Abstract
Aim:
RNA-binding proteins are a large group of regulators (800–1000 in humans), some of which play significant roles in mRNA local translation. In this study, we analyzed the functions of the protein RNP-1, which was previously discovered in a genetic selection screen for nocodazole suppression.
Methods:
The growth rates and the microtubule networks of Dictyostelium cells were assessed with or without nocodazole (10 μmol/L) in suspension culture. Fluorescent images of RNP-1-GFP and RFP-tubulin were captured when cells were undergoing cytokinesis, then the GFP signal intensity and distance to the nearest centrosome were analyzed by using a computer program written in Matlab®. The RNP-1-GFP-expresseding cells were polarized, and the time-lapse images of cells were captured when cells were chemotaxing to a cAMP source.
Results:
Over-expression of RNP-1 rescued the growth defects caused by the microtubule-destabilizing agent nocodazole. Over-expression of RNP-1 protected microtubules from nocodazole treatment. In cells undergoing cytokinesis, the RNP-1 protein was localized to the polar regions of the cell cortex, and protein levels decreased proportionally as the power of the distance from the cell cortex to the nearest centrosome. In chemotactic cells, the RNP-1 protein localized to the leading edge of moving cells. Sequence analysis revealed that RNP-1 has two RNA-binding domains and is related to cytosolic poly(A)-binding proteins (PABPCs) in humans.
Conclusion:
RNP-1 has roles in protecting microtubules and in directing cortical movement during cytokinesis and cell migration in Dictyostelium cells. The sequence similarity of RNP-1 to human PABPCs suggests that PABPCs may have similar functions in mammalian cells, perhaps in regulating microtubule dynamics and functions during cortical movement in cytokinesis and cell migration.
Similar content being viewed by others
Introduction
Research using the model organism Dictyostelium discoideum has led to many conceptual advances in our understanding of cytoskeleton dynamics during cytokinesis and cell migration. Dictyostelium cells, with a combination of robust genetic screening methods with chemical inhibitors and a variety of cell biological approaches1,2, comprise an excellent system for the study of cytoskeleton regulation in different cell states.
Nocodazole is a compound that blocks microtubule polymerization by sequestering α/β tubulin dimers and inhibits cell growth. Nocodazole has been used for several decades as a chemotherapy reagent in cancer patients to inhibit cancer cell growth3. In this study, we analyzed the function of the protein RNP-1 (protein ID: DDB0233340), which was previously discovered in a genetic selection screen in which library plasmid-transformed Dictyostelium cells were used to select for genetic suppressors of nocodazole-induced growth defects4.
Microtubules are one of the major cytoskeletal filament systems found in the cell. Microtubules are polarized molecules composed of α/β-tubulin dimers, and they emanate from the centrosome, where the microtubule minus ends are embedded, with the plus ends growing radially toward the cell cortex5. The dynamics of microtubules are largely regulated at the plus ends through extension, shrinkage, or bending6. In addition to their structural dynamics, microtubules also provide binding surfaces for many cellular proteins, including motor proteins and microtubule-associated proteins (MAPs). Motor proteins traffic materials to the cell periphery along microtubule tracks. Most MAPs interact with microtubules to either stabilize or destabilize them and, in particular, some MAPs bind to the plus ends of microtubules to prevent catastrophic collapse or to interact with cortical proteins7,8,9. Due to their unique structural dynamics and the many types of MAPs and other MAP-associated proteins, microtubules have the ability to deposit signaling proteins at specific regions of the cell cortex and direct cell movement, particularly through the generation of cell protrusions. Recently, many studies have shown that microtubule networks play fundamental regulatory roles in cell migration. In one study of breast cancer cell motility, the formin protein mDia1 was shown to be a microtubule regulator that is required for the cortical localization of Rab6IP2, helping to tether microtubules to the leading edge10. Another study showed that the deubiquitinase cylindromatosis interacts with EB1 to regulate microtubule dynamics and stimulate cell migration11. Furthermore, pregnenolone (P5), which binds to the microtubule plus-end tracking protein CLIP-170, was shown to stimulate cell migration12. In Dictyostelium cells, chemotaxis was impaired when TsuA, a protein associated with the microtubule network, was lost13. Taken together, these observations suggest that the microtubule network interacts with a variety of proteins within the cell cortex to regulate directional cell movement. However, how microtubule plus ends are protected from disassembly while regulating proteins at the cell cortex is still not well understood.
Local protein biogenesis is an important mechanism for intracellular protein targeting, which is often accompanied by mRNA transportation and localization. Local mRNA translation is frequently observed during synaptic neuronal development14 and has been shown to be important for synaptic plasticity and neurological diseases15. In migrating cells, more than 700 protrusion-enriched transcripts were identified using direct RNA sequencing in an MDA-MB-231 cancer cell line16. The β-actin mRNA is the best-studied transcript with respect to local translation in migrating cells, and this mRNA is localized to protrusions in fibroblasts and myoblasts as well as to the growth cones of neurons17,18. Other examples of protrusion-localized mRNAs include transcripts encoding the actin polymerization nucleator Arp2/3 complex proteins. Both β-actin and Arp2/3 mRNAs contain a zip code fragment in their 3'UTRs, and these molecules are localized to the leading edge of migrating cells by the zip code-binding protein 1, also called IGF2BP319,20,21. These observations of local translation concurrent with cell cortex dynamics suggest that local translation may be important during other cellular events, such as cytokinesis, a cellular event characterized by dramatic cytoskeletal extension and remodeling.
RNA-binding proteins are a large group of regulators (800–1000 in humans), some of which play significant roles in mRNA local translation22,23. Indeed, the importance of maintaining protein homeostasis by RNA-binding proteins is highlighted by the numerous Mendelian genetic diseases caused by mutations in RNA-binding proteins24. In this study, we analyzed the function of RNP-1 and identified a role for this protein in protecting microtubule ends and directing cortical movement during cytokinesis and cell migration.
Materials and methods
cDNA library screening
cDNA library screening was performed as previously described4. Briefly, a Dictyostelium discoideum cDNA library prepared from vegetative cells was transformed into Ax3 (Rep orf+) cells as previously described25. One hundred pools of transformants (1000 transformants/pool) were cultured in suspension with nocodazole at the IC50 concentration (10 μmol/L) for approximately two weeks while monitoring growth rate. Pools with growth rates that were significantly higher than average (P<0.1) were extracted for plasmid DNA using a Wizard Plus Miniprep kit (Promega, Madison, WI, USA). The isolated plasmid DNA was then transformed into STBL2 cells (Invitrogen, Carlsbad, CA, USA). Once isolated, the plasmid DNA was then sequenced and identified using dictyBase BLAST. The plasmid was then reintroduced into Ax3 (Rep orf+) cells to confirm that the rescued growth rate in the presence of nocodazole was due to the isolated plasmid.
Growth rate measurement
Cells were transferred from plates to suspension culture at an initial concentration of ∼2×105 cells/mL. Cell density was determined every 24 h using a hemocytometer. After 4–5 d of monitoring, the cell densities were plotted against time, and the resulting log phase curve was fit to a single exponential equation. The wild-type control strain transfected with the empty vector pLD1 (labeled as EV in the figures) was normalized to 1, and the growth rates of the other strains within each experiment were normalized to the control cells.
Plasmid construction
The RNP-1 over-expression plasmid was constructed using the pLD1A15SN cassette containing the Ddp2-based origin of replication25. RNP-1-GFP was constructed from an in-house GFP-tagging cassette and cloned into pLD1A15SN25. When selecting the transformed cells, G418 was used at a concentration of 10 μg/mL, which was the same concentration used to maintain the cell line.
Live cell imaging
GFP- or RFP-labeled log phase cells were plated in imaging chambers. Immediately before imaging, the regular medium was replaced with MES (2-(N-morpholino)ethanesulfonic acid) starvation buffer (50 mmol/L MES pH 6.8, 2 mmol/L MgCl2 and 0.2 mmol/L CaCl2). Cells were imaged using a 40× (NA1.3) oil immersion objective and a 1.6× optovar on a Zeiss Axiovert microscope. Images were captured using a CCD camera operated using the MetaMorph Automation and Image Analysis software program. To image GFP-labeled proteins, z-stack images were taken of dividing or interphase cells, and the images were corrected using the MetaMorph 3D Deconvolution function. For GFP-microtubules, the 3D views were generated using the MetaMorph 3D reconstruction function.
Microtubule analysis
GFP images were obtained from live cells expressing GFP-tubulin and corrected by subtracting the background fluorescence. Control cells were transformed with the empty vector pLD1 (EV). Filamentous structures were identified manually and average intensity was measured using ImageJ. Cytosolic regions were identified manually as the areas without filamentous structures, and the average intensity was measured using ImageJ.
Cell migration
Cell movements toward a micropipette containing cAMP were performed as described previously26. Briefly, D discoideum cells were rinsed with development buffer (10 mmol/L phosphate buffer, 2 mmol/L MgSO4, and 0.2 mmol/L CaCl2, pH 6.5), resuspended in development buffer (2×107 cells/mL), and rotated on a shaker at 100 rounds per minute. After 1 h, the cell suspension was pulsed with doses of 50 nmol/L cAMP every 6 min. After 6 h of cAMP pulsing, cells were plated onto chambered coverslips. Once the cells had attached, the chambers were filled with 3 mL of development buffer, and a micropipette filled with 10 μmol/L cAMP was positioned near the cells. Images of moving cells were recorded using an Olympus inverted microscope equipped with a CCD camera. GFP intensity in the migrating cells was measured using ImageJ. The regions of leading edge were defined manually within each time frame.
Image processing and analysis
A program titled XM was written in Matlab® 2003a for image analysis. Briefly, images of GFP-labeled cells with marked centrosome locations were imported into Matlab and processed using the following morphological operations. First, the gray scale image was converted to a binary image based on a preset background threshold, such that the target cells appeared white on a black background. Second, if necessary, the background was cleaned by removing small objects (not the target cell) and filling in black holes within the target cell that were relatively dark before thresholding. Images 1 and 2 were then overlaid to verify the proper placement of the cell boundary. Third, an edge mask was generated by subtracting the target cell image from an eroded version of the image. Fourth, the pixels under the edge mask were selected, and their brightness values and corresponding distances toward the two centrosomes were extracted. Fifth, the distance values were regularized and reorganized into bins with a distance unit of 0.75 pixels, and the mean brightness values within each bin were calculated. Finally, the mean brightness values were plotted with increasing distance (minimal unit of 0.75 pixels), and their relationship was fitted using both the power model  and the exponential model
and the exponential model  , where y is the brightness value, x is the distance, and a1, b1, a2, b2 are the corresponding model parameters.
, where y is the brightness value, x is the distance, and a1, b1, a2, b2 are the corresponding model parameters.
Results
RNP-1 is a genetic suppressor of nocodazole
The inhibitory effects of nocodazole on microtubules in Dictyostelium cells were analyzed in cells expressing GFP-tubulin. Cells were incubated with 10 μmol/L nocodazole for 10 min, and microtubule structures were visualized using live cell imaging (Figure 1). In control (DMSO-treated) Dictyostelium cells, microtubule structures were extensive and reached to the cell cortex (cell edge; Figure 1A, left panel). A 3D reconstruction also illustrates the extensive microtubule array (Supplementary Movie S1). Upon treatment with 10 μmol/L nocodazole, these microtubule structures were greatly reduced (Figure 1A, right panel; Supplementary Movie S2).
Over-expression of RNP-1 rescues cells from nocodazole treatment. (A) Nocodazole caused microtubule structural changes in Dictyostelium cells. Ax3 cells expressing GFP-tubulin were treated with 10 μmol/L DMSO (Control, left) or 10 μmol/L nocodazole (Treatment, right) for 10 min. Scale bar, 5 μm. The 3D-reconstructed images of the microtubule network were taken immediately after the 10-min incubation (see Supplementary Movie S1 and S2). (B) Recapitulation of RNP-1 suppression of nocodazole treatment. Control cells carrying the empty vector (EV) or cells over-expressing RNP-1 (RNP-1-OE) were challenged with 10 μmol/L nocodazole (the IC50) or DMSO. Data are presented as mean±SEM (n>=6); **P<0.01, nsP<0.1. (C) Growth curves of control cells (EV) and RNP-1-OE cells with or without nocodazole over the course of 4 d.
In our previous study4, we performed a genetic selection screen to identify novel proteins that regulate microtubule dynamics in which nocodazole was used to inhibit the growth of library plasmid-transformed Dictyostelium cells. The transformed colonies were grown in suspension culture and challenged with 10 μmol/L nocodazole, the concentration that inhibits the cell growth rate by 50%4. During selection, most pools failed to show increased growth. However, in pools containing cells that received a suppressor plasmid (ie, a plasmid that rescues growth in nocodazole-challenged cells), the rescued cells eventually overtake the culture. In addition to 14-3-34 and a truncated version of enlazin27, the selection process also recovered plasmids containing full-length cDNAs for the rnp-1 gene and sterol-24-c methyltransferase (Table 1). The rnp-1 gene was recovered in 9 individual pools from the 100 pools of cDNA-transformed cells (each pool contained 100–1000 cDNA transformed colonies), whereas the other positive hits were each recovered one time (Table 1). To confirm that RNP-1 can rescue cells from the effects of nocodazole treatment, an RNP-1 over-expression plasmid and an RNP-1-GFP (C-terminus tag) construct were introduced into wild-type Dictyostelium cells, which were then challenged with 10 μmol/L nocodazole in suspension culture. The growth rates of these cells were then measured. When treated with nocodazole, control cells grew at 50% of the rate of DMSO-treated cells, whereas cells over-expressing RNP-1 and RNP-1-GFP grew at 87% and 62%, respectively, of the DMSO-treated cells (Figure 1B, 1C). The RNP-1-GFP construct did not rescue the growth inhibition from nocodazole to the same extent as wild-type RNP-1, but when it was compared with cells transfected with EV, RNP-1-GFP-expressing cells showed significantly increased growth in the presence of nocodazole. These results confirm that RNP-1 suppresses the growth inhibition induced by microtubule disruption due to nocodazole treatment.
RNP-1 protects microtubule structures during nocodazole treatment
We next generated a hairpin RNAi construct for RNP-1 knockdown, but upon transformation into wild-type Dictyostelium cells, no colonies grew, even when the selecting drug was titrated to a minimal concentration. These observations indicated that RNP-1 is necessary for cell survival and that wild-type expression levels are essential. Therefore, to determine the mechanism through which RNP-1 rescues cells from nocodazole treatment, we determined whether over-expression of RNP-1 could stabilize microtubules in the presence of nocodazole, as nocodazole is a microtubule destabilizer. We first tested the effect of RNP-1 on microtubules in cells cultured on surfaces in petri dishes. GFP-tubulin-expressing cells were transformed with empty vector or the RNP-1 over-expression vector and then incubated with DMSO or 10 μmol/L nocodazole overnight. Nocodazole treatment induced noticeable microtubule damage compared with DMSO treatment, but no significant microtubule changes were observed between cells transformed with empty vector and cells transformed with the RNP-1 over-expression (OE) vector upon nocodazole treatment (Supplementary Figure S1). As cells grown on surfaces have a different cytoskeletal composition (more polymerized actin) compared with cells grown in suspension28, a different growth context could add additional stress on the microtubule cytoskeleton. Therefore, we next tested the effects of RNP-1 on microtubules in suspension-grown cells. GFP-tubulin signals revealed that suspension-grown cells had fewer assembled microtubules than cells cultured on surfaces (Figure 1A and 2A). When treated with DMSO alone, both control cells (EV) and RNP-1-OE cells showed intact microtubule structures (Figure 2A), and the ratios of filamentous GFP intensity over cytosolic GFP intensity were 2.3 and 2.8, respectively (Figure 2B, n=6). However, after nocodazole treatment, control cells showed significant microtubule structural damage (Figure 2A), and the ratio of filamentous GFP intensity over cytosolic GFP intensity decreased to 1.4 (Figure 2B, n=6). When cells over-expressed RNP-1, their microtubules were more resistant to nocodazole treatment (Figure 2A), and the ratio of filamentous GFP intensity over cytosolic intensity remained significantly higher (2.0) compared with the control cells (EV) (Figure 2B, P<0.05, n≥6). Based on these results, we concluded that RNP-1 protects microtubules from nocodazole.
RNP-1 over-expression protects microtubules from nocodazole. (A) Control cells carrying an empty vector (EV) or RNP-1-OE cells expressing GFP-tubulin were treated with 10 μmol/L nocodazole or DMSO overnight and then grown in suspension culture. Images of microtubules were collected after incubation. A representative image of each condition is shown. Top left, control cells incubated with DMSO. Scale bar, 5 μm for all panels. Top right, control cells incubated with nocodazole. Bottom left, RNP-1-OE cells incubated with DMSO. Bottom right, RNP-1-OE cells incubated with nocodazole. (B) Measured filamentous GFP intensity normalized to cytosolic GFP intensity in control (EV) or RNP-1-OE cells treated with DMSO or nocodazole. **P<0.05, n>=6.
Localization of RNP-1-GFP in dividing cells
As microtubule structures are vital to cytokinesis and cell migration, we next assessed the role of RNP-1 in cytokinesis and cell migration. A plasmid encoding a C-terminal GFP-tagged RNP-1 was constructed, and it showed an ability to rescue cells from nocodazole treatment (Figure 1B). We then co-expressed RNP-1-GFP along with RFP-tubulin in cells. Live cell imaging of dividing cells showed that RNP-1-GFP was concentrated at the polar region when cells were undergoing cytokinesis (Figure 3A). By contrast, in interphase cells, RNP-1-GFP was primarily found in the cytosol. In addition, when the mitotic spindle was not centered in the cell, RNP-1-GFP concentration was increased at the polar cortex closest to the spindle (Figure 3A, Cell 1). Subsequently, the mitotic spindle re-centered, producing symmetrical cell division. To systematically evaluate whether the RNP-1-GFP signal was correlated with the cortex-centrosome distance, we developed a specific computer program to calculate the correlation between cortical GFP signal and the cortex-centrosome distance in dividing cells (Figure 3B). A total of five cells were analyzed, and the results are summarized in Figure 3C and Table 2. The power model  and the exponential model
and the exponential model  were used to fit the trends of GFP signal over the distance between the cortex and the centrosome. The power function fit relatively better than the exponential function, with overall higher R2 values, suggesting that the signal regulating RNP-1 diffuses from the centrosome and decreases proportionally as the power of the distance.
were used to fit the trends of GFP signal over the distance between the cortex and the centrosome. The power function fit relatively better than the exponential function, with overall higher R2 values, suggesting that the signal regulating RNP-1 diffuses from the centrosome and decreases proportionally as the power of the distance.
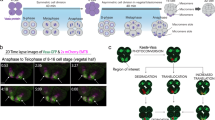
Localization of RNP-1-GFP in dividing cells. (A) WT (Ax3(Rep orf+)) cells expressing RFP-tubulin (left panel) and RNP-1-GFP (right panel) at the different stages of cytokinesis and interphase. (B) Representative images from each step of analysis in the Matlab® program XM. Arrowheads indicate centrosome positions, as marked by black dots. (C) The power function curve fitting of RNP-1-GFP signal versus cortical distance to the nearest centrosome in five analyzed cells.
Localization of RNP-1-GFP in migrating cells
The polar cortex of a dividing cell and the leading edge of a motile cell share many proteins in common, and they represent the boundaries of extensive cell shape changes. As RNP-1-GFP is highly localized to the polar regions of dividing cells, particularly in the cellular protrusions of these regions, we next examined the cellular localization of RNP-1-GFP in Dictyostelium cells during chemotactic migration. During cell migration, the microtubule network was a dynamic three-dimensional structure: in the anterior region, microtubules radiated toward the front and sides of the cell, whereas in the rear they appeared stretched and under tension13. We first analyzed the cellular distribution of RNP-1-GFP in chemotaxing cells. The top panels of Figure 4A and 4B show two images from a time series of chemotaxing cells expressing RNP-1-GFP. They both show that RNP-1-GFP was concentrated at the leading edges of the cells, suggesting that it may be regulated or localized by radiating microtubules at the leading edge (Figure 4, Supplementary Movie S1). To further investigate this hypothesis, we next measured the relative levels of GFP at the leading edges, which were normalized to the GFP signal in the rest of the cell body. The lower panels of Figure 4A and 4B show that there was an overall increased concentration of GFP (>1) at the leading edge compared with the rest of the cell body. Overall, we conclude that RNP-1 protects microtubules from the microtubule-destabilizing agent nocodazole and that it functions with microtubules to direct cell movement during cytokinesis and cell migration.
RNP-1-GFP localizes to the leading edge of chemotactic cells. (A) The top panel shows images from a time series of polarized, chemotaxing WT (Ax3(Rep orf+)) cells expressing RNP-1-GFP. Cells were chemotaxing towards cAMP, and the asterisk “*” indicates the location of the cAMP needle tip. The bottom panel shows changes in GFP distribution over time in one migrating cell (the white arrow in the top panel indicates the cell). The Y axis represents the ratio of GFP intensity (leading edge over the rest of the cell body). (B) A second set of example images. Top panel: a time series of chemotaxing cells; bottom panel: analyzed GFP distribution in one migrating cell. (A larger view with the location of the cAMP needle tip is included in panel B). The images shown here were taken from two representative recordings (see Supplementary Movies S3, S4).
Discussion
To investigate the cytokinesis machinery regulated by microtubules, we used a cDNA library to screen for suppressors of nocodazole treatment at a drug concentration that inhibits the cell growth rate by 50% (EC50). Using the same cDNA library selection for suppressors of nocodazole4, we identified several winner genes, which encoded 14-3-3, a fragment of Enlazin27, RNP-1, and sterol-24-c methyltransferase. Indeed, RNP-1 was recovered several times in the screen, indicating a robust ability to suppress nocodazole. However, the rescuing capability of a RNP-1-GFP construct was not as great as that of the non-GFP-tagged form of RNP-1, although the RNP-1-GFP did significantly rescue growth compared with control cells treated with nocodazole. Most likely, the 20 kDa GFP domain interfered with the natural structure of the RNP-1 protein, despite the fact that the RNP-1 and GFP were separated by a glycine/serine linkage. Another reason for this discrepancy could be that the expression levels of the GFP-tagged protein are lower than the wild type, as the overall molecular weight of the GFP-tagged RNP-1 is higher than RNP-1 alone. Based on other studies, the 14-3-3 and Enlazin proteins play major roles in regulating tension at the cell cortex; for example, cells in which 14-3-3 and enlazin were silenced exhibited reduced cortical tension, and 14-3-3 over-expression partially rescued mechanical defects in racE null cells4,27. RNP-1 may play a role in regulating the cell cortex by serving as a signal linkage between microtubules and the cell cortex.
Recently, using a proteomics approach, the RNP-1 protein was shown to be localized to the macropinocytosis compartment in Dictyostelium cells, suggesting that this protein plays a role in regulating cytoskeletal dynamics29. A bioinformatics analysis of the amino acid sequence using a motif scan revealed that the RNP-1 protein contains multiple kinase phosphorylation sites in a threonine-rich motif located at position 100–200 of the amino acid sequence (Figure 5), indicating that this protein could serve as a substrate for multiple kinases, such as PKA, PKC and Casein kinase II, which are critical regulators of cell migration30,31,32.
Proposed functional domains and sequence motifs of RNP-1. The protein sequence was analyzed using Expasy Prosite (Database of protein domains, families and functional sites) to obtain potential functional domains and sequence motifs.
In addition, sequence analysis revealed that this protein contains two eukaryotic RNA Recognition Motifs (RRMs) at positions 15–95 and 200–270 (Figure 5). A growing body of evidence suggests that mRNA metabolism is controlled by RNA-binding proteins33. Containing two RRMs, the RNP-1 protein could participate in the transport of mRNAs and related proteins to the cell periphery along microtubules, which is likely critical when cells are challenged with nocodazole. The idea of spatial regulation of protein translation has gained attention as an important process in a variety of cell biological processes. For example, local translation of mRNAs has been observed during neuronal development and in migrating cells14,16. The observation that local translation is often concurrent with dynamic changes at the cell cortex suggests that regional protein translation may occur during cytokinesis; in particular, mRNA directionally localizes to the peripheral region during cytokinesis, and protein translation may take place locally where those proteins are required. Thus, the presence of RNP-1 at the leading edges of cells undergoing cytokinesis and migrating cells suggests an important role for mRNA localization and local protein translation in these cellular processes. Finally, RNP-1-protected microtubules from nocodazole, which could involve RNP-1-mediated stabilization of tubulin mRNAs and related proteins to facilitate local protein translation.
We observed that RNP-1 localized to the cell cortex in dividing cells. The relationship between RNP-1 signaling and the distance to the nearest centrosome was modeled using both power models and exponential models. However, by comparison, we found that the power model significantly improved the fit to the relationship between the RNP-1 signals and the distance to the centrosome in most cases, indicating that the RNP-1 signal decreased proportionally with the power of the distance. The only exception was a single cell (Cell 5) that was in the later stages of cell separation. This observation appears reasonable, as it abides the general physical law that diffusive signals attenuate during propagation as a function of power of distance. In Table 2, “b1” is the power coefficient that represents the attenuation level with distance, ranging from -0.8865 to -0.4131, whereas “a1” represents the reference signal or the brightness value when distant from the 'center' based on a standard unit of distance, which ranged from 67.35 to 269.
During chemotaxis, RNP-1-GFP was concentrated at the leading edge of migrating cells. One could argue that this increased signal could have been due to increased volume (depth) at the leading edge of migrating cells. However, based on other studies, GFP or RFP controls were more uniformly distributed in migrating cells34, ruling out the possibility that the increased RNP-1 signal was due to increased volume alone. Nevertheless, it does appear that the RNP-1 signal is more cytosolic at the leading edge compared with membrane-associated proteins during chemotaxis, such as PTEN and PKC35. The distribution of RNP-1 during chemotaxis is very similar to that of actin in the pseudopod of a migrating cell36, further supporting the idea that the RNP-1 protein may be co-localizing with actin at the leading edge to facilitate actin protein translation. Other actin-associated proteins, such as coronin, were also found to have similar distribution patterns as RNP-1 in migrating cells.
When searching for RNP-1 homologs in other organisms, including humans and yeast, we identified a group of proteins called cytoplasmic polyadenylate-binding proteins (PABPCs) with high sequence similarities to RNP-1, particularly PABPC4 (BLAST expect value 5e-11, 24% sequence identities), PABPC3 (BLAST expect value 6e-11, 25% sequence identities) and PABPC1 (BLAST expect value 2e-10, 24% sequence identities) in humans. The PABPC proteins bind to the poly(A) tail of eukaryotic mRNAs to regulate mRNA dynamics and protein translation37. A recent study showed that PABPC can be enriched at sites of localized translation, such as the leading edges of migrating fibroblasts38 or neuronal dendrites39. In another study using atomic force microscopy, PABPC mediated the binding of mRNAs to microtubules40. Our sequence analysis also revealed that both PABPCs and RNP-1 proteins contain multiple RRMs with high pI values close to 9.5, indicating that they may have high affinities to poly-anionic molecules, such as RNA, microtubules and actin filaments41.
Taken together, we discovered that RNP-1, a PABPC-related protein, protected microtubules from nocodazole-induced damage and localized to leading edges in dividing and migrating cells. The RNP-1 protein may serve as part of the linkage between microtubules and the cell cortex by stabilizing microtubules and delivering regulatory information to the cell cortex. RNP-1 may help to stabilize and transport mRNAs to the peripheral region where localized protein translation is required. Our findings are consistent with previous studies of PABPC proteins, and they provide a functional demonstration of a relationship between RNP-1 and the microtubule network. Perhaps most interestingly, the unique localization of RNP-1 in Dictyostelium cells when cells are undergoing cytokinesis or chemotaxis suggests an important role for PABPC proteins in local protein translation when cells are undergoing dramatic cell shape changes in mammalian systems.
Author contribution
Thu NGO performed the sequence analysis of the RNP-1 protein and drafted Figure 5; Xin MIAO wrote the computer program XM in Matlab® and performed the analysis of GPF signals and their relationship with the cortical distance to the nearest centrosome; Douglas N ROBINSON designed and supervised the biological research, and he also helped with proofreading of the manuscript; Qiong-qiong ZHOU performed most of the biological research, carried out the data analysis, and wrote the manuscript.
References
Kuspa A, Dingermann T, Nellen W . Analysis of gene function in Dictyostelium. Experientia 1995; 51: 1116–23.
Torija P, Robles A, Escalante R . Optimization of a large-scale gene disruption protocol in Dictyostelium and analysis of conserved genes of unknown function. BMC Microbiol 2006; 6: 75.
De Brabander MJ, Van de Veire RM, Aerts FE, Borgers M, Janssen PA . The effects of methyl (5-(2-thienylcarbonyl)-1H-benzimidazol-2-yl) carbamate, (R 17934; NSC 238159), a new synthetic antitumoral drug interfering with microtubules, on mammalian cells cultured in vitro. Cancer Res 1976; 36: 905–16.
Zhou Q, Kee YS, Poirier CC, Jelinek C, Osborne J, Divi S, et al. 14-3-3 coordinates microtubules, Rac, and myosin II to control cell mechanics and cytokinesis. Curr Biol 2010; 20: 1881–9.
Babinec P, Babincova M . Self-organized criticality and dynamic instability of microtubule growth. Z Naturforsch C 1995; 50: 739–40.
Mitchison T, Kirschner M . Dynamic instability of microtubule growth. Nature 1984; 312: 237–42.
Akhmanova A, Steinmetz MO . Tracking the ends: a dynamic protein network controls the fate of microtubule tips. Nat Rev Mol Cell Biol 2008; 9: 309–22.
Lansbergen G, Akhmanova A . Microtubule plus end: a hub of cellular activities. Traffic 2006; 7: 499–507.
Etienne-Manneville S . Microtubules in cell migration. Annu Rev Cell Dev Biol 2013; 29: 471–99.
Daou P, Hasan S, Breitsprecher D, Baudelet E, Camoin L, Audebert S, et al. Essential and nonredundant roles for Diaphanous formins in cortical microtubule capture and directed cell migration. Mol Biol Cell 2014; 25: 658–68.
Li D, Gao J, Yang Y, Sun L, Suo S, Luo Y, et al. CYLD coordinates with EB1 to regulate microtubule dynamics and cell migration. Cell Cycle 2014; 13: 974–83.
Weng JH, Liang MR, Chen CH, Tong SK, Huang TC, Lee SP, et al. Pregnenolone activates CLIP-170 to promote microtubule growth and cell migration. Nat Chem Biol 2013; 9: 636–42.
Tang L, Franca-Koh J, Xiong Y, Chen MY, Long Y, Bickford RM, et al. tsunami, the Dictyostelium homolog of the Fused kinase, is required for polarization and chemotaxis. Genes Dev 2008; 22: 2278–90.
Dynes JL, Steward O . Arc mRNA docks precisely at the base of individual dendritic spines indicating the existence of a specialized microdomain for synapse-specific mRNA translation. J Comp Neurol 2012; 520: 3105–19.
Liu-Yesucevitz L, Bassell GJ, Gitler AD, Hart AC, Klann E, Richter JD, et al. Local RNA translation at the synapse and in disease. J Neurosci 2011; 31: 16086–93.
Jakobsen KR, Sorensen E, Brondum KK, Daugaard TF, Thomsen R, Nielsen AL . Direct RNA sequencing mediated identification of mRNA localized in protrusions of human MDA-MB-231 metastatic breast cancer cells. J Mol Signal 2013; 8: 9.
Lawrence JB, Singer RH . Intracellular localization of messenger RNAs for cytoskeletal proteins. Cell 1986; 45: 407–15.
Bassell GJ, Singer RH . Neuronal RNA localization and the cytoskeleton. Results Probl Cell Differ 2001; 34: 41–56.
Huttelmaier S, Zenklusen D, Lederer M, Dictenberg J, Lorenz M, Meng X, et al. Spatial regulation of beta-actin translation by Src-dependent phosphorylation of ZBP1. Nature 2005; 438: 512–5.
Rodriguez AJ, Shenoy SM, Singer RH, Condeelis J . Visualization of mRNA translation in living cells. J Cell Biol 2006; 175: 67–76.
Gu W, Katz Z, Wu B, Park HY, Li D, Lin S, et al. Regulation of local expression of cell adhesion and motility-related mRNAs in breast cancer cells by IMP1/ZBP1. J Cell Sci 2012; 125: 81–91.
Castello A, Fischer B, Eichelbaum K, Horos R, Beckmann BM, Strein C, et al. Insights into RNA biology from an atlas of mammalian mRNA-binding proteins. Cell 2012; 149: 1393–406.
Glisovic T, Bachorik JL, Yong J, Dreyfuss G . RNA-binding proteins and post-transcriptional gene regulation. FEBS Lett 2008; 582: 1977–86.
Castello A, Fischer B, Hentze MW, Preiss T . RNA-binding proteins in Mendelian disease. Trends Genet 2013; 29: 318–27.
Robinson DN, Spudich JA . Dynacortin, a genetic link between equatorial contractility and global shape control discovered by library complementation of a Dictyostelium discoideum cytokinesis mutant. J Cell Biol 2000; 150: 823–38.
Parent CA, Blacklock BJ, Froehlich WM, Murphy DB, Devreotes PN . G protein signaling events are activated at the leading edge of chemotactic cells. Cell 1998; 95: 81–91.
Octtaviani E, Effler JC, Robinson DN . Enlazin, a natural fusion of two classes of canonical cytoskeletal proteins, contributes to cytokinesis dynamics. Mol Biol Cell 2006; 17: 5275–86.
Gerald N, Dai J, Ting-Beall HP, De Lozanne A . A role for Dictyostelium racE in cortical tension and cleavage furrow progression. J Cell Biol 1998; 141: 483–92.
Journet A, Klein G, Brugiere S, Vandenbrouck Y, Chapel A, Kieffer S, et al. Investigating the macropinocytic proteome of Dictyostelium amoebae by high-resolution mass spectrometry. Proteomics 2012; 12: 241–5.
Newell-Litwa KA, Horwitz AR . Cell migration: PKA and RhoA set the pace. Curr Biol 2011; 21: R596–8.
Kermorgant S, Zicha D, Parker PJ . PKC controls HGF-dependent c-Met traffic, signalling and cell migration. EMBO J 2004; 23: 3721–34.
Delorme V, Cayla X, Faure G, Garcia A, Tardieux I . Actin dynamics is controlled by a casein kinase II and phosphatase 2C interplay on Toxoplasma gondii Toxofilin. Mol Biol Cell 2003; 14: 1900–12.
Pichon X, Wilson LA, Stoneley M, Bastide A, King HA . Somers J, et al. RNA binding protein/RNA element interactions and the control of translation. Curr Protein Pept Sci 2012; 13: 294–304.
Skoge M, Adler M, Groisman A, Levine H, Loomis WF, Rappel WJ . Gradient sensing in defined chemotactic fields. Integr Biol (Camb) 2010; 2: 659–68.
Iijima M, Devreotes P . Tumor suppressor PTEN mediates sensing of chemoattractant gradients. Cell 2002; 109: 599–610.
Yumura S, Fukui Y . Spatiotemporal dynamics of actin concentration during cytokinesis and locomotion in Dictyostelium. J Cell Sci 1998; 111: 2097–108.
Smith RW, Blee TK, Gray NK . Poly(A)-binding proteins are required for diverse biological processes in metazoans. Biochem Soc Trans 2014; 42: 1229–37.
Woods AJ, Roberts MS, Choudhary J, Barry ST, Mazaki Y . Sabe H, et al. Paxillin associates with poly(A)-binding protein 1 at the dense endoplasmic reticulum and the leading edge of migrating cells. J Biol Chem 2002; 277: 6428–37.
Muddashetty R, Khanam T, Kondrashov A, Bundman M, Iacoangeli A . Kremerskothen J, et al. Poly(A)-binding protein is associated with neuronal BC1 and BC200 ribonucleoprotein particles. J Mol Biol 2002; 321: 433–45.
Chernov KG, Curmi PA, Hamon L, Mechulam A, Ovchinnikov LP, Pastre D . Atomic force microscopy reveals binding of mRNA to microtubules mediated by two major mRNP proteins YB-1 and PABP. FEBS Lett 2008; 582: 2875–81.
Jones LS, Yazzie B, Middaugh CR . Polyanions and the proteome. Mol Cell Proteomics 2004; 3: 746–69.
Acknowledgements
The majority of the bench experiments were performed in the laboratory of Dr Douglas N ROBINSON, and we thank him for his generous support. We also thank the NIH (R01 GM66817 to Douglas N ROBINSON) and AHA (Qiong-qiong ZHOU's fellowship) for support.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary information is available on the website of Acta Pharmacologica Sinica.
Supplementary information
Supplementary Movie S1
GFP-tubulin labeled 3D microtubule network in a Dictyostelium cell treated with 10 μmol/L of DMSO. (AVI 321 kb)
Supplementary Movie S2
GFP-tubulin labeled 3D microtubule network in a Dictyostelium cell treated with 10 μmol/L of nocodazole. (AVI 456 kb)
Supplementary Movie S3
RNP-1-GFP localization in Dictyostelium cells during chemotaxing toward cAMP needles. (AVI 171 kb)
Supplementary Movie S4
RNP-1-GFP localization in Dictyostelium cells during chemotaxing toward cAMP needles. (AVI 168 kb)
Supplementary Figure S1
Microtubule networks in the surface cultured cells after overnight nacodazole treatment. (JPG 1135 kb)
Rights and permissions
About this article
Cite this article
Ngo, T., Miao, X., Robinson, D. et al. An RNA-binding protein, RNP-1, protects microtubules from nocodazole and localizes to the leading edge during cytokinesis and cell migration in Dictyostelium cells. Acta Pharmacol Sin 37, 1449–1457 (2016). https://doi.org/10.1038/aps.2016.57
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/aps.2016.57