplant. A total of 160 patients received standard induction chemotherapy with cytarabine × 7 days +3 days of anthracycline, 53 received a hypomethylating agent +/− combination, 26 were treated on a clinical trial, 19 received low dose cytarabine, 42 received other treatment modalities (such as hydroxyurea and supportive care only) and 168 TCGA patients whom their treatments was not reported in the database.1
Statistical analysis
Continuous and categorical variables were compared using Wilcoxon rank sum test and Fisher’s exact test. OS was calculated from the date of diagnosis to date of last follow-up or death. A logistic regression and Cox regression multivariate analyses that included all clinical variables and significant mutations were used whichever appropriate to compare the mutation distribution and the impact of mutations on OS between pAML and sAML, respectively. P-values were two-sided and considered significant at <0.05. All analyses were conducted using R package language.
Results
Patient characteristics
A total of 468 patients were included in the final analysis, 247 with pAML and 221 with sAML (Table 1). The median age for the entire cohort was 64 years (range, 18–100) and 222 patients (47.4%) had NK. Compared with patients with pAML, those with sAML were older (68 vs 60 years, P<0.001), had a lower WBC at presentation (3.85 vs 13 × 109 g/l, P<0.001), less likely to have NK cytogenetics (35.7 vs 57.9%, P<0.001), and more likely to have unfavorable cytogenetics (36.7 vs 23.9%, P=0.002), respectively (Table 1). The median follow-up for the entire cohort was 18.6 months (range, 0–114.1).
Mutation distribution in primary versus secondary AML
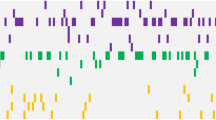
An average of two mutations/sample was observed (range, 0–18). Comparing the genetic landscape between pAML and sAML, distinct genomic alterations specific for each phenotype were observed (Figure 1a). Overall, mutations in FLT3, DNMT3A, NPM1 and IDH2 were more specific for pAML whereas 11 mutations (in UTAF1, STAG2, BCORL1, BCOR, EZH2, JAK2, CBL, PRPF8, SF3B1, ASXL1 and DHX29) were more specific for sAML (Figures 1a and b, Supplementary Data).
When the analysis was restricted to distinct cytogenetic subgroups, a different spectrum of mutations was found. In patients with NK, FLT3, DNMT3A and NPM1 remained specific for pAML, but only UTAF1, STAG2, BCORL1, BCOR, EZH2, JAK2, CBL, SF3B1 and ASXL1 retained their specificity for sAML, with mutations in MECOM, SETBP1 and RAD21 becoming significant for the sAML phenotype (Figure 2, Supplementary Data). Similarly, mutations specific to each AML phenotype differed in patients with complex karyotype, and in those with intermediate risk and unfavorable risk cytogenentics (Figure 2, Supplementary Data).
Similar results were obtained when other clinical variables, such as age (⩽60 years vs >60 years) and WBC (⩽15 000 vs >15 000) were taken into account (Figure 2, Supplementary Data).
In a logistic regression multivariate analysis that included clinical and mutational variables, only two mutations—FLT3 and DNMT3A—remained specific for pAML, whereas only four mutations—EZH2, BCOR, SF3B1 and ASXL1—were specific for sAML (Table 2).
Impact of mutations on overall survival
The median OS for patients with sAML was shorter compared with those with pAML (7.4 (range, 0–82.2) vs 11.2 (range, 0–114.1) months, respectively, P 0.001).
When the impact of mutations on OS was evaluated in the entire cohort, mutations in DNMT3A (hazard ratio (HR), 1.49, P.003), PRPF8 (HR, 1.69, P.04), ASXL1 (HR, 1.83, P<0.001), CBL (HR, 1.97, P.017), EZH2 (HR, 2.37, P.001) and TP53 (HR, 3.05, P<0.001) had a deleterious effect on OS, and no mutation was associated with improved OS (Figure A–C, Supplementary Data).
When cytogenetic grouping was taken into account, different mutations had a different impact on OS (Figure 3, Supplementary Data). In patients with NK, mutations in FLT3, DNMT3A, APC, ASXL1, SETBP1 and CBL had a negative impact on OS, whereas only DNMT3A mutations had a negative impact on OS in patients with unfavorable cytogenetics (Figure 3). Interestingly, mutations in NPM1 and CEBPA lost their favorable impact on OS in patients with NK, likely because the analysis included patients with both pAML and sAML. Similarly, the impact of mutations on OS changed in patients ⩽60 year-old vs >60 and patients with WBC ⩽15000 vs >15000 (Figure 3, Supplementary Data).
In a cox multivariate analyses that included clinical and mutational data, only mutations in DNMT3A, ASXL1, EZH2 and TP53 retained their prognostic impact (Table 3). However, when the analysis was restricted to the patients with available treatment data and treatment history (intensive vs non-intensive) was added to the multivariate analysis, all mutations retained their prognostic impact on OS (Supplementary Data) suggesting that intensive chemotherapy did not overcome the negative impact of these mutations on OS.
Discussion
AML is a heterogeneous clonal neoplasm characterized by complex genomic alterations that drive disease progression and resistance to therapy. Traditionally, sAML had been defined in part by the existence of an antecedent hematologic disorder such as myelodysplastic syndromes or myelproliferative neoplasm, whereas pAML arises de novo. Genomic discoveries have revealed the extent of genetic heterogenecity in AML, and have enabled better risk stratification, particularly in patients with NK. Further, certain somatic mutations are highly specific for disease phenotype (pAML vs sAML) and can in some cases define disease ontogeny, distinguishing sAML from pAML even in the absence of a confirmed preceding hematologic condition. Genomic abnormalities cannot be used in a vacuum, however, as been highlighted by divergent results regarding the impact of some mutations on OS in a subgroup of patients with AML, and the lack of reproducibility of several models that were developed to risk stratify AML patients. This raises the issue of the optimal way to interpret genomic information in AML in conjunction with traditional risk stratification tools that use clinical variables, such as age, conventional cytogenetics and treatment intensity especially when the analysis include older adults with AML who are not eligible to receive intensive chemotherapy.
In this study, we explored the interplay of established prognostic clinical variables and somatic mutations that are associated with disease phenotype and prognosis. We identified 11 mutations that are highly specific for sAML and four specific for pAML. These mutations were largely consistent with prior reports.9 However, when we investigated the specificity of these mutations to define disease phenotype (sAML vs pAML) in AML subtypes characterized by age, conventional cytogenetics and WBC at presentation the specificity of the mutations was lost. For example, in patients with a complex karytope at presentation, none of the somatic mutations were specific for either sAML or pAML, suggesting morphological assessment of the bone marrow biopsy by experienced hematopathologist and a history of an antecedent hematologic disorder is still needed to define disease phenotype. Moreover, when mutations were combined with clinical variables in a logistic regression multivariate analysis, only four mutations (EZH2, BCOR, SF3B1 and ASXL1) remained specific for sAML whereas three mutations (FLT3, NPM1 and DNMT3A) remained specific from pAML. In a study of 194 patients with secondary and therapy-related AML, Lindsley et al.9 showed that the presence of mutations in SRSF2, SF3B1, U2AF1, ZRSR2, ASXL1, EZH2, BCOR or STAG2 were >95% specific for the diagnosis of sAML, and only NPM1 mutations were specific for pAML. Incorporation of clinical variables, and inherent differences in patients enrolled in a formal clinical trial vs capture of all-comers, likely contributed to differences between the studies.
When we investigated the impact of recurrent somatic mutation on OS in the entire group, including patients with pAML and sAML, we found that six mutations (DNMT3A, PRPF8, ASXL1, CBL, EZH2 and TP53) had a negative impact on OS, and none had a favorable effect. Similarly, the impact of mutations on OS changed when clinical variables were taken into account. In a Cox regression multivariate analysis that included significant mutations and clinical variables, only four mutations (DNMT3A, ASXL1, EZH2 and TP53) retained their negative impact on OS even when treatment intensity was added as a covariate. This suggests that treatment with intensive chemotherapy may not overcome the negative impact of these mutations.
Prior evidence suggests that the impact of mutations on OS differ in subgroups of patients with AML. For example, NPM1 mutations have a positive impact on OS in patients with NK AML, whereas their impact on OS is lost in patients with unfavorable risk cytogenetics. Similarly, FLT3-ITD mutations have a negative impact on OS in patients with NK, but their impact on survival is lost in patients with unfavorable risk cytogenetics. The impact of multiple mutations on overall outcome differ between younger and elder patients with AML. To add to the complexity of interpreting this information, mutations occur in the context of other mutations, which may affect outcome. For example, Patel et al.12 showed that NPM1 mutations have a positive impact on OS only when they occur with IDH1 or IDH2 mutations and in the absence of FLT3-ITD mutations. Although the authors developed a genomic model that could be used to risk stratify patients with pAML a subsequent study could not validate it in a cohort of patients treated at a single institution.13 To date, there is no accepted and validated model that includes mutational data, and most clinical decisions are based on clinical variables along with only three mutations: NPM1, FLT-3ITD and CEBPa in patients with NK. This may evolve as novel methods such as machine learning and the ability to take advantage of large sets of databases are explored in the future.
In conclusion, clear genomic variations exist between sAML and pAML. The interpretation of genomic information should take into account clinical information along with coexistent mutations. A novel method to incorporate all this information is needed to further accurately predict AML phenotype and prognosis.
References
Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N Eng J Med 2013; 368: 2059–2074.
Ding L, Ley TJ, Larson DE, Miller CA, Koboldt DC, Welch JS et al. Clonal evolution in relapsed acute myeloid leukaemia revealed by whole-genome sequencing. Nature 2012; 481: 506–510.
Mrozek K, Heerema NA, Bloomfield CD . Cytogenetics in acute leukemia. Blood Rev 2004; 18: 115–136.
Byrd JC, Mrozek K, Dodge RK, Carroll AJ, Edwards CG, Arthur DC et al. Pretreatment cytogenetic abnormalities are predictive of induction success, cumulative incidence of relapse, and overall survival in adult patients with de novo acute myeloid leukemia: results from Cancer and Leukemia Group B (CALGB 8461). Blood 2002; 100: 4325–4336.
Walter MJ, Payton JE, Ries RE, Shannon WD, Deshmukh H, Zhao Y et al. Acquired copy number alterations in adult acute myeloid leukemia genomes. Proc Natl Acad Sci USA 2009; 106: 12950–12955.
Bullinger L, Kronke J, Schon C, Radtke I, Urlbauer K, Botzenhardt U et al. Identification of acquired copy number alterations and uniparental disomies in cytogenetically normal acute myeloid leukemia using high-resolution single-nucleotide polymorphism analysis. Leukemia 2010; 24: 438–449.
Suela J, Alvarez S, Cigudosa JC . DNA profiling by arrayCGH in acute myeloid leukemia and myelodysplastic syndromes. Cytogenet Genome Res 2007; 118: 304–309.
Grimwade D, Ivey A, Huntly BJ . Molecular landscape of acute myeloid leukemia in younger adults and its clinical relevance. Blood 2016; 127: 29–41.
Lindsley RC, Ebert BL . The biology and clinical impact of genetic lesions in myeloid malignancies. Blood 2013; 122: 3741–3748.
Vardiman JW, Thiele J, Arber DA, Brunning RD, Borowitz MJ, Porwit A et al. The 2008 revision of the World Health Organization (WHO) classification of myeloid neoplasms and acute leukemia: rationale and important changes. Blood 2009; 114: 937–951.
International Standing Committee on Human Cytogenetic Nomenclature., Shaffer LG, Tommerup N ISCN 2005. an international system for human cytogenetic nomenclature (2005): recommendations of the International Standing Committee on Human Cytogenetic Nomenclature. Basel Farmington, CT: Karger.
Patel JP, Gonen M, Figueroa ME, Fernandez H, Sun Z, Racevskis J et al. Prognostic relevance of integrated genetic profiling in acute myeloid leukemia. N Engl J Med 2012; 366: 1079–1089.
Sloan CE, Luskin MR, Boccuti AM, Sehgal AR, Zhao J, Daber RD et al. A modified integrated genetic model for risk prediction in younger patients with acute myeloid leukemia. PLoS One 2016; 11: e0153016.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
The study was presented as an oral presentation at the 57th Annual Meeting of the American Society of Hematology, Orlando, 2015.
Author contributions
AN, JPM and MAS designed the study, analysed and interpreted the data, wrote and approved the manuscript. AZ, KA, TR, CH, BP and MC collected the data, analysed the samples, performed statistical analysis, reviewed and approved the manuscript. MK, HEC and SRS contributed patients, analysed the data, reviewed and approved the manuscript.
Supplementary Information accompanies this paper on Blood Cancer Journal website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Nazha, A., Zarzour, A., Al-Issa, K. et al. The complexity of interpreting genomic data in patients with acute myeloid leukemia. Blood Cancer Journal 6, e510 (2016). https://doi.org/10.1038/bcj.2016.115
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bcj.2016.115
This article is cited by
-
Acute myeloid leukemia in a father and son with a germline mutation of ASXL1
Biomarker Research (2018)
-
Herausforderungen in der Behandlung älterer Patienten mit akuter myeloischer Leukämie
Der Onkologe (2017)