Abstract
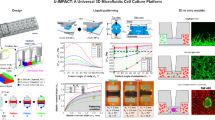
Both two-dimensional (2D) and three-dimensional (3D) biomaterial-based culture platforms that are capable of mimicking the in vivo microenvironment to recapitulate the physiological conditions are vital tools in a wide range of cellular and clinical research. Here we report tissue cultures in a microfluidic chip that allows deterministic patterning of cells in 2D/3D. The chip contains a cell-supporting membrane bioengineered to attain either 2D or 3D cell patterns by selectable deposition of extracellular matrix molecules. Results show a cell-trapping rate as high as 97% in our microchip. Tuning of the surface enables not only highly controlled geometry of the monolayer (2D) cell mass but also 3D culture of uniformly sized multicellular spheroids. The 3D spheroid culture of human epithelial ovarian cancer cells in the microfluidic chip resulted in acquisition of mesenchymal traits—increased expressions of N-cadherin, vimentin and fibronectin—and lowered expression of epithelial marker (CD326/epithelial cell adhesion molecule) compared with that in traditional 2D cultures, which is indicative of epithelial–mesenchymal transition in the spheres. In conclusion, these results offer new opportunities to achieve active control of 2D cellular patterns and 3D multicellular spheroids on demand, and may be amenable toward the study of the metastatic processes by in vitro modeling.
Similar content being viewed by others
Introduction
Over the last decade, microengineering has emerged as invaluable surrogate tools to probe a wide range of biomedical issues.1, 2, 3, 4, 5, 6 Microsystems open doors to new high-throughput and high-content approaches, and provide systematic insights into cellular functions influenced by microenvironmental stimuli, which include the interactions between neighboring cells, extracellular matrices, chemical factors and physical forces.7, 8, 9 To create and recapitulate the cellular in vivo conditions, biomaterial-based surfaces capable of generating precise patterning of cells for two-dimensional (2D) or three-dimensional (3D) cell culture, and for controlling microenvironmental stimuli at the single-cell level (approximately 10 μm) in microsystems are crucial. These surfaces have served as versatile tools to explore fundamental cell biology, tissue engineering and drug development, and thereby became key components in a wide range of applications.10
Various techniques for patterning cells in 2D or 3D tissue culture have demonstrated the ability to direct cells onto selected areas of a surface, including microcontact printing8 or soft lithography,1 microfluidic- or microstructure-based methods,11, 12 active control of cell attachment or suspension by physical forces13, 14, 15 and layer-by-layer assembly.16, 17 However, challenges still remain. For example, some techniques typically require a substantial amount of cells to seed onto open surfaces and wash away numerous unattached cells after the cell-patterning step has been executed.1, 8, 14, 18 These result in cell waste and restrict potential applications, for example, rare cells19 and patient samples.10 In addition, complexity arises from the use of external systems, such as syringe pumps or electric power, which is less attractive than more simplistic approaches.4, 14 Other challenges include the use of nanoparticles and magnetic fields to suspend cells, which may complicate the patterning of cells in enclosed microfluidic channels.4, 15
Here we present a multifunctional microchip that can produce deterministic cell trapping not only for 2D cellular patterning but also for 3D spheroid culture on demand. This chip contains two-layer microchannels sandwiching a bioengineered membrane that supports the patterned cells. The membrane was first microfabricated with through-holes and then tuned by precise control of extracellular matrix (ECM; collagen type I) deposition. By selectively tuning the heights of the solution level using manual pipetting, subsequent cell patterning in 2D was fast and deterministic, and the culture of 3D multicellular spheroids was robust. Taken together, the proposed simplistic microchip has the advantages of fast patterning of cells, viable culture of cellular microenvironment either in 2D or 3D, and pipette-friendly usage, which may be further integrated into biological assays.
Materials and methods
Operational principle of the cell trapping
Figure 1a presents the overall configuration of the microchip, including top and bottom channels that sandwich the perforated membrane. Cells are transported hydrodynamically in the top channel of the two-channel structure via a pressure gradient along the top channel. A suitable pressure difference across the membrane (via the through-holes) is achieved by fine-tuning the pressure gradient across the membrane, which results in suction of suspended cells toward the top surface of the membrane. As the cells approach the through-holes, this pressure difference immobilizes the cells onto the membrane to achieve the desirable cell patterning (see Supplementary Movie S1). Precise control of the pressure gradients both along the top channel and across the membrane is critical to successful and efficient cell patterning.
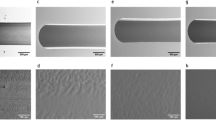
Design of the microfluidic chip. (a) Illustration of the overall configuration with top and bottom channels sandwiching the cell-supporting membrane with fabricated through-holes. (b) Photography of the fabricated chip. Four wells were used as reservoirs to control the flow rates by selectively tuning the solution levels, in which h indicates the solution height in the well. Blue and yellow colors represent the top and bottom channels, respectively. The inset shows the cell-trapping area in the top channel. The scale bar is 5 mm; the scale bar in the inset is 200 μm.
Operationally, the parameters used in this study to characterize the pressure drop along the top channel is Δh_top=0.25, 0.5, 0.75 and 1 mm, and that across the membrane is Δh_bottom=2 and 4 mm, in which Δh_top=h1–h2 and Δh_bottom=h1–h3 or h1–h4 (because h3=h4); h1 to h4 are the heights of the solution inside the corresponding wells (see Figure 2). Conceptually, Δh_top corresponds to the flow rate along the top channel, and Δh_bottom corresponds to the pressure difference across the membrane. Further details are provided in subsequent sections.
Cell-trapping rates in the microfluidic chip. The top, left photograph shows the microchip, where h1–4 represent the column heights in the four wells. The top, right micrograph presents cells trapped on three circular regions, each with 61 through-holes. The bottom figure shows the calculated cell-trapping rates based on Δh_top and Δh_bottom, in which Δh_top=h1–h2, Δh_bottom=h1–h3 (because h3=h4), and h1 remains unchanged, that is, h1=5 mm, during each test. The trapping rate is defined as the averaged percentage of the trapped cells in each pattern divided by 61. SKOV3 cells (approximately 1.5 × 103 cells) were introduced from the inlet in well 1, and each test was completed within 10 min. The asterisked value (red) indicates values obtained by tuning the solution heights in all four wells at the 5 min mark (total trapping time is 10 min) to that corresponding to the initial setting. The data are presented as the mean±s.d., n=3. CV: coefficient of variation. The scale bars in the top-left and top-right photographs are 5 mm and 200 μm, respectively.
Fabrication of the microfluidic chip
Fabrication began with alignment and permanent bonding of the two polydimethylsiloxane (PDMS) microchannels and a patterned perforated membrane with a diameter of 10 μm (Figure 1a). The top and bottom channels were fabricated by casting PDMS (Sylgard 184, Dow Corning, Midland, MI, USA) prepolymer against two photoresist-made (THB-151N, JSR, Sunnyvale, CA, USA) masters that contain two positive reliefs of orthogonal microchannels. The weight ratio of base to curing agent was 10:1. The dimensional size was 600 μm width × 85 μm height for the top channel and 8000 μm width × 40 μm height for the bottom channel. Slender posts, with a width of 50 μm and separated by 1100 μm, were designed into the bottom channel to prevent the collapse between the two channels. Holes of 1 mm in diameter were mechanically drilled through the cured channels to serve as fluidic inlet/outlet connections. The thin perforated membrane was fabricated by spin-coating PDMS prepolymer (15:1) on a glass substrate with a patterned array of 40-μm tall circular posts (with a diameter of 10 μm) that were constructed from photoresist (SU8-25, MicroChem, Newton, MA, USA). The thickness of the membrane was 10 μm after spin-coating at 2500 r.p.m. for 3 min. After curing at 70 °C for 1 h, the top channel was aligned and bonded with the membrane via an oxygen plasma treatment (15 sccm, 50 W, 30 s); subsequently, the bottom surface of the membrane was treated with oxygen plasma and bonded to the bottom channel. Polymethylmethacrylate (PMMA) nuts (5 mm in both diameter and height, inner volume of approximately 100 μl) were used as solution reservoirs and adhered onto the inlet/outlet holes of the chip via PDMS prepolymer cured at 70 °C for 1 h to achieve irreversible bonding. Figure 1b shows the top features of the microchip that contains four wells. The blue wells served as the inlet (well 1) and outlet (well 2) for the top channel, and the yellow wells (wells 3 and 4) served as the pressure sink for the bottom channel, which allowed the pressure difference across the perforated membrane to be tuned to optimize cell trapping.
Cell culture
Human ovarian cancer cell line SKOV3 (HTB-77, ATCC) was maintained in Dulbecco’s modified Eagle medium (31600-034, GIBCO, Carlsbad, CA, USA), supplemented with 10% fetal bovine serum (SV30014, Hyclone, South Logan, UT, USA), 1% penicillin/streptomycin (15140, GIBCO) and 1.5 g l−1 D-(+)-glucose (G5400-250G, SIGMA, St Louis, MO, USA), in a humidified 5% CO2 incubator at 37 °C.
Configurable 2D and 3D spheroid tissue cultures
Before loading of cells, the microfluidic chip was first sterilized using UV light in a laminar-flow hood for 1 h. The chip was then placed in the vacuum chamber to degas for 10 min and subsequently filled with 70% ethanol via manual pipetting. Afterwards, the chip was washed twice with phosphate-buffered saline (PBS) and stored in a sterilized dish by sealing with parafilm at 4 °C until needed.
For 2D cell patterning (Figure 3a), type I collagen gel (BD Biosciences, Franklin Lakes, NJ, USA; 100 μg ml−1 in PBS) was loaded into the bottom channel and placed in a humidified incubator at 37 °C for 1 h to allow binding to the membrane. Loading of collagen was performed via fluid height difference along the bottom channel (well 3 and well 4). Upon equalization to a medium-level height, local diffusion of collagen commenced across the perforated membrane onto the cell-supporting top surface. The chip was then coated with Pluronic copolymers (1% F108 in PBS, BASF Corp., Florham Park, NJ, USA) for 30 min at room temperature, followed by washing twice with PBS. Then, PBS and culture medium were introduced to wash away unbound Pluronic copolymers. To generate cell patterns, 100 μl of SKOV3 cells (approximately 1.5 × 103 cells) in culture medium were loaded into the top channel by tuning the solution-level height (that is, Δh_top=0.5 mm); meanwhile, a solution height in the bottom channel (that is, Δh_bottom=4 mm) was produced to create a suction to trap cells onto the microfabricated holes (see Supplementary Movie S1). Afterward, untrapped cells were washed away with 100 μl of medium solution, and the solution heights remained the same to enable cell attachment and spread on the perforated membrane in a 5% CO2 incubator at 37 °C. The patterned cells were observed under a microscope (DM IL, Leica, Wetzlar, Germany) and replaced with fresh culture medium daily.
Configurable two-dimensional (2D) cell patterning. (a) Schematic showing the cell-patterning procedure. Step 1: cross-sectional view of the membrane from A-A′ in Figure 1a. A cell-adhesive extracellular matrix (ECM) molecule is diffused from the bottom channel and absorbed onto the top surface of the perforated membrane. Step 2: Pluronic copolymers are then coated on the top membrane surface to prevent cell attachment. Step 3: SKOV3 cells are introduced into the top channel and then trapped on the through-holes. Step 4: cells are incubated at 37 °C and 5% CO2 to achieve the desired cell pattern. (b) Results obtained with ECM and Pluronic coating (via the procedure illustrated in a) and those obtained with only the ECM coating on the entire top surface; the results indicate the ability to pattern cells on demand and sustain cell viability (green) at day 2. Scale bar, 100 μm. (c) Characteristics of 2D cell patterning in the microchip, including patterning of single cells, different sizes of circular patterns and L-shaped symbols. Scale bars, 100 μm.
For 3D multicellular spheroids culture (see Figure 4a), the procedure was similar to that described for 2D culture, except that ECM protein coating was not performed. The chip was coated with Pluronic copolymers only to prevent cell attachment onto the perforated membrane. Subsequent to cell patterning, medium was also replaced daily to ensure freshness.
Three-dimensional (3D) spheroid culture in the microfluidic chip. (a) Schematics showing the procedure for 3D on-chip culture. The chip was coated with Pluronic copolymers to prevent the attachment of cells to the perforated membrane. The trapped cells (SKOV3 cells) will aggregate into cellular clusters. (b) Photographs of 3D spheroid culture in the microchip over 4 days. Cells conjoined within hours and definitively formed a spheroid at day 2. The white arrows indicate the location of the formed spheroid (the slender post is located in the bottom channel and optical transparent occurs due to the thinness of the membrane). (c) Immunofluorescence detection of epithelial marker (CD326/EpCAM (epithelial cell adhesion molecule), red), live cell staining (green) and Hoechst nuclear staining (blue) in the multicellular spheroid in the chip. (d) Multiple uniformly shaped cell spheroids were grown in the chip. The designed pattern (dash circles), which is 150 μm in diameter, will generate spheroids with a mean diameter of 78±10 μm (±s.d., n=8; coefficient of variation: 12.8%). Scale bars, 100 μm.
3D anchor-independent culture on dish
Before cell loading, plastic Petri dishes in 6-cm diameter were pre-coated with Pluronic copolymers (1% F108 in PBS) for 30 min at room temperature. After being washed twice with PBS, human ovarian cancer cells were plated by 1 × 104 cells with culture medium and incubated under 5% CO2 in an incubator at 37 °C. Cellular spheroids were observed by microscopy every day and replaced with fresh medium every week.
Live-cell labeling and imaging
To examine live cells, either 2D or 3D cultured cells in the microfluidic chip were stained with a concentration of 7 μM calcein AM (acetoxymethyl ester; C3100MP, Invitrogen, Carlsbad, CA, USA) in cell culture medium without serum. The chip was incubated under 5% CO2 at 37 °C for 30 min, and then washed with medium that contained serum; afterwards, bright-field and fluorescent images of the live cells were captured using a charge-coupled device camera (DP-70, Olympus, Shinjuku-ku, Tokyo, Japan) on an inverted microscope.
Evaluation of cell spheroid size
The size of the 3D tumor spheroids was calculated based on measurements of their diameters using imaging software (ImageJ, 1.42q, National Institutes of Health, Bethesda, MD, USA). As some spheroids were oval shaped, as shown in Figure 4, the mean diameter (l) of the cell spheroids was determined by the following equation:11 l=(a × b)1/2, where a and b represent two orthogonal diameters of the spheroids.
mRNA expression analysis
Total RNA of cells from different passages and chips were individually extracted and isolated using TRIzol reagent (Invitrogen). Complementary DNA (cDNA) was synthesized from 1 μg total RNA with a reverse-transcription PCR (RT-PCR). Primer sequences for real-time PCR were: GAPDH, E-cadherin (forward: 5′-TGCCCAGAAAATGAAAAAGG-3′; reverse: 5′-GTGTATGTGGCAATGCGTTC-3′), N-cadherin (forward: 5′-GACAATGCCCCTCAAGTGTT-3′; reverse: 5′-CCATTAAGCCGAGTGATGGT-3′), vimentin (forward: 5′-GAGAACTTTGCCGTTGAAGC-3′; reverse: 5′-GCTTCCTGTAGGTGGCAATC-3′); and fibronectin (forward: 5′-CAGTGGGAGACCTCGAGAAG-3′; reverse: 5′-TCCCTCGGAACATCAGAAAC-3′).
Real-time PCR using the mixture reagent SYBR-Green I (Thermo Scientific, San Diego, CA, USA) was performed in an iCycler iQ real-time detection system (Bio-Rad, Hercules, CA, USA). The specificity of the primers was confirmed from a single peak of the melting curve. Each target mRNA level was evaluated from the real-time threshold cycle and compared with the amount of GAPDH as an internal control.
Scratch-wound assays
Scratch wounds were made with a p1000 pipette tip on confluent cells, derived from both 2D dish culture and 3D on-chip culture, in 3.5-cm cell culture dishes (11035, SPL Lifescience, Gyeonggi-do, Korea). Wound closure was imaged using microscopy at × 10 magnification at 0 and 9 h. Three independent migration assays were performed for each condition. The area of the scratch wound was determined using image software (ImageJ, 1.42q), and the percentage of wound closure was defined as the area of the scratch wound at t=0 h divided by the area of wound closure at t=9 h.
Immunofluorescence
Human ovarian cancer cells used for 2D cell culture were prepared by plating approximately 1 × 104 cells in a 3.5-cm cell culture dish and incubating them overnight to enable cell attachment. The 3D cellular spheroids from on-dish or on-chip culture were collected, trypsinized into single cells, plated in a cell culture dish and incubated overnight. For epithelial cell adhesion molecule (EpCAM; anti-EpCAM-PE, 130-091-253, MACS, Bergisch Gladbach, Germany; 1:100 dilution) staining, cells were subsequently washed twice with PBS and fixed in 4% paraformaldehyde in PBS for 20 min at room temperature. For vimentin (anti-vimentin (V9), sc-6260 AF488, Santa Cruz Biotechnology, Santa Cruz, CA, USA; 1:200 dilution) staining, cells were fixed in 4% paraformaldehyde and permeabilized in 0.1% Triton X-100 in PBS for 30 min. Blocking was performed with PBS that contained 1% bovine serum albumin for 30 min, followed by two washings with PBS. The cells were then incubated with antibodies in PBS that contained 1% bovine serum albumin, and nuclei staining was subsequently performed using Hoechst (33342, Invitrogen) in PBS for 10 min. The dishes were extensively washed with PBS and mounted on a fluorescent microscopy stage (DM IL, Leica). Imaging was performed as previously described. Image acquisition and analysis were controlled using the MetaMorph software (Molecular Devices, Sunnyvale, CA, USA) using identical exposure times.
Statistical analysis
Statistical results from RT-PCR were presented as the mean±s.e.m., and the others were presented as the mean±s.d. The Student’s t-test was used to compare data from two groups of data (P<0.05 was considered statistically significant).
Results and discussion
The efficacy of the microchip in enabling cell patterning, deterministic 2D culture and subsequent viability, and the formation of 3D spheroids and their molecular characterization will be presented below. Additional information is also available in the Supplementary Material.
Controllable cell trapping in the microfluidic chip
Figure 2 presents results of 2D cell trapping in the microchip. The cell-trapping rates—the number of cells trapped in a circular region divided by 61 (the total number of through-holes in the circular region)—show no significant difference within the investigated range of Δh_top and Δh_bottom. In other words, both the flow rate along the top channel (characterized by Δh_top) and the pressure difference across the membrane (characterized by Δh_bottom) show no statistical significance for cell trapping. This result indicates that the net number of cells trapped in a particular circular region are approximately the same, although cells are trapped more rapidly in some cases (for example, Δh_bottom=4 mm). Note that, first, for all cases over the 10 min trapping period, the velocity of cells gradually decreased due to equilibrium of the solution heights in the corresponding wells. Second, the asterisked data represented in red text (that is, Δh_top=0.5 and Δh_bottom=4 mm in Figure 2) resulted in a trapping rate of 97%, which was obtained by tuning the solution heights in all four wells at the 5-min mark (total trapping time was 10 min) to that corresponding to the initial setting. This testing parameter—Δh_top=0.5 mm, Δh_bottom=4 mm and all solution heights reset at the 5-min mark—was used in all experiments thereafter. Third, the coefficient of variation (CV) of the cell count with respect to this parameter was 7.6%, which is comparable to that in our previous design for cell patterning (5.4%).4
The diameter of the through-holes was 10 μm, which is smaller than SKOV3 cells used in this study (approximately 18.3±3.8 μm based on our observation and those of others20). Cells are being trapped onto the membrane with none passing through into the bottom channel, as confirmed by detail image interrogation. With respect to the trapping efficiency, approximately 1500 cells in a cell suspension of 100 μl were injected into the chip with 999 holes and resulted in an average of 963 cells being trapped. Therefore, the trapping rate was 96% (963/999). If the conventional definition of trapping efficiency is considered, which is based on cells trapped per total cell input, the trapping efficiency was 64% (963/1500). However, this efficiency is a fictitious interpretation because a value of 100% is meaningless; the chip contained only 999 holes.
One important issue on cell viability during the trapping process is the consideration of fluidic shear stress. The shear stress on the cells within the channels can be estimated using the equation21

where  is the maximum shear stress on cells, μ is the viscosity of the solution (water=0.01 dyn s cm−2),
is the maximum shear stress on cells, μ is the viscosity of the solution (water=0.01 dyn s cm−2),  is the flow rate per unit width, and H is the height of the channel. The maximum shear stress for a flow velocity of 240 μm s−1 measured in the beginning of the cell injection was calculated to be 0.5 dyn cm−2, which is in the low range of shear stress present at different areas of systemic microvasculature.21 This result indicates that the shear stress adopted in this study by the simple passive microfluidic method could mitigate unwanted physical stimulus to the trapped cells.
is the flow rate per unit width, and H is the height of the channel. The maximum shear stress for a flow velocity of 240 μm s−1 measured in the beginning of the cell injection was calculated to be 0.5 dyn cm−2, which is in the low range of shear stress present at different areas of systemic microvasculature.21 This result indicates that the shear stress adopted in this study by the simple passive microfluidic method could mitigate unwanted physical stimulus to the trapped cells.
Configurable 2D cell patterning
Particular features of the microchip enabled deterministic patterning of cells in 2D via the coating procedure described earlier and illustrated in Figure 3a. One key feature is the perforated membrane, which allows ECM to diffuse from the bottom channel to cover the cell-supporting top surface of the membrane. Figure 3b compares the results of surface patterning between ECM plus the Pluronic coating, and that achieved only with ECM (coated on the top surface of the entire membrane) with the through-holes arranged in a circular configuration. Several results are noteworthy. First, for ECM plus the Pluronic coating, the trapped cells adhered onto the perforated membrane were restricted within the circular pattern and adjusted their morphology to closely follow the boundary (dash line). As the gap between neighboring through-holes is 40 μm, sufficient area remains to permit cell spreading. Second, for both surface-coating procedures, cell viability is demonstrated by calcein AM (green) of proliferated cells (from day 2). Third, in the case with only ECM coating, the trapped cells within the circles proliferated and spread to the entire top channel beyond the original circular pattern (with cells on through-holes only). Moreover, Figure 3c shows this chip enables configurable cellular patterns, including single cells, circular and L-shaped patterns, or any desirable pattern (see Supplementary Figure S2), through the redesign of the location of through-holes and suitable surface patterning. As shown in Supplementary Figure S2b and c, not only does the cell patterning approach have the ability to perform cell patterns down to single-cell dimensions, but it can also achieve a sharp interface between cell-binding and cell-resistant areas. In some cases, as shown in Supplementary Figure S2a, edge sharpness is compromised, possibly because of the yield of some fabricated through-holes, and the resulting lack of appropriate control over diffusion and spreading of ECM protein and F108 copolymer. However, the slight equivocality in sharpness is comparable to those of others12, 22, 23 and does not affect the presented results in any way.
The patterned cells adhered and were confined only to the region over the selected through-holes with the Pluronic coating. This result can be understood on the basis that, at the second step of the cell patterning process (Figure 3a), a monolayer of Pluronic copolymers was formed on the membrane. This monolayer prevents cell attachment because the cells’ hydrophobic polypropylene oxide (PPO) chains bind to the hydrophobic PDMS surface through hydrophobic–hydrophobic interactions and swing their hydrophilic chains freely in the medium. In general, because of the high surface density of the Pluronic copolymers employed in our method, large-sized cells were repelled from the particular surface (no through-holes distributed) by Pluronic’s steric repulsion, as reported elsewhere.14, 22, 24 Indeed, from the standpoint of wetting behavior, we also observed the ability of the copolymer-coated PDMS surface to repel proteins (see Supplementary Figure S1). Supplementary Figure S1 indicates that, because of PDMS’s hydrophobic nature, it tends to absorb small hydrophobic molecules in cell culture medium. In contrast, the copolymer-coated PDMS prevents the absorption of molecules or proteins and thus can potentially prevent cell attachment.
Further results (see Supplementary Figure S3) indicate the ability of the coating procedure to sustain the viability of the patterned cells. The trapped cells continued to spread for up to 6 days and remained viable and restricted within the patterned area during the entire duration. These results indicate that the microchip has the ability to sustain cell viability for long-term experiments.
Configurable 3D spheroid culture
Figure 4b–d presents particular results of the same epithelial SKOV3 cell line to enable 3D culturing in the microchip with the Pluronic coating procedure illustrated in Figure 4a, as previously discussed. First, Figure 4b shows the observation of the formation rate and size over a culture period of 4 days to allow an assessment of the growth of the spheroid in the microchip. Cells started to conjoin within hours and underwent multicellular assembly at 48 h; moreover, the spheroid reached a mean diameter of approximately 140 μm at 96 h. The spheroid was observed to be suspended in the microchannel because of its slight motion when the microchip was lightly disturbed. Even so, the spheroid is spatially confined during medium exchange within the pattern arranged by the through-holes due to the pressure drop across the perforated membrane. The cells likely became trapped on the perforated membrane due to cell–cell interactions and the anchor-independent culture condition. Second, Figure 4c shows staining results with epithelial cell marker (CD326), calcein AM for live cells and Hoechst for staining of the nucleus. The merged image demonstrates that this microchip has the potential for 3D culture and sustains the cell viability up to at least 5 days of culture. Finally, Figure 4d shows multiple spheroids performed on the same microchip at 96 h. Notably, the size of each spheroid was substantially uniform—approximately 78±10 μm in diameter—arranged by through-holes (150 μm pattern diameter). The CV for the spheroids was approximately 13%, whereas that formed in straight channels11 was approximately 20%.
Compared with the cellular spheroids generated via the microchip, several resembling traits derived from the on-dish culture with the Pluronic copolymer coating are evident (see the Materials and Methods section and Supplementary Figure S4). Supplementary Figure S4b presents a sample of an on-dish culture of spheroids. Such spheroids exhibit a large variation in their mean diameter (CV=107%), which is substantially larger than that from the microchip (CV=13%). The microenvironment presented by the microchip enables fewer cells to be handled and generates significantly more uniformly sized spheroids than the on-dish environment. Notably, the statistics for the CV values based on the spheroid volume would be more disparate; thus, this calculated CV will be larger than that based on the diameter. However, the CV values may be optimized and decreased by a redesign of the top channel as well as by the spatial control of uniform side chambers in a microchannel.11 Importantly, the improved CV in the present study is likely resulted from the deterministic control of equal cell numbers trapped on each perforated membrane, which indicates that this approach may be applied not only for tumor spheroids in cancer studies but also for uniformly sized embryoid bodies in stem cell research.
On-chip 3D spheroid culture generates and enhances the EMT properties on cells
Figure 5 shows the particular biological attributes (phenotypic/genotypic characterization) afforded by 3D spheroid culture in the microchip according to the previously described procedures in normal cell culture medium without any supplements of growth factors. The results show that (1) the monolayer cells derived from the cell spheroids cultured in the microchip reduced cell–cell interactions and acquired fibroblast-like mesenchymal morphology compared with 2D on-dish culture (Figure 5a); (2) downregulated mRNA encoding epithelial marker (E-cadherin, approximately 0.4-fold) and upregulated mRNAs encoding mesenchymal markers (N-cadherin, approximately 1.8-fold; vimentin, approximately 1.8-fold; fibronectin, approximately 198-fold; Figure 5b); and (3) an increased ability of migration compared with parental cancer cells (Figure 5c). These results also show that the cells derived from the microchip were reprogrammed toward epithelial–mesenchymal transition (EMT), similar to the hypoxic tumoral microenvironment.25 The measurements of the relative mRNAs from 3D on-chip and on-dish cultures (see Supplementary Figure S5) provided further indication that these cells derived from the chip exist in a mesenchymal-like cell state, which more closely resembles those that have undergone EMT than those from 3D on-dish culture. Moreover, Figure 5d presents results of expressions of CD326/EpCAM (broadly expressed on cells of epithelial origin and on epithelium-derived tumor cells26) and vimentin (often used as a marker of mesenchymally derived cells27). For 3D on-chip culture, the reduction in epithelial cell adhesion is evident by a marked decrease in CD326 expression and a gain in vimentin expression, which are two classic features of EMT, compared with that obtained from 2D on-dish culture.
Three-dimensional (3D) on-chip spheroid culture generates cells with EMT properties. (a) Comparison between the cell morphology with native SKOV3 cells (two-dimensional (2D) on-dish) and that derived from the microchip and replaced in tissue culture dishes (3D on-chip; spheroids cultured for 5 days). (b) Relative expression of the mRNAs that encode E-cadherin, N-cadherin, vimentin and fibronectin in the two cells from a, as evaluated by real-time reverse-transcription PCR. GAPDH was used as an internal control and to normalize the variability in sample loading. The data are presented as the mean±s.e.m., n=3. (c) Scratch-wound migration assays. Representative panels (top row) from a scratch wound assay show slow migration of native SKOV3 cells, whereas cells derived from spheroids (3D on-chip) migrate faster (bottom row). The enclosure areas were estimated using the ImageJ software package. The data are representative of mean±s.d., n=3. (d) Immunofluorescence detection of CD326/EpCAM (red), vimentin (green), and Hoechst nuclear staining (blue) on cell monolayers derived from 3D on-chip spheroid growth and 2D standard culture for 5 days. Total area of CD326 and vimentin signal intensities in the cell monolayers were measured using the MetaMorph image software. Each bar (right column) represents the mean±s.d. of relative intensity obtained from randomly captured images (n=5), *P<0.05, **P<0.01, ***P<0.001 compared to the control. Scale bars, 100 μm.
EMT has a crucial role during cancer invasion and metastasis, and it purportedly generates cells with properties of stem cells;25, 27, 28, 29 the present microfluidic platform appears to be unique in this regard. Several studies have demonstrated that EMT may be induced in cellular spheres cultured in serum-free medium supplemented with adequate mitogens, such as the basic fibroblast growth factor and epidermal growth factor, using a low-attachment culture dish; however, this approach appears to be costly and ineffective.27, 28, 30 These drawbacks are partially overcome in this study, in which the 3D microscale on-chip environment selects clones that are EMT transmitted. Moreover, the present approach does not require growth factors and possesses the potential to handle rare cells, which is especially important for clinical samples and circulating tumor cells.31 For comparison with conventional cultures, microscale cultures in the chip generally use significantly less reagent, which results in a greater volume density (1605 cells μl−1), that is, more cells in a specific volume of reagent, than in macrocultures in cell culture dishes (500 cells μl−1). The surface density4 of cells in the chip was 5.4 × 10−4 cells μm−2, which is comparable to that in dishes (with a surface density of 5.2 × 10−4 cells μm−2). In addition, the results show that epithelial cancer cells cultured by the 3D on-dish culture also confer an increased drug resistance for chemotherapeutic drugs (data not shown).
Sphere culture has been widely adopted for the enrichment of cancer stem cells, and there is increasing evidence that suggests a correlation exists between EMT and cancer stem cells that involves cell phenotypic and genotypic alternation.27, 28, 30, 32 This correlation is supported by our data, which shows that 3D on-chip culture conditions induced EMT from spheroids cultured using the epithelial SKOV3 ovarian cancer cell line. Similar evidence with the MCF7 breast cancer cell line27 and the human oral squamous cancer cell line30 has also been reported. In addition, our data are qualitatively consistent with those recently reported,27, 28 in which epithelial breast cancer cells grown in vitro also yielded differential EMT properties in 3D culture relative to 2D culture. As EMT has been recognized to have an important role in the metastatic process in primary tumors, the reverted process—mesenchymal to epithelial transition—is also crucial and recognized during tumor progress in the secondary organ environment.33, 34, 35 The measurements of relative mRNAs from cells in on-chip and off-chip cultures (see Supplementary Figure S6) further indicated that the cells from off-chip culture reduced mesenchymal expressions but increased epithelial traits, and that the cells underwent mesenchymal to epithelial transition, that is, the phenotypic plasticity observed in the results suggested that the microenvironment is crucial for inducing epigenetic changes.
Limitations to the present method include: (1) microfluidic patterning is incompatible with certain complex 3D structures, such as layer-by-layer assembly to mimic in vivo-like tissues;16, 17 (2) dynamic and continuous nutrient supply and waste removal to mimic the physiological microenvironments could not be easily achieved with the current proposed design (although this drawback could be overcome by integrating with an evaporation-based micropump36). However, the new capabilities allowed by the combination of techniques reported here (for example, the perforated membrane deterministically enabling cell trapping and in-situ generation of uniformly sized spheroids) may enable studies in 3D modeling of complex metastatic cascade (see results for the migration of tumor cells in 3D matrices in Supplementary Figure S7) and drug screening for metastatic cancers in 3D (via delivery of drugs diffused across the perforated cell culture areas9).
In summary, we presented a microfluidic chip to deterministically enable configurable 2D monolayer and 3D spheroid cell cultures on demand. Toward this end, ECM proteins and Pluronic copolymers were used to selectively coat the perforated membrane. One key result of this study was that cells cultured in the microchip configured into a spherical geometry and acquired EMT properties due to the microscale 3D in vivo-like environments. This feature might be useful for the study of cancer metastatic cascade in vitro and thereby potentially achieve a long-term goal of realizing ‘organotypic cultures’ that represent a tumor microenvironment for personalized drug testing.5, 25, 29 Furthermore, the passive method that enables the pumping of multiple fluids using pipettes has been widely adopted, which has the advantages of ease of use, accessibility and a reduction of the complexity of working with pumps and tubing.11, 12, 16, 37 Here, we successfully adapt these methods in the microchip, which indicates that the microchip could be adopted for use in existing, widely used high-throughput screening instruments, such as liquid-handling robots.3 This easy, flexible and effective approach may be used for a range of applications in drug discovery, tissue engineering or stem cell research, and should be feasible to integrate with other biological assay components into a monolithic system.
References
Khetani, S. R. & Bhatia, S. N. Microscale culture of human liver cells for drug development. Nat. Biotechnol. 26, 120–126 (2008).
Lindstron, S., Larsson, R. & Svahn, H. A. Towards high-throughput single cell/clone cultivation and analysis. Electrophoresis 29, 1219–1227 (2008).
Tung, Y. C., Hsiao, A. Y., Allen, S. G., Torisawa, Y. -S., Ho, M. & Takayama, S. High-throughput 3D spheroid culture and drug testing using a 384 hanging drop array. Analyst 136, 473–478 (2011).
Hsiung, L. -C., Chiang, C. -L., Wang, C. -H., Huang, Y. -H., Kuo, C. -T., Cheng, J. -Y., Lin, C. -H., Wu, V., Chou, H. -Y., Jong, D. -S., Lee, H. & Wo, A. M. Dielectrophoresis-based cellular microarray chip for anticancer drug screening in perfusion microenvironments. Lab Chip 11, 2333–2342 (2011).
Benbrook, D. M. Organotypic cultures represent tumor microenvironment for drug testing. Drug Discov. Today Dis. Models 3, 143–148 (2006).
Lutolf, M. P., Gilbert, P. M. & Blau, H. M. Designing materials to direct stem-cell fate. Nature 462, 433–441 (2009).
Choi, N. W., Cabodi, M., Held, B., Gleghorn, J. P., Bonassar, L. J. & Stroock, A. D. Microfluidic scaffolds for tissue engineering. Nat. Mater. 6, 908–915 (2007).
Wan, L. Q., Ronaldson, K., Park, M., Taylor, G., Zhang, Y., Gimble, J. M. & Vunjak-Novakovic, G. Micropatterned mammalial cells exhibit phenotype-specific left-right asymmetry. PNAS 108, 12295–12300 (2011).
VanDersarl, J. J., Xu, A. M. & Melosh, N. A. Rapid spatial and temporal controlled signal delivery over large cell culture areas. Lab Chip 11, 3057–3063 (2011).
Jamil, E. -A., Sorger, P. K. & Jensen, K. F. Cells on chips. Nature 442, 403–441 (2006).
Torisawa, Y. -S., Chueh, B. -H., Huh, D., Ramamurthy, P., Roth, T. M., Barald, K. F. & Takayama, S. Efficient formation of uniform-sized embryoid bodies using a compartmentalized microchannel device. Lab Chip 7, 770–776 (2007).
Berthier, E., Warrick, J., Casavant, B. & Beebe, D. J. Pipette-friendly laminar flow patterning for cell-based assays. Lab Chip 11, 2060–2065 (2011).
Ho, C. -T., Lin, R. -Z., Chang, W. -Y., Chang, H. -Y. & Liu, C. -H. Rapid heterogeneous liver-cell on-chip patterning via the enhanced filed-induced dielectrophoresis trap. Lab Chip 6, 724–734 (2006).
Fan, C. Y., Yung, Y. -C., Takayama, S., Meyhofer, E. & Kurabayashi, K. Electrically programmable surfaces for configurable patterning of cells. Adv. Mater. 20, 1418–1423 (2008).
Souza, G. R., Molina, J. R., Raphael, R. M., Ozawa, M. G., Stark, D. J., Levin, C. S., Bronk, L. F., Ananta, J. S., Mandelin, J., Georgescu, M. -M., Bankson, J. A., Gelovani, J. G., Killian, T. C., Arap, W. & Pasqualini, R. Three-dimensional tissue culture based on magnetic cell levitation. Nat. Nanotechnol. 5, 291–296 (2010).
Derda, R., Laromaine, A., Mammoto, A., Tang, S. K. Y., Mammoto, T., Ingber, D. E. & Whitesides, G. M. Paper-supported 3D cell culture for tissue-based bioassays. PNAS 106, 18457–18462 (2009).
Yuan, B., Jin, Y., Sun, Y., Wang, D., Sun, J., Wang, Z., Zhang, W. & Jiang, X. A strategy for depositing different types of cells in three dimensions to mimic tubular structures in tissues. Adv. Mater. 24, 890–896 (2012).
Hardelauf, H., Frimat, J. -P., Stewart, J. D., Schormann, W., Chiang, Y. -Y., Lampen, P., Franzke, J., Hengstler, J. G., Cadenas, C., Kunz-Schughart, L. A. & West, J. Microarrays for the scalable production of metabolically relevant tumor spheroids: a tool for modulating chemosensitivity traits. Lab Chip 11, 419–428 (2011).
Chen, C. -L., Chen, K. -C., Pan, Y. -C., Lee, T. -P., Hsiung, L. C., Lin, C. -M., Chen, C. -Y., Lin, C. -H., Chiang, B. -L. & Wo, A. M. Separation and detection of rare cells in a microfluidic disk via negative selection. Lab Chip 11, 474–483 (2011).
Nikitin, M. P., Zdobnova, T. A., Lukash, S. V., Stremovskiy, O. A. & Deyev, S. M. Protein-assisted self-assembly of multifunctional nanoparticles. PNAS 107, 5827–5832 (2010).
Farokhzad, O. C., Khademhosseini, A., Jon, S., Hermmann, A., Cheng, J., Chin, C., Kiselyuk, A., Teply, B., Eng, G. & Langer, R. Microfluidic system for studying the interaction of nanoparticles and microparticles with cells. Anal. Chem. 77, 5453–5459 (2005).
Liu, V. A., Jastromb, W. E. & Bhatia, S. N. Engineering protein and cell adhesivity using PEO-terminated triblock polymers. J. Biomed. Mater. Res. 60, 126–134 (2002).
Corey, J. M., Gertz, C. C., Sutton, T. J., Chen, Q., Mycek, K. B., Wang, B. -S., Martin, A. A., Johnson, S. L. & Feldman, E. L. Patterning N-type and S-type neuroblastoma cells with Pluronic F108 and ECM proteins. J. Biomed. Mater. Res. A 93, 673–686 (2010).
Cheng, Q. & Komvopoulos, K. Integration of plasma-assisted surface chemical modification, soft lithography, and protein surface activation for single-cell patterning. Appl. Phys. Lett. 97, 043705 (2010).
Thiery, J. P., Acloque, H., Huang, R. Y. & Nieto, M. A. Epithelial-mesenchymal transitions in development and disease. Cell 139, 871–890 (2009).
Mikolajczyk, S. D., Millar, L. S., Tsinberg, P., Coutts, S. M., Zomorrodi, M., Pham, T., Bischoff, F. Z. & Pircher, T. J. Detection of EpCAM-negative and cytokeratin-negative circulating tumor cells in peripheral blood. J. Oncol. 2011, 252361 (2011).
Guttilla, I. K., Phoenix, K. N., Hong, X., Tirnauer, J. S., Claffey, K. P. & White, B. A. Prolonged mammosphere culture of MCF-7 cells induces an EMT and repression of the estrogen receptor by microRNAs. Breast Cancer Res. Treat. 132, 75–85 (2011).
Mani, S. A., Guo, W., Liao, M. -J., Eaton, E. N., Ayyanan, A., Zhou, A. Y., Books, M., Reinhard, F., Zhang, C. C., Shipitsin, M., Campbell, L. L., Polyak, K., Brisken, C., Yang, J. & Weinberg, R. A. The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell 133, 704–715 (2008).
Chaffer, C. L. & Weinberg, R. A. A perspective on cancer cell metastasis. Science 331, 1559–1564 (2011).
Chen, S. -F., Chang, Y. -C., Nieh, S., Liu, C. -L., Yang, C. -Y. & Lin, Y. -S. Nonadhesive culture system as a model of rapid sphere formation with cancer stem cell properties. PLOS One 7, e31864 (2012).
Pecot, C. V., Bischoff, F. Z., Mayer, J. A., Wong, K. L., Pham, T., Bottsford-Miller, J., Stone, R. L., Lin, Y. G., Jaladurgam, P., Roh, J. W., Goodman, B. W., Merritt, W. M., Pircher, T. J., Mikolajczyk, S. D., Nick, A. M., Celestino, J., Eng, C., Ellis, L. M., Deavers, M. T. & Sood, A. K. A. Novel platform for detection of CK+ and CK− CTCs. Cancer Discov. 1, 580–586 (2011).
Chen, C., Wei, Y., Hummel, M., Hoffmann, T. K., Gross, M., Kaufmann, A. M. & Albers, A. E. Evidence for epithelial-mesenchymal transition in cancer stem cells of head and neck squamous cell carcinoma. PLOS One 6, e16466 (2011).
Chao, Y. L., Shepard, C. R. & Wells, A. Breast carcinoma cells re-express E-cadherin during mesenchymal to epithelial reverting transition. Mol. Cancer 9, 179 (2010).
Aokage, K., Ishii, G., Ohtaki, Y., Yamaguchi, Y., Hishida, T., Yoshida, J., Nichimura, M., Nagai, K. & Ochiai, A. Dynamic molecular changes associated with epithelial-mesenchymal transition and subsequent mesenchymal-epithelial transition in the early phase of metastatic tumor formation. Int. J. Cancer 128, 1585–1595 (2011).
Chen, J., Wang, L., Matyunina, L. V., Hill, C. G. & McDonald, J. F. Overexpression of miR-429 induces mesenchymal-to-epithelial transition (MET) in metastatic ovarian cancer cells. Gynecol. Oncol. 121, 200–205 (2011).
Effenhauser, C. S., Harttig, H. & Kramer, P. An evaporation-based disposable micropump concept for continuous monitoring applications. Biomed. Microdevices 4, 27–32 (2002).
Kuo, C. -T., Chiang, C. -L., Ruby, Y. J. H., Lee, H. & Wo, A. M. Probing the traits of epithelial-mesenchymal transition in a microfluidic device in 15th International Conference on Miniaturized Systems for Chemistry and Life Sciences 145–147 Seattle: Washington, USA, (2011).
Acknowledgements
We are grateful to Professor Jiahn-Chun Wu for providing SKOV3 cancer cells and Mr Chen-Tse Wu for assistance with real time RT-PCR analyses. Financial support from the National Science Council, Taiwan, under grant NSC 100-2120-M-002-001 is gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the NPG Asia Materials website
Supplementary information
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Kuo, CT., Chiang, CL., Yun-Ju Huang, R. et al. Configurable 2D and 3D spheroid tissue cultures on bioengineered surfaces with acquisition of epithelial–mesenchymal transition characteristics. NPG Asia Mater 4, e27 (2012). https://doi.org/10.1038/am.2012.50
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/am.2012.50
Keywords
This article is cited by
-
Scaffold-based 3D cell culture models in cancer research
Journal of Biomedical Science (2024)
-
Prevascularized Micro-/Nano-Sized Spheroid/Bead Aggregates for Vascular Tissue Engineering
Nano-Micro Letters (2021)
-
Unravelling Receptor and RGD Motif Dependence of Retargeted Adenoviral Vectors using Advanced Tumor Model Systems
Scientific Reports (2019)
-
Engineering 3D approaches to model the dynamic microenvironments of cancer bone metastasis
Bone Research (2018)
-
Three-dimensional spheroid culture targeting versatile tissue bioassays using a PDMS-based hanging drop array
Scientific Reports (2017)