Abstract
Recent advancements in genome-wide analyses and RNA-sequencing technologies led to the discovery of small noncoding RNAs, such as microRNAs (miRs), as well as both linear long noncoding RNAs (lncRNAs) and circular long noncoding RNAs (circRNAs). The importance of miRs and lncRNAs in the treatment, prognosis and diagnosis of cardiovascular diseases (CVDs) has been extensively reported. We also previously reviewed their implications in therapies and as biomarkers for CVDs. More recently, circRNAs have also emerged as important regulators in CVDs. CircRNAs are circular genome products that are generated by back splicing of specific regions of pre-messenger RNAs (pre-mRNAs). Growing interest in circRNAs led to the discovery of a wide array of their pathophysiological functions. CircRNAs have been shown to be key regulators of CVDs such as myocardial infarction, atherosclerosis, cardiomyopathy and cardiac fibrosis. Accordingly, circRNAs have been recently proposed as potential therapeutic targets and biomarkers for CVDs. In this review, we summarize the current state of the literature on circRNAs, starting with their biogenesis and global mechanisms of actions. We then provide a synopsis of their involvement in various CVDs. Lastly, we emphasize the great potential of circRNAs as biomarkers for the early detection of CVDs, and discuss several patents and recent papers that highlight the utilization of circRNAs as promising biomarkers.
Similar content being viewed by others
Introduction
Cardiovascular disease (CVD) is a major killer of the human population in the USA and the world. It is estimated that approximately 100 million American adults (>1 in 3) have ≥1 type of CVD. A total of 11.5% of American adults (27.6 million) have been diagnosed with heart disease. By the year 2030, 43.9% of the US population is predicted to have some form of CVD1,2. With CVD being such a huge burden, advancements in disease management have been directed towards not only the treatment of such diseases, but also the development of platforms for early detection and the possible prevention of CVDs. Over the years, multiple CVD-related biomarkers have been discovered and utilized in clinical settings. According to the American Association for Clinical Chemistry (AACC), common cardiac biomarkers extensively used in the clinic are cardiac troponin and creatine kinase (CK). Cardiac troponin diagnoses myocardial infarction; its levels are elevated in the blood within 3–4 h upon cardiac injury and remain elevated for 10 to 14 d3. CK is mostly used to diagnose a second myocardial infarction occurring shortly after a first one. Blood CK levels are elevated 3–6 h after cardiac injury and remain elevated for 48 to 72 h unless a new injury occurs4.
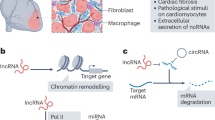
With the advancements in technology and the development of deep RNA-sequencing, new components of the genome have been unveiled, including noncoding RNAs. These are regions of the genome that do not code for proteins, but can regulate the function of genes and therefore modulate a vast number of physiological and pathological processes. Based on their function, length and structure, noncoding RNAs are divided into transfer RNAs (tRNAs; 74–95 bp), ribosomal RNAs (rRNAs; 121–5000 bp), small nuclear RNAs (snRNAs; 100–300 bp), small nucleolar RNAs (snoRNAs; 100–300 bp), guide RNAs (gRNAs; 55–70 bp), microRNAs (miRNAs or miRs; 19–23 bp), piwi-interacting RNAs (piRNAs; 24–30 bp), small interfering RNAs (siRNAs; 21–25 bp), long noncoding RNAs (lncRNAs; >200 bp and linear), and circular noncoding RNAs (circRNAs; >200 bp and circular)5. tRNAs were discovered to transfer amino acids to ribosomes during protein synthesis. rRNAs are ribosome components and are directly involved in protein synthesis. snRNAs process mRNA precursors (pre-mRNAs), leading to pre-mRNA splicing and maturation. snoRNAs mediate chemical modifications of other RNAs. gRNAs participate in RNA editing. piRNAs play roles in gametogenesis, transposon silencing, translation suppression, and epigenetic regulation. siRNAs silence complementary target mRNAs. miRs negatively regulate target genes by translational inhibition or promoting mRNA degradation. lncRNAs activate or inhibit gene expression by transcriptional, post-transcriptional, and miR sponge mechanisms, as well as through epigenetic mechanisms to regulate chromatin remodeling. circRNAs regulate alternative splicing and parental gene expression, while also functioning as competing endogenous RNAs or miR sponges6. Among the multiple noncoding RNAs, miRs, lncRNAs and circRNAs, in this listed order, have been extensively studied in various pathophysiological processes. Since their discovery, they have become the interest of a large group of researchers and health providers for their tremendous potential as future therapeutics and disease biomarkers.
We have reported some cardioprotective miRs, such as miR-199a-3p, miR-2147, miR-1508, miR-125b-5p9, and miR-532-5p10. We have also published multiple reviews on the roles of miRs and lncRNAs, including a review discussing the roles of lncRNAs in CVDs11 and another review summarizing the literature on the crosstalk between lncRNAs and miRs in health and disease12.
Viereck et al also extensively reviewed the literature and provided the current state of knowledge on circulating noncoding RNAs as biomarkers for cardiovascular disease and injury. Even though the authors mainly focused on miRs and lncRNAs, they briefly discussed circRNAs that could be used as biomarkers for CVDs13. This could be attributed to the fact that circRNAs were more recently identified as important regulators of health and disease; thus, there has not been enough literature showing their involvement in cardiovascular health and disease. The involvement of circRNAs in cardiovascular functions and diseases has been increasingly reported14. For example, a recent study by Tan et al performed RNA-sequencing on human and mouse hearts as well as on cardiomyocytes derived from embryonic stem cells, and identified over 10 000 circRNAs in the heart15. CircRNAs have also been shown to be abundant in tissues and bodily fluids, and are more stable than other noncoding RNAs because their circularization protects them from endonuclease activities16. Many review articles including the three described above11,12,13 have summarized the importance of miRs and lncRNAs in CVDs. Thus, in this manuscript, we will mainly focus on reviewing studies that identify circRNAs as potential therapies and biomarkers for CVDs; we will also report on the importance of crosstalk between circRNAs and miRs in CVDs.
Circular RNAs as potential therapies and biomarkers for cardiovascular diseases
Circular RNA biogenesis and mechanisms of actions
Biogenesis of circular RNAs
Pre-mRNAs are the precursors of circRNAs, similar to linear mRNAs. Pre-mRNAs are generated by RNA polymerase II (RNA Pol II). If the pre-mRNA is destined to form a linear mRNA, the 2′-OH of a branch point located on an adenine near the 3′-end of an intron performs a nucleophilic attack on the 5′-splice site, forming an intronic lariat and the free 3′-OH of the upstream exon. The free 3′-OH then initiates a nucleophilic attack on the 3′-splice site, joining the exons in the genomic order and releasing the intronic lariat17.
However, if the pre-mRNA is destined to become a circRNA, the pre-mRNA undergoes a process called “back splicing”. Back splicing is different from the “canonical splicing” that forms linear mRNAs (Figure 1). The exact process of back splicing and the formation of circRNAs are not clearly understood. However, a study proposed that the branch point in the upstream intron attacks the 5′-splicing site in the downstream intron, forming a lariat containing the exons18,19,20.
Biogenesis of circular RNAs. Different from linear mRNAs, which are formed by canonical splicing and cutting away introns of the pre-mRNAs using small nuclear ribonucleoproteins (snRNPs), circRNAs are formed by back splicing of the pre-mRNAs and circularization of the cut segment, where the 5' end joins the 3' end.
Another study suggested the possibility of co-transcriptional back splicing attributed to the following two reasons: (i) nascent chromatin-bound circRNAs17 and (ii) a change in the ratio of linear mRNAs to circRNAs for individual genes due to changes in the co-transcriptional splicing ratio of RNA Pol II17. Although it is not very well understood what could be the trigger for back splicing, two hypotheses have been proposed: (i) Back splicing is associated with Alu elements (approximately 2-fold more) within the intronic regions flanking the exons that form circRNAs. These intronic regions are not only rich in Alu elements, but they are also relatively longer than introns flanking linear mRNA exons21. (ii) More recently, a study showed that for back splicing, the 5'-splice site should be rich in the 7-nucleotide GU element, while the branch point should be rich in the 11-nucleotide C element22.
Depending on their genomic locations, circRNAs are divided into one exonic, multiple exonic, intronic, exon-intronic, intergenic and antisense circRNAs. The expression of circRNAs is tissue- and developmental stage-specific as well as is relatively stable because of their closed ring structure, which prevents them from being degraded by RNA exonucleases23. Distinct from classical protein-coding linear mRNAs, circRNAs play regulatory roles by acting as miR sponges24. Moreover, circRNAs compete with the linear splicing of pre-mRNAs and regulate the transcription of parental genes18,25. In the following section, we will further discuss the unique roles of circRNAs and how they regulate cardiovascular pathology.
Global mechanisms of circular RNA function
MiRNA sponge:
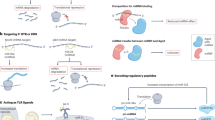
Cytoplasmic circRNAs are shown to contain binding sites for miRNAs, which are also called miRNA response elements (MREs). This makes circRNAs function as competing endogenous RNAs (ceRNAs)26. Because of the ability of circRNAs to bind to miRNAs, they are considered miRNA sponges23. Once bound to circRNAs, miRNAs would not be able to bind to their target genes (Figure 2). Because miRNAs exert an inhibitory effect on their target genes, when miRNAs are sponged by circRNAs, their target genes would be upregulated. CircRNAs can possess MREs for multiple miRNAs and can also possess multiple binding sites for an individual miRNA. For example, murine sex-determining region Y (Sry) possesses more than 70 binding sites for miR-7 and has 16 binding sites for miR-13824.
Mechanisms of actions of circular RNAs. CircRNAs exert their functions through different mechanisms of actions. Some circRNAs bind to miRNAs acting as miRNA sponges, while others bind to proteins acting as protein sponges. Some nuclear circRNAs regulate the transcription of host genes. Lastly, some circRNAs can even code for proteins. IRES: internal ribosome entry site.
Here, we seek to summarize the current knowledge on the interactions of circular RNAs and downstream miRNAs, and to discuss how they work together to regulate the pathogenesis of various CVDs (Table 1). In a paper by Geng et al, it was shown that the circRNA CDRlas regulated apoptosis by acting as a sponge for anti-apoptotic miR-7, leading to increased apoptosis27. For cardiac fibrosis, circRNAs CircRNA_000203 and CircRNA_010567 have been shown to regulate cardiac fibrosis by sponging miR-26b-5p and miR-141, respectively (Table 1). It is also known that CircRNA_000203 activates cardiac remodeling and that CircRNA_010567 increases the rate of myocardial fibrosis by inhibiting miR functions28,29. CircRNA HRCR can also sponge the deleterious miR-223, leading to cardioprotection against heart failure30. A more recent study showed that mitochondrial fission and apoptosis-related circRNA (MFACR) regulates mitochondrial fission and apoptosis in the heart by downregulating miR-652-3p and subsequently upregulating mitochondrial protein 18 kDa (MTP18)31.
The interactions between circRNAs and miRs are not only important for cardiac diseases but also play a role in vascular diseases (Table 1). For example, it was reported that circRNA CircR-284 regulates the pathogenesis of atherosclerosis by targeting miR-221. By this miR sponging mechanism, CircR-284 increases the risk for plaque rupture32. A recent study also showed that circRNAs promote the expression of transient receptor potential cation channel subfamily M member 3 (TRPM3) by inhibiting miR-130a-3p in coronary artery disease patients35. Interestingly, a systematic study on circRNA expression profiles in human adult and fetal tissues suggests that circRNAs exhibit their roles in a tissue- and development-specific manner and that investigation of the circRNA-miR-mRNA network will provide important insights into therapeutic and diagnostic targets36.
Protein sponge:
CircRNAs not only possess miRNA-binding elements but also possess binding sites for proteins. When circRNAs bind to their target proteins, this will lead to the inhibition of protein function (Figure 2). An example is muscle blind (MBL) protein, which is a critical factor for the alternative splicing of mbl pre-mRNA. CircMbl possesses binding sites for MBL protein and can bind to that protein, leading to inhibition of mbl pre-mRNA alternative splicing. It is worth noting that there is a feedback loop mechanism in this process as proposed by Ashwal-Fluss et al. This feedback loop entails that when MBL protein is abundant, mbl pre-mRNA is spliced into circMbl, which binds to MBL and decreases its bioavailability18.
Transcriptional host gene regulation:
CircRNAs are released into the cytoplasm and even extracellularly to have the potential to serve as biomarkers. However, to a much lesser extent, a few circRNAs remain in the nucleus. These nuclear circRNAs can react with RNA Pol II in the promoter region of host genes (Figure 2) and regulate transcription22,37. Examples of these circRNAs are exon-intron circRNAs (EIciRNAs) and circular intronic RNAs (ciRNAs). EIciRNAs such as CircEIF3J and CircPAIP2 possess binding sites for the U1 small nuclear ribonucleoprotein (snRNP) in the nucleus and have the ability to bind to U1 snRNP, forming the EIciRNA-U1 snRNP complex. This complex can then interact with RNA Pol II in the gene promoter region, which enhances the transcription of their parental genes25,37.
Protein coding:
For a gene to be translated into a protein, the formation of a translation initiation complex is critical. The assembly of this complex requires the recognition of the 5'-cap structure at the 5'-end of mRNAs38. CircRNAs do not possess a 5'-cap, and therefore do not normally code for proteins. However, CircZNF609 in human muscle cells39 and CircMbl in fruit fly head40 have been recently reported to encode proteins. Both of these circRNAs were able to interact with ribosomes and code for proteins in a 5'-cap-independent manner (Figure 2). The mechanism for 5'-cap-independent protein coding was explained by Yang et al as occurring as a result of N6-methyladenosine (m6A) modifications near the circRNA start codon. This m6A modification can act as an internal ribosome entry site (IRES) and lead to translation initiation41.
Identification of circular RNAs as important regulators of cardiovascular development and disease
CircRNAs were initially discovered in the early 1990s42 and were thought to be generated by RNA splicing errors or be splicing process by-products at that time. In 2013, circRNAs were revealed to function as molecular sponges for miRs24, making circRNAs a new hotspot in the noncoding RNA field after miRs and lncRNAs.
Due to advanced sequencing technologies and data analysis methods, a growing number of circRNAs have been identified in human and murine hearts15,43,44. The first circRNA profiling study discovered 575 circRNAs from adult murine hearts and demonstrated that several circRNAs are involved in CVDs43. The systemic analysis of circRNA expression from rats (neonatal vs adult), mice (sham vs transverse aortic constriction) and humans (non-failing vs failing) identified over 9,000 circRNAs from each species44. More recently, a detailed annotation and analysis of genome-wide circRNA expression reported 15 318 circRNAs in human hearts15. Even though the aforementioned studies are descriptive and somehow coarse in their designs, they have provided a very important basis for the discovery of novel circRNAs that could be translated clinically as either disease biomarkers or therapeutic agents for CVDs.
There has been significant effort in establishing bioinformatics tools to obtain insights into the roles of the identified cardiac circRNAs. For example, Liu et al reported a circRNA database (http://circnet.mbc.nctu.edu.tw/)45, in which researchers could access information about tissue-specific circRNA expression profiles and target information for circRNAs of interest45,46. Several other circRNA databases have also been constructed, including circBase (http://circbase.org/)47, CircRNABase (http://starbase.sysu.edu.cn/mirCircRNA.php)48, Circ2Traits (http://gyanxet-β.com/circdb/)49, and CircInteractome (http://circinteractome.nia.nih.gov)50. These databases provide excellent platforms to facilitate further functional research on circRNAs. However, the predictive bioinformatics data herein should be experimentally validated before pursuing any functional studies. For example, a novel circRNA should be subjected to extensive validation of its circularity using RNase R digestion, assessment by PCR using divergent primers and Sanger sequencing of the junction region. Quantitative PCR analysis remains the most widely used technique to assess the levels of circRNAs. In the next section, we will discuss how circRNAs are identified and assessed in various CVDs by distinct diseases.
Circular RNAs in cardiac development
The importance of noncoding RNAs in the gene regulatory network of cardiac development has been studied and demonstrated in multiple studies51,52,53. It has been recently shown that not only does alternative splicing regulate cardiac development, but back splicing can also play a critical role in cardiac development54. A study by Werfel et al also identified that circRNAs are highly expressed during cardiac development using cardiac tissues from humans and rats as well as human embryonic stem cells that were differentiated to cardiomyocytes. Even though this study was descriptive in nature and lacked mechanistic details, it was able to identify some circRNAs that potentially play important roles in cardiac development. According to the study, the most abundantly expressed circRNAs in the heart tissues were circRNA slc8a1 (Table 2) and circRNA rhobtb344. Another study by Tan et al showed similar results15. Interestingly, Slc8a1 is known to be a sodium/calcium exchanger that is critical for heart development and contraction55. Rhobtb3 is a Rab9-regulated ATPase required for endosome to Golgi transport and is highly expressed in the heart56. A study focused on a human induced pluripotent stem cell-derived cardiomyocyte (hiPSC-CM) model of cardiomyopathy also uncovered a new signature of disease-relevant circRNAs, including circRNAs from genes that are known regulators in cardiac development and function (eg, atxn10, chd7, dnajc6 and slc8a1)57. Lastly, a recent functional enrichment study showed that circRNAs play important roles in pathways that are specifically activated during cardiac differentiation58.
Circular RNAs in vascular development
CircRNAs' roles in embryonic development extend to vascular development. A study by Boeckel et al demonstrated that circRNA cZNF292 has a critical role in promoting angiogenesis (Table 2). CircRNA cZNF292 is expressed in endothelial cells in the human umbilical vein, microvessels and aorta. When endothelial cells were exposed to hypoxia, the expression of circRNA cZNF292 was increased, while in vitro silencing of cZNF292 inhibited the normal functional characteristics of endothelial cells, such as tube formation and spheroid sprouting, suggesting pro-angiogenic roles for circRNA cZNF29259.
Roles of circular RNAs in myocardial infarction
Myocardial infarction (MI) is defined as damage and remodeling of cardiac tissue in response to decreased blood flow in coronary arteries. It is usually predisposed by atherosclerosis and leads to the occlusion of coronaries. MI eventually leads to cardiac dysfunction, heart failure and ultimately death. MI treatment is directed towards reducing post-infarction cardiac remodeling, which prevents heart failure64. Since the discovery of noncoding RNAs, they have been regarded as potential therapeutics for MI65. Using microarray analysis, Wu et al 66 discovered 63 circRNA candidates, including circRNA CDR1as, that are differentially expressed in normal murine myocardial tissues compared to failing myocardial tissues due to MI. Simultaneously, another study by Geng et al 27 reported the circRNA CDR1as as a mechanism of action in MI injury (Table 2) and showed that MI induces the upregulation of CDR1as, which in turn promotes cardiomyocyte apoptosis by acting as a sponge for cardioprotective miR-7a (Table 1).
Roles of circular RNAs in ischemia-reperfusion injury
Ischemia-reperfusion injury is damage occurring to vital organs, such as heart, kidney and brain, due to the transient decrease in blood flow to these organs67. In a study by Mehta et al 68, the authors subjected mice to transient middle cerebral artery occlusion and performed circRNA expression profiling in the penumbral cortex at 6, 12, and 24 h of reperfusion using circRNA microarray analysis. They first found over 1000 circRNAs to be expressed at detectable levels from mostly exonic regions of genes in the cerebral cortex of sham animals. Among them, 283 circRNAs were expressed higher than 2-fold during at least at one of the reperfusion time points, whereas 16 were altered at all 3 reperfusion time points after the transient middle cerebral artery occlusion. The authors also used bioinformatics analysis to show that these 16 circRNAs contain binding sites for many miRs, although they did not experimentally validate the target miRs.
Roles of circular RNAs in cardiomyopathy
Cardiomyopathy, including hypertrophic and dilated cardiomyopathy, is a cardiac pathology that affects the myocardium69. Hypertrophic cardiomyopathy (HCM) occurs when the left ventricular myocardium becomes thicker than normal, leading to incomplete filling at diastole. This eventually leads to heart failure and death. Dilated cardiomyopathy (DCM) occurs when the left or right ventricles are impaired, leading to ventricular dilation, heart failure and death. Wang et al reported that the heart-related circRNA HRCR confers protective effects against myocardial hypertrophy and heart failure30. In this study, when mice were infused with isoproterenol and subjected to transverse aortic constriction, HRCR expression was downregulated (Table 2). Using bioinformatics prediction, co-immunoprecipitation and AGO2 immunoprecipitation analyses, the study demonstrated that HRCR interacts with miR-223, retains this miR in the cytoplasm, and inhibits the pro-hypertrophic activity of miR-223 (Table 1). Another study by Khan et al 61 reported abnormal circRNA splicing events during cardiomyopathy. Using RNA-sequencing analysis on heart samples from HCM and DCM patients, the authors identified over 800 back splicing junctions. They validated the profiling results using QRT-PCR analyses and confirmed that 10 exonic calcium/calmodulin-dependent protein kinase type II delta (camk2d) and titin circRNAs were associated with cardiomyopathy. Camk2d circRNAs were downregulated in both HCM and DCM patients, while titin circRNAs were downregulated only in DCM. A profiling and validation study also provided a comprehensive catalogue of RNase R-resistant 575 circRNAs in adult murine hearts that may play important roles in cardiomyopathy43.
Roles of circular RNAs in cardiac fibrosis
Cardiac fibrosis is a cardiac pathology characterized by the activation of cardiac fibroblasts, leading to the replacement of healthy myocardium with non-beating fibrous tissues70. Cardiac fibrosis occurs as a part in the pathogenesis of various CVDs, such as MI and diabetes mellitus. A study by Tang et al showed that CircRNA_000203 back spliced from the mouse myo9a gene is associated with cardiac fibrosis28. Diabetic mouse myocardium and angiotensin II-induced cardiac fibroblasts showed CircRNA_000203 upregulation (Table 2). The study also identified anti-fibrotic miR-26b-5p to be a direct target for circRNA_000203 (Table 1). If miR-26b-5p is sponged by CircRNA_000203, cardiac fibroblast proliferation is enhanced via the activation of collagen type I a2 (Col1a2) and connective tissue growth factor. CircActa2 has also been reported to regulate the expression and function of alpha smooth muscle actin (α-SMA; the main player in myocardial and endocardial fibrosis) via sponging miR-548f-5p71,72.
Roles of circular RNAs in cardiac senescence
The turnover of adult cardiomyocytes is very low and is not sufficient to overcome cardiac damage once they are differentiated into their mature forms. Cardiomyocytes also undergo pathological aging termed cardiac senescence. Du et al reported that exonic CircFoxo3 could induce cardiac senescence34. Exogenous CircFoxo3 could aggravate heart senescence and doxorubicin-induced cardiomyopathy. This could be reversed by knockdown of CircFoxo3 (Table 2). The study also provided mechanistic insights showing that CircFoxo3 promotes cardiac senescence by binding to inhibitor of differentiation 1 (ID1), transcription factor E2F1, focal adhesion kinase (FAK) and hypoxia-inducible factor 1α (HIF1α). Once bound to CircFoxo3, the aforementioned factors remain in the cytoplasm, leading to cardiac stress (Table 1).
Roles of circular RNAs in atherosclerosis
Atherosclerosis is a vascular disease caused by injury to the vascular endothelium that leads to the accumulation of cholesterol that eventually blocks the vessel. MI usually occurs when a blood clot blocks blood flow to the heart. Atherosclerosis could have a genetic predisposition due to its association with the ink4/arf locus on human chromosome 9p21, where several tumor suppressor genes (p16ink4a, p15ink4b and arf) and the lncRNA ANRIL are transcribed. Several circular transcripts of anril have been reported, and their expression is shown to correlate with atherosclerosis risk14,33. Holdt et al showed the molecular mechanism of one of the circular ANRIL isoforms62. CircANRIL impaired the maturation of ribosomes by binding to the 60S ribosomal assembly factor PES1, leading to nuclear stress and p53 activation (Table 2). This eventually results in the inactivation of smooth muscle cells and macrophages while inducing apoptosis. Thus, CircANRIL could be protective against atherosclerosis.
Circular RNAs as novel biomarkers for cardiovascular diseases
Of all noncoding RNAs, circRNAs could be the best candidates as future disease biomarkers for the following reasons. First, the circular 3'- and 5'-ends of circRNAs are not exposed to degradation by exoribonucleases (RNases), and thus their half-lives are longer upon extracellular release. This makes circRNAs more stable and abundant in extracellular fluids for easy detection. Second, deep RNA-sequencing studies have identified hundreds to thousands of cell- and/or tissue-specific circRNAs in human and mouse73. Third, circRNAs are present in the whole blood, plasma and extracellular vesicles such as exosomes60,74,75, indicating that most components of the blood (eg, peripheral blood mononuclear cells, exosomes, platelets, plasma and serum) can be used for biomarker studies.
For example, an interesting study on human platelets showed that circRNAs are enriched in platelets 14- to 26-fold relative to samples digested with RNase R to selectively remove linear RNAs, suggesting that linear RNAs decay more rapidly than circRNAs in platelets76. CircRNAs are also abundant in exosomes or microvesicles, which are secreted into the extracellular fluid by cells. The abundance of circRNAs in exosomes is positively correlated with their cellular expression77. Many studies have already reported circRNAs as biomarkers in some diseases, mainly cancer. For example, a study by Chen et al showed Circ_0000190 to be downregulated in human gastric cancers78, while Xu et al identified CircRNA ciRs-7 (CDR1as) as a risk factor for microvascular invasion in hepatocellular carcinoma79. CircRNAs not only serve as disease biomarkers for cancers but can also act as biomarkers for other diseases such as neurological disorders. For example, a study by Floris et al showed CircRNA_103636 to be downregulated in major human neurological disorders80. Another study by Iparraguirre et al showed that Circ_0005402 and Circ_0035560 are back spliced from anxa2 and downregulated in multiple sclerosis81. More recent work by Maass et al also highlighted the promise of circRNAs as tissue- and disease-specific markers82.
Like the aforementioned diseases, CVDs are associated with the dysregulated expression of circRNAs. Accordingly, there has been increasing interest in circRNAs as potential biomarkers for CVDs. A study by Zhao et al reported Circ_0124644 to be upregulated in the bloodstream in patients with coronary artery disease and proposed Circ_0124644 as a potential clinical biomarker for coronary artery disease63 (Table 2). Another study showed that a circular RNA called myocardial infarction-associated circular RNA (MICRA) back spliced from exon 1 of zinc finger protein 609 as a predictor of left ventricular dysfunction after acute MI60 (Table 2). In this case, the blood levels of MICRA in MI patients were lower than in healthy people. MI patients with relatively low levels of MICRA are considered at high risk of left ventricular dysfunction. A study by Bazan et al also described the role of CircRNA-284 as a biomarker for carotid plaque rupture32 (Table 1). CircRNA-284 is an inhibitor of miR-221 and miR-222, which are indicators of carotid plaque rupture. Thus, increased blood levels of CircRNA-284, and decreased blood levels of miR-221 and miR-222 could be utilized as a biomarker for carotid plaque rupture. Although there have been not many reports on circRNAs as potential biomarkers for CVDs, the utility of circRNAs as CVD biomarkers will significantly increase due to their superior stability compared with other noncoding RNAs.
Review of recent patents
Even though the identification of circRNAs as biomarkers for CVDs is pretty recent, one can already find multiple patents filed on multiple circRNAs as potential CVD biomarkers. In this section, we review some of these patents that present methods and techniques for the detection of aberrantly expressed circRNAs as well as their utilities for the diagnosis and prognosis of heart diseases. These circRNAs could be used for new types of biomarkers that lead to novel diagnostic, prognostic and predictive platforms.
Patents EP3054017 A1 and WO2016124655 A1 reported 16 circRNA transcripts that are correlated with CVDs, in particular, acute myocardial infarction (Table 3). It is proposed that the circRNA transcripts in these patents could serve as biomarkers for the diagnosis of cardiovascular disorders. The patents also provide diagnostic kits for the dysregulated circRNAs in patients with CVDs. The patents also discuss the effects of these circRNAs on endothelial cell sprouting and pathological angiogenesis. Lastly, the patents discuss that compounds or vascular tissue materials, which have been used to treat CVDs and endothelial dysfunction, could regulate the identified 16 circRNAs83,84. Patent WO2017046203 A1 (Table 3) also demonstrated Circ_0000615 (MICRA), Circ_0000005, Circ_0000673, Circ_0000585, Circ_0000816, Circ_0000817, Circ_0000917, Circ_0001423, Circ_0000540, Circ_0001844, Circ_0000994 and CircNPPA to be dysregulated in heart failure and myocardial hypertrophy, and thus can be used as biomarkers for these diseases. This patent claims the utility of the identified circRNAs in the diagnosis of heart failure and describes kits designed to diagnose MI using quantitative expression data for one or more of the identified circRNAs. Lastly, the patent discusses the current pharmacotherapy that regulates the identified circRNAs85. Finally, patent WO2012050975 A2 discovered circRNAs generated from one or more anril exons (Table 3). The patent also provided methods for detecting Circ-ANRIL in multiple disorders, such as atherosclerosis, and information about related kits. Lastly, the patent discusses methods for screening for compounds to prevent or treat such disorders86.
Conclusions and current & future developments
Very little is known about circRNAs and how they function and regulate multiple physiological and pathological processes. This certainly would delay the utilization of circRNAs in the clinical setting until enough knowledge is established about them16,87. All of the previous studies involving human subjects contain only a small cohort of patients or healthy subjects, making the studies inconclusive. Accordingly, the field is in need for studies with much larger cohorts that can provide more convincing data on pathophysiological roles of circRNAs and their potential utilities as biomarkers. Moreover, there is often a large discrepancy in the results between different groups and a lack of data reproducibility, which is mainly attributed to the absence of standardized methodologies for measuring the levels of circRNAs in body fluids and tissues. The aforementioned limitations make the translation of circRNAs as biomarkers for CVDs to the clinic somehow risky and not currently favorable for FDA approval.
However, circular RNAs are increasingly being recognized to play important roles in the evolving strategies for tackling CVDs as described above. The recent studies and patents in the circular RNA field described in this review also illustrate the tremendous promise of circRNAs in biomarker research. Although some circular RNA biomarkers for heart disease have been identified, there remain significant hurdles with respect to clinical use due to diagnostic limitations and off-target effects. More work is needed to better validate and integrate circRNAs into clinical practice. For example, many research directions may help deepen our knowledge of the roles of circRNAs in CVDs and addressing their value as biomarkers. (i) A detailed characterization of circRNA biogenesis is warranted. (ii) Profiling analyses focused on the temporal-, cell- and tissue-specific expression of circRNAs in healthy and diseased tissues would provide important insights, as the originating tissues for circulating circRNAs are unknown and it is possible that they can be taken up by other tissues, leading to secondary effects. (iii) A characterization of the full spectrum of circulating circRNAs in the bloodstream would guide in selecting candidate circRNAs for future biomarker studies. (iv) The identification of the best components of blood for circRNA detection would be needed for developing clinically applicable diagnostics for circRNAs. In conclusion, collaborative approaches between basic scientists and clinicians will be required to develop circRNA diagnostic or therapeutic modalities.
References
Benjamin EJ, Blaha MJ, Chiuve SE, Cushman M, Das SR, Deo R, et al. Heart disease and stroke statistics-2017 update: A report from the American Heart Association. Circulation 2017; 135: e146–e603.
Heidenreich PA, Albert NM, Allen LA, Bluemke DA, Butler J, Fonarow GC, et al. Forecasting the impact of heart failure in the United States: a policy statement from the American Heart Association. Circ Heart Fail 2013; 6: 606–19.
Wu AH, Christenson RH. Analytical and assay issues for use of cardiac troponin testing for risk stratification in primary care. Clin Biochem 2013; 46: 969–78.
Gohbara M, Hayakawa A, Akazawa Y, Furihata S, Kondo A, Fukushima Y, et al. Association between acidosis soon after reperfusion and contrast-induced nephropathy in patients with a first-time ST-segment elevation myocardial infarction. J Am Heart Assoc 2017; 6: e006380.
Esteller M. Non-coding RNAs in human disease. Nat Rev Genet 2011; 12: 861–74.
Gao J, Xu W, Wang J, Wang K, Li P. The role and molecular mechanism of non-coding RNAs in pathological cardiac remodeling. Int J Mol Sci 2017; 18: E608.
Park KM, Teoh JP, Wang Y, Broskova Z, Bayoumi AS, Tang Y, et al. Carvedilol-responsive microRNAs, miR-199a-3p and -214 protect cardiomyocytes from simulated ischemia-reperfusion injury. Am J Physiol Heart Circ Physiol 2016; 311: H371–83.
Tang Y, Wang Y, Park KM, Hu Q, Teoh JP, Broskova Z, et al. MicroRNA-150 protects the mouse heart from ischaemic injury by regulating cell death. Cardiovasc Res 2015; 106: 387–97.
Bayoumi AS, Park KM, Wang Y, Teoh JP, Aonuma T, Tang Y, et al. A carvedilol-responsive microRNA, miR-125b-5p protects the heart from acute myocardial infarction by repressing pro-apoptotic bak1 and klf13 in cardiomyocytes. J Mol Cell Cardiol 2018; 114: 72–82.
Bayoumi AS, Teoh JP, Aonuma T, Yuan Z, Ruan X, Tang Y, et al. MicroRNA-532 protects the heart in acute myocardial infarction, and represses prss23, a positive regulator of endothelial-to-mesenchymal transition. Cardiovasc Res 2017; 113: 1603–14.
Archer K, Broskova Z, Bayoumi AS, Teoh JP, Davila A, Tang Y, et al. Long non-coding RNAs as master regulators in cardiovascular diseases. Int J Mol Sci 2015; 16: 23651–67.
Bayoumi AS, Sayed A, Broskova Z, Teoh JP, Wilson J, Su H, et al. Crosstalk between long noncoding RNAs and microRNAs in health and disease. Int J Mol Sci 2016; 17: 356.
Viereck J, Thum T. Circulating noncoding RNAs as biomarkers of cardiovascular disease and injury. Circ Res 2017; 120: 381–99.
Li M, Ding W, Sun T, Tariq MA, Xu T, Li P, et al. Biogenesis of circular RNAs and their roles in cardiovascular development and pathology. FEBS J 2018; 285: 220–32.
Tan WL, Lim BT, Anene-Nzelu CG, Ackers-Johnson M, Dashi A, See K, et al. A landscape of circular RNA expression in the human heart. Cardiovasc Res 2017; 113: 298–309.
Memczak S, Papavasileiou P, Peters O, Rajewsky N. Identification and characterization of circular RNAs as a new class of putative biomarkers in human blood. PLoS One 2015; 10: e0141214.
Jurica MS, Moore MJ. Pre-mRNA splicing: awash in a sea of proteins. Mol Cell 2003; 12: 5–14.
Ashwal-Fluss R, Meyer M, Pamudurti NR, Ivanov A, Bartok O, Hanan M, et al. circRNA biogenesis competes with pre-mRNA splicing. Mol Cell 2014; 56: 55–66.
Liu J, Liu T, Wang X, He A. Circles reshaping the RNA world: from waste to treasure. Mol Cancer 2017; 16: 58.
Starke S, Jost I, Rossbach O, Schneider T, Schreiner S, Hung LH, et al. Exon circularization requires canonical splice signals. Cell Rep 2015; 10: 103–11.
Jeck WR, Sorrentino JA, Wang K, Slevin MK, Burd CE, Liu J, et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 2013; 19: 141–57.
Zhang Y, Zhang XO, Chen T, Xiang JF, Yin QF, Xing YH, et al. Circular intronic long noncoding RNAs. Mol Cell 2013; 51: 792–806.
Memczak S, Jens M, Elefsinioti A, Torti F, Krueger J, Rybak A, et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 2013; 495: 333–8.
Hansen TB, Jensen TI, Clausen BH, Bramsen JB, Finsen B, Damgaard CK, et al. Natural RNA circles function as efficient microRNA sponges. Nature 2013; 495: 384–8.
Li Z, Huang C, Bao C, Chen L, Lin M, Wang X, et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat Struct Mol Biol 2015; 22: 256–64.
Salmena L, Poliseno L, Tay Y, Kats L, Pandolfi PP. A ceRNA hypothesis: the Rosetta Stone of a hidden RNA language? Cell 2011; 146: 353–8.
Geng HH, Li R, Su YM, Xiao J, Pan M, Cai XX, et al. The circular RNA Cdr1as promotes myocardial infarction by mediating the regulation of miR-7a on its target genes expression. PLoS One 2016; 11: e0151753.
Tang CM, Zhang M, Huang L, Hu ZQ, Zhu JN, Xiao Z, et al. CircRNA_000203 enhances the expression of fibrosis-associated genes by derepressing targets of miR-26b-5p, Col1a2 and CTGF, in cardiac fibroblasts. Sci Rep 2017; 7: 40342.
Zhou B, Yu JW. A novel identified circular RNA, circRNA_010567, promotes myocardial fibrosis via suppressing miR-141 by targeting TGF-beta1. Biochem Biophys Res Commun 2017; 487: 769–75.
Wang K, Long B, Liu F, Wang JX, Liu CY, Zhao B, et al. A circular RNA protects the heart from pathological hypertrophy and heart failure by targeting miR-223. Eur Heart J 2016; 37: 2602–11.
Wang K, Gan TY, Li N, Liu CY, Zhou LY, Gao JN, et al. Circular RNA mediates cardiomyocyte death via miRNA-dependent upregulation of MTP18 expression. Cell Death Differ 2017; 24: 1111–20.
Bazan HA, Hatfield SA, Brug A, Brooks AJ. Lightell DJ Jr, Woods TC. Carotid plaque rupture is accompanied by an increase in the ratio of serum circR-284 to miR-221 levels. Circ Cardiovasc Genet 2017; 10: e001720.
Burd CE, Jeck WR, Liu Y, Sanoff HK, Wang Z, Sharpless NE. Expression of linear and novel circular forms of an INK4/ARF-associated non-coding RNA correlates with atherosclerosis risk. PLoS Genet 2010; 6: e1001233.
Du WW, Yang W, Chen Y, Wu ZK, Foster FS, Yang Z, et al. Foxo3 circular RNA promotes cardiac senescence by modulating multiple factors associated with stress and senescence responses. Eur Heart J 2017; 38: 1402–12.
Pan RY, Liu P, Zhou HT, Sun WX, Song J, Shu J, et al. Circular RNAs promote TRPM3 expression by inhibiting hsa-miR-130a-3p in coronary artery disease patients. Oncotarget 2017; 8: 60280–90.
Xu T, Wu J, Han P, Zhao Z, Song X. Circular RNA expression profiles and features in human tissues: a study using RNA-seq data. BMC Genomics 2017; 18: 680.
Chen LL. The biogenesis and emerging roles of circular RNAs. Nat Rev Mol Cell Biol 2016; 17: 205–11.
Chen CY, Sarnow P. Initiation of protein synthesis by the eukaryotic translational apparatus on circular RNAs. Science 1995; 268: 415–7.
Legnini I, Di Timoteo G, Rossi F, Morlando M, Briganti F, Sthandier O, et al. Circ-ZNF609 is a circular RNA that can be translated and functions in myogenesis. Mol Cell 2017; 66: 22–37 e9.
Pamudurti NR, Bartok O, Jens M, Ashwal-Fluss R, Stottmeister C, Ruhe L, et al. Translation of CircRNAs. Mol Cell 2017; 66: 9–21 e7.
Yang Y, Fan X, Mao M, Song X, Wu P, Zhang Y, et al. Extensive translation of circular RNAs driven by N6-methyladenosine. Cell Res 2017; 27: 626–41.
Capel B, Swain A, Nicolis S, Hacker A, Walter M, Koopman P, et al. Circular transcripts of the testis-determining gene Sry in adult mouse testis. Cell 1993; 73: 1019–30.
Jakobi T, Czaja-Hasse LF, Reinhardt R, Dieterich C. Profiling and validation of the circular RNA repertoire in adult murine hearts. Genomics Proteomics Bioinformatics 2016; 14: 216–23.
Werfel S, Nothjunge S, Schwarzmayr T, Strom TM, Meitinger T, Engelhardt S. Characterization of circular RNAs in human, mouse and rat hearts. J Mol Cell Cardiol 2016; 98: 103–7.
Liu YC, Li JR, Sun CH, Andrews E, Chao RF, Lin FM, et al. CircNet: a database of circular RNAs derived from transcriptome sequencing data. Nucleic Acids Res 2016; 44: D209–15.
Fan X, Weng X, Zhao Y, Chen W, Gan T, Xu D. Circular RNAs in cardiovascular disease: An overview. Biomed Res Int 2017; 2017: 5135781.
Glazar P, Papavasileiou P, Rajewsky N. circBase: a database for circular RNAs. RNA 2014; 20: 1666–70.
Li JH, Liu S, Zhou H, Qu LH, Yang JH. starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res 2014; 42: D92–7.
Ghosal S, Das S, Sen R, Basak P, Chakrabarti J. Circ2Traits: a comprehensive database for circular RNA potentially associated with disease and traits. Front Genet 2013; 4: 283.
Dudekula DB, Panda AC, Grammatikakis I, De S, Abdelmohsen K, Gorospe M. CircInteractome: A web tool for exploring circular RNAs and their interacting proteins and microRNAs. RNA Biol 2016; 13: 34–42.
Kalsotra A, Xiao X, Ward AJ, Castle JC, Johnson JM, Burge CB, et al. A postnatal switch of CELF and MBNL proteins reprograms alternative splicing in the developing heart. Proc Natl Acad Sci U S A 2008; 105: 20333–8.
Kalsotra A, Wang K, Li PF, Cooper TA. MicroRNAs coordinate an alternative splicing network during mouse postnatal heart development. Genes Dev 2010; 24: 653–8.
Giudice J, Xia Z, Wang ET, Scavuzzo MA, Ward AJ, Kalsotra A, et al. Alternative splicing regulates vesicular trafficking genes in cardiomyocytes during postnatal heart development. Nat Commun 2014; 5: 3603.
Szabo L, Morey R, Palpant NJ, Wang PL, Afari N, Jiang C, et al. Statistically based splicing detection reveals neural enrichment and tissue-specific induction of circular RNA during human fetal development. Genome Biol 2015; 16: 126.
Jordan MC, Henderson SA, Han T, Fishbein MC, Philipson KD, Roos KP. Myocardial function with reduced expression of the sodium-calcium exchanger. J Card Fail 2010; 16: 786–96.
Ramos S, Khademi F, Somesh BP, Rivero F. Genomic organization and expression profile of the small GTPases of the RhoBTB family in human and mouse. Gene 2002; 298: 147–57.
Siede D, Rapti K, Gorska AA, Katus HA, Altmuller J, Boeckel JN, et al. Identification of circular RNAs with host gene-independent expression in human model systems for cardiac differentiation and disease. J Mol Cell Cardiol 2017; 109: 48–56.
Li Y, Zhang J, Huo C, Ding N, Li J, Xiao J, et al. Dynamic organization of lncRNA and circular RNA regulators collectively controlled cardiac differentiation in humans. EBioMedicine 2017; 24: 137–46.
Boeckel JN, Jae N, Heumuller AW, Chen W, Boon RA, Stellos K, et al. Identification and characterization of hypoxia-regulated endothelial circular RNA. Circ Res 2015; 117: 884–90.
Vausort M, Salgado-Somoza A, Zhang L, Leszek P, Scholz M, Teren A, et al. Myocardial infarction-associated circular RNA predicting left ventricular dysfunction. J Am Coll Cardiol 2016; 68: 1247–8.
Khan MA, Reckman YJ, Aufiero S, van den Hoogenhof MM, van der Made I, Beqqali A, et al. RBM20 regulates circular RNA production from the titin gene. Circ Res 2016; 119: 996–1003.
Holdt LM, Stahringer A, Sass K, Pichler G, Kulak NA, Wilfert W, et al. Circular non-coding RNA ANRIL modulates ribosomal RNA maturation and atherosclerosis in humans. Nat Commun 2016; 7: 12429.
Zhao Z, Li X, Gao C, Jian D, Hao P, Rao L, et al. Peripheral blood circular RNA hsa_circ_0124644 can be used as a diagnostic biomarker of coronary artery disease. Sci Rep 2017; 7: 39918.
Puymirat E, Simon T, Cayla G, Cottin Y, Elbaz M, Coste P, et al. Acute myocardial infarction: Changes in patient characteristics, management, and 6-month outcomes over a period of 20 years in the FAST-MI program (French Registry of Acute ST-Elevation or Non-ST-elevation Myocardial Infarction) 1995 to 2015. Circulation 2017; 136: 1908–19.
Gomes CPC, Spencer H, Ford KL, Michel LYM, Baker AH, Emanueli C, et al. The function and therapeutic potential of long non-coding RNAs in cardiovascular development and disease. Mol Ther Nucleic Acids 2017; 8: 494–507.
Wu HJ, Zhang CY, Zhang S, Chang M, Wang HY. Microarray expression profile of circular RNAs in heart tissue of mice with myocardial infarction-induced heart failure. Cell Physiol Biochem 2016; 39: 205–16.
Zhang JH, Obenaus A, Liebeskind DS, Tang J, Hartman R, Pearce WJ. Recanalization, reperfusion, and recirculation in stroke. J Cereb Blood Flow Metab 2017; 37: 3818–23.
Mehta SL, Pandi G, Vemuganti R. Circular RNA expression profiles alter significantly in mouse brain after transient focal ischemia. Stroke 2017; 48: 2541–8.
Steinberg BA, Mulpuru SK, Fang JC, Gersh BJ. Sudden death mechanisms in non-ischemic cardiomyopathies; Insights gleaned from clinical implantable cardioverter-defibrillator trials. Heart Rhythm 2017; 14: 1839–48.
Wu QQ, Xiao Y, Yuan Y, Ma ZG, Liao HH, Liu C, et al. Mechanisms contributing to cardiac remodelling. Clin Sci (Lond) 2017; 131: 2319–45.
Sun Y, Yang Z, Zheng B, Zhang XH, Zhang ML, Zhao XS, et al. A novel regulatory mechanism of smooth muscle alpha-actin expression by NRG-1/circACTA2/miR-548f-5p axis. Circ Res 2017; 121: 628–35.
Weiser-Evans MCM. Smooth muscle differentiation control comes full circle: The circular noncoding RNA, circActa2, functions as a miRNA sponge to fine-tune alpha-SMA expression. Circ Res 2017; 121: 591–3.
Salzman J, Chen RE, Olsen MN, Wang PL, Brown PO. Cell-type specific features of circular RNA expression. PLoS Genet 2013; 9: e1003777.
Koh W, Pan W, Gawad C, Fan HC, Kerchner GA, Wyss-Coray T, et al. Noninvasive in vivo monitoring of tissue-specific global gene expression in humans. Proc Natl Acad Sci U S A 2014; 111: 7361–6.
Li P, Chen S, Chen H, Mo X, Li T, Shao Y, et al. Using circular RNA as a novel type of biomarker in the screening of gastric cancer. Clin Chim Acta 2015; 444: 132–6.
Alhasan AA, Izuogu OG, Al-Balool HH, Steyn JS, Evans A, Colzani M, et al. Circular RNA enrichment in platelets is a signature of transcriptome degradation. Blood 2016; 127: e1–e11.
Li Y, Zheng Q, Bao C, Li S, Guo W, Zhao J, et al. Circular RNA is enriched and stable in exosomes: a promising biomarker for cancer diagnosis. Cell Res 2015; 25: 981–4.
Chen S, Li T, Zhao Q, Xiao B, Guo J. Using circular RNA hsa_circ_0000190 as a new biomarker in the diagnosis of gastric cancer. Clin Chim Acta 2017; 466: 167–71.
Xu L, Zhang M, Zheng X, Yi P, Lan C, Xu M. The circular RNA ciRS-7 (Cdr1as) acts as a risk factor of hepatic microvascular invasion in hepatocellular carcinoma. J Cancer Res Clin Oncol 2017; 143: 17–27.
Floris G, Zhang L, Follesa P, Sun T. Regulatory role of circular RNAs and neurological disorders. Mol Neurobiol 2017; 54: 5156–65.
Iparraguirre L, Munoz-Culla M, Prada-Luengo I, Castillo-Trivino T, Olascoaga J, Otaegui D. Circular RNA profiling reveals that circular RNAs from ANXA2 can be used as new biomarkers for multiple sclerosis. Hum Mol Genet 2017; 26: 3564–72.
Maass PG, Glazar P, Memczak S, Dittmar G, Hollfinger I, Schreyer L, et al. A map of human circular RNAs in clinically relevant tissues. J Mol Med (Berl) 2017; 95: 1179–89.
Boeckel N, Dimmeler S, Zeiher AM, Jaé N. (Google Patents, 2016).
Dimmeler S, Jaé N, Boeckel N, Zeiher AM. (Google Patents, 2016).
Devaux Y, Vausort M, Zhang L. (Google Patents, 2017).
Sharpless NE, Wang Z, Burd CE, Jeck WR, Siebold AP. (Google Patents, 2012).
Byron SA, Van Keuren-Jensen KR, Engelthaler DM, Carpten JD, Craig DW. Translating RNA sequencing into clinical diagnostics: opportunities and challenges. Nat Rev Genet 2016; 17: 257–71.
Acknowledgements
We thank the editors for inviting us to write this review. Due to space restrictions, the authors cannot cite all the relevant literature in the field. The authors apologize to those colleagues whose work contributed significantly. This work was supported by an American Heart Association Predoctoral Fellowship 16PRE30210016 to Jian-peng TEOH; National Institutes of Health R01s HL134354 and AR070029 to Yao-liang TANG; and an American Physiological Society Shih-Chun WANG Young Investigator Award, an American Heart Association Grant-in-Aid 12GRNT12100048 and Scientist Development Grant 14SDG18970040, and a National Institutes of Health R01 HL124251 to Il-man KIM.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bayoumi, A.S., Aonuma, T., Teoh, Jp. et al. Circular noncoding RNAs as potential therapies and circulating biomarkers for cardiovascular diseases. Acta Pharmacol Sin 39, 1100–1109 (2018). https://doi.org/10.1038/aps.2017.196
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/aps.2017.196
Keywords
This article is cited by
-
CircCAMTA1 facilitates atrial fibrosis by regulating the miR-214-3p/TGFBR1 axis in atrial fibrillation
Journal of Molecular Histology (2023)
-
Epigenetic regulation in cardiovascular disease: mechanisms and advances in clinical trials
Signal Transduction and Targeted Therapy (2022)
-
The Mechanisms Underlying the Beneficial Effects of Stem Cell-Derived Exosomes in Repairing Ischemic Tissue Injury
Journal of Cardiovascular Translational Research (2022)
-
Expression patterns of miR-34a, miR-125b, miR-221 and antioxidant gene NRF2 in plasma samples of patients with atherosclerosis
Journal of Biosciences (2022)
-
circCELF1 Inhibits Myocardial Fibrosis by Regulating the Expression of DKK2 Through FTO/m6A and miR-636
Journal of Cardiovascular Translational Research (2022)