Abstract
Aim:
To evaluate the effects of UDP-glucuronosyltransferases (UGTs) polymorphisms on the pharmacokinetics of the immunosuppressant mycophenolate mofetil (MMF) in Chinese renal transplant recipients.
Methods:
A total of 127 renal transplant patients receiving MMF were genotyped for polymorphisms in UGT1A9 −1818T>C, I399C>T, −118T9/10, −440C>T, −331T>C, UGT2B7 IVS1+985A>G, 211G>T, −900A>G, UGT1A8 518C>G and UGT1A7 622T>C. The plasma concentrations of the MMF active moiety mycophenolic acid (MPA) and main metabolite 7-O-MPA-glucuronide (MPAG) were analyzed using HPLC. Univariate and multivariate analyses were used to assess the effects of UGT-related gene polymorphisms on MPA pharmacokinetics.
Results:
The dose-adjusted MPA AUC0–12 h of the patients with the UGT2B7 IVS1+985AG genotype was 48% higher than that of the patients with the IVS1+985AA genotype, which could explain 11.2% of the inter-individual variation in MPA pharmacokinetics. The dose-adjusted MPAG AUC0–12 h of the patients with the UGT1A7 622CC and UGT1A9 −440CT/−331TC genotypes, respectively, was significantly higher than that of the patients with 622T homozygotes and −440C/−331T homozygotes. Furthermore, the genotypes UGT1A9 −1818T>C and UGT1A8 518C>G were associated with a low dose-adjusted MPAG AUC0–12 h.
Conclusion:
The UGT2B7 11+985A>G genotype is associated with the pharmacokinetics of MPA in Chinese renal transplant patients, which demonstrates the usefulness of this SNP for individualizing MMF dosing.
Similar content being viewed by others
Introduction
Mycophenolic acid (MPA) is the active moiety of mycophenolate mofetil (MMF), which is a potent immunosuppressive agent that is widely used in renal transplant patients1. MPA with a recommended AUC0–12 h of 30–60 mg·h·L−1 has been shown to be effective in preventing allograft rejection after solid organ transplantation1. The pharmacokinetics of MPA vary by approximately 7–10-fold among individuals, resulting in different states of immunosuppression2. Insufficient immunosuppression might result in the rejection of transplanted organs. Excessive immunosuppression might lead to side effects, such as diarrhea, anemia, leukopenia and infection3. It is important to understand the factors that cause this large pharmacokinetic variation to achieve the necessary drug concentration and avoid potential drug toxicity.
After administration, MMF is quickly hydrolyzed into MPA by esterase and then converted by UDP-glucuronosyltransferases (UGTs), predominantly in the liver, into phenolic glucuronide metabolite (7-O-MPA-glucuronide, MPAG, more than 90%) and acyl-MPAG (AcMPAG, less than 10%), which are excreted by the kidneys1,2. MPAG is characterized by enterohepatic circulation, which might affect MPA exposure, and MPA and MPAG exposure are important indicators of UGT activity in vivo. In addition, MPAG is reported to be related to the occurrence of diarrhea4. UGT isoforms that are involved in the metabolism of MPA include UGT1A9, 2B7, 1A8, and 1A7, among which UGT1A9 is the principal enzyme responsible for MPAG and 2B7 for AcMPAG5,6.
Many UGT-related polymorphisms have been reported to affect UGT activity and thus affect drug metabolism7,8. UGT1A9 −440C>T, −331T>C, I399C>T, and −118T10 were shown to be associated with higher activity in vitro9,10,11,12. UGT1A9 −1818T>C was reported to be related to slightly elevated UGT1A9 activity, without a significant difference13. UGT1A8*2 (518C>G) led to the decreased activity of UGT1A814,15,16. UGT2B7*2 (802C>T) was reported to have a limited effect on UGT2B7 activity. However, UGT2B7 −900A>G, which was in complete linkage disequilibrium (LD) with UGT2B7*2, has been shown to result in lower activity of the enzyme15. UGT2B7*3 (211G>T) was associated with higher enzyme activity10. The UGT1A7 622T>C genotype was reported to abolish the activity of UGT1A7 in vitro17.
All of these single nucleotide polymorphism (SNPs) have been investigated in MPA pharmacokinetics in vivo, with conflicting results that might be due to the limitations of the sample size, the differences in the ethnicities and co-administered drugs, or the confounding effects of other genetic polymorphisms. Unlike other rare UGT1A9 SNPs, the UGT1A9 −1818T>C genotype is a common variation found in the Chinese population, with a mutant frequency of 52.4% and a distinct ethnic difference13. It is necessary to investigate the effect of the UGT1A9 −1818T>C genotype on MPA pharmacokinetics in Chinese patients. The influence of the UGT1A7 622T>C genotype on MPA metabolism in the Chinese population has not been studied. Innocenti et al recently discovered a novel SNP in the intron region, UGT2B7 IVS1+985G>A, which could affect UGT2B7 activity. No in vivo study has examined the influence of the IVS1+985G>A polymorphism on drug metabolism. We hypothesized that the UGT2B7 IVS1+985G>A polymorphism might affect the metabolism of MPA.
To evaluate the genetic factors influencing MPA pharmacokinetics, we systematically investigated the relationship between UGT-related polymorphisms and MPA pharmacokinetics in Chinese renal transplant patients, including well-understood SNPs that have been poorly examined in the Chinese population as well as SNPs that have not been studied in any population.
Materials and methods
Ethics statement
The study was performed according to the Declaration of Helsinki and with ethical approval from the Ethics Committee of the First Affiliated Hospital of Sun Yat-sen University (No [2008]23). Written informed consent was obtained from all of the subjects.
Patients and therapy
A total of 127 renal transplant recipients (82 males and 45 females) from the Kidney Transplant Department, Transplant Center, the First Affiliated Hospital, Sun Yat-sen University were enrolled in this study. The average age of the patients was 37.05±10.6 years, whereas the average weight was 58.53±11.0 kg.
All of the patients received a triple immunosuppressive regimen consisting of MMF(Cellcept®, Roche, Basel, Switzerland), tacrolimus (Prograf®, Fujisawa Ireland Ltd, Killorglin, Ireland) and steroids (prednisone, Guangdong Huanan Pharmacy, Ltd, Dongguan, China). Except for tacrolimus and prednisone, any medication that might affect the MPA concentration used during treatment, such as proton pump inhibitors, were prohibited. Patients with impaired hepatic function, combined organ transplantation or a smoking habit were excluded. All of the patients received maintenance therapy with a fixed dose of MMF for at least 2 weeks before blood sample collection. There should be no adverse reaction at least 1 week before or after blood sample collection. The MMF dose regimens ranged from 0.5 g q 12 h to 1 g q 12 h according to the clinical symptoms and side effects.
Collection of the blood samples
Venous blood samples were collected at 0.5, 1.5, 4, and 9 h after the morning dose of MMF. We used a validated abbreviated sampling strategy to estimate the AUC0–12 h of MPA and MPAG instead of a full time-concentration profile18,19. Briefly, blood samples before and 0.5, 1.0, 1.5, 2.0, 3.0, 4.0, 6.0, 9.0, and 12.0 h after the MMF morning dose were obtained from 42 adult renal transplant patients who used a fixed MMF dose combined with tacrolimus for at least 2 weeks. The best-fit description of the MPA AUC0–12 h was decided by multiple linear regression analysis with r2=92.8%, model P<1.0×10−6 and D–W=1.917. The MPAG AUC0–12 h was determined by the multiple linear regression analysis with r2=95.6%, model P<1.0×10−8 and D–W=2.24520, all of which were within the acceptable range. We calculated the dose-adjusted AUC0–12 h of both MPA and of MPAG.
MPA and MPAG concentration detection
The plasma MPA and MPAG concentrations were simultaneously analyzed using a validated high performance liquid chromatography (HPLC) method21,22,23,24. Briefly, the plasma was extracted from whole blood after centrifugation at 4000 rpm/10 min. The plasma was stored at −80 °C until detection. A mixture of 5% ZnSO4 and methanol (30:70, v/v) was used as the protein precipitator, and naproxen was used as the internal standard. A mixture of 100 μL of plasma, 10 μL of naproxen and 100 μL of the precipitator was fully blended by a 1-min vortex before centrifugation at 14 000 rpm for 10 min to eliminate the denatured protein. The supernatant was separated on a solid reversed-column (4.6 mm×250 mm, 5 μm particle size; Xtimate™ C18, Welch Materials, Inc, Shanghai, China) using a Waters 1525-717-2487 HPLC system with ultraviolet detection at 215 nm. The mobile phase consisted of 0.1% trifluoroacetic acid and methanol (35:65, v/v), and the flow rate was 0.8 mL/min. All of the material used was of HPLC grade.
DNA extraction and genotyping
Total genomic DNA extraction was conducted following a previously described method25. The UGT1A8*2 polymorphism was detected using previously reported methods of polymerase chain reaction restriction-fragment length polymorphism (PCR-RFLP)16. The polymorphisms of UGT1A9 −118dT9/10 and I399C>T were genotyped by PCR-sequencing10. The other polymorphisms were genotyped by a MassArray® SNP genotyping system (Sequenom, Inc, San Diego, CA, USA)26.
Statistical analysis
Nonparametric tests were used for the univariate analysis using SPSS (Statistical Package for the Social Sciences) statistical software (version 21.0). The Mann-Whitney U-test was used for the comparison of two groups, whereas the Kruskal-Wallis test was used for the comparison of more than two groups. The Hardy-Weinberg equilibrium was tested in the frequencies of all the genotypes using a chi-square test procedure. The LD between the SNPs was tested using SHEsis27. A multivariate analysis by stepwise linear regression was applied to examine the association between MPA pharmacokinetics and various genotypes. The pharmacokinetic parameters were logarithmically transformed prior to the multivariate analysis. Only the variables with a P value of less than 0.1 in the univariate analysis were included in the multivariate analysis. The differences were considered to be statistically significant with P<0.05.
Results
Genotype frequencies
All of the 127 patients were genotyped for polymorphisms in UGT1A9 −1818T>C, I399C>T, −118T9/10, −440C>T, −331T>C, UGT2B7*3, IVS1+985A>G, −900A>G, UGT1A8*2 and UGT1A7 622T>C. All of the allele and genotype frequencies complied with the Hardy-Weinberg equilibrium and previous reports in Chinese Han populations (Table 1). A complete LD was observed between UGT1A9 −440C>T and 331T>C (D'=1). No significant link was found between the other individual combinations of SNPs.
The influence of the UGT2B7 polymorphisms on MPA pharmacokinetics
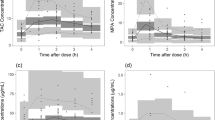
The MPA dose-adjusted AUC0–12 h was significantly higher in the recipients with the UGT2B7 IVS1+985AG genotype than that in the UGT2B7 IVS1+985AA carriers (47.93±28.92 vs32.42±24.35 mg·h·L−1·g−1, P=0.002) (Figure 1). We found that other UGT2B7-related polymorphisms did not affect the dose-adjusted MPA AUC0–12 h (data not shown).
Correlation of the UGT2B7 IVS1+985A>G genotype with the dose-adjusted AUC0–12 h of MPA in Chinese renal transplant patients.
The influence of the UGT1A7, UGT1A8, and UGT1A9 polymorphisms on MPA/MPAG pharmacokinetics
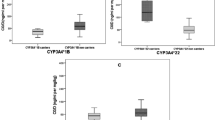
The dose-adjusted AUC0–12 h of MPAG was significantly greater in the patients with the UGT1A9 −1818CT genotype than that in the carriers of the UGT1A9 −1818CC genotype (671.39±338.62 vs 548.98±330.11 mg·h·L−1·g−1, P=0.002) (Figure 2A), and the dose-adjusted MPA AUC0–12 h was greater in the patients with the UGT1A9 −1818CT genotype than that in the carriers of the UGT1A9 −1818CC genotype, without a statistically significant difference (47.14±26.12 vs 38.18±18.14 mg·h·L−1·g−1, P=0.061). Although UGT1A9 −440C>T/−331T>C mutant carriers were rare in this study group (n=5), our data showed that these two variant alleles were related to an increase in the dose-adjusted AUC0–12 h of MPAG in comparison to that of the wild-type carriers (870.4±431.3 vs 622.1±363.5 mg·h·L−1·g−1, P=0.028) (Figure 2B). Compared to the carriers of UGT1A8*1/*1, the UGT1A8*1/*2 and *2/*2 subjects presented a 22% lower dose-adjusted AUC0–12 h of MPAG (597.6±337.6 vs 726.5±349.7 mg·h·L−1·g−1, P=0.004) (Figure 2C). The dose-adjusted AUC0–12 h of MPAG in the individuals with the UGT1A7 622CC genotype was significantly higher than that in the patients with the 622TT genotype (826.60±118.60 vs 539.20±156.40 mg·h·L−1·g−1, P=0.012) (Figure 2D). No association was observed between the polymorphisms of UGT1A9 I399C>T and −118T9/10 and MPA pharmacokinetics (data not shown).
Correlation of the UGT1A9 −1818T>C genotype (A), the UGT1A9 −440C>T/−331T>C genotype (B), the UGT1A8*2 genotype (C) and the UGT1A7 622C>T genotype (D) with the dose-adjusted AUC0–12 h of MPAG in Chinese renal transplant patients.
Multivariate analysis of MPA pharmacokinetics
The results of the multivariate analysis are listed in Table 2. Two polymorphisms (UGT2B7 IVS1+985A>G and UGT1A9 −1818T>C) as well as the age, weight, albumin level and gender were included in building the model. In the results, the UGT2B7 IVS1+985A>G and UGT1A9 −1818T>C polymorphisms were included in the final model (model P=0.001). The UGT2B7 IVS1+985AG genotype was independently predictive of a higher dose-adjusted MPA AUC0–12 h, which explained 11.2% of the inter-individual variations, whereas the UGT1A9 −1818T>C polymorphism was independently predictive of a lower dose-adjusted MPA AUC0–12 h. In addition to the genetic factors, age and gender were included in the final model (P=0.004 and P=0.019, respectively), the total of which led to a 23.1% explanation of the variation of MPA pharmacokinetics.
Discussion
The large inter-subject variations in MPA pharmacokinetics are characterized by large differences in the MPA/MPAG plasma concentrations. In this work, we found that the UGT2B7 IVS1+985A>G, UGT1A9 −440C>T/−331T>C, −1818C>T, UGT1A8*2 and UGT1A7 −622T>C polymorphisms were associated with MPA pharmacokinetics in Chinese renal transplant patients.
In our research, the IVS1+985AG genotype was found to be associated with an elevated dose-adjusted MPA AUC0–12 h (P=0.002), which could explain 11.2% of the inter-individual variations in MPA pharmacokinetics (P<0.001). Innocenti et al discovered a haplotype consisting of the UGT2B7 IVS1+985A>G polymorphism and two other SNPs, which were in high LD with each other, to be associated with an increasing enzyme activity by regulating mRNA expression of UGT2B7 in vitro28. However, no in vivo data focusing on the effect of the IVS1+985A>G polymorphism on any drug metabolism has been reported. Despite a tendency for the up-regulation of the UGT2B7 activity by the IVS1+985A>G polymorphism in vitro, we found a negative effect of this polymorphism on transforming the MPA. This discordancy between in vivo and in vitro studies might have many causes. First, alternative splicing mechanisms are very complex, and the in vitro data could not exactly predict the actual situation in vivo. The UGT2B7 IVS1+985A>G polymorphism was reported to favor the formation of splicing variant 1, which is expressed most frequently in the liver and intestinal tract29,30. Additionally, other splicing variants naturally occur in other tissues, including the kidneys, in which the effects of the IVS1+985A>G polymorphism on these splicing variants were unknown. Moreover, it was reported that different splicing variants could interact with each other by competitive binding, which further complicates the process. It is possible that this polymorphism not only influences splicing variant 1 formation but also affects the formation of other splicing variants, which in turn could affect the formation of splicing variant 1. Second, the effect of the Y483D polymorphism on UGT1A9 activity was substrate dependent31,32. The IVS1+985A>G mutation might affect the substrate specificity of UGT2B7 in the similar way, which might lead to unexpected responses in MPA pharmacokinetics. Third, all of the patients in this study were co-administered with tacrolimus, which might inhibit UGT enzyme activity, counteracting the influence of this polymorphism on MPA pharmacokinetics. However, the clear mechanisms by which this SNP leads to an increased exposure to MPA remain to be explained.
We found that individuals with the UGT1A9 −440CT/−331TC genotypes had higher dose-adjusted AUC0–12 h of MPAG than those of the individuals with the wild type (P=0.028), whereas the UGT1A9 −1818CC genotype was associated with a lower dose-adjusted AUC0–12 h of MPAG (P=0.002). The most well-studied gene involved in the metabolism of MPA is UGT1A9, which accounts for more than 50% of MPAG production. The UGT1A9 −440C>T/−331T>C polymorphism is associated with enhanced UGT activity in human liver microsomes6,33, a finding that was consistent with our result. Very few in vivo studies demonstrated the effect of the UGT1A9 −1818T>C polymorphism on MPA pharmacokinetics. An in vitro study showed that the −1818C allele might lead to slightly evaluated enzyme activity; however, the result was inconclusive and without significant differences. We showed, for the first time, that the UGT1A9 −1818T>C polymorphism was associated with a lower dose-adjusted AUC0–12 h of MPAG in the Chinese population.
In addition, the UGT1A9, UGT1A8, and UGT1A7 polymorphisms play important roles in regulating the metabolism of MPA. We found a positive relationship between the dose-adjusted AUC0–12 h of MPAG and the UGT1A7 622C allele in Chinese renal transplant patients (P=0.030), and UGT1A8*2 was found to be associated with a lower dose-adjusted AUC0–12 h of MPAG (P=0.004). The influence of the UGT1A7 −622T>C polymorphism on MPA metabolism in vivo had only been investigated in populations other than Chinese populations, and conflicting results have been observed. Patients with the UGT1A7 622TC genotype demonstrated increased oral clearance and a decreased Cmax of MPA8, which resulted in an accumulation of MPAG. These findings were in accordance with our results. Compared to the wild type, the UGT1A8*2 polymorphism was found to have a limited effect on MPAG production in vitro5,6. The MPA dose-adjusted trough concentrations were 60% higher in the *1/*2 and *2/*2 carriers than those in the patients with the wild type in vivo34, from which we could assume an inhibitive effect of the UGT1A8*2 polymorphism on UGT1A8 activity. Correspondingly, our result that the UGT1A8*2 polymorphism had a negative effect on MPAG formation in vivo was consistent with this assumption.
No association was observed between the UGT1A9 I399C>T, −118T9/10, UGT2B7*3, −900A>G polymorphisms and MPA pharmacokinetics in our research. Conflicting results concerning these polymorphisms have been observed in other studies15,35,36, which might have resulted from the specific binding of different substrates to UGT1A9 or UGT2B7 affected by mutant alleles, different co-administered drugs and limitation of the sample size. Thus, a larger sample size is needed to evaluate the effect of these polymorphisms on MPA pharmacokinetics, whereas co-medication should be taken into consideration as well.
In this study, we systematically evaluated the potential influence of the UGTs-related polymorphisms on MPA pharmacokinetics in Chinese renal transplant patients. We found that the UGT2B7 IVS1+985A>G polymorphism was the most profound factor in the large variation in MPA pharmacokinetics. Patients who carry the UGT2B7 IVS1+985AG genotype might be at a greater risk of a higher dose-adjusted MPA AUC0–12 h, which would lead to infection and other side effects that might require reducing the MMF dosage. In addition, the UGT1A9 −440C>T/−331T>C, −1818C>T, and UGT1A7 −622T>C and UGT1A8*2 polymorphisms were related to the dose-adjusted AUC0–12 h of MPAG, which might influence MPA exposure and thus clinical outcomes. The UGTs-related polymorphisms might be useful for the individualization of MMF dosing.
Author contribution
Xiao-chun XIE, Jia-li LI, Min HUANG, and Chang-xi WANG designed the research. Xiao-chun XIE, Jun LI, Qian FU, Hong-yang WANG, and Ping-ping SUN performed the research. Xiao-chun XIE, Xue-ding WANG, Jing LIU, Jia-wen HUANG, and Hong-liang LI contributed new reagents and analytic tools. Xiao-chun XIE, Guo-ping ZHONG, and Chen ZHU analyzed the data. Xiao-chun XIE and Jia-li LI wrote the paper.
References
Staatz CE, Tett SE . Clinical pharmacokinetics and pharmacodynamics of mycophenolate in solid organ transplant recipients. Clin Pharmacokinet 2007; 46: 13–58.
Bullingham RE, Nicholls AJ, Kamm BR . Clinical pharmacokinetics of mycophenolate mofetil. Clin Pharmacokinet 1998; 34: 429–55.
Staatz CE, Tett SE . Pharmacology and toxicology of mycophenolate in organ transplant recipients: an update. Arch Toxicol 2014; 88: 1351–89.
Woillard JB, Rerolle JP, Picard N, Rousseau A, Drouet M, Munteanu E, et al. Risk of diarrhoea in a long-term cohort of renal transplant patients given mycophenolate mofetil: the significant role of the UGT1A8 2 variant allele. Br J Clin Pharmacol 2010; 69: 675–83.
Bernard O, Tojcic J, Journault K, Perusse L, Guillemette C . Influence of nonsynonymous polymorphisms of UGT1A8 and UGT2B7 metabolizing enzymes on the formation of phenolic and acyl glucuronides of mycophenolic acid. Drug Metab Dispos 2006; 34: 1539–45.
Bernard O, Guillemette C . The main role of UGT1A9 in the hepatic metabolism of mycophenolic acid and the effects of naturally occurring variants. Drug Metab Dispos 2004; 32: 775–8.
Mazidi T, Rouini MR, Ghahremani MH, Dashti-Khavidaki S, Lessan-Pezeshki M, Ahmadi FL, et al. Impact of UGT1A9 polymorphism on mycophenolic acid pharmacokinetic parameters in stable renal transplant patients. Iran J Pharm Res 2013; 12: 547–56.
Joy MS, Boyette T, Hu Y, Wang J, La M, Hogan SL, et al. Effects of uridine diphosphate glucuronosyltransferase 2B7 and 1A7 pharmacogenomics and patient clinical parameters on steady-state mycophenolic acid pharmacokinetics in glomerulonephritis. Eur J Clin Pharmacol 2010; 66: 1119–30.
Girard H, Villeneuve L, Court MH, Fortier LC, Caron P, Hao Q, et al. The novel UGT1A9 intronic I399 polymorphism appears as a predictor of 7-ethyl-10-hydroxycamptothecin glucuronidation levels in the liver. Drug Metab Dispos 2006; 34: 1220–8.
Guo D, Pang LF, Han Y, Yang H, Wang G, Tan ZR, et al. Polymorphisms of UGT1A9 and UGT2B7 influence the pharmacokinetics of mycophenolic acid after a single oral dose in healthy Chinese volunteers. Eur J Clin Pharmacol 2013; 69: 843–9.
Jiao Z, Ding JJ, Shen J, Liang HQ, Zhong LJ, Wang Y, et al. Population pharmacokinetic modelling for enterohepatic circulation of mycophenolic acid in healthy Chinese and the influence of polymorphisms in UGT1A9. Br J Clin Pharmacol 2008; 65: 893–907.
Yamanaka H, Nakajima M, Katoh M, Hara Y, Tachibana O, Yamashita J, et al. A novel polymorphism in the promoter region of human UGT1A9 gene (UGT1A9*22) and its effects on the transcriptional activity. Pharmacogenetics 2004; 14: 329–32.
Girard H, Court MH, Bernard O, Fortier LC, Villeneuve L, Hao Q, et al. Identification of common polymorphisms in the promoter of the UGT1A9 gene: evidence that UGT1A9 protein and activity levels are strongly genetically controlled in the liver. Pharmacogenetics 2004; 14: 501–15.
Geng F, Jiao Z, Dao YJ, Qiu XY, Ding JJ, Shi XJ, et al. The association of the UGT1A8, SLCO1B3 and ABCC2/ABCG2 genetic polymorphisms with the pharmacokinetics of mycophenolic acid and its phenolic glucuronide metabolite in Chinese individuals. Clin Chim Acta 2012; 413: 683–90.
Levesque E, Delage R, Benoit-Biancamano MO, Caron P, Bernard O, Couture F, et al. The impact of UGT1A8, UGT1A9, and UGT2B7 genetic polymorphisms on the pharmacokinetic profile of mycophenolic acid after a single oral dose in healthy volunteers. Clin Pharmacol Ther 2007; 81: 392–400.
Kagaya H, Inoue K, Miura M, Satoh S, Saito M, Tada H, et al. Influence of UGT1A8 and UGT2B7 genetic polymorphisms on mycophenolic acid pharmacokinetics in Japanese renal transplant recipients. Eur J Clin Pharmacol 2007; 63: 279–88.
Guillemette C, Ritter JK, Auyeung DJ, Kessler FK, Housman DE . Structural heterogeneity at the UDP-glucuronosyltransferase 1 locus: functional consequences of three novel missense mutations in the human UGT1A7 gene. Pharmacogenetics 2000; 10: 629–44.
Yamaguchi K, Fukuoka N, Kimura S, Watanabe M, Tani K, Tanaka H, et al. Limited sampling strategy for the estimation of mycophenolic acid area under the concentration-time curve treated in Japanese living-related renal transplant recipients with concomitant extended-release tacrolimus. Biol Pharm Bull 2013; 36: 1036–9.
Abd Rahman AN, Tett SE, Staatz CE . How accurate and precise are limited sampling strategies in estimating exposure to mycophenolic acid in people with autoimmune disease? Clin Pharmacokinet 2014; 53: 227–45.
Liu YF, Li J, Huang JW, Fu Q, Liu LS, Chen E, et al. Evaluation of mycophenolic acid exposure by limited sampling strategy in Chinese adult renal transplant recipients receiving mycophenolate mofeil and tacrolimus. Chin J Organ Transplant 2012; 33: 101–4.
Teshima D, Otsubo K, Kitagawa N, Yoshimura S, Itoh Y, Oishi R . High-performance liquid chromatographic method for mycophenolic acid and its glucuronide in serum and urine. J Clin Pharm Ther 2003; 28: 17–22.
Shipkova M, Niedmann PD, Armstrong VW, Schutz E, Wieland E, Shaw LM, et al. Simultaneous determination of mycophenolic acid and its glucuronide in human plasma using a simple high-performance liquid chromatography procedure. Clin Chem 1998; 44: 1481–8.
Wiwattanawongsa K, Heinzen EL, Kemp DC, Dupuis RE, Smith PC . Determination of mycophenolic acid and its phenol glucuronide metabolite in human plasma and urine by high-performance liquid chromatography. J Chromatogr B Biomed Sci Appl 2001; 763: 35–45.
Huang JW, Liu YF, Li JL, Wang L, Wang CX, Zhong GP, et al. Development and application of HPLC method in simultaneous determination of mycophenolic acid and its 7-O-glucuronide metabolite in human plasma. Chin J Prim Med Pharm 2011; 18: 1729–31.
Loparev VN, Cartas MA, Monken CE, Velpandi A, Srinivasan A . An efficient and simple method of DNA extraction from whole blood and cell lines to identify infectious agents. J Virol Methods 1991; 34: 105–12.
Basu P, Chandna P, Bamezai RN, Siddiqi M, Saranath D, Lear A, et al. MassARRAY spectrometry is more sensitive than PreTect HPV-Proofer and consensus PCR for type-specific detection of high-risk oncogenic human papillomavirus genotypes in cervical cancer. J Clin Microbiol 2011; 49: 3537–44.
Shi YY, He L . SHEsis, a powerful software platform for analyses of linkage disequilibrium, haplotype construction, and genetic association at polymorphism loci. Cell Res 2005; 15: 97–8.
Innocenti F, Liu W, Fackenthal D, Ramirez J, Chen P, Ye X, et al. Single nucleotide polymorphism discovery and functional assessment of variation in the UDP-glucuronosyltransferase 2B7 gene. Pharmacogenet Genomics 2008; 18: 683–97.
Guillemette C, Levesque E, Rouleau M . Pharmacogenomics of human uridine diphospho-glucuronosyltransferases and clinical implications. Clin Pharmacol Ther 2014; 96: 324–39.
Menard V, Levesque E, Chen S, Eap O, Joy MS, Ekstrom L, et al. Expression of UGT2B7 is driven by two mutually exclusive promoters and alternative splicing in human tissues: changes from prenatal life to adulthood and in kidney cancer. Pharmacogenet Genomics 2013; 23: 684–96.
Holthe M, Rakvag TN, Klepstad P, Idle JR, Kaasa S, Krokan HE, et al. Sequence variations in the UDP-glucuronosyltransferase 2B7 (UGT2B7) gene: identification of 10 novel single nucleotide polymorphisms (SNPs) and analysis of their relevance to morphine glucuronidation in cancer patients. Pharmacogenomics J 2003; 3: 17–26.
Takahashi H, Maruo Y, Mori A, Iwai M, Sato H, Takeuchi Y . Effect of D256N and Y483D on propofol glucuronidation by human uridine 5′-diphosphate glucuronosyltransferase (UGT1A9). Basic Clin Pharmacol Toxicol 2008; 103: 131–6.
Girard H, Court MH, Bernard O, Fortier LC, Villeneuve L, Hao Q, et al. Identification of common polymorphisms in the promoter of the UGT1A9 gene. Pharmacogenetics 2004; 14: 501–15.
Johnson LA, Oetting WS, Basu S, Prausa S, Matas A, Jacobson PA . Pharmacogenetic effect of the UGT polymorphisms on mycophenolate is modified by calcineurin inhibitors. Eur J Clin Pharmacol 2008; 64: 1047–56.
Zhang WX, Chen B, Jin Z, Yu Z, Wang X, Chen H, et al. Influence of uridine diphosphate (UDP)-glucuronosyltransferases and ABCC2 genetic polymorphisms on the pharmacokinetics of mycophenolic acid and its metabolites in Chinese renal transplant recipients. Xenobiotica 2008; 38: 1422–36.
Inoue K, Miura M, Satoh S, Kagaya H, Saito M, Habuchi T, et al. Influence of UGT1A7 and UGT1A9 intronic I399 genetic polymorphisms on mycophenolic acid pharmacokinetics in Japanese renal transplant recipients. Ther Drug Monit 2007; 29: 299–304.
Acknowledgements
We appreciate the financial support provided by the National Major Projects for Science and Technology Development from Science and Technology Ministry of China (Grant No 2012ZX09506001-004), the National Natural Science Foundations of China (No 81102515, 81320108027 and 81173131), the Key Laboratory Foundation of Guangdong Province (No 2011A060901014) and the Major Scientific and Technological Project of Guangdong Province (No 2011A080300001 and 2012A080202013).
Author information
Authors and Affiliations
Corresponding author
PowerPoint slides
Rights and permissions
About this article
Cite this article
Xie, Xc., Li, J., Wang, Hy. et al. Associations of UDP-glucuronosyltransferases polymorphisms with mycophenolate mofetil pharmacokinetics in Chinese renal transplant patients. Acta Pharmacol Sin 36, 644–650 (2015). https://doi.org/10.1038/aps.2015.7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/aps.2015.7
Keywords
This article is cited by
-
A Systematic Review of Multiple Linear Regression-Based Limited Sampling Strategies for Mycophenolic Acid Area Under the Concentration–Time Curve Estimation
European Journal of Drug Metabolism and Pharmacokinetics (2021)
-
Comparison of the Pharmacokinetics of RIPK1 Inhibitor GSK2982772 in Healthy Western and Japanese Subjects
European Journal of Drug Metabolism and Pharmacokinetics (2021)
-
The Impact of Genetic Polymorphisms on the Pharmacokinetics and Pharmacodynamics of Mycophenolic Acid: Systematic Review and Meta-analysis
Clinical Pharmacokinetics (2021)
-
Population pharmacokinetics and Bayesian estimation of mycophenolic acid concentrations in Chinese adult renal transplant recipients
Acta Pharmacologica Sinica (2017)
-
Inter-isoform Hetero-dimerization of Human UDP-Glucuronosyltransferases (UGTs) 1A1, 1A9 and 2B7 and Impacts on Glucuronidation Activity
Scientific Reports (2016)