Abstract
Dendritic cell nuclear protein-1 (DCNP1) is a protein associated with major depression. In the brains of depression patients, DCNP1 is up-regulated. However, how DCNP1 participates in the pathogenesis of major depression remains unknown. In this study, we first transfected HEK293 cells with EGFP-DCNP1 and demonstrated that the full-length DCNP1 protein was localized in the nucleus, and RRK (the residues 117-119) composed its nuclear localization signal (NLS). An RRK-deletion form of DCNP1 (DCNP1ΔRRK) and truncated form (DCNP11-116), each lacking the RRK residues, did not show the specific nuclear localization like full-length DCNP1 in the cells. A rat glioma cell line C6 can synthesize melatonin, a hormone that plays important roles in both sleep and depression. We then revealed that transfection of C6 cells with full-length DCNP1 but not DCNP1ΔRRK or DCNP11-116 significantly decreased the levels of melatonin. Furthermore, overexpression of full-length DCNP1, but not DCNP1ΔRRK or DCNP11-116, in C6 cells significantly decreased both the mRNA and protein levels of N-acetyltransferase (NAT), a key enzyme in melatonin synthesis. Full-length DCNP1 but not DCNP1ΔRRK or DCNP11-116 was detected to interact with the Nat promoter and inhibited its activity through its E-box motif. Furthermore, full-length DCNP1 but not the mutants interacted with and repressed the transcriptional activity of BMAL1, a transcription factor that transactivates Nat through the E-box motif. In conclusion, we have shown that RRK (the residues 117-119) are the NLS responsible for DCNP1 nuclear localization. Nuclear DCNP1 represses NAT expression and melatonin biosynthesis by interacting with BMAL1 and repressing its transcriptional activity. Our study reveals a connection between the major depression candidate protein DCNP1, circadian system and melatonin biosynthesis, which may contribute to the pathogenesis of depression.
Similar content being viewed by others
Introduction
Major depression is a mental disorder that is characterized by a continuous low mood, low self-esteem, insomnia, loss of interest in normally enjoyable activities, and suicide1,2. The circadian system, which controls the sleep-wake cycles, hormonal secretion patterns and mood fluctuations, plays important roles in the pathogenesis and symptoms of depression3,4. Polymorphisms in several core clock genes, such as PER2, BMAL1, CLOCK and NPAS2, whose products are necessary components for the generation and regulation of circadian rhythm, appear to be associated with susceptibility to depression5,6,7. However, the molecular mechanisms linking major depression and circadian rhythm are poorly understood.
Melatonin plays important roles in many physiological and pathological conditions, such as circadian rhythm and depression, which may explain the link between them8,9,10. Clinically, 90% of individuals with major depression suffer from insomnia with an abnormal melatonin pathway function11,12,13,14. Many studies have demonstrated that melatonin decreases with a decreased or phase-shifted peak at night are found in patients with depressive disorders15,16,17,18,19. The biosynthetic pathway of pineal melatonin has been thoroughly studied and is controlled by the circadian system9,10. It is initially synthesized from L-tryptophan, which is first converted to 5-hydroxy-L-tryptophan (5-HTP) by tryptophan hydroxylase. 5-HTP is then decarboxylated by 5-hydroxytryptophan decarboxylase to produce serotonin20,21. In the darkness, the key enzyme N-acetyltransferase (NAT) is activated and then converts serotonin to N-acetyl-5-hydroxytryptamine, which is subsequently converted to melatonin by hydroxyindole-o-methyltransferase (HIOMT)22,23. NAT is the rate-limiting enzyme in the synthesis of melatonin from L-tryptophan. The Nat gene contains an E-box element in its promoter, which is the site of BMAL1/CLOCK heterodimer binding and transactivation24,25.
BMAL1 is a gene that encodes a bHLH-PAS domain transcription factor26. It plays a key role in generating circadian rhythms27. BMAL1 knock-out mice show a complete loss of circadian rhythm and alterations of locomotion and other behaviors27. Furthermore, individuals harboring polymorphisms in BMAL1 are susceptible to depression7,28, implying a possible association between BMAL1 and this disorder. BMAL1 and CLOCK form heterodimers and bind to E-box components in the promoters of PER, CRY, REV-ERB and other clock-controlled genes (CCGs), such as Nat to drive their transcription8,29.
Dendritic cell nuclear protein-1 (DCNP1) was first discovered in mature or immature dendritic cells, and in humans, DCNP1 is mainly expressed in the brain and skeletal muscle30. Recent studies have shown that DCNP1 is closely associated with major depression31,32. The DCNP1 mRNA levels are dramatically increased in the paraventricular nucleus (PVN) of depressed patients compared with control subjects33. Furthermore, a truncated form of DCNP1 (DCNP11-116) encoded by the T allele of DCNP1 has been reported to lead to an increased risk of major depression31,32. These results indicate that increased DCNP1 expression or DCNP1 mutation-induced dysfunction possibly participate in the pathogenesis of depression. However, the function of DCNP1 as well as its role in depression are largely unknown.
In the present study, we identified a nuclear localization signal (NLS) (RRK, amino acids 117-119) in DCNP1 and demonstrated that full-length DCNP1, but not the truncated form of DCNP1 (DCNP11-116) or an NLS deletion mutant (DCNP1ΔRRK), represses NAT expression to down-regulate melatonin levels. Full-length DCNP1 binds to and represses BMAL1, a transcription factor that transactivates NAT expression.
Materials and methods
Plasmid constructs
Full-length DCNP1 cDNA was amplified by PCR using the primers 5′-GAGTCGACACTATGCATTACGGAGCA-3′ and 5′-TTGGATCCAACTCAGGCACGTGGGCTG-3′ with human fetal brain cDNA library as the template (Clontech). The PCR product was then inserted into the pEGFP-N1 (Clontech) vector via its Sal I/BamH I sites. pEGFP-DCNP11-116 and pEGFP-DCNP1ΔRRK were generated with the primer pairs 5′-GAGTCGACACTATGCATTACGGAGCA-3′ and 5′-CTGGATCCTTGCTGCTATGCAGTTC-3′ or 5′-GCTGCTATGCAGTTCATCCTG-3′ and 5′-ACAGGCCAGACCAGGCGGGAG-3′, respectively, using pEGFP-DCNP1 as the template. Rat Nat promoter DNA (-2276 bp to +0 bp) was obtained using PCR with the primers 5′-CCGCTCGAGGAGACCTTCCTGTTCTCCTGGTAC-3′ and 5′-CCCAAGCTTGGGTATCTGGCCACTGACCCCTCC-3′ and C6 cell genomic DNA as the template; the PCR products were then inserted into the pGL3-Basic vector (Promega) via its Xho I/Hind III sites. pGL3-NatΔE-box, lacking nucleotides -1192 to -1137, was generated by PCR with the primers 5′-AGAGGACTGCTGTGGGAGTCTGTTCTCTC-3′ and 5′-TTAAATGGGCGATGGCGATGGCTCAAGCAC-3′ using pGL3-Nat as the template. The HA-BMAL1 and HA-CLOCK plasmids were kind gifts from Dr Ying XU (Soochow University, China).
Cell culture and transfection
HEK293 cells or C6 cells were cultured in Dulbecco's modified Eagle's medium (DMEM) (Life Technologies) containing 10% fetal bovine serum (Life Technologies). Cells were transfected with expression plasmids using the Lipofectamine 2000 reagent (Life Technologies) according to the manufacturer's instructions. Forty-eight hours after transfection, cells were observed using an inverted system microscope IX71 (Olympus), or harvested for immunoblot or immunoprecipitation analyses.
Immunoblot analysis
Proteins were separated by 12% SDS-PAGE and then transferred onto polyvinylidene difluoride membranes (Millipore). The following monoclonal primary antibodies were used: anti-BMAL1 (Santa Cruz Biotech), anti-GAPDH (Millipore), anti-GFP (Santa Cruz Biotech), and anti-NAT (Sigma). The secondary sheep anti-mouse IgG-HRP antibody was from Jackson ImmunoResearch. Proteins were visualized using an ECL detection kit (Thermo Fisher Scientific).
To re-probe membranes with another primary antibody, each membrane was first incubated with stripping buffer (50 mmol/L Tris-HCl, pH 6.8, 0.1 mmol/L β-Mercaptoethanol, 20 mmol/L SDS) for 30 min at 50 °C. Then the stripped membrane was subjected to immunoblot analysis following the standard protocol.
Nuclear and cytoplasmic fractionation assay
HEK293 cells transfected with EGFP, EGFP-DCNP1, EGFP-DCNP11-116 or EGFP-DCNP1ΔRRK were lysed in fractionation buffer (320 mmol/L sucrose, 3 mmol/L CaCl2, 2 mmol/L MgAc, 0.1 mmol/L EDTA, 1 mmol/L DTT, 0.5 mmol/L phenylmethylsulfonyl fluoride (PMSF) and 0.5% NP-40) for 20 min on ice. After centrifugation at 600×g for 15 min at 4 °C, the supernatant was collected as the cytoplasmic fraction. The pellet was washed once with fractionation buffer without NP-40 and lysed in cell lysis buffer (150 mmol/L NaCl, 50 mmol/L Tris-HCl pH 7.5, 0.5% deoxycholate, 1% NP-40 and protease inhibitor cocktail (Roche)) as the nuclear fraction. Nuclear poly (ADP-ribose) polymerase (PARP) was used as a nuclear marker, and GAPDH was used as a cytoplasmic marker.
Immunoprecipitation
HEK293 cells co-transfected with HA-BMAL1 along with EGFP, EGFP-DCNP1, EGFP-DCNP11-116 or EGFP-DCNP1ΔRRK were harvested 48 h after transfection and sonicated in TSPI buffer containing 50 mmol/L Tris-HCl pH 7.5, 150 mmol/L sodium chloride, 1 mmol/L EDTA, and 1% NP-40 supplemented with cOmplete Mini Protease Inhibitor Cocktail (Roche). Cell lysates were centrifuged at 12 000×g for 30 min at 4 °C to remove cellular debris. Supernatants were incubated with a monoclonal anti-GFP antibody (Roche) for 4 h at 4 °C. After incubation, protein G Sepharose beads were used for precipitation. Beads were washed with TSPI buffer six times and then eluted with SDS sample buffer for immunoblot analysis.
RNA isolation, reverse transcription, and real-time quantitative PCR
Total RNA was extracted from C6 cells by Trizol reagent (Life Technologies) as previously described33 and then reverse-transcribed into cDNA with a TransScript First-Stand cDNA Synthesis Kit (Takara). Real-time quantitative PCR (qRT-PCR) was performed using SYBR Green Master Mix (Applied Biosystems) on a 7500 Real-Time PCR System (Applied Biosystems). The primer pairs were as follows: 5′-TGCTGTGGCGATACCTTCACCA-3′ and 5′-CAGCTCAGTGAAGGTGAGAGAT-3′ for rat Nat; 5′-TACGGGGACAGGAAGTTTTG-3′ and 5′-GTGCCACTTCTGGGTTCATT-3′ for rat Hiomt; 5′-GCCAGGAACGGAAATTTGTA-3′ and 5′-TCTCAGGTGGAAGCTCTGGT-3′ for rat Maoa; 5′-TGGGAAGATTCCAGAGGATG-3′ and 5′-GCTGACAAGATGGTGGTCAA-3′ for rat Maob; 5′-TTGCTGACAGGATGCAGAA-3′ and 5′-ACCAATCCACACAGAGTACTT-3′ for rat β-actin. The relative target gene mRNA levels were determined using the 2−ΔΔCT method.
Chromatin immunoprecipitation and immunoblot analysis
Formaldehyde cross-linking and chromatin immunoprecipitation (ChIP) were performed as previously described33. Immunoprecipitation assays were performed with anti-GFP antibodies (Roche) and protein G Sepharose beads (Roche). Immunoprecipitates obtained by ChIP were subjected to PCR with the primers 5′-GGAGACCTTCCTGTTCTCCTG-3′ and 5′-TAGGGAGAGCATGGGCTAAG-3′. The amplification conditions were as follows: 5 min at 95 °C; then 40 cycles of 95 °C for 30 s, 60 °C for 30 s and 72 °C for 30 s.
Dual-luciferase reporter gene assay
HEK293 cells were transfected with a Nat- or NatΔE-box-luciferase reporter construct along with the EGFP, EGFP-DCNP1, EGFP-DCNP11-116 or EGFP-DCNP1ΔRRK plasmids or siRNAs against BMAL1, as indicated. The renilla luciferase expression plasmid pRL-CMV was co-transfected to normalize for variations in the transfection efficiency. Cells were lysed with passive lysis buffer (Promega). The activities of both firefly and Renilla luciferase were measured with a Dual Luciferase Reporter Assay System (Promega) using a Veritas Microplate Luminometer according to the manufacturer's instructions. The absolute values of firefly luminescence were normalized to those of Renilla, and the ratios are presented as the median of three transfection experiments. The activity of pGL3-Basic luciferase was normalized to 1.
Immunocytochemistry
HEK293 cells were washed with PBS and fixed with 4% paraformaldehyde for 5 min at room temperature. Then, cells were treated with 0.25% Triton X-100 for 15 min and blocked with 4% FBS in PBS overnight at 4 °C with anti-BMAL1 antibodies (Santa Cruz). Next, cells were incubated with Alexa Fluor 594 donkey anti-mouse secondary antibodies (Life Technologies). Nuclei were stained with DAPI (Sigma).
Melatonin determination
The melatonin concentrations in the culture medium of C6 cells expressing EGFP, EGFP-DCNP1, EGFP-DCNP11-116 or EGFP-DCNP1ΔRRK were measured by ELISA (IBL International) according to the manufacturer's instructions.
RNA interference
Double-stranded oligonucleotides targeting 5′-TAGGCACATCGTGTTATGAAT-3′ of human BMAL1 mRNA were synthesized by Shanghai GenePharma (Shanghai, China), and an irrelevant oligonucleotide was used as the negative control. Transfection was performed with Lipofectamine RNAiMAX reagent (Life Technologies) according to manufacturer's instructions. Briefly, siRNA and Lipofectamine RNAiMAX reagent (Life Technologies) were mixed in Opti-MEM medium (Invitrogen), incubated for 30 min at room temperature to allow for complex formation, and then added to culture medium. Cells were harvested 72 h after transfection for further analyses.
Statistical analysis
Statistical comparisons between groups and treatments were performed using one-way or two-way analysis of variance (ANOVA) for comparisons among multiple groups, followed by Student's t-tests for analyzing significance, as indicated. P values <0.05 were considered statistically significant. Data are presented as the mean±SEM.
Results
Identification of the DCNP1 NLS
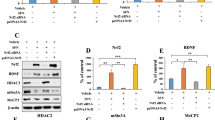
Our previous study showed that endogenous DCNP1 was probably localized in the nucleus33. To confirm the subcellular distribution of DCNP1, we transfected EGFP or EGFP-DCNP1 into HEK293 cells and examined their subcellular localization. Unlike EGFP, which was diffusely distributed throughout whole cells, EGFP-DCNP1 was predominately localized in the nucleus (Figure 1A). By examining the DCNP1 protein sequence, we found that it has a potential NLS (RRK) at amino acid positions 117-119 (Figure 1B). We therefore created two deletion mutants of DCNP1, DCNP1ΔRRK, in which the putative NLS (RRK, amino acids 117-119) was deleted, and DCNP11-116, a truncated form of DCNP1 lacking amino acids 117-224, including the RRK motif, which has been reported to increase the risk of major depression31,32 (Figure 1C). Full-length DCNP1 showed specific nuclear localization. However, like EGFP, both EGFP-DCNP1ΔRRK and EGFP-DCNP11-116 showed a diffuse distribution with no clear nuclear pattern (Figure 1D). We also performed nuclear and cytoplasmic fractionation assays to further verify differences in the distribution patterns between full-length DCNP1 and its mutants. As shown in Figure 1E and 1F, full-length DCNP1 was mainly present in the nuclear fraction. However, although EGFP-DCNP1ΔRRK and EGFP-DCNP11-116 were partially detected in the nuclear fraction, a large proportion of each protein was distributed in the cytoplasm (Figure 1E and 1F). These results suggest that the amino acids RRK at positions 117-119 are essential for DCNP1 nuclear localization.
Nuclear localization of DCNP1. (A) Subcellular localization of EGFP-DCNP1 (green). HEK293 cells were transfected with EGFP or EGFP-DCNP1 and visualized using a fluorescence microscope (magnification ×400). Nuclei were stained with DAPI (1 μg/mL) (blue). Scale bars, 5 μm. (B) Schematic illustration of the predicted NLS (aa 117-119) of DCNP1. (C) Schematic illustration of EGFP-tagged expression constructs. (D) RRK is a functional NLS of DCNP1. HEK293 cells transfected with EGFP-DCNP1 or its mutants (1-116 and ΔRRK) were visualized using a fluorescence microscope (magnification ×400). Nuclei were stained with DAPI (1 μg/mL) (blue). Scale bars, 5 μm. (E) HEK293 cells were transfected with EGFP, EGFP-DCNP1 or its mutants (1-116 and ΔRRK). Forty-eight hours later, cells were subjected to nuclear and cytoplasmic fractionation. (F) Percentages of EGFP, EGFP-DCNP1 or its mutants (1-116 and ΔRRK) in nuclear and cytoplasmic fractions relative to their total protein levels in (E) were quantified.
Regulation of melatonin levels and Nat mRNA levels by full-length DCNP1
As a significant clinical feature of major depression is insomnia, with low levels of melatonin in patients11,12,13,14,34, we investigated whether DCNP1 or it mutants regulate melatonin production. To address this issue, we transfected EGFP, EGFP-DCNP1, EGFP-DCNP11-116 or EGFP-DCNP1ΔRRK into C6 cells, a rat glioma cell line that can synthesize melatonin35. Overexpression of EGFP-DCNP1 significantly decreased the melatonin levels, whereas its mutants (1-116, ΔRRK) did not (Figure 2A). To identify which factors are involved in the regulation of melatonin by DCNP1, we examined the expression of two key enzymes in the melatonin synthesis pathway, HIOMT and NAT, and two other key enzymes, monoamine oxidase A and B (MAOA and MAOB), for 5-HT metabolism in the bypass of melatonin synthesis. Significantly decreased levels of Nat mRNA were observed in EGFP-DCNP1-transfected C6 cells, but the Hiomt, Maoa and Maob mRNA levels were not changed (Figure 2B). We next examined whether DCNP1 mutants affect NAT expression. In C6 cells overexpressing EGFP, EGFP-DCNP1, EGFP-DCNP11-116 or EGFP-DCNP1ΔRRK, EGFP-DCNP1 repressed the Nat mRNA and protein levels, while EGFP-DCNP11-116 and EGFP-DCNP1ΔRRK failed to repress Nat mRNA or protein expression (Figure 3A and 3B). These data suggest that full-length DCNP1, unlike its mutants, regulates melatonin synthesis by repressing NAT transcription.
DCNP1 regulates melatonin and Nat mRNA levels. (A) Melatonin levels in culture medium of C6 cells transfected with EGFP-DCNP1 or its mutants (1-116 and ΔRRK) were measured by ELISA. The values are the mean±SEM from three independent experiments. **P<0.01, ΔP>0.05, one-way ANOVA, followed by the t-test. (B) qRT-PCR assays were performed to measure the mRNA levels of Nat, Hiomt, Maoa and Maob in C6 cells expressing EGFP or EGFP-DCNP1. The values are the mean±SEM of three independent experiments. **P<0.01, ΔP>0.05, t-test.
Nuclear DCNP1 regulates Nat mRNA and protein levels. qRT-PCR analysis (A) and immunoblot analysis (B) showing the Nat mRNA and protein levels in C6 cells expressing EGFP-DCNP1 or its mutants (1-116 and ΔRRK). Antibodies were used as indicated. The values are the mean±SEM of three independent experiments. **P<0.01, ΔP>0.05, one-way ANOVA.
Interactions between DCNP1 and the Nat promoter or BMAL1
As full-length DCNP1 localizes to the nucleus and regulates Nat mRNA expression, we investigated whether DCNP1 interacts with the Nat promoter. We performed a chromatin immunoprecipitation assay to verify this interaction. In cells overexpressing wild-type or mutant DCNP1, the Nat promoter co-immunoprecipitated when EGFP-DCNP1, but not its mutants (1-116, ΔRRK), was precipitated using anti-GFP antibodies (Figure 4A), suggesting that DCNP1 interacts with the Nat promoter and that this interaction is dependent on DCNP1's nuclear localization. Considering that BMAL1 binds to the E-box in the Nat promoter to regulate its transcription24,25, we next examined whether regulation of NAT by DCNP1 is mediated by BMAL1. We first tested whether DCNP1 could bind to BMAL1 to regulate the transcription of Nat. In HEK293 cells expressing HA-BMAL1 with EGFP or EGFP-DCNP1 or EGFP-DCNP1 mutants (1-116, ΔRRK), HA-BMAL1 co-precipitated with EGFP-DCNP1 using anti-GFP antibodies, but not with EGFP alone or EGFP-DCNP1 mutants (Figure 4B). These immunocytochemical assays also showed that both EGFP-DCNP1 and BMAL1 were nuclear-localized, whereas DCNP1 mutant proteins were diffusely distributed throughout whole cells (Figure 4C).
DCNP1 binds to the Nat promoter and BMAL1 in the nucleus. (A) HEK293 cells were transfected with EGFP-DCNP1 or its mutants (1-116 and ΔRRK) for chromatin immunoprecipitation assays. Inputs and chromatin immunoprecipitation products were amplified by PCR. (B) HEK293 cells expressing EGFP-DCNP1 or its mutants (1-116 and ΔRRK) were subjected to immunoprecipitation using anti-GFP antibodies. Immunoprecipitates and inputs were detected with the antibodies as indicated. (C) Co-localization of DCNP1 and BMAL1 in the nucleus. HEK293 cells expressing EGFP-DCNP1 or its mutants (1-116 and ΔRRK) were stained with anti-BMAL1 antibodies and visualized using a fluorescence microscope (magnification ×400). Nuclei were stained with DAPI (1 μg/mL) (blue). Scale bars, 5 μm.
DCNP1 regulates the activity of the Nat promoter through BMAL1
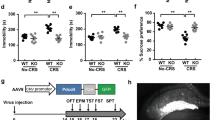
As DCNP1 interacts with BMAL1 and suppresses the transcription of Nat, we performed a luciferase reporter gene assay to further examine whether DCNP1 regulates Nat promoter activity and to determine whether this regulation is BMAL1-dependent. We transfected HEK293 cells with a Nat-luciferase construct along with expression plasmids for EGFP, EGFP-DCNP1, and EGFP-DCNP1 mutants (1-116, ΔRRK); HA; or HA-BMAL1/HA-CLOCK. Cells were harvested 48 h after transfection, and the supernatants were used for luciferase activity assays. Overexpression of HA-BMAL/HA-CLOCK heterodimer significantly increased Nat promoter activity. EGFP-DCNP1 significantly inhibited the basal Nat promoter activity as well as BMAL/CLOCK-induced Nat promoter activity (Figure 5A). As BMAL1/CLOCK binds to the E-box in the Nat promoter to transactivate Nat transcription, we deleted the E-box in the Nat-luciferase plasmid (pGL3-NatΔE-box promoter) and used it for co-transfection with the EGFP or EGFP-DCNP1 expression plasmids. Although DCNP1 repressed wild-type Nat promoter activity, it failed to influence Nat promoter activity when the E-box motif was deleted (Figure 5B), further suggesting that DCNP1 regulation of Nat transcription is dependent on BMAL1. To further investigate the dependence of DCNP1-regulated NAT expression on BMAL1, we examined the effects of DCNP1 on Nat-luciferase reporter activity in BMAL1-knockdown cells. The knockdown efficiency of Bmal1 siRNA is shown in Figure 5C. Full-length DCNP1 repressed Nat reporter activity in the presence of BMAL1, while it did not affect Nat reporter activity when BMAL1 had been knocked down (Figure 5D). We also found that mutants of DCNP1 (1-116, ΔRRK) did not influence Nat promoter activity (Figure 5D). These results indicate that inhibition of Nat transcription by DCNP1 is dependent on BMAL1. Together, these data suggest that the regulation of melatonin biosynthesis by DCNP1 is dependent on its binding to BMAL1 to repress NAT expression.
DCNP1 regulates Nat promoter activity through BMAL1. (A) HEK293 cells were co-transfected with equal amounts (400 ng) of a Nat promoter construct and luciferase reporter, along with plasmids that expressed EGFP, EGFP-DCNP1, HA or HA-BMAL1/HA-CLOCK. Forty-eight hours later, the luciferase activities were quantified and relative ratios were calculated. The data shown are the mean±SEM of three independent experiments. **P<0.01, two-way ANOVA. (B) HEK293 cells were co-transfected with equal amounts (400 ng) of a Nat promoter luciferase reporter construct or an E-box deletion construct along with plasmids that expressed EGFP or EGFP-DCNP1. Forty-eight hours later, the luciferase activities were quantified and relative ratios were calculated. The values are the mean±SEM of three independent experiments. **P<0.01, ΔP>0.05, two-way ANOVA, followed by the t-test. (C) RNA oligonucleotides against BMAL1 (BMAL1 siRNA) or negative control (NC) were transfected into HEK293 cells. Seventy-two hours later, cells were harvested and subjected to immunoblot analysis. The BMAL1 protein levels were quantified relative to GAPDH. The values are the mean±SEM of three independent experiments. ***P<0.001, t-test. (D) HEK293 cells were treated with NC or BMAL1 siRNA. Then cells were co-transfected with equal amounts (400 ng) of a Nat promoter luciferase reporter construct along with plasmids that expressed EGFP, EGFP-DCNP1, EGFP-DCNP11-116 or EGFP-DCNP1ΔRRK. Forty-eight hours later, the luciferase activities were quantified and relative ratios were calculated. The data shown are the mean±SEM of three independent experiments. **P<0.01, ΔP>0.05, two-way ANOVA followed by t-test.
In conclusion, we found that the amino acids RRK at residues 117-119 are responsible for nuclear localization of full-length DCNP1. Nuclear-localized DCNP1 binds to BMAL1 to repress its transcriptional activity, thus inhibiting Nat transcription and melatonin biosynthesis. However, the NLS-deletion mutants DCNP1ΔRRK and truncated DCNP11-116 lack specific nuclear localization and fail to interact with BMAL1 to inhibit BMAL1/NAT-mediated melatonin biosynthesis (Figure 6). Thus, our results suggest that the involvement of nuclear DCNP1 in the pathogenesis of depression may depend on its regulation of the circadian clock and melatonin biosynthesis.
Schematic diagram representing the function of DCNP1 or its mutants (1-116 and ΔRRK) in melatonin biosynthesis. Full-length DCNP1 localizes to the nucleus and binds to BMAL1, thus repressing NAT transcription and subsequently inhibiting melatonin biosynthesis. However, DCNP1 mutants (1-116 and ΔRRK) lack the NLS and cannot inhibit the BMAL1/NAT/melatonin pathway.
Discussion
In the present study, we determined that the product of DCNP1, a candidate gene for major depression, plays an important role in melatonin biosynthesis by regulating NAT expression via BMAL1.
Melatonin is mainly produced in the pineal gland and is also produced by various other tissues and cells36, such as astrocytes37, macrophages38, fibroblasts and skin cells39. Melatonin is involved in many biological processes, including sleep-wake timing and blood pressure regulation, mainly through activation of the melatonin receptors MT1 and MT2, which are expressed in many tissues and cells36,40. Melatonin is also closely associated with depression as its levels are dramatically decreased in depression patients and is considered to be a significant marker of depression15,16,17,18,19. In addition, a melatonin analog, agomelatine, has been approved for major depression treatment41. Agomelatine acts as an agonist of MT1, MT2 and the 5-hydroxytryptamine 2C (5-HT2C) receptor to improve both depressive symptoms and sleep disorders in patients with depression41. Our previous study showed that the mRNA levels of DCNP1 were dramatically increased in patients with depression33. In the present study, we further characterized the relationship between DCNP1 and the melatonin biosynthesis pathway. We identified a novel NLS, amino acids RRK at residues 117-119, that is critical for DCNP1 nuclear localization. Full-length DCNP1, but not mutants lacking this NLS, represses melatonin biosynthesis. As nuclear DCNP1 is increased in depression patients33, our present study shows that the negative regulation of melatonin biosynthesis by DCNP1 may contribute to depression pathogenesis.
Although the truncated form DCNP11-116 has been reported to increase the risk of major depression31,32, it did not seem to influence melatonin biosynthesis in our experiments. The premature DCNP11-116 encoded by the T allele of DCNP1 lacks an NLS and shows a diffuse distribution pattern. We speculate that the truncated DCNP11-116 may increase the risk for major depression through other signaling pathways rather than by regulating melatonin biosynthesis in the nucleus.
Melatonin biosynthesis is initiated from serotonin, which is first converted to N-acetylserotonin by NAT and then further converted to melatonin by HIOMT22,23,36. Serotonin can also be oxidized by MAOA and MAOB to 5-hydroxyindoleacetic acid (5-HIAA), bypassing melatonin synthesis10,20. In the present study, we found that the levels of melatonin were significantly lower in DCNP1-overexpressing C6 cells. The decrease in melatonin levels induced by DCNP1 are mediated by NAT, but not other enzymes, such as HIOMT, MAOA and MAOB, that are involved in melatonin biosynthesis or breakdown, as only the NAT mRNA and protein levels are significantly down-regulated by overexpression of DCNP1.
The circadian clock protein BMAL1 plays core roles in circadian rhythms and controls the daily rhythms of melatonin8. It forms heterodimers with CLOCK to bind to E-box response elements to activate transcription of downstream genes, including circadian clock genes42,43. BMAL1/CLOCK heterodimers also transactivate Nat transcription via the E-box in its promoter24,25. In our observations, full-length DCNP1 co-localized and interacted with BMAL1 in the nucleus. In addition, full-length DCNP1 interacted with the Nat promoter. DCNP1 mutants (1-116 and ΔRRK) lack specific nuclear localization and instead show diffuse distribution throughout the whole cell and fail to interact with BMAL1 or the Nat promoter, suggesting that the RRK motif may be important for DCNP1 to bind to BMAL1 and regulate its function. Moreover, luciferase assays showed that full-length DCNP1, but not its mutants, significantly inhibits Nat reporter activity, which is dependent on the E-box in the Nat promoter, further suggesting that the effects of DCNP1 on NAT expression are mediated by BMAL1.
Interestingly, genetic variants of NAT, the key enzyme in melatonin synthesis, are associated with susceptibility to major depression44. In addition, mice harboring Nat mutations show depression-like behaviors45, strongly suggesting an association between the melatonin pathway and depression. Our study also reveals that DCNP1, a major depression candidate gene product, is involved in the regulation of melatonin biosynthesis, implying a role in depression pathogenesis.
In summary, we determined that full-length DCNP1 is involved in regulation of melatonin biosynthesis, dependent on its interaction with BMAL1. Full-length DCNP1 represses BMAL1-mediated Nat transcription via an E-box motif and subsequently decreases NAT expression and melatonin biosynthesis. However, mutants of DCNP1 (1-116 and ΔRRK) that lack an NLS fail to interact with BMAL1, leading to a failure to regulate melatonin biosynthesis. Thus, our findings provide linkages between a depression-associated gene product, the circadian system and melatonin metabolism.
Author contribution
Dong CHEN and Yi-pei LI performed most of the experiments and analyzed the data; Yan-xia YU performed the ELISA assays; Tian ZHOU, Chao LIU, Er-kang FEI, Feng GAO and Chen-chen MU performed some of the immunoblot analyses; Dong CHEN and Hai-gang REN drafted the manuscript and discussed the experiments; Dong CHEN and Guang-hui WANG designed the experiments; Guang-hui WANG and Hai-gang REN interpreted the experiments and revised the manuscript. All of the authors read and approved the final manuscript.
References
Belmaker RH, Agam G . Major depressive disorder. N Engl J Med 2008; 358: 55–68.
Cai S, Huang S, Hao W . New hypothesis and treatment targets of depression: an integrated view of key findings. Neurosci Bull 2015; 31: 61–74.
Nechita F, Pirlog MC, ChiriTa AL . Circadian malfunctions in depression - neurobiological and psychosocial approaches. Rom J Morphol Embryol 2015; 56: 949–55.
Courtet P, Olie E . Circadian dimension and severity of depression. Eur Neuropsychopharmacol 2012; 22: S476–81.
Benedetti F, Dallaspezia S, Fulgosi MC, Lorenzi C, Serretti A, Barbini B, et al. Actimetric evidence that CLOCK 3111 T/C SNP influences sleep and activity patterns in patients affected by bipolar depression. Am J Med Genet B Neuropsychiatr Genet 2007; 144B: 631–5.
Johansson C, Willeit M, Smedh C, Ekholm J, Paunio T, Kieseppa T, et al. Circadian clock-related polymorphisms in seasonal affective disorder and their relevance to diurnal preference. Neuropsychopharmacology 2003; 28: 734–9.
Partonen T, Treutlein J, Alpman A, Frank J, Johansson C, Depner M, et al. Three circadian clock genes Per2, Arntl, and Npas2 contribute to winter depression. Ann Med 2007; 39: 229–38.
Vriend J, Reiter RJ . Melatonin feedback on clock genes: a theory involving the proteasome. J Pineal Res 2015; 58: 1–11.
Stehle JH, Saade A, Rawashdeh O, Ackermann K, Jilg A, Sebesteny T, et al. A survey of molecular details in the human pineal gland in the light of phylogeny, structure, function and chronobiological diseases. J Pineal Res 2011; 51: 17–43.
Reiter RJ . Pineal melatonin: cell biology of its synthesis and of its physiological interactions. Endocr Rev 1991; 12: 151–80.
Tsuno N, Besset A, Ritchie K . Sleep and depression. J Clin Psychiatry 2005; 66: 1254–69.
Avery D, Wildschiodtz G, Rafaelsen O . REM latency and temperature in affective disorder before and after treatment. Biol Psychiatry 1982; 17: 463–70.
Boivin DB . Influence of sleep-wake and circadian rhythm disturbances in psychiatric disorders. J Psychiatry Neurosci 2000; 25: 446–58.
Wu YH, Ursinus J, Zhou JN, Scheer FA, Ai-Min B, Jockers R, et al. Alterations of melatonin receptors MT1 and MT2 in the hypothalamic suprachiasmatic nucleus during depression. J Affect Disord 2013; 148: 357–67.
Crasson M, Kjiri S, Colin A, Kjiri K, L'Hermite-Baleriaux M, Ansseau M, et al. Serum melatonin and urinary 6-sulfatoxymelatonin in major depression. Psychoneuroendocrinology 2004; 29: 1–12.
Fountoulakis KN, Karamouzis M, Iacovides A, Nimatoudis J, Diakogiannis J, Kaprinis G, et al. Morning and evening plasma melatonin and dexamethasone suppression test in patients with nonseasonal major depressive disorder from northern Greece (latitude 40-41.5 degrees). Neuropsychobiology 2001; 44: 113–7.
Paparrigopoulos T . Melatonin response to atenolol administration in depression: indication of beta-adrenoceptor dysfunction in a subtype of depression. Acta Psychiatr Scand 2002; 106: 440–5.
Buckley TM, Schatzberg AF . A pilot study of the phase angle between cortisol and melatonin in major depression – a potential biomarker? J Psychiatr Res 2010; 44: 69–74.
Oglodek EA, Just MJ, Szromek AR, Araszkiewicz A . Melatonin and neurotrophins NT-3, BDNF, NGF in patients with varying levels of depression severity. Pharmacol Rep 2016; 68: 945–51.
Calvo JR, Gonzalez-Yanes C, Maldonado MD . The role of melatonin in the cells of the innate immunity: a review. J Pineal Res 2013; 55: 103–20.
Tan DX, Zheng X, Kong J, Manchester LC, Hardeland R, Kim SJ, et al. Fundamental issues related to the origin of melatonin and melatonin isomers during evolution: relation to their biological functions. Int J Mol Sci 2014; 15: 15858–90.
Arendt J . Melatonin, circadian rhythms, and sleep. N Engl J Med 2000; 343: 1114–6.
Amaral FG, Turati AO, Barone M, Scialfa JH, do Carmo Buonfiglio D, Peres R, et al. Melatonin synthesis impairment as a new deleterious outcome of diabetes-derived hyperglycemia. J Pineal Res 2014; 57: 67–79.
Chong NW, Bernard M, Klein DC . Characterization of the chicken serotonin N-acetyltransferase gene. Activation via clock gene heterodimer/E box interaction. J Biol Chem 2000; 275: 32991–8.
Chen W, Baler R . The rat arylalkylamine N-acetyltransferase E-box: differential use in a master vs. a slave oscillator. Brain Res Mol Brain Res 2000; 81: 43–50.
Hogenesch JB, Chan WK, Jackiw VH, Brown RC, Gu YZ, Pray-Grant M, et al. Characterization of a subset of the basic-helix-loop-helix-PAS superfamily that interacts with components of the dioxin signaling pathway. J Biol Chem 1997; 272: 8581–93.
Bunger MK, Wilsbacher LD, Moran SM, Clendenin C, Radcliffe LA, Hogenesch JB, et al. Mop3 is an essential component of the master circadian pacemaker in mammals. Cell 2000; 103: 1009–17.
Nievergelt CM, Kripke DF, Barrett TB, Burg E, Remick RA, Sadovnick AD, et al. Suggestive evidence for association of the circadian genes PERIOD3 and ARNTL with bipolar disorder. Am J Med Genet B 2006; 141B: 234–41.
Lima-Cabello E, Diaz-Casado ME, Guerrero JA, Otalora BB, Escames G, Lopez LC, et al. A review of the melatonin functions in zebrafish physiology. J Pineal Res 2014; 57: 1–9.
Masuda M, Senju S, Fujii Si S, Terasaki Y, Takeya M, Hashimoto Si S, et al. Identification and immunocytochemical analysis of DCNP1, a dendritic cell-associated nuclear protein. Biochem Biophys Res Commun 2002; 290: 1022–9.
Bosker FJ, Hartman CA, Nolte IM, Prins BP, Terpstra P, Posthuma D, et al. Poor replication of candidate genes for major depressive disorder using genome-wide association data. Mol Psychiatry 2011; 16: 516–32.
Willis-Owen SA, Shifman S, Copley RR, Flint J . DCNP1: a novel candidate gene for major depression. Mol Psychiatry 2006; 11: 121–2.
Zhou T, Wang S, Ren H, Qi XR, Luchetti S, Kamphuis W, et al. Dendritic cell nuclear protein-1, a novel depression-related protein, upregulates corticotropin-releasing hormone expression. Brain 2010; 133: 3069–79.
Ramirez-Rodriguez G, Vega-Rivera NM, Oikawa-Sala J, Gomez-Sanchez A, Ortiz-Lopez L, Estrada-Camarena E . Melatonin synergizes with citalopram to induce antidepressant-like behavior and to promote hippocampal neurogenesis in adult mice. J Pineal Res 2014; 56: 450–61.
Liu YJ, Meng FT, Wang LL, Zhang LF, Cheng XP, Zhou JN . Apolipoprotein E influences melatonin biosynthesis by regulating NAT and MAOA expression in C6 cells. J Pineal Res 2012; 52: 397–402.
Hardeland R, Cardinali DP, Srinivasan V, Spence DW, Brown GM, Pandi-Perumal SR . Melatonin-a pleiotropic, orchestrating regulator molecule. Prog Neurobiol 2011; 93: 350–84.
Liu YJ, Zhuang J, Zhu HY, Shen YX, Tan ZL, Zhou JN . Cultured rat cortical astrocytes synthesize melatonin: absence of a diurnal rhythm. J Pineal Res 2007; 43: 232–8.
Muxel SM, Pires-Lapa MA, Monteiro AW, Cecon E, Tamura EK, Floeter-Winter LM, et al. NF-kappaB drives the synthesis of melatonin in RAW 264.7 macrophages by inducing the transcription of the arylalkylamine-N-acetyltransferase (AA-NAT) gene. PLoS One 2012; 7: e52010.
Liu YJ, Meng FT, Wu L, Zhou JN . Serotoninergic and melatoninergic systems are expressed in mouse embryonic fibroblasts NIH3T3 cells. Neuro Endocrinol Lett 2013; 34: 236–40.
Boutin JA, Audinot V, Ferry G, Delagrange P . Molecular tools to study melatonin pathways and actions. Trends Pharmacol Sci 2005; 26: 412–9.
Cardinali DP, Srinivasan V, Brzezinski A, Brown GM . Melatonin and its analogs in insomnia and depression. J Pineal Res 2012; 52: 365–75.
Hogenesch JB, Gu YZ, Jain S, Bradfield CA . The basic-helix-loop-helix-PAS orphan MOP3 forms transcriptionally active complexes with circadian and hypoxia factors. Proc Natl Acad Sci U S A 1998; 95: 5474–9.
Jin X, Shearman LP, Weaver DR, Zylka MJ, de Vries GJ, Reppert SM . A molecular mechanism regulating rhythmic output from the suprachiasmatic circadian clock. Cell 1999; 96: 57–68.
Soria V, Martinez-Amoros E, Escaramis G, Valero J, Crespo JM, Gutierrez-Zotes A, et al. Resequencing and association analysis of arylalkylamine N-acetyltransferase (AANAT) gene and its contribution to major depression susceptibility. J Pineal Res 2010; 49: 35–44.
Uz T, Manev H . Prolonged swim-test immobility of serotonin N-acetyltransferase (AANAT)-mutant mice. J Pineal Res 2001; 30: 166–70.
Acknowledgements
This work was supported in part by the National R&D Program of China (No 2016YFC1306000), the National Natural Science Foundation of China (No 31330030, 81371393, 31471012 and 81761148024), the National High-Tech Research and Development Program of China (973-projects, No 2012CB947602), and a project funded by the Priority Academic Program Development of Jiangsu Higher Education Institutions.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Chen, D., Li, Yp., Yu, Yx. et al. Dendritic cell nuclear protein-1 regulates melatonin biosynthesis by binding to BMAL1 and inhibiting the transcription of N-acetyltransferase in C6 cells. Acta Pharmacol Sin 39, 597–606 (2018). https://doi.org/10.1038/aps.2017.163
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/aps.2017.163
Keywords
This article is cited by
-
DJ-1 inhibits microglial activation and protects dopaminergic neurons in vitro and in vivo through interacting with microglial p65
Cell Death & Disease (2021)
-
Poly-PR in C9ORF72-Related Amyotrophic Lateral Sclerosis/Frontotemporal Dementia Causes Neurotoxicity by Clathrin-Dependent Endocytosis
Neuroscience Bulletin (2019)