Abstract
Aim:
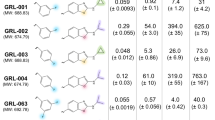
To characterize the functional and pharmacological features of a symmetrical 1-pyrrolidineacetamide, N,N′-(methylene-di-4,1-phenylene) bis-1-pyrrolidineacetamide, as a new anti-HIV compound which could competitively inhibit HIV-1 integrase (IN) binding to viral DNA.
Methods:
A surface plasma resonance (SPR)-based competitive assay was employed to determine the compound's inhibitory activity, and the 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide cell assay was used to qualify the antiviral activity. The potential binding sites were predicted by molecular modeling and determined by site-directed mutagenesis and a SPR binding assay.
Results:
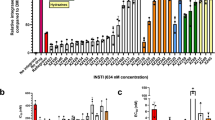
1-pyrrolidineacetamide, N,N′-(methylene-di-4,1-phenylene) bis- 1-pyrrolidineacetamide could competitively inhibit IN binding to viral DNA with a 50% inhibitory concentration (IC50) value of 7.29±0.68 μmol/L as investigated by SPR-based investigation. Another antiretroviral activity assay showed that this compound exhibited inhibition against HIV-1(IIIB) replication with a 50% effective concentration (EC50) value of 40.54 μmol/L in C8166 cells, and cytotoxicity with a cytotoxic concentration value of 173.84 μmol/L in mock-infected C8166 cells. Molecular docking predicted 3 potential residues as 1-pyrrolidineacetamide, N,N′-(methylene-di-4,1-phenylene)bis-1-pyrrolidineacetamide binding sites. The importance of 3 key amino acid residues (Lys 103, Lys 173, and Thr 174) involved in the binding was further identified by site-directed mutagenesis and a SPR binding assay.
Conclusion:
This present work identified a new anti-HIV compound through a new IN-binding site which is expected to supply new potential drug-binding site information for HIV-1 integrase inhibitor discovery and development.
Similar content being viewed by others
Article PDF
References
Asante-Appiah E, Skalka AM . HIV-1 integrase: structural organization, conformational changes, and catalysis. Adv Virus Res 1999; 52: 351–69.
Cara A, Guarnaccia F, Reitz MS, Gallo RC, Lori F . Self-limiting, cell type-dependent replication of an integrase-defective human immunodeficiency virus type 1 in human primary macrophages but not T lymphocytes. Virology 1995; 208: 242–8.
Pommier Y, Neamati N . Inhibitors of human immunodeficiency virus integrase. Adv Virus Res 1999; 52: 427–58.
Sherman PA, Fyfe JA . Human immunodeficiency virus integration protein expressed in Escherichia coli possesses selective DNA cleaving activity. Proc Natl Acad Sci USA 1990; 87: 5119–23.
LaFemina RL, Callahan PL, Cordingley MG . Substrate specificity of recombinant human immunodeficiency virus integrase protein. J Virol 1991; 65: 5624–30.
Bushman FD, Craigie R . Activities of human immunodeficiency virus (HIV) integration protein in vitro: specific cleavage and integration of HIV DNA. Proc Natl Acad Sci USA 1991; 88: 1339–43.
Pommier Y, Johnson AA, Marchand C . Integrase inhibitors to treat HIV/AIDS. Nat Rev Drug Discov 2005; 4: 236–48.
Chiu TK, Davies DR . Structure and function of HIV-1 integrase. Curr Top Med Chem 2004; 4: 965–77.
Cai M, Zheng R, Caffrey M, Craigie R, Clore GM, Gronenborn AM . Solution structure of the N-terminal zinc binding domain of HIV-1 integrase. Nat Struct Biol 1997; 4: 567–77.
Wang JY, Ling H, Yang W, Craigie R . Structure of a two-domain fragment of HIV-1 integrase: implications for domain organization in the intact protein. Embo J 2001; 20: 7333–43.
Dayam R, Deng J, Neamati N . HIV-1 integrase inhibitors: 2003–2004 update. Med Res Rev 2006; 26: 271–309.
Hazuda DJ, Felock P, Witmer M, Wolfe A, Stillmock K, Grobler JA, et al. Inhibitors of strand transfer that prevent integration and inhibit HIV-1 replication in cells. Science 2000; 287: 646–50.
Cotelle P . Patented HIV-1 integrase inhibitors (1998–2005). Recent Pat Anti-infect Drug Disc 2006; 1: 1–15.
Johnson AA, Marchand C, Pommier Y . HIV-1 integrase inhibitors: a decade of research and two drugs in clinical trial. Curr Top Med Chem 2004; 4: 1059–77.
Drake RR, Neamati N, Hong H, Pilon AA, Sunthankar P, Hume SD, et al. Identification of a nucleotide binding site in HIV-1 integrase. Proc Natl Acad Sci USA 1998; 95: 4170–5.
Shimura K, Kodama E, Sakagami Y, Matsuzaki Y, Watanabe W, Yamataka K, et al. Broad antiretroviral activity and resistance profile of the novel human immunodeficiency virus integrase inhibitor elvitegravir (JTK-303/GS-9137). J Virol 2008; 82: 764–74.
Savarino A . In-silico docking of HIV-1 integrase inhibitors reveals a novel drug type acting on an enzyme/DNA reaction intermediate. Retrovirology 2007; 4: 21.
Jenkins TM, Engelman A, Ghirlando R, Craigie R . A soluble active mutant of HIV-1 integrase: involvement of both the core and carboxyl-terminal domains in multimerization. J Biol Chem 1996; 271: 7712–8.
Du L, Shen L, Yu Z, Chen J, Guo Y, Tang Y, et al. Hyrtiosal, from the marine sponge Hyrtios erectus, inhibits HIV-1 integrase binding to viral DNA by a new inhibitor binding site. Chem Med Chem 2008; 3: 173–80.
Pauwels R, Balzarini J, Baba M, Snoeck R, Schols D . Rapid and automated tetrazolium-based colorimetric assay for the detection of anti-HIV compounds. J Virol Methods 1988; 20: 309–21.
Johnson VA, Byington RE . Quantitative assays for virus infectivity. New York: Srockton Press; 1990.
Wang Q, Wang YT, Pu SP, Zheng YT . Zinc coupling potentiates anti-HIV- 1 activity of baicalin. Biochem Biophys Res Commun 2004; 324: 605–10.
Gasteiger J, Marsili M . Iterative partial equalization of orbital electronegativity: a rapid access to atomic charges. Tetrahedron 1980; 36: 3219–28.
Clark M, Cramer R, van Opdebosch N . Validation of the general purpose Tripos 5.2 force field. J Comput Chem 1989; 10: 982–1012.
Sybyl (Computer program). Version 7.0. St Louis, MO: Tripos Associates Inc; 2004.
Berman HM, Bhat TN, Bourne PE, Feng Z, Gilliland G, Weissig H, et al. The Protein Data Bank and the challenge of structural genomics. Nat Struct Biol 2000; 7 Suppl: 957–9.
Goldgur Y, Craigie R, Cohen GH, Fujiwara T, Yoshinaga T, Fujishita T, et al. Structure of the HIV-1 integrase catalytic domain complexed with an inhibitor: a platform for antiviral drug design. Proc Natl Acad Sci USA 1999; 96: 13 040–3.
Weiner SJ, Kollman PA, Nguyen DT, Case DA . An all atom force field for simulations of proteins and nucleic acid. J Comput Chem 1986; 11: 431–9.
Weiner SJ, Kollman PA, Case DA, Singh C, Ghio G, Alagona S, et al. New force field for molecular mechanical simulation of nucleic acids and proteins. J Am Chem Soc 1984; 106: 765–84.
Rosin CD, Belew RK, Morris GM, Olson AJ, Goodsell DS . Computational coevolution of antiviral drug resistance. Artif Life 1998; 4: 41–59.
Verdonk ML, Cole JC, Hartshorn MJ, Murray CW, Taylor RD . Improved protein-ligand docking using GOLD. Proteins 2003; 52: 609–23.
Jones G, Willett P, Glen RC, Leach AR, Taylor R . Development and validation of a genetic algorithm for flexible docking. J Mol Biol 1997; 267: 727–48.
Shkriabai N, Patil SS, Hess S, Budihas SR, Craigie R, Burke TR Jr, et al. Identification of an inhibitor-binding site to HIV-1 integrase with affinity acetylation and mass spectrometry. Proc Natl Acad Sci USA 2004; 101: 6894–9.
Marchand C, Zhang X, Pais GC, Cowansage K, Neamati N, Burke TR Jr, et al. Structural determinants for HIV-1 integrase inhibition by beta-diketo acids. J Biol Chem 2002; 277: 12 596–603.
Merck . Monogram and Merck to collaborate on phase III trials of Merck's investigational HIV-1 integrase inhibitor. [Cited 07 Sep 2007] Available from URL: http://www.hivandhepatitis.com/recent/2007/090707_a.html.
Li HY, Zawahir Z, Song LD, Long YQ, Neamati N . Sequence-based design and discovery of peptide inhibitors of HIV-1 integrase: insight into the binding mode of the enzyme. J Med Chem 2006; 49: 4477–86.
Author information
Authors and Affiliations
Corresponding authors
Additional information
This work was financially supported by the National Natural Science Foundation of China (No 30525024, 20472095, and 20572023), Shanghai Pujiang Program (No 05PJ14034), Shanghai Key Basic Research Project (No 06JC14080 and 05JC14092), the State Key Program of Basic Research of China (No 2004CB58905 and 2006AA09Z447), and a grant from the Chinese Academy of Sciences (No KSCX2-YW-R-18).
Rights and permissions
About this article
Cite this article
Du, L., Zhao, Yx., Yang, Lm. et al. Symmetrical 1-pyrrolidineacetamide showing anti-HIV activity through a new binding site on HIV-1 integrase. Acta Pharmacol Sin 29, 1261–1267 (2008). https://doi.org/10.1111/j.1745-7254.2008.00863.x
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1111/j.1745-7254.2008.00863.x