Abstract
Aim:
Increasing evidence shows that mRNAs exert regulatory function along with coding proteins. Recently we report that a hairpin within YAP mRNA 3′UTR can modulate the Hippo signaling pathway. PTEN is a tumor suppressor, and is mutated in human cancers. In this study we examined whether PTEN mRNA 3′UTR contained a hairpin structure that could regulate gene regulation at the post-transcriptional level.
Methods:
The secondary structure of PTEN mRNA 3′UTR was analyzed using RNAdraw and RNAstructure. Function of hairpin structure derived from the PTEN mRNA 3′UTR was examined using luciferase reporter assay, RT-PCR and Western blotting. RNA-immunoprecipitation (RIP) assay was used to analyze the interaction between PTEN mRNA and microprocessor Drosha and DGCR8. Endogenous siRNA (esiRNA) derived from PTEN mRNA 3′UTR was identified by RT-PCR and rt-PCR, and its target genes were predicted using RNAhybrid.
Results:
A bioinformatics analysis revealed that PTEN mRNA contained a hairpin structure (termed PTEN-sh) within 3′UTR, which markedly increased the reporter activities of AP-1 and NF-κB in 293T cells. Moreover, treatment with PTEN-sh (1 and 2 μg) dose-dependently inhibited the expression of PTEN in human liver L-O2 cells. RIP assay demonstrated that the microprocessor Drosha and DGCR8 was bound to PTEN-sh in L-O2 cells, leading to the cleavage of PTEN-sh from PTEN mRNA 3′UTR. In addition, microprocessor Dicer was involved in the processing of PTEN-sh. Interestingly, esiRNA (termed PTEN-sh-3p21) cleaved from PTEN-sh was identified in 293T cells and human liver tissues, which was found to target the mRNA 3′UTRs of protein phosphatase PPP2CA and PTEN in L-O2 cells. Treatment of L-O2 or Chang liver cells with PTEN-sh-3p21 (50, 100 nmol/L) promoted the cell proliferation in dose- and time-dependent manners.
Conclusion:
The endogenous siRNA (PTEN-sh-3p21) cleaved from PTEN-sh within PTEN mRNA 3′UTR modulates PPP2CA and PTEN at the post-transcriptional level in liver cells.
Similar content being viewed by others
Introduction
The regulatory function of RNAs in gene expression has been widely studied1,2 to assess the ways in which network regulation plays crucial roles in development, stem cell maintenance and memory formation3,4. MicroRNAs are small regulatory RNAs that are processed from stem-loop regions of primary transcripts. The nuclear microprocessor complex (composed of Drosha and DGCR8) and Dicer are double strand RNA binding proteins that are critical in microRNA maturation5. Drosha and DGCR8 directly modulate gene expression and RNA metabolism at different stages, such as transcriptional initiation and termination and the processing of various RNA species, including pre-mRNAs6. Dicer isoforms are involved in apoptosis, development and diseases7,8,9. Recently, we have reported that a hairpin within the YAP mRNA 3′UTR can serve as a regulatory element modulating the Hippo signaling pathway10. However, the regulatory role of the hairpin structure within the 3′UTR is poorly understood.
PTEN is an important tumor suppressor, and loss of PTEN function has been implicated in a wide variety of tumor types11. The PTEN-PI3K-AKT-mTOR pathway plays an essential role in the regulation of glucose metabolism, owing to its position downstream of the insulin receptor (INSR) and IRS adaptor molecules. Increasing evidence supports the hypothesis that mRNAs can serve as regulatory RNAs. PTEN mRNA functions in the regulation of gene expression by competing with common microRNAs12. PTEN expression can be post-transcriptionally modulated by a PTEN pseudogene, whose transcripts exert regulatory control of their ancestral cancer gene's expression levels by competing with microRNAs that target sequences common to the mRNA and the pseudo-mRNA13,14. However, whether the PTEN mRNA 3′UTR contains hairpin structure and functions in regulation are not well documented.
In this study, we sought to determine whether PTEN mRNA functions in regulation at the transcriptional level. Interestingly, we identified an endogenous siRNA cleaved from a hairpin within the PTEN mRNA 3′UTR in liver cells. This endogenous siRNA regulates target genes, such as PPP2CA and PTEN, at the post-transcriptional level. Our finding provides new insights into the mechanism by which PTEN mRNA functions in regulation.
Materials and methods
Cell culture and transfection
The human embryonic kidney cell line 293T was maintained in Dulbecco's modified Eagle's medium (Gibco, Grand Island, NY, USA)15. Hepatocytes from human immortalized normal liver L-O2 and Chang liver cell lines were cultured in RPMI Medium 1640 (Gibco) supplemented with 10% fetal calf serum (FCS), 100 U/mL penicillin, and 100 μg/mL streptomycin in 5% CO2 at 37 °C. The cells were cultured in 6-well or 24-well plates for 12 h and were then transfected with plasmid or siRNA. All transfections were conducted using Lipofectamine 2000 reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer's protocol.
Plasmid construction
The complementary DNA sequences of PTEN-sh (5′-CGCGGATCCAAAAAAAAGACATTTGATTTTTCAGTAGAAATTGTCCTACATGTGCTTTATTGATTTGCTATTGAAAGAATAGGGTTTTTTTTTTGGTACCCCG-3′), PTEN-sh-non (5′-CGCGGATCCAGCGTTTTTTTTCTTTGAAGATTTATGATGCACTTATTCAATAGCTGTCAGCCGTTCCACCCTTTTGACCTTACACATTCTATTAGGTACCCCG-3′) and PTEN-sh-mut (5′-CGCGGATCCAAATATTACATTAAGGGTTAAGTGATTTGATTCAGTAATTGTGCTATTCCTACATGTGCTTTATAGATTAAAGAATTGATTTTTTGGTACCCCG-3′) were synthesized by Augct (Beijing, China) and were then cloned into a pRNAT-U6.1/neo vector via BamH I and Kpn I sites to evaluate the effects of the hairpin on cell activity. The PTEN 3′UTR fragments containing PTEN-sh and PPP2CA 3′UTR were inserted downstream of the pGL3-control vector (Promega, USA) via Fse I and Xba I sites to measure the effects of PTEN-sh or PTEN-sh-3p21 on PTEN and PPP2CA at the post-transcriptional level. Mutant constructs of PPP2CA 3′UTR, carrying a substitution of four nucleotides, were cloned into a pGL3-control vector using overlapping extension PCR to evaluate the binding ability of esiRNA. The primers for plasmid construction are listed in Supplementary Table S1. The PTEN-sh-3p21 PCR products derived from 293T cells and 14 samples were inserted into a pEASY-T1 vector and sequenced (BGI, Beijing, China).
Total RNA isolation, RT-PCR, real-time PCR and walking PCR
Total RNA was extracted from the cells (or liver tissues) using TRIzol (Invitrogen, USA) according to the manufacturer's protocol. To test small RNA derived from PTEN-sh, total RNA was polyadenylated by poly (A) polymerase (Ambion, Austin, TX, USA), as previously described10. Reverse transcription was performed using poly (A)-tailed total RNA and reverse transcription primer (5′-GCGAGCACAGAATTAATACGACTACTATAGGTTTTTTTTTTTTTTTTTTVN-3′) with ImPro-II Reverse Transcriptase (Promega, USA) according to the manufacturer's protocol. Real-time PCR was conducted using a Bio-Rad sequence detection system according to the manufacturer's instructions, with double-stranded DNA-specific SYBR GreenPremix Ex TaqTM II Kit (TaKaRa Bio, Dalian, China). Relative transcriptional fold changes were calculated as 2-ΔΔCt. U6 was used as an internal control to normalize small RNA levels. GAPDH was used as an internal control to normalize PTEN mRNA levels. The primers for RT-PCR, real-time PCR and walking PCR are listed in Supplementary Table S1. More details are described in the supplementary materials.
Luciferase reporter assay
A luciferase reporter assay was conducted using the Dual-Luciferase Reporter Assay System (Promega, USA) according to the manufacturer's instructions. The 293T cells (3×104/per well) were seeded into 24-well plates. After 12 h, the cells were transiently co-transfected with 0.1 μg/well of pRL-TK plasmid (Promega) containing the Renilla luciferase gene used for internal normalization and various constructs containing pGL3-Ap-1, pGL3-NF-κB, pGL3-PTEN 3′UTR, pGL3-PPP2CA 3′UTR-wt and pGL3-PPP2CA 3′UTR-mut. Cells were lysed and assayed for luciferase activity 36 h after transfection. One hundred microliters of protein extracts was analyzed in a luminometer. To evaluate the response of cells to the over-expression of hairpin structures, the AP-1 and NF-κB reporter systems were used in 293T cells10. The luciferase activities were measured as previously described16. All experiments were performed at least three times.
RNA-immunoprecipitation (RIP)
An RIP assay was conducted in native conditions as previously described17. Briefly, L-O2 cells were pelleted and lysed. The lysates were passed through a 27.5 gauge needle 3 times for nuclear lysis. The supernatant was incubated with 2 μg of primary rabbit anti-Drosha antibody (Proteintech, Chicago, IL, USA), rabbit anti-DGCR8 (Proteintech, Chicago, IL, USA) or IgG (Sigma-Aldrich, St Louis, MO, USA) with 50 μL protein G-conjugated agarose beads (Millipore). The RNA/antibody complex was washed with NT2 buffer (50 mmol/L Tris-HCl pH 7.4, 150 mmol/L NaCl, 1 mmol/L MgCl2, 0.05% NP-40). The RNA was extracted with TRIzol (Invitrogen) per the manufacturer's protocol and subjected to RT-PCR analysis.
Western blot analysis
Total protein lysate was extracted from L-O2 cells with RIPA buffer. Protein concentrations were calculated using the Bradford assay, and 20–40 μg of protein extracts was subjected to SDS-PAGE. Then, proteins were transferred to a nitrocellulose membrane, blocked with 5% non-fat milk and incubated with primary antibodies. Primary antibodies were rabbit anti-PPP2CA (Proteintech, Chicago, IL, USA), rabbit anti-Akt (Santa Cruz, CA, USA), mouse anti-PTEN (Santa Cruz, USA), mouse anti-phosphorylated Akt (Cell Signaling, USA), rabbit anti-Dicer (Boster, Wuhan, China) and mouse anti-β-actin (Sigma-Aldrich, St Louis, MO, USA). The intensity for each band was quantified by software Image J. Each experiment was repeated three times.
Tissue samples
Thirty non-tumorous, human liver tissues from HCC patients were used in this study. The tissues were obtained from Tianjin First Center Hospital (Tianjin, China) immediately after surgical resection. Detailed information about the patients was obtained from patient records. Written consent approving the use of their tissues for research purposes was obtained from each patient after the operation. The study protocol was approved by the Institutional Research Ethics Committee at Nankai University (Tianjin, China).
Oligonucleotides
Small interfering RNAs (siDicer: 5′-UUUGUUGCGAGGCUGAUUC-3′; siDrosha: 5′-AACGAGUAGGCUUCGUGACUU-3′; siDGCR8: 5′-AACGAUGAUGACCAAGAUUAA-3′) and negative control siRNA (siNC) were synthesized by RiboBio (Guangzhou, China)18,19. The PTEN-sh-3p21 mimics (5′-GAUUUGCUAUUGAAAGAAUA-3′) and mimics negative control (mimics NC) were purchased from RiboBio (Guangzhou, China).
Analysis of cell proliferation
L-O2 cells and Chang liver cells were seeded on 96-well plates (1000 cells/well) for 12 h before transfection, and a 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay was used to assess cell proliferation every day from the first day until the third day after transfection. The protocol has been described previously16. A five-ethynyl-2'-deoxyuridine (EdU) incorporation assay was performed using the Cell-LightTM EdU imaging detecting kit according to the manufacturer's instructions (RiboBio, Guangzhou, China).
Statistical analysis
Statistical significance was assessed by comparing mean values (±standard deviation; SD), using an unpaired Student's two-tailed t-test for independent groups. P<0.05 and P<0.01 were considered to be significant. Pearson's correlation coefficient was used to evaluate the relationship between the levels of PTEN-sh-3p21 and PTEN in non-tumorous, human liver tissues. Each experiment was repeated at least three times.
Results
A hairpin within the PTEN mRNA 3′UTR is responsible for regulatory function
To evaluate whether PTEN mRNA has a regulatory function, we analyzed the secondary structure of PTEN mRNA by bioinformatics analysis using RNAdraw20 and RNAstructure21. Interestingly, we observed that the secondary structure of PTEN 3′UTR was complicated at different free energy levels, and the secondary structure analysis displayed a hairpin structure (termed PTEN-sh) that was relatively stable as compared with other hairpin structures, such as the PTEN non-standard hairpin (PTEN-non-sh) (Figure 1A). Then, we constructed a vector using the U6 promoter for over-expression of PTEN-sh. It has been reported that the AP-1 and NF-κB transcription factors modulate the expression of numerous target genes related to lipid metabolism, proliferation, apoptosis, angiogenesis and metastasis in the cells, and the AP-1 and NF-κB reporter system can be used to evaluate the response to over-expression of YAP-sh in 293T cells10,22,23. Accordingly, in this study we used this system to evaluate the response of cells to the over-expression of PTEN-sh. Interestingly, a luciferase reporter assay showed that PTEN-sh increased the activities of AP-1 and NF-κB in 293T cells, whereas the PTEN-non-sh did not (Figure 1B). Next, we cloned the mutant of PTEN-sh, termed PTEN-sh-mut (Figure 1C). Interestingly, PTEN-sh-mut did not activate the AP-1 and NF-κB reporters (Figure 1D). To better understand the biological significance of this observation, we aligned the PTEN-sh sequence in different species and observed that the sequence was highly conserved in higher mammals (Figure 1E), suggesting that PTEN-sh is a conserved regulatory element. Thus, we conclude that PTEN-sh, a hairpin within the PTEN mRNA 3′UTR, is responsible for its regulatory function.
A hairpin within the PTEN mRNA 3′UTR is responsible for its regulatory function. (A) Diagram of the secondary structure of PTEN 3′UTR, including PTEN-sh and PTEN-non-sh, analyzed by using RNAstructure and RNAdraw. (B) The effects of PTEN-sh and PTEN-non-sh on AP-1 and NF-κB promoter activities were examined with a luciferase reporter assay in 293T cells. (C) The secondary structure of PTEN-sh-mut with a crippled PTEN-sh sequence was analyzed using RNAstructure and RNAdraw. (D) The effect of PTEN-sh and PTEN-sh-mut on the promoter activities of AP-1 and NF-κB was examined with a luciferase reporter assay in 293T cells. Vector indicates empty plasmid DNA. (E) The sequence alignment of PTEN-sh in different species. The sequence alignment was performed with the CLC sequence viewer 6.3 software . Each experiment was repeated three times. Mean±SD. n=3. **P<0.01 vs vector, unpaired Student's two-tailed t-test.
PTEN-sh is processed by micro-processor Drosha/DGCR8 and Dicer and functions at the post-transcriptional level
To better understand the function of PTEN-sh, we evaluated the effect of PTEN-sh from the PTEN mRNA 3′UTR on PTEN expression regulation with a luciferase reporter assay, because the luciferase reporter assay has been shown to reflect the interaction of microRNAs with their target genes24. Strikingly, we observed that the luciferase activities of pGL3-PTEN-3′UTR were attenuated by treatment with PTEN-sh in 293T cells in a dose-dependent manner (Figure 2A). Moreover, a Western blot analysis showed that the over-expression of PTEN-sh led to down-regulation of PTEN at the protein level in a dose-dependent manner. In addition, we validated that the levels of phosphorylated-Akt (p-Akt), a downstream effector of PTEN, were also increased in the system, but not by PTEN-sh-mut (Figure 2B), thus suggesting that PTEN-sh functions in regulating PTEN. Previous studies have shown that Drosha and DGCR8 are required for the generation of the heterotetramer in hairpin structure processing25,26. Interestingly, the luciferase reporter assay showed that the activities of pGL3-PTEN-3′UTR increased in 293T cells when Drosha or DGCR8 was knocked down by siRNA (Figure 2C). Moreover, an RT-PCR assay showed that the levels of PTEN mRNA were increased in L-O2 cells treated with Drosha or DGCR8 siRNA (Figure 2D). An RIP analysis showed that Drosha or DGCR8 bound to PTEN mRNA in cells (Figure 2E), thus suggesting that PTEN-sh is processed by micro-processor Drosha and DGCR8.
PTEN-sh is processed by micro-processor Drosha/DGCR8 and Dicer and functions at the post-transcriptional level. (A) The effect of PTEN-sh on the luciferase activities of pGL3-PTEN-3′UTR was examined with a luciferase reporter assay in 293T cells. Vector indicates empty plasmid DNA. (B) The effect of PTEN-sh and PTEN-sh-mut on the expression of PTEN and p-Akt level was assessed with a Western blot analysis in L-O2 cells. (C) The effects of Drosha and DGCR8 on luciferase activities of pGL3-PTEN-3′UTR were measured with a luciferase reporter assay in 293T cells. (D) PTEN mRNA levels were examined by RT-PCR in L-O2 cells when Drosha and DGCR8 were knocked down by siRNA. (E) The interaction between PTEN mRNA and Drosha or DGCR8 was examined with an RIP analysis. (F) The effects of 100 nmol/L siDicer on the luciferase activities of pGL3-PTEN-3′UTR induced by PTEN-sh were detected in 293T cells. Mean±SD. n=3. Each experiment was repeated three times. *P<0.05, **P<0.01 vs vector. NSP>0.05, #P<0.05, ##P<0.01 vs siNC, unpaired Student's two-tailed t-test.
Given that Dicer can recognize stem-loop or double-strand RNA and cleave it into ∼22 nucleotide long fragments27,28,29, we evaluated the effect of siDicer on PTEN-sh processing. Our data showed that PTEN-sh-decreased luciferase activities of pGL3-PTEN-3′UTR were rescued by siDicer in 293T cells (Figure 2F), in which the efficiency of siDicer was validated (Supplementary Figure 1A), thus suggesting that Dicer is involved in the processing of PTEN-sh. When Dicer is knocked down, the expression of microRNAs is subsequently affected. Therefore, we used bioinformatics to analyze the microRNAs targeting the 3′UTR of PTEN mRNA, using the DIANA, TargetScan and Miranda databases. It has been reported that miR-19a/b and 26a/b target the PTEN mRNA 3′UTR30. Interestingly, miR-19a/b and 26a/b targeted the region (from 1013–1497 nt), which we cloned for further study of the mechanisms of the hairpin structure within the PTEN mRNA 3′UTR (Supplementary Figure 1B). However, our data showed that the expression levels of miR-19a/b and miR-26a/b were not changed when Dicer was knocked down in 293T cells (Supplementary Figure 1C), thus suggesting that the stability of the PTEN mRNA 3′UTR is affected by Dicer-mediated PTEN-sh, rather than by miR-19 and miR-26. Therefore, we concluded that PTEN-sh is processed by the micro-processor Drosha/DGCR8 and Dicer and functions at the post-transcriptional level.
An esiRNA, derived from PTEN-sh, in 293T cells and human liver tissues
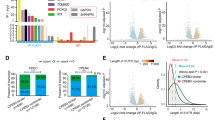
We have reported that a hairpin within the YAP mRNA 3′UTR (YAP-sh) is processed into an endogenous siRNA10. Accordingly, we hypothesized that an endogenous siRNA might be cleaved from PTEN-sh. Different walking PCR primers for PTEN-sh were designed (Figure 3A). Interestingly, walking PCR using PTEN-sh-3p21 primers detected a fragment of 20 nucleotides in 293T cells transfected with PTEN-sh (Figure 3B). The PTEN-sh-3p21 was found to be located within the PTEN mRNA 3′UTR (Supplementary Figure 2). Real-time PCR analysis validated that the expression levels of PTEN-sh-3p21 were elevated in the PTEN-sh over-expressing 293T cells (Figure 3C). Furthermore, a fragment of PTEN-sh-3p21 was detected in 14 out of 20 human paratumor liver samples (Figure 3D). All PCR products from 293T cells and 14 samples were validated by sequencing, thus suggesting that PTEN-sh-3p21 is an esiRNA cleaved from PTEN-sh. Moreover, real-time PCR validated that the PTEN-sh-3p21 level was inversely correlated with that of PTEN expression in 30 human non-tumorous liver tissues (Figure 3E). Thus, we conclude that an esiRNA, PTEN-sh-3p21, is formed from cleavage of PTEN-sh in the cells.
An esiRNA, derived from PTEN-sh was identified in 293T cells and human liver tissues. (A) The diagram shows the designed primers for esiRNAs derived from PTEN-sh. (B) The esiRNAs were tested by walking RT-PCR using designed primers in PTEN-sh-overexpressing 293T cells. (C) The identified PTEN-sh-21, an esiRNA, was validated by real-time PCR in the cells. (D) PTEN-sh-3p21 was validated with RT-PCR in 14 out of 20 human non-tumorous liver tissues. (E) The correlation between PTEN-sh-3p21 and PTEN expression was examined by using real-time PCR in human non-tumorous liver tissues. The expression of PTEN was normalized to that of GAPDH. The PTEN-sh-3p21 expression was normalized to that of U6. Statistical analysis was performed using Pearson's correlation coefficient (r=−0.5546, P<0.01). Error bars represent SD. n=3. NC, purified water; Mock, without plasmid DNA; Vector, empty plasmid DNA. Each experiment was repeated three times. **P<0.01 vs vector, unpaired Student's two-tailed t-test.
PTEN-sh-3p21 targets the 3′UTR of protein phosphatase PPP2CA or PTEN
Next, we predicted that protein phosphatase PPP2CA is one of the target genes of PTEN-sh-3p21 by using RNAhybrid software31,32 (Figure 4A). However, we also predicted that the PTEN mRNA is a target of PTEN-sh-3p21. It has been reported that both PPP2CA and PTEN are members of the PI3K/PTEN/Akt pathway33. Accordingly, we cloned the 3′UTR of PPP2CA mRNA (termed 3′UTR-wt) and a mutant PPP2CA 3′UTR (termed 3′UTR-mut) into the pGL3-control vector (Figure 4B). Interestingly, PTEN-sh-3p21 attenuated the luciferase activities of PPP2CA 3′UTR-wt, but not the PPP2CA 3′UTR-mut in 293T cells (Figure 4C). Furthermore, Western blot analysis showed that PTEN-sh-3p21 down-regulated PPP2CA and PTEN at the protein level (Figure 4D), supporting the hypothesis that PTEN-sh-3p21 is an esiRNA that is able to silence target genes, such as PPP2CA and PTEN. Therefore, we conclude that the esiRNA, PTEN-sh-3p21, targets the mRNA 3′UTR of PPP2CA or PTEN.
PTEN-sh-3p21 targets the 3′UTR of protein phosphatase PPP2CA or PTEN. (A) A model of the PPP2CA locus, showing putative target sites for PTEN-sh-3p21. (B) Luciferase reporter constructs showing the wild type and mutant PPP2CA mRNA 3′UTRs. (C) The effect of PTEN-sh-3p21 on the PPP2CA mRNA 3′UTR was examined by luciferase reporter assay in 293T cells. (D) The expression of PPP2CA and PTEN was tested by Western blot analysis in L-O2 cells treated with PTEN-sh-3p21. Mean±SD. n=3. Each experiment was repeated three times. NSP>0.05, **P<0.01 vs mimics NC, unpaired Student's two-tailed t-test.
PTEN-sh-3p21 enhances cell proliferation in vitro
To better understand the significance of PTEN-sh-3p21 function, we tested the effect of PTEN-sh-3p21 on cell proliferation. Using an MTT assay, we showed that PTEN-sh-3p21 enhanced the proliferation of L-O2 cells (Figure 5A). An EdU incorporation assay also confirmed that PTEN-sh-3p21 promoted the proliferation of L-O2 cells (Figure 5B). Moreover, using MTT and EdU incorporation assays in the Chang liver cell line, we validated that PTEN-sh-3p21 promoted liver cell proliferation (Figure 5C and 5D). These findings suggest that PTEN-sh-3p21 functions as an esiRNA. Therefore, we conclude that PTEN-sh-3p21, an esiRNA derived from PTEN-sh, enhances cell proliferation in vitro.
PTEN-sh-3p21 enhances cell proliferation in vitro. (A) The effect of PTEN-sh-3p21 on cell proliferation was determined with an MTT assay in L-O2 cells. (B) The effect of PTEN-sh-3p21 on cell proliferation was examined with an EdU incorporation assay in L-O2 cells. The EdU positive cell population was analyzed with the software Image J. (C and D) The effect of PTEN-sh-3p21 on cell proliferation in Chang liver cells was examined by using an MTT assay and EdU incorporation assay. The EdU positive cell population was analyzed with the software Image J. Mean±SD. n=3. Each experiment was repeated three times. *P<0.05, **P<0.01 vs mimics NC, unpaired Student's two-tailed t-test. (E) A model of the functions of the hairpin within the PTEN mRNA 3′UTR in gene regulation at the post-transcriptional level in liver cells. Bioinformatics analysis indicated the secondary structure of the PTEN mRNA 3′UTR, revealing a hairpin (PTEN-sh). PTEN-sh is recognized and processed by Drosha and DGCR8. PTEN-sh generates esiRNAs (PTEN-sh-3p21) after cleavage by Dicer, resulting in instability and degradation of the PTEN mRNA 3′UTR. PTEN-sh-3p21 targeting the 3′UTRs of PTEN or PPP2CA leads to an increase in the Akt phosphorylation level, thereby promoting cell proliferation.
Discussion
Coding RNAs with regulatory functions are still not completely understood. Our laboratory has provided evidence that a hairpin within the YAP mRNA 3′UTR functions in regulation10. PTEN, which functions as a tumor suppressor, is commonly mutated in human cancers34. However, the underlying mechanism of action is not well documented. Therefore, we hypothesized that PTEN mRNA might have a regulatory function. In this study, we investigated the significance of the PTEN mRNA 3′UTR in liver cell regulation.
We identified a hairpin structure within the PTEN mRNA 3′UTR through bioinformatics analysis; this hairpin may act as a regulatory element in gene expression regulation. Our data showed that the PTEN-sh sequence is highly conserved in higher mammals. Interestingly, PTEN-sh increased the activities of AP-1 and NF-κB in 293T cells, thus suggesting that the hairpin functions in regulation. This finding is consistent with our previous findings that a hairpin within the YAP mRNA 3′UTR functions in regulation10. Next, we sought to identify the mechanism by which PTEN-sh is processed in the cells. It has been reported that the Drosha and DGCR8 complex is widely involved in RNA hairpin structure recognition and processing35. Therefore, we speculated that the hairpin within PTEN mRNA (PTEN-sh) might be cleaved from the PTEN mRNA 3′UTR by Drosha and DGCR8. Interestingly, we observed that PTEN mRNA levels were increased when Drosha or DGCR8 was knocked down in L-O2 cells. Moreover, using an RIP analysis, we validated that PTEN-sh interacts with Drosha or DGCR8 in L-O2 cells. These results suggest that Drosha and DGCR8 contribute to the PTEN-sh processing cells. Dicer recognizes stem-loop or double-stranded RNA and processes them into ∼22 nucleotide long fragments27; we hypothesized that the cleavage of PTEN-sh within PTEN mRNA might be processed by Dicer as well. As expected, we observed that Dicer was involved in the cleavage of the hairpin within the PTEN mRNA by using a luciferase reporter assay.
It has been reported that esiRNAs can be globally predicted by deep sequencing from transcriptome (also known as RNA-seq)36. RNA-seq is extensively improving the understanding of gene regulation and signaling networks37. In this study, through walking PCR analysis, we screened an esiRNA termed PTEN-sh-3p21, which is cleaved from a specific hairpin structure (PTEN-sh). There are three developed computational approaches: RNAhybrid, miRanda and TargetScan, that have been used to identify esiRNA target sites within the conserved regions of the 3′UTR of genes38. In addition, to assess the esiRNA targets at scale, StarScan, CleaveLand, SeqTar, sPARTA, PAREsnip, StarBase and sPARTA have been developed to predict esiRNA targets from degradome sequencing data39,40,41,42,43,44. We identified target genes of PTEN-sh-3p21, such as PPP2CA and PTEN, by using RNAhybrid. It has been reported that PPP2CA is part of the PI3K/PTEN/Akt pathway45. In this study, our findings suggested that PTEN-sh-3p21 cleaved from PTEN mRNA contributes to the precise regulation of PI3K/PTEN/Akt signaling. Functionally, the esiRNA, PTEN-sh-3p21, enhanced the proliferation of L-O2 cells and Chang liver cells in vitro. Thus, we conclude that an esiRNA from the hairpin within the PTEN mRNA 3′UTR post-transcriptionally modulates PPP2CA and PTEN in liver cells.
We propose a model in which a hairpin structure within the PTEN mRNA 3′UTR post-transcriptionally functions in liver cells (Figure 5E). A bioinformatics analysis showed that the secondary structure of PTEN mRNA 3′UTR has a hairpin structure. The element can be recognized by Drosha and DGCR8 and cleaved from the PTEN mRNA 3′UTR. Dicer contributes to the generation of esiRNA, such as PTEN-sh-3p21 targeting the 3′UTR of PPP2CA or PTEN. PTEN-sh-3p21 results in an increase of phosphorylated Akt, thereby promoting cell proliferation. Our findings provide new insights into the mechanism by which a hairpin structure within the PTEN mRNA 3′UTR functions post-transcriptionally in liver cells.
Author contribution
Xiao-dong ZHANG and Li-hong YE conceived the projects, designed the experiments and drafted the manuscript. Yu-en GAO designed the experiments, drafted the manuscript and performed the experiments. Yuan WANG, Fu-quan CHEN, Jin-yan FENG, Gang YANG, Guo-xing FENG, and Zhe YANG performed the experiments.
References
Battle A, Khan Z, Wang SH, Mitrano A, Ford MJ, Pritchard JK, et al. Genomic variation. Impact of regulatory variation from RNA to protein. Science 2015; 347: 664–7.
Pek JW, Okamura K . Regulatory RNAs discovered in unexpected places. Wiley Interdiscip Rev RNA 2015; 6: 671–86.
Busto GU, Guven-Ozkan T, Fulga TA, Van Vactor D, Davis RL . microRNAs that promote or inhibit memory formation in Drosophila melanogaster. Genetics 2015; 200: 569–80.
Stefani G, Slack FJ . Small non-coding RNAs in animal development. Nat Rev Mol Cell Biol 2008; 9: 219–30.
Cullen BR . Transcription and processing of human microRNA precursors. Mol Cell 2004; 16: 861–5.
Burger K, Gullerova M . Swiss army knives: non-canonical functions of nuclear Drosha and Dicer. Nat Rev Mol Cell Biol 2015; 16: 417–30.
Chu JYS, Sims-Lucas S, Bushnell DS, Bodnar AJ, Kreidberg JA, Ho J . Dicer function is required in the metanephric mesenchyme for early kidney development. Am J Physiol-Renal Physiol 2014; 306: 764–72.
Zhang B, Chen H, Zhang L, Dakhova O, Zhang Y, Lewis MT, et al. A dosage-dependent pleiotropic role of Dicer in prostate cancer growth and metastasis. Oncogene 2014; 33: 3099–108.
Jiang Z, Kong C, Zhang Z, Zhu Y, Zhang Y, Chen X . Reduction of protein kinase C alpha (PKC-alpha) promotes apoptosis via down-regulation of Dicer in bladder cancer. J Cell Mol Med 2015; 19: 1085–93.
Gao Y, Wang Y, Feng J, Feng G, Zheng M, Yang Z, et al. A hairpin within YAP mRNA 3′UTR functions in regulation at post-transcription level. Biochem Biophys Res Commun 2015; 459: 306–12.
Song MS, Salmena L, Pandolfi PP . The functions and regulation of the PTEN tumour suppressor. Nat Rev Mol Cell Biol 2012; 13: 283–96.
Tay Y, Kats L, Salmena L, Weiss D, Tan SM, Ala U, et al. Coding-independent regulation of the tumor suppressor PTEN by competing endogenous mRNAs. Cell 2011; 147: 344–57.
Poliseno L, Salmena L, Zhang JW, Carver B, Haveman WJ, Pandolfi PP . A coding-independent function of gene and pseudogene mRNAs regulates tumour biology. Nature 2010; 465: 1033–90.
Johnsson P, Ackley A, Vidarsdottir L, Lui WO, Corcoran M, Grander D, et al. A pseudogene long-noncoding-RNA network regulates PTEN transcription and translation in human cells. Nat Struct Mol Biol 2013; 20: 440–6.
Kong G, Zhang J, Zhang S, Shan C, Ye L, Zhang X . Upregulated microRNA-29a by hepatitis B virus X protein enhances hepatoma cell migration by targeting PTEN in cell culture model. PLoS One 2011; 6: e19518.
Shan C, Xu F, Zhang S, You J, You X, Qiu L, et al. Hepatitis B virus X protein promotes liver cell proliferation via a positive cascade loop involving arachidonic acid metabolism and p-ERK1/2. Cell Res 2010; 20: 563–75.
Keene JD, Komisarow JM, Friedersdorf MB . RIP-Chip: the isolation and identification of mRNAs, microRNAs and protein components of ribonucleoprotein complexes from cell extracts. Nat Protocols 2006; 1: 302–7.
Kuehbacher A, Urbich C, Zeiher AM, Dimmeler S . Role of Dicer and Drosha for endothelial microRNA expression and angiogenesis. Circ Res 2007; 101: 59–68.
Han J, Lee Y, Yeom KH, Kim YK, Jin H, Kim VN . The Drosha-DGCR8 complex in primary microRNA processing. Genes Dev 2004; 18: 3016–27.
Matzura O, Wennborg A . RNAdraw: an integrated program for RNA secondary structure calculation and analysis under 32-bit Microsoft Windows. Comput Appl Biosci 1996; 12: 247–9.
Bellaousov S, Reuter JS, Seetin MG, Mathews DH . RNA structure: Web servers for RNA secondary structure prediction and analysis. Nucleic Acids Res 2013; 41: 471–4.
You X, Liu F, Zhang T, Li Y, Ye L, Zhang X . Hepatitis B virus X protein upregulates oncogene Rab18 to result in the dysregulation of lipogenesis and proliferation of hepatoma cells. Carcinogenesis 2013; 34: 1644–52.
Zhao Y, Wang W, Wang Q, Zhang X, Ye L . Lipid metabolism enzyme 5-LOX and its metabolite LTB4 are capable of activating transcription factor NF-kappaB in hepatoma cells. Biochem Biophys Res Commun 2012; 418: 647–51.
Zhang S, Shan C, Kong G, Du Y, Ye L, Zhang X . MicroRNA-520e suppresses growth of hepatoma cells by targeting the NF-kappaB-inducing kinase (NIK). Oncogene 2012; 31: 3607–20.
Havens MA, Reich AA, Hastings ML . Drosha promotes splicing of a pre-microRNA-like alternative exon. PloS Genet 2014; 10: e1004312.
Quick-Cleveland J, Jacob JP, Weitz SH, Shoffner G, Senturia R, Guo F . The DGCR8 RNA-binding heme domain recognizes primary microRNAs by clamping the hairpin. Cell Rep 2014; 7: 1994–2005.
Bartel DP . MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004; 116: 281–97.
Macrae IJ, Zhou K, Li F, Repic A, Brooks AN, Cande WZ, et al. Structural basis for double-stranded RNA processing by Dicer. Science 2006; 311: 195–8.
Park JE, Heo I, Tian Y, Simanshu DK, Chang H, Jee D, et al. Dicer recognizes the 5′ end of RNA for efficient and accurate processing. Nature 2011; 475: 201–5.
Poliseno L, Haimovic A, Christos PJ, Vega Y Saenz de Miera EC, Shapiro R, et al. Deletion of PTENP1 pseudogene in human melanoma. J Invest Dermatol 2011; 131: 2497–500.
Bartel DP . MicroRNAs: target recognition and regulatory functions. Cell 2009; 136: 215–33.
Kruger J, Rehmsmeier M . RNAhybrid: microRNA target prediction easy, fast and flexible. Nucleic Acids Res 2006; 34: 451–4.
Hales EC, Orr SM, Larson Gedman A, Taub JW, Matherly LH . Notch1 receptor regulates AKT protein activation loop (Thr308) dephosphorylation through modulation of the PP2A phosphatase in phosphatase and tensin homolog (PTEN)-null T-cell acute lymphoblastic leukemia cells. J Biol Chem 2013; 288: 22836–48.
Schwartz S, Carver BS, Wongvipat J, Rodrik-Outmezguine V, Stanchina ED, Trigwell C, et al. The antitumor effects of PI3K beta inhibitors in PTEN negative prostate cancer are enhanced by inhibition of reactivated PI3K alpha signaling. Cancer Res 2014; 74: 4774.
Han J, Lee Y, Yeom KH, Nam JW, Heo I, Rhee JK, et al. Molecular basis for the recognition of primary microRNAs by the Drosha-DGCR8 complex. Cell 2006; 125: 887–901.
Liao JY, Guo YH, Zheng LL, Li Y, Xu WL, Zhang YC, et al. Both endo-siRNAs and tRNA-derived small RNAs are involved in the differentiation of primitive eukaryote Giardia lamblia. Proc Natl Acad Sci U S A 2014; 111: 14159–64.
Spies D, Ciaudo C . Dynamics in transcriptomics: advancements in RNA-seq time course and downstream analysis. Comput Struct Biotechnol J 2015; 13: 469–77.
Chan WL, You CY, Yang WK, Hung SY, Chang YS, Chiu CC, et al. Transcribed pseudogene psiPPM1K generates endogenous siRNA to suppress oncogenic cell growth in hepatocellular carcinoma. Nucleic Acids Res 2013; 41: 3734–47.
Liu S, Li JH, Wu J, Zhou KR, Zhou H, Yang JH, et al. StarScan: a web server for scanning small RNA targets from degradome sequencing data. Nucleic Acids Res 2015; 43: 480–6.
Yang JH, Li JH, Shao P, Zhou H, Chen YQ, Qu LH . starBase: a database for exploring microRNA-mRNA interaction maps from Argonaute CLIP-Seq and Degradome-Seq data. Nucleic Acids Res 2011; 39: 202–9.
Addo-Quaye C, Miller W, Axtell MJ . CleaveLand: a pipeline for using degradome data to find cleaved small RNA targets. Bioinformatics 2009; 25: 130–1.
Zheng Y, Li YF, Sunkar R, Zhang W . SeqTar: an effective method for identifying microRNA guided cleavage sites from degradome of polyadenylated transcripts in plants. Nucleic Acids Res 2012; 40: e28.
Kakrana A, Hammond R, Patel P, Nakano M, Meyers BC . sPARTA: a parallelized pipeline for integrated analysis of plant miRNA and cleaved mRNA data sets, including new miRNA target-identification software. Nucleic Acids Res 2014; 42: e139.
Folkes L, Moxon S, Woolfenden HC, Stocks MB, Szittya G, Dalmay T, et al. PAREsnip: a tool for rapid genome-wide discovery of small RNA/target interactions evidenced through degradome sequencing. Nucleic Acids Res 2012; 40: e103.
Bhardwaj A, Singh S, Srivastava SK, Arora S, Hyde SJ, Andrews J, et al. Restoration of PPP2CA expression reverses epithelial-to-mesenchymal transition and suppresses prostate tumour growth and metastasis in an orthotopic mouse model. Br J Cancer 2014; 110: 2000–10.
Acknowledgements
This work was supported by grants from the National Basic Research Program of China (973 Program, No 2015CB553703, No 2015CB553905), the National Natural Science Foundation of China (No 81272218, No 31470756, No 81372186) and the Project of Prevention and Treatment of Key Infectious Diseases (No 2014ZX0002002-005).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Supplementary information is available at the Acta Pharmacologica Sinica's website.
Supplementary information
Supplementary Information Figure S1
The effect of Dicer on miR-19a/b and miR-26a/b is measured in 293T cells. (JPG 431 kb)
Supplementary Information Figure S2
The location of PTEN-sh-3p21 within PTEN mRNA 3′UTR was displayed. (JPG 115 kb)
Supplementary Information Table S1
List of primers used in this paper. (PDF 188 kb)
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Gao, Ye., Wang, Y., Chen, Fq. et al. Post-transcriptional modulation of protein phosphatase PPP2CA and tumor suppressor PTEN by endogenous siRNA cleaved from hairpin within PTEN mRNA 3′UTR in human liver cells. Acta Pharmacol Sin 37, 898–907 (2016). https://doi.org/10.1038/aps.2016.18
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/aps.2016.18
Keywords
This article is cited by
-
The reasonable development technology policy in sweet spot of tight gas reservoir
Journal of Petroleum Exploration and Production Technology (2018)
-
The Fragment HMGA2-sh-3p20 from HMGA2 mRNA 3′UTR Promotes the Growth of Hepatoma Cells by Upregulating HMGA2
Scientific Reports (2017)
-
miR-511 promotes the proliferation of human hepatoma cells by targeting the 3′UTR of B cell translocation gene 1 (BTG1) mRNA
Acta Pharmacologica Sinica (2017)